Abstract
Background:
The emergence of early-onset colorectal cancer (EOCRC), particularly among populations with disproportionate health burdens, has exposed critical gaps in our understanding of age- and ancestry-specific oncogenic processes. Aberrations in the mitogen-activated protein kinase (MAPK) pathway—encompassing KRAS, NRAS, BRAF, MAP2K1, MAPK3, and others—are central to tumorigenesis, yet their distribution, prognostic impact, and therapeutic implications in EOCRC remain poorly defined.
Methods:
We developed AI-HOPE-MAPK, a conversational artificial intelligence platform built upon a fine-tuned biomedical large language model (LLaMA 3) to enable real-time, natural language-driven analysis of clinical and genomic data. The platform integrates harmonized datasets from TCGA, cBioPortal, and AACR Project GENIE and supports cohort construction, mutation frequency comparisons, survival modeling, and odds ratio estimation across diverse subgroups. We applied AI-HOPE-MAPK to characterize MAPK pathway alterations in colorectal cancer by age, race/ethnicity, microsatellite instability (MSI), tumor mutational burden (TMB), treatment history, and anatomical site.
Results:
AI-HOPE-MAPK identified significant age- and ancestry-specific patterns in MAPK alterations. NF1 and MAP2K1 mutations were enriched in Hispanic/Latino (H/L) EOCRC cases, with NF1 mutations reaching statistical significance (p = 0.045). Comparative analyses revealed elevated MAPK3 and NF1 mutation frequencies in H/L and non-Hispanic White EOCRC patients. MAP2K1-mutated stage I–III tumors exhibited worse survival and lower FOLFOX treatment rates (p = 0.0447 and p = 0.015, respectively). Kaplan–Meier and odds ratio analyses showed that younger patients with low-TMB MAPK-altered tumors had significantly poorer outcomes and were overrepresented in aggressive molecular subtypes. Furthermore, MAPK alterations conferred significantly reduced survival in microsatellite-stable tumors but not in MSI-H cases, underscoring context-specific prognostic effects. Anatomical stratification showed site-specific impacts of AKT1 mutations, with poorer outcomes in colon versus rectal cancers.
Conclusions:
AI-HOPE-MAPK provides a promising framework for AI-driven precision oncology by enabling intuitive, scalable, and population-aware exploration of MAPK pathway dysregulation in colorectal cancer. In this study, the platform identified differences in mutation prevalence, survival outcomes, and treatment access, particularly among patients with disproportionate health burdens. By integrating language-based user interfaces with rigorous genomic analytics, AI-HOPE-MAPK facilitates biomarker exploration, supports more equitable research practices, and offers a tool for generating hypotheses that may inform future efforts to refine precision oncology across diverse patient populations.
Introduction
Colorectal cancer (CRC) remains a major global health challenge, ranking as the third most common malignancy and the second leading cause of cancer-related deaths worldwide.1,2 Although CRC incidence in older adults has plateaued or declined in high-income countries, the global rise in early-onset colorectal cancer (EOCRC)—defined as diagnosis before age 50—has garnered increasing attention.3–6 In the United States, this trend is especially pronounced among Hispanic/Latino (H/L) populations, who face the most significant increase in EOCRC incidence as well as disproportionately high mortality rates compared with non-Hispanic White (NHW) individuals.7–12 These disparities are compounded by later-stage diagnoses resulting from current screening guidelines that historically initiated routine surveillance at age 50. 13
Beyond delayed detection, EOCRC displays distinct molecular and immunological features when compared with late-onset CRC (LOCRC). Studies have identified a higher prevalence of microsatellite instability (MSI), increased tumor mutation burden, and overexpression of immune checkpoint proteins such as PD-L1 in EOCRC cases.12–17 Epigenetic modifications, such as LINE-1 hypomethylation, have also emerged as potential molecular signatures distinguishing EOCRC from LOCRC.18–21 These findings suggest that EOCRC may be driven by unique, age- and ancestry-specific oncogenic mechanisms that remain poorly understood in racially and ethnically diverse populations.
Among the most critical pathways implicated in CRC development is the mitogen-activated protein kinase (MAPK) signaling cascade. Comprising ERK, BMK-1, JNK, and p38 branches, the MAPK pathway orchestrates key cellular processes such as proliferation, differentiation, and apoptosis.22–25 In CRC, aberrant MAPK activation—often through RAS and BRAF mutations—leads to uncontrolled tumor growth, metastasis, and resistance to therapy.23,26,27 Importantly, MAPK reactivation via EGFR signaling has been shown to confer resistance to BRAF inhibitors like vemurafenib, underscoring the clinical relevance of pathway-specific therapeutic vulnerabilities. 20 Despite extensive research into MAPK-driven CRC biology,28–32 its role in shaping EOCRC disparities—particularly among H/L individuals—remains largely unexplored.
To address this critical knowledge gap, we developed AI-HOPE-MAPK, a conversational artificial intelligence platform designed to identify, analyze, and interpret MAPK pathway alterations in CRC using clinical and genomic data. AI-HOPE-MAPK integrates publicly available datasets, supports natural language-driven analysis, and enables population-aware interrogation of pathway-specific alterations. Here, we apply AI-HOPE-MAPK to characterize the molecular landscape of MAPK signaling in EOCRC across diverse patient groups. By linking genomic alterations to clinical outcomes, this study aims to uncover actionable targets and support the development of tailored precision oncology strategies for different communities.
