Abstract
Machine learning (ML) methods have demonstrated strong predictive capabilities when trained on large datasets. However, in domains where data is scarce or sensitive, ML models often exhibit sub-optimal performance. Our hypothesis is that semantically enriching the available training dataset can enhance the predictive power of ML models, particularly in data-scarce scenarios. To investigate this hypothesis, we propose novel neuro-symbolic approaches that augment tabular data with knowledge graph (KG) information, providing additional context and structure to improve model performance. Concretely, we introduce and examine several integration techniques of KG information through embeddings and explore how different KG embedding algorithms affect model performance, with a specific focus on accuracy and F2 scores. Our evaluation involves four distinct ML algorithms and four KG embedding techniques. We apply our approach to binary classification tasks on tabular data, including heart disease and chronic kidney disease. Our experimental results show improvements in performance particularly when tabular data is augmented with distance features computed in the embedding space. Notably, we achieve gains in F2 scores, such as an increase in XGBoost performance from 75.19% to 90.85% for heart disease prediction. These findings demonstrate the potential of KG-based augmentation to enhance ML performance.
Introduction
Machine learning (ML) has revolutionised various domains by providing powerful tools for pattern recognition, predictive analytics and data-driven decision-making. Techniques such as deep learning have achieved remarkable success in fields ranging from computer vision (Deng et al., 2009; Wortsman et al., 2022) to natural language processing (Kenton & Toutanova, 2019; Ramesh et al., 2022). These advancements have been largely driven by the availability of large datasets and the computational power to process them.
However, ML methods often face significant challenges related to data quality and availability. Data sparsity, imbalance and sensitivity can severely hinder the performance of ML models (Poulinakis et al., 2023). In the medical domain, one important task is predicting patient outcomes, for instance, determining the presence or absence of a disease based on clinical observations. This task often suffers from an insufficient amount of labelled data due to privacy concerns (