Methods
AI-HOPE-MAPK was developed as an artificial intelligence—powered system designed to translate natural language inquiries into rigorous, clinically meaningful genomic analyses, with a specific focus on alterations in the MAPK signaling pathway in CRC (Fig. 1). The system is built upon a fine-tuned biomedical large language model (LLM) based on the LLaMA 3 architecture, which enables semantic understanding of user queries and automatic generation of analytical workflows. Researchers can input complex, multilayered questions—for example, assessing survival outcomes associated with specific mutations in microsatellite-stable (MSS), early-onset, ethnicity-specific CRC cases—and receive comprehensive statistical outputs without the need for computational programming expertise.
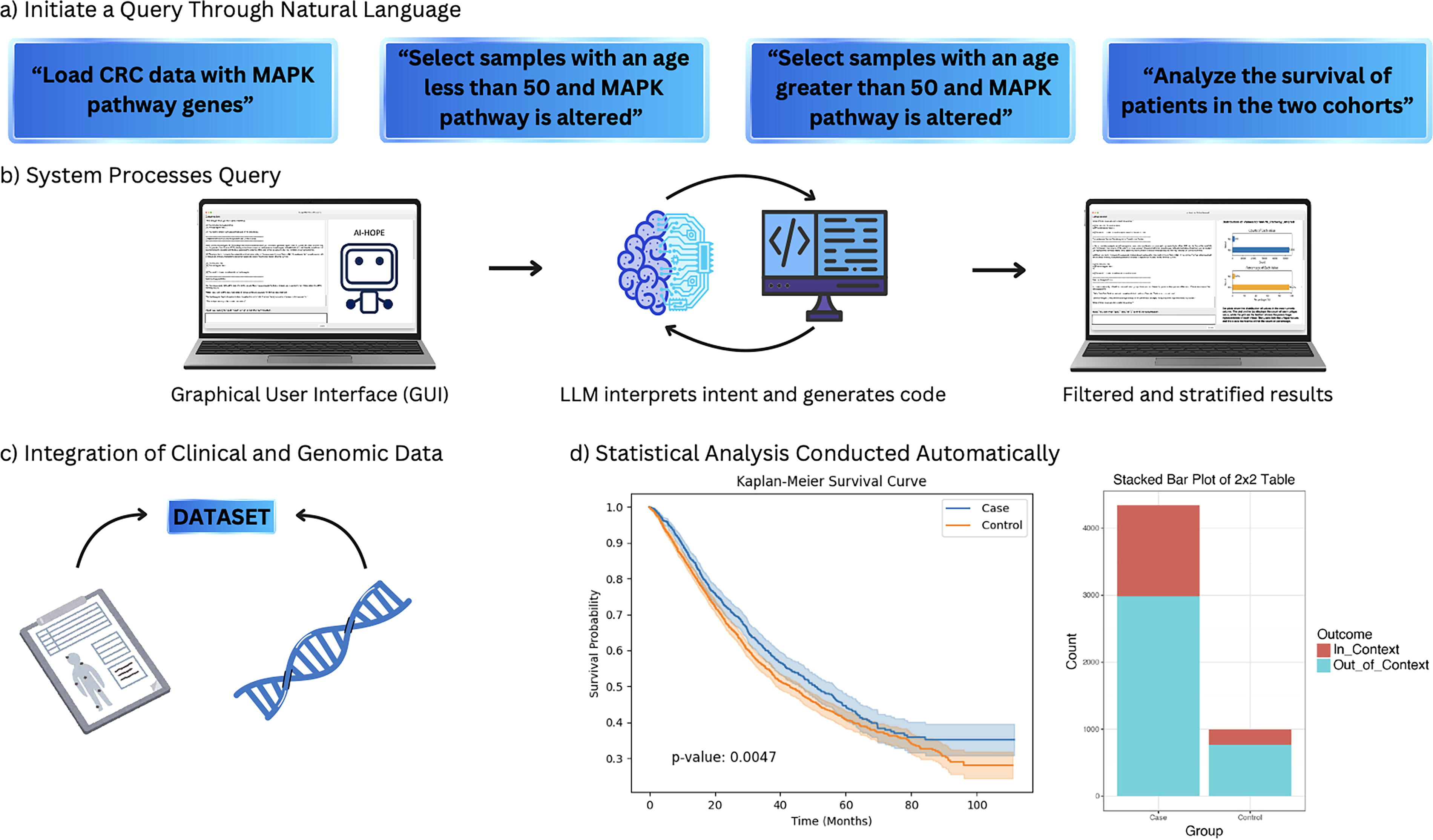
AI-HOPE-MAPK workflow for discovering clinically relevant MAPK alterations in colorectal cancer. This figure presents a visual summary of the AI-HOPE-MAPK system, an interactive artificial intelligence platform designed to explore MAPK pathway alterations in colorectal cancer (CRC) through natural language queries.
The platform integrates harmonized clinical and genomic data from cBioPortal and the AACR Project GENIE. These datasets were filtered to include colorectal adenocarcinomas with complete annotations for somatic mutations, copy number alterations, MSI status, tumor site, clinical stage, patient race/ethnicity, age at diagnosis, treatment history, and survival time. A standardized preprocessing pipeline was employed to normalize patient identifiers, resolve inconsistent annotations, and map all gene alterations to predefined MAPK pathway components, including KRAS, NRAS, BRAF, MAP2K1, MAPK1, and MAPK14. Tumors were stratified by anatomical location (left colon, right colon, rectum), MSI phenotype, and racial or ethnic background, with particular emphasis on distinguishing EOCRC (diagnosed before age 50) from later-onset disease.
Upon receiving a user query, AI-HOPE-MAPK applies a semantic parsing engine to extract analytical intent, identify the genes and pathways involved, and apply relevant cohort filters. This automated translation process initiates a statistical pipeline tailored to the nature of the question. The system supports frequency analysis of mutations and copy number alterations across clinical subgroups using chi-square or Fisher’s exact tests, calculates odds ratios and 95% confidence intervals to assess enrichment of alterations by demographic or molecular category, and conducts time-to-event analyses including Kaplan–Meier estimation, log-rank testing, and Cox proportional hazards regression. Pathway interaction modeling is also enabled, including co-occurrence and mutual exclusivity analysis with other relevant oncogenic pathways such as TP53, TGF-β, and PI3K.
To validate the system’s performance, we reproduced known associations between MAPK pathway mutations and clinical outcomes in CRC, including the poor prognosis of BRAF V600E mutations and the differential impact of RAS mutations in MSS tumors. The platform was benchmarked against publicly available bioinformatics tools such as cBioPortal and UCSC Xena, with evaluation criteria including cohort construction accuracy, response time, and clarity of output. In all test cases, AI-HOPE-MAPK demonstrated strong reproducibility and superior flexibility in handling queries that involved multiple intersecting variables, including ethnicity, tumor location, and MSI status.
All analytical outputs are returned as high-resolution visualizations and narrative summaries. These include Kaplan–Meier survival curves annotated with p-values and sample sizes, forest plots of estimated effect sizes, co-mutation heatmaps, and subgroup-specific mutation frequency charts. Result tables are exportable in multiple formats for downstream use in article, presentations, or supplemental materials. The conversational interface also supports iterative querying, enabling users to refine hypotheses or generate stratified comparisons across subgroups in real time.
By bridging the gap between biomedical language understanding and pathway-specific clinical-genomic analysis, AI-HOPE-MAPK provides a powerful, accessible tool for investigating the molecular underpinnings of CRC and identifying disparities in disease presentation and outcomes across diverse populations.
Results
AI-HOPE-MAPK provides an intuitive, conversation-based framework for investigating clinically relevant alterations in the MAPK signaling pathway among CRC patients from multiple populations. Leveraging natural language inputs, the system constructs cohort-specific analyses spanning mutation frequency comparisons, survival modeling, and odds ratio testing, while incorporating key clinical variables such as age, ethnicity, tumor site, and treatment history. The platform enables hypothesis-driven exploration of CRC disparities, particularly among early-onset and H/L subgroups, through real-time, data-integrated insights.
Age-associated molecular trends in population-specific CRC
AI-HOPE-MAPK was first applied to explore molecular distinctions between EOCRC and LOCRC in H/L patients. Although infrequent, ACVR1 mutations appeared more often in EOCRC H/L cases (1.96%) than in LOCRC H/L counterparts (0.49%), yielding an odds ratio of 4.06 (95% CI: 0.418–39.416; p = 0.425) (Fig. 2). While not statistically significant, this pattern hints at age-specific enrichment requiring validation. In contrast, NF1 mutations showed a statistically significant difference, with a 10.46% prevalence in EOCRC H/L samples compared with 4.41% in LOCRC H/L (OR = 2.53; 95% CI: 1.087–5.893; p = 0.045) (Supplementary Fig. S1), suggesting a potentially pathogenic role in early-onset disease.
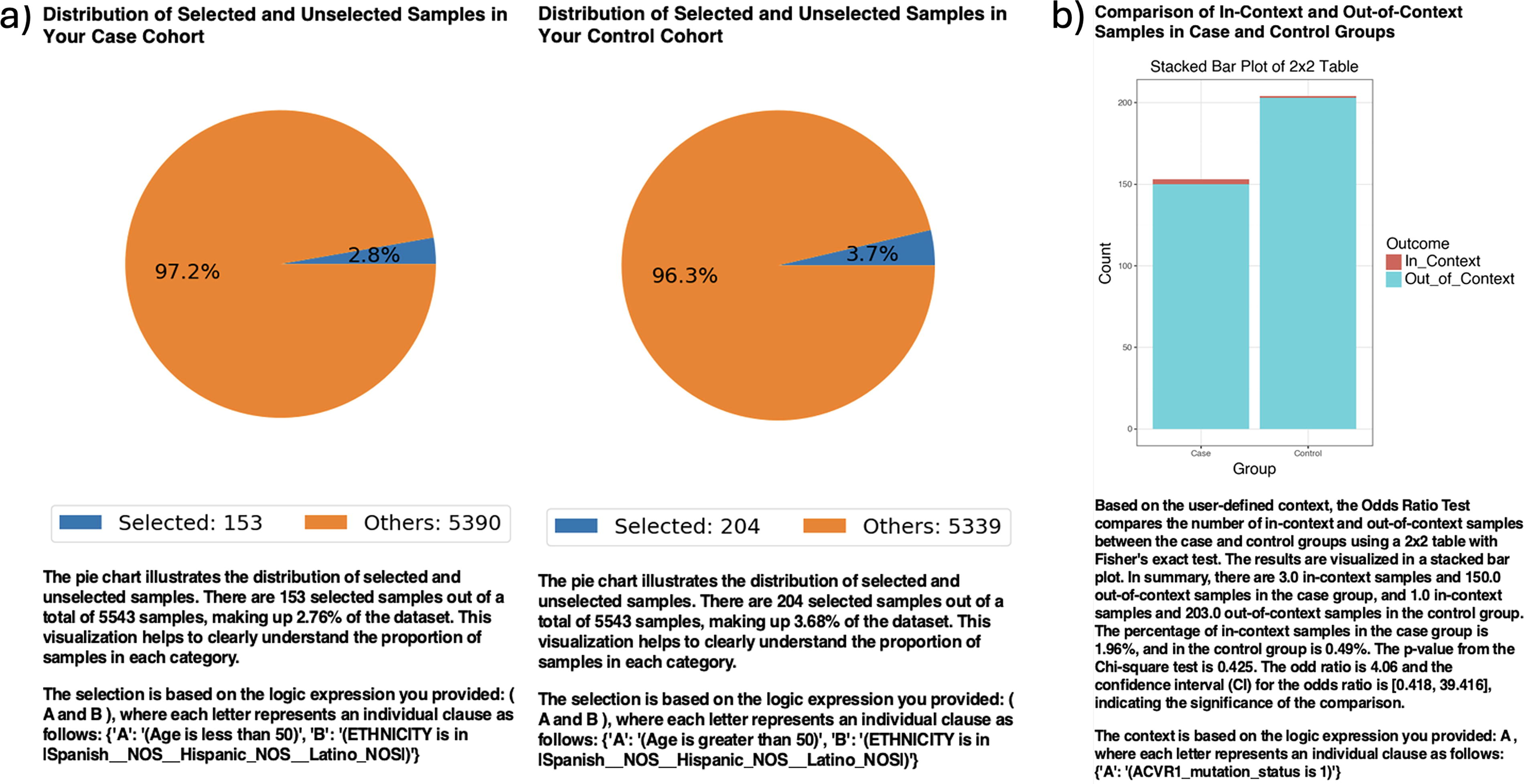
AI-HOPE-MAPK analysis of ACVR1 mutation frequency in early-onset versus late-onset Hispanic/Latino colorectal cancer patients.
MAP2K1 alterations—although rare—were restricted almost entirely to the EOCRC H/L group (2.61% vs. 0.24% in LOCRC H/L), producing an infinite odds ratio (95% CI: 0.575–208.75; p = 0.07) due to absence in the older cohort (Supplementary Fig. S2). In contrast, BRAF mutations were more prominent in LOCRC H/L samples (17.65%) than EOCRC H/L (5.23%) and achieved statistical significance (OR = 3.88; 95% CI: 1.749–8.624; p = 0.001) (Supplementary Fig. S3), underscoring the potential age-dependent divergence in MAPK pathway biology.
Ancestry-informed differences in MAPK gene alterations
To assess population-level disparities, AI-HOPE-MAPK compared mutation patterns between H/L and NHW EOCRC patients. While AKT1 mutation rates showed a modest, nonsignificant increase in H/L cases (3.27% vs. 1.52%, OR = 2.19; 95% CI: 0.795–6.013; p = 0.222) (Supplementary Fig. S4), MAPK3 mutations were significantly more common in the H/L group (2.61% vs. 0.63%; OR = 4.26; 95% CI: 1.232–14.715; p = 0.043) (Supplementary Fig. S5). Similarly, NF1 mutations occurred at a higher rate in H/L patients (10.46%) compared with NHW (5.37%) with statistical significance (OR = 2.06; 95% CI: 1.153–3.673; p = 0.021) (Supplementary Fig. S6). PDGFRB mutations showed a similar trend (3.92% vs. 1.97%, OR = 2.03; 95% CI: 0.81–5.092; p = 0.212), though they did not reach significance (Supplementary Fig. S7). These observations reinforce the relevance of ancestry-stratified analyses in identifying population-specific MAPK alterations.
Therapeutic disparities and treatment context
AI-HOPE-MAPK further examined age-specific responses to standard FOLFOX chemotherapy in H/L CRC patients. Kaplan–Meier analysis demonstrated significantly worse survival among early-onset cases compared with late-onset patients (p = 0.0047) (Fig. 3). Odds ratio testing revealed that H/L individuals comprised a disproportionately higher fraction of the early-onset group (10.29%) than the late-onset group (5.35%) (OR = 1.957; 95% CI: 1.534–2.497; p < 0.001), suggesting a potential convergence of ancestry and age-related vulnerability in CRC.
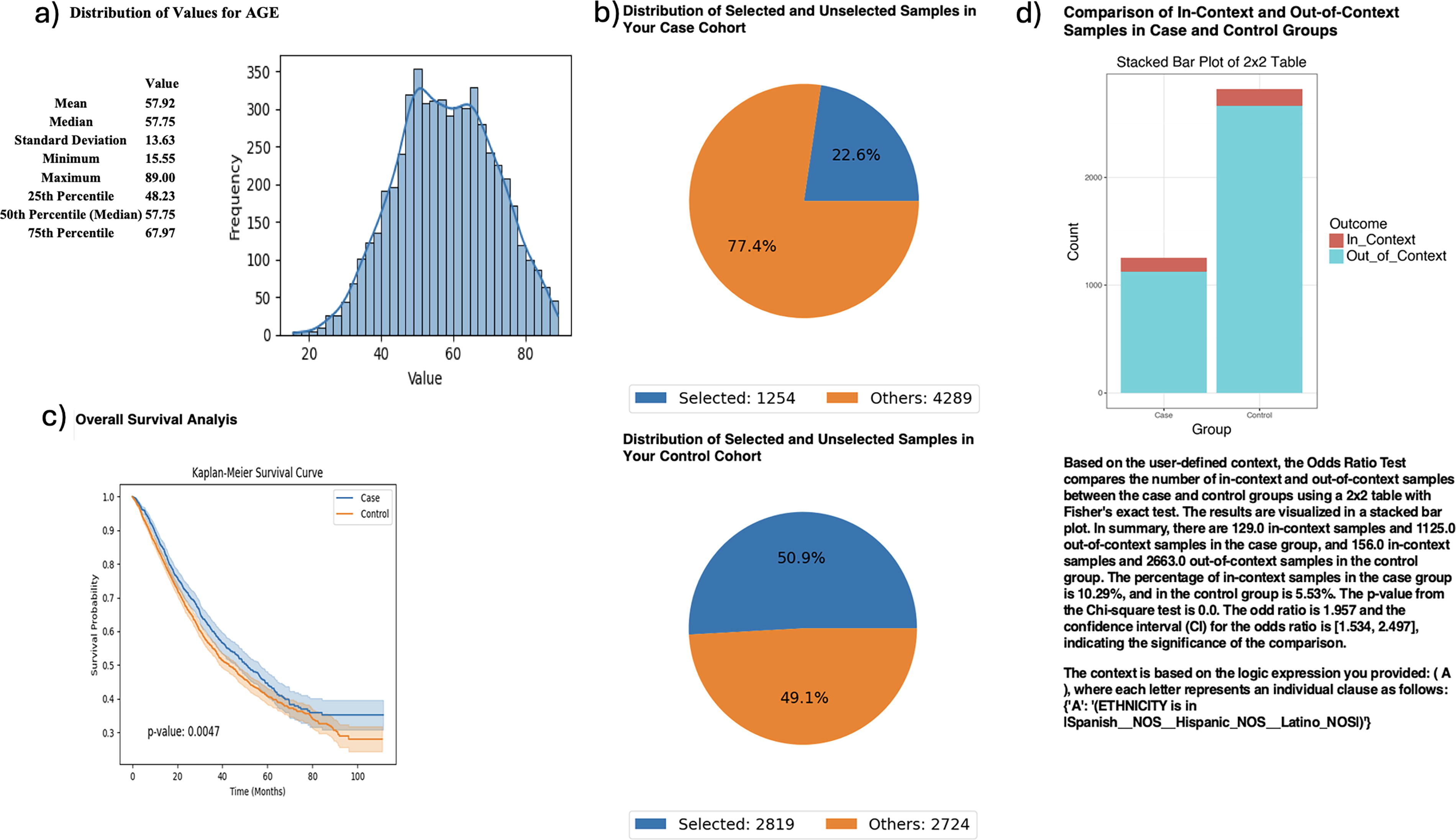
AI-HOPE-MAPK analysis of age-stratified Hispanic/Latino colorectal cancer patients treated with FOLFOX chemotherapy. This figure demonstrates the use of AI-HOPE-MAPK to investigate age-related survival patterns and ethnic disparities among colorectal cancer (CRC) patients treated with the FOLFOX regimen (Fluorouracil, Leucovorin, Oxaliplatin).
Survival differences by MSI and MAPK interaction
AI-HOPE-MAPK was used to assess how MAPK alterations intersect with MSI status in FOLFOX-treated patients. Although age-based enrichment of younger patients did not differ significantly between MSS and MSI-H groups (OR = 1.206; 95% CI: 0.893–1.629; p = 0.251), survival analysis showed a striking disadvantage for MSS tumors with MAPK alterations compared with their MSI-H counterparts (p < 0.0001) (Fig. 4). This divergence underscores the importance of MSI context in stratifying risk among MAPK-driven CRC.
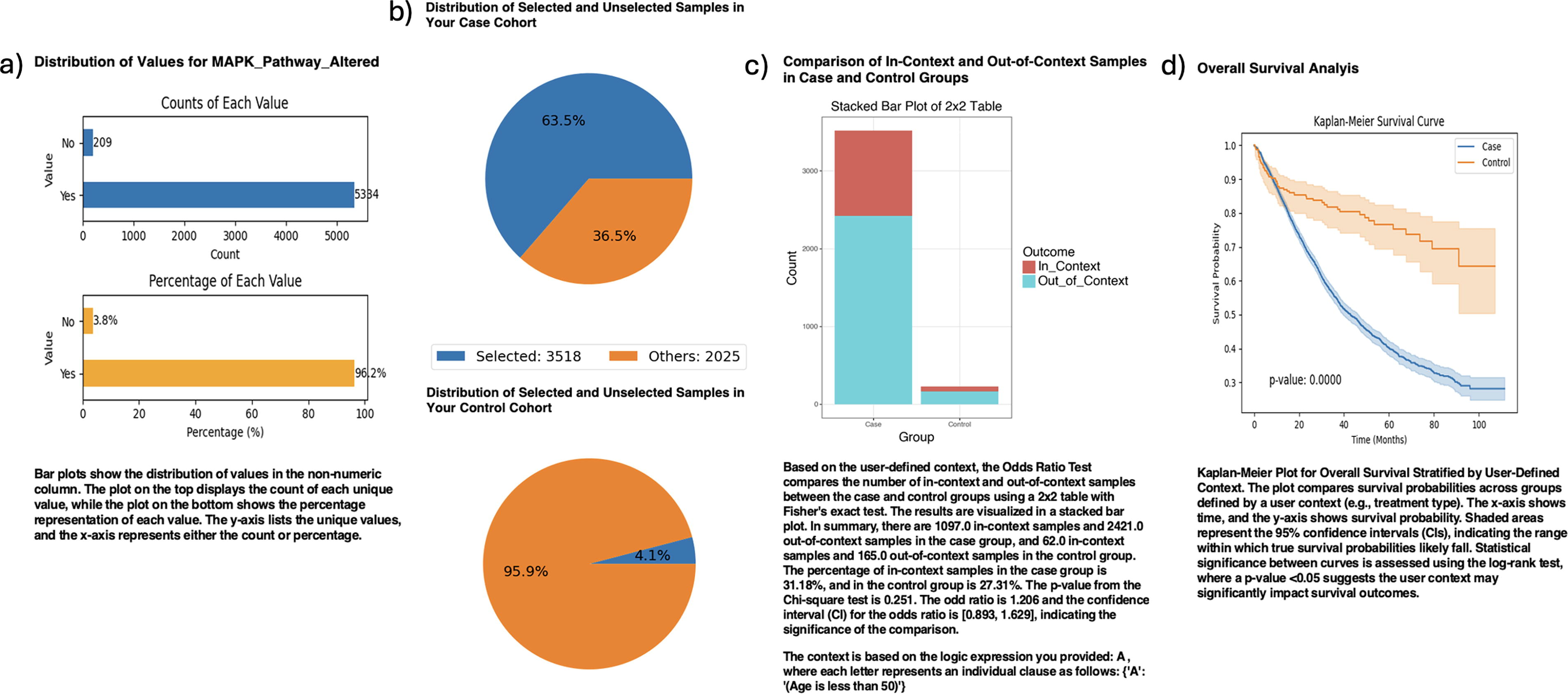
AI-HOPE-MAPK analysis of survival and enrichment patterns in MAPK-altered colorectal cancer stratified by MSI status and chemotherapy exposure. This figure showcases the application of AI-HOPE-MAPK to investigate survival outcomes and age-based enrichment in microsatellite-stable (MSS) versus microsatellite-instability-high (MSI-H) colorectal cancer (CRC) patients with MAPK pathway alterations who were treated with FOLFOX chemotherapy (fluorouracil, leucovorin, and oxaliplatin).
Anatomical and mutational site interactions
In a site-specific analysis, patients with AKT1-mutated colon adenocarcinomas showed significantly poorer overall survival compared with those with rectal tumors (p = 0.0483) (Fig. 5), suggesting the modifying influence of tumor location on mutational prognosis.
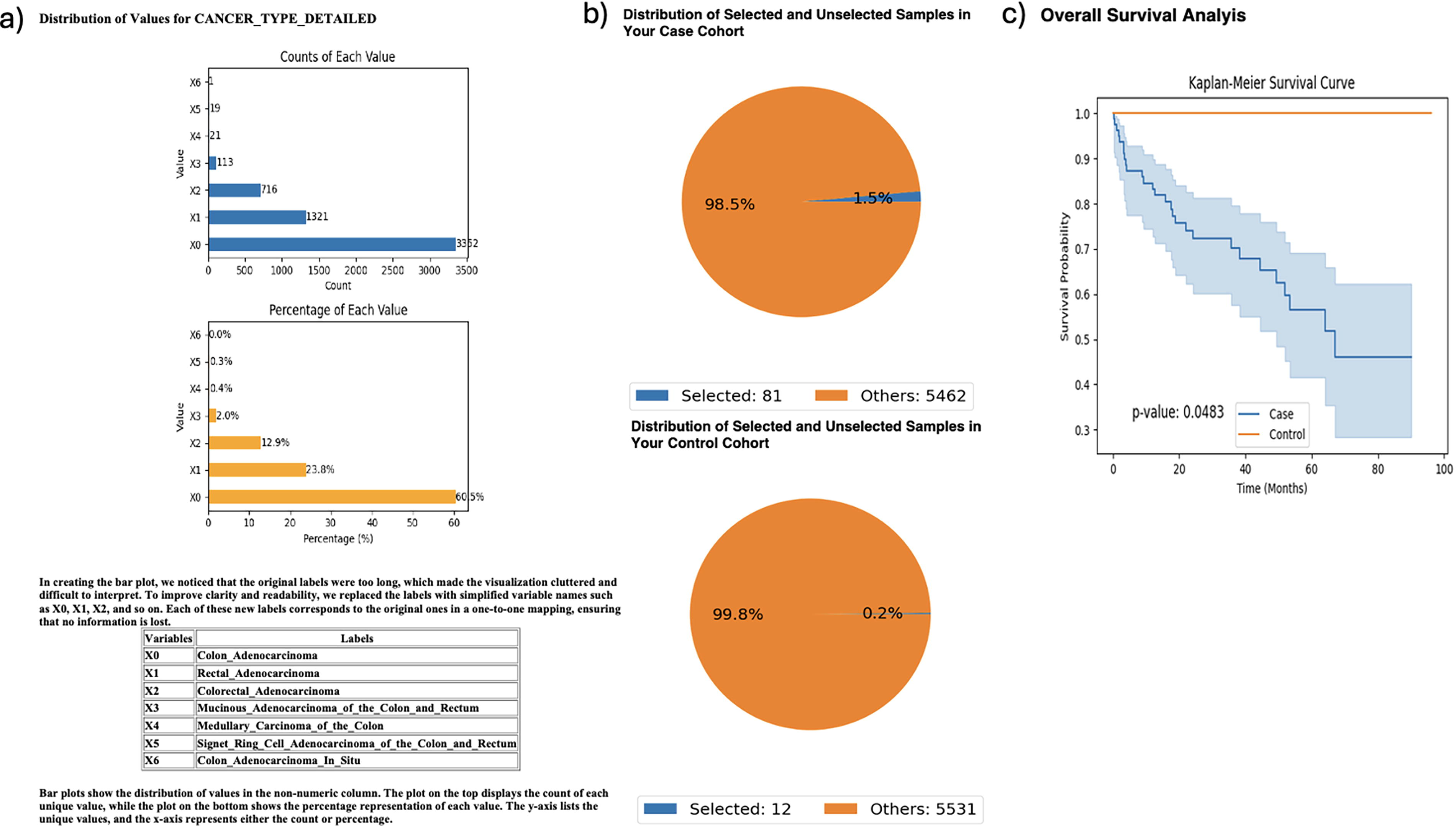
AI-HOPE-MAPK comparison of survival outcomes in AKT1-mutated colon versus rectal adenocarcinomas. This figure illustrates the use of AI-HOPE-MAPK to evaluate differences in clinical behavior among colorectal cancer (CRC) patients with AKT1 mutations, comparing those with colon adenocarcinoma (case group) to those with rectal adenocarcinoma (control group).
TMB-defined molecular subtypes in MAPK-driven CRC
AI-HOPE-MAPK also examined tumor mutational burden (TMB) in relation to MAPK status. Patients with MAPK pathway alterations and low TMB (<10 mutations/Mb) showed significantly worse survival than high-TMB, MAPK-wildtype controls (p < 0.0001) (Supplementary Fig. S8). Odds ratio analysis revealed a significantly greater proportion of younger patients (<50) in the low-TMB MAPK-altered group (31.8%) than in controls (23.29%) (OR = 1.501; 95% CI: 1.278–1.762; p < 0.001), suggesting that EOCRC may harbor a unique, MAPK-driven, low-TMB molecular phenotype.
Prognostic impact of MAP2K1 in early-stage disease
Finally, MAP2K1 mutations were evaluated in Stage I–III CRC patients. Mutant cases showed significantly poorer survival (p = 0.0447) and were less likely to receive FOLFOX (46.15% vs. 63.55%, OR = 0.491; 95% CI: 0.283–0.851; p = 0.015) (Supplementary Fig. S9). These findings suggest a prognostic and potentially treatment-modifying role for MAP2K1 in early-stage CRC.
Discussion
In this study, we describe the development and application of AI-HOPE-MAPK, a conversational artificial intelligence platform designed to examine MAPK pathway alterations in CRC through integrative, population-aware analysis. Built upon a fine-tuned LLM, AI-HOPE-MAPK combines natural language-based querying with clinical-genomic computation, allowing researchers to stratify cohorts, identify mutation patterns, and evaluate survival outcomes without requiring programming expertise. By supporting real-time, multiparameter interrogation of large-scale datasets, the platform may help reduce barriers to conducting precision oncology research, particularly those related to equity, accessibility, and representation in cancer genomics.
Applying AI-HOPE-MAPK to EOCRC, with a focus on H/L populations, yielded key insights into age- and ancestry-specific differences in MAPK signaling alterations. Notably, NF1 mutations were significantly more prevalent in H/L EOCRC cases compared with their late-onset counterparts, suggesting a potentially underappreciated role for NF1 in age-associated tumorigenesis. Although ACVR1 and MAP2K1 mutations did not reach statistical significance due to low frequency, their higher prevalence in EOCRC H/L patients indicates potential biological relevance that warrants further investigation. The consistent presence of these alterations in younger patients, in combination with the absence or lower frequency in late-onset cases, raises important questions about distinct oncogenic trajectories in EOCRC—particularly among underserved populations.
Our analysis also revealed significant ancestry-informed disparities in the mutation landscape. MAPK3 and NF1 mutations were significantly enriched in H/L EOCRC patients relative to NHW individuals, with MAPK3 displaying a fourfold increase in frequency. These results underscore the critical importance of ancestry-stratified analyses in genomic research, which have historically relied on predominantly NHW datasets. Without tools like AI-HOPE-MAPK, such population-specific mutation patterns—especially in genes less frequently studied outside canonical drivers like KRAS or BRAF—may remain undetected. These findings contribute to a growing body of evidence suggesting that certain MAPK pathway components may play divergent roles across racially and ethnically distinct CRC subtypes and may offer novel therapeutic targets tailored to these groups.
In addition to uncovering mutation patterns, AI-HOPE-MAPK facilitated deeper exploration of therapeutic response and survival outcomes. We observed significantly worse survival in H/L EOCRC patients receiving FOLFOX chemotherapy compared with their late-onset counterparts, despite standardized treatment. This survival gap raises the possibility that younger patients, particularly from minoritized populations, may harbor distinct tumor biology—potentially characterized by immune evasion, chemotherapy resistance, or MAPK-mediated signaling resilience. These observations are consistent with earlier studies showing suboptimal responses in EOCRC despite aggressive disease management and highlight the need for more nuanced, biology-informed treatment strategies.
Furthermore, our integrative analyses demonstrate the prognostic importance of combining MAPK pathway status with molecular markers such as MSI and TMB. Specifically, MAPK-altered tumors in MSS patients exhibited significantly worse survival than MSI-high (MSI-H) tumors, reinforcing MSI status as a critical contextual modifier of pathway-driven outcomes. Notably, CRC cases with low TMB and concurrent MAPK alterations were significantly enriched in younger patients and exhibited inferior prognosis, suggesting a unique molecular subtype of EOCRC that may be poorly responsive to both immunotherapy and conventional cytotoxic regimens. These findings highlight an urgent need for molecular profiling in young patients and support the development of alternative therapeutic approaches, such as MEK inhibitors or EGFR blockade tailored to specific MAPK pathway mutations.
Anatomical context also emerged as a relevant modifier of MAPK-associated prognosis. Patients with AKT1-mutated tumors in the colon experienced significantly poorer survival than those with similar mutations in rectal tumors, consistent with prior evidence suggesting that tumor site influences molecular evolution, treatment response, and disease progression. Such findings further underscore the need for integrated models of CRC biology that consider not only genomic alterations but also anatomical, demographic, and treatment-related variables.
The prognostic implications of MAP2K1 mutations in early-stage disease were noteworthy. Although uncommon, these mutations were associated with worse survival outcomes and a significantly lower likelihood of receiving FOLFOX chemotherapy. This pattern may indicate underlying biological aggressiveness and/or differences in treatment utilization within this subgroup. The contributing factors—whether clinical decision-making, comorbidities, physician perception of benefit, or unmeasured social determinants—remain uncertain. Importantly, AI-HOPE-MAPK facilitated the identification of these disparities, underscoring its potential utility for generating hypotheses and informing future clinical investigation.
Although MAP2K1 and ACVR1 mutations were detected at low frequencies, their distribution patterns provide important exploratory insights. In particular, MAP2K1 alterations were nearly exclusive to H/L EOCRC patients, yielding an infinite odds ratio due to their absence in the LOCRC comparator group, while ACVR1 mutations also appeared more often in EOCRC than LOCRC cases. Although these associations did not achieve statistical significance—reflecting the limited statistical power inherent to rare events—the magnitude of effect and consistency of enrichment in younger patients suggest potential biological relevance. Importantly, the prognostic impact of MAP2K1 mutations in stage I–III CRC was significant, with worse survival and reduced FOLFOX exposure, reinforcing that even low-frequency events may carry clinically actionable consequences. These findings highlight both the promise and limitations of AI-driven rare variant detection: while such analyses can uncover underappreciated signals, careful interpretation and future validation in larger, multi-institutional cohorts will be essential to confirm their role in EOCRC biology.
This study leverages retrospectively assembled, harmonized datasets and is therefore subject to selection bias, site- and platform-specific differences, variable sequencing coverage, and incomplete or heterogeneous clinical annotations (e.g., treatment exposure and timing). Although we applied uniform harmonization, residual confounding cannot be excluded, and causal inference is not warranted. Analyses of rare alterations (e.g., MAP2K1, ACVR1) are underpowered, yielding wide confidence intervals and occasional zero-cell estimates. Findings may also depend on modeling choices within AI-HOPE-MAPK—including the fine-tuned LLM, prompt schemas, mapping rules, and analytic parameters—so reproducibility across alternative pipelines merits evaluation. Finally, the platform has not been deployed in clinical workflows, and our results have not undergone prospective validation. Accordingly, all signals should be viewed as hypothesis-generating and require confirmation in larger, ancestry-informed cohorts and prospective or interventional studies.
Collectively, these findings demonstrate that AI-HOPE-MAPK not only recapitulates known CRC biology—such as BRAF enrichment in LOCRC—but also expands the landscape of actionable discoveries by uncovering under-recognized alterations in early-onset and H/L patient groups. The platform’s conversational interface, robust analytical engine, and harmonized integration of clinical-genomic data represent a potential scalable solution for precision oncology research and equitable biomarker discovery.
Future directions for AI-HOPE-MAPK include expansion to additional molecular pathways (e.g., TP53, TGF-β, WNT), integration of multi-omics 33 and spatial transcriptomics/proteomics, 34 and deployment in prospective clinical trial design. Critically, our work supports the use of AI-powered tools to accelerate the identification of population-specific vulnerabilities, enhance translational research in diverse communities, and ultimately inform therapeutic decision-making for CRC patients who have historically been excluded from precision medicine pipelines. As the field moves toward increasingly data-driven, inclusive models of cancer care, platforms like AI-HOPE-MAPK will be essential in translating complexity into clinical action.
Conclusion
AI-HOPE-MAPK represents a promising advancement in the application of artificial intelligence to precision oncology, enabling dynamic, population-aware exploration of MAPK pathway alterations in CRC. Through natural language-driven workflows, the platform integrates genomic and clinical data to reveal age- and ancestry-specific patterns that may be underappreciated in traditional analyses. In this study, AI-HOPE-MAPK identified differential mutation burdens, prognostic associations of low TMB MAPK-altered subtypes, and treatment-related differences in H/L EOCRC patients. These findings recapitulate known biology while also pointing to novel disparities that warrant further investigation. While prospective validation will be critical to confirm clinical applicability, AI-HOPE-MAPK provides a scalable framework to support hypothesis generation, guide future studies, and refine our molecular understanding of CRC across diverse patient populations.
Footnotes
Acknowledgments
The authors would like to express their gratitude to the Department of Integrative Translational Sciences at City of Hope, the Cancer Control and Population Sciences Program (NIH P30 CA033572) at the City of Hope Comprehensive Cancer Center, and the Drug Development and Capacity Building: A UCR/CoH-CCC Partnership project (NIH U54 CA285116).
Authors’ Contributions
The contributions of the authors are as follows: Conceptualization, E.-V.V., E.-W.Y., and B.W.; methodology, E.-V.V. and B.W.; software, E.-V.V. and E.-W.Y.; validation, B.W., E.-W.Y., and E.V.; formal analysis, E.-V.V. and B.W.; investigation, E.-V.V. and B.W.; resources, E.-V.V.; data curation, B.W. and E.-V.V.; writing—original draft preparation, E.-V.V. and E.-W.Y.; writing—review and editing, E.-V.V., B.W., and E.-W.Y.; visualization, E.-V.V. and B.W.; supervision, E.-V.V.; project administration, E.-V.V.; funding acquisition, E.-V.V. All authors have read and agreed to the published version of the article.
Data Availability Statement
Author Disclosure Statement
The authors declare no conflicts of interest
Funding Information
This research was supported by the Department of Integrative Translational Sciences at City of Hope, the City of Hope Cancer Control and Population Sciences Program, National Institutes of Health (NIH), National Cancer Institute (NCI), award number NIH/NCI P30-CA033572; NIH, NCI, Cancer Moonshot project, PE-CGS: Optimizing Engagement of Hispanic Colorectal Cancer Patients in Cancer Genomic Characterization Studies, award number NIH/NCI U2C-CA252971; and NIH, NCI, U54 University of California Riverside (UCR) and City of Hope Comprehensive Cancer Center (COH-CCC) partnership, Drug Development and Capacity Building, award number NIH/NCI U54-CA285116.
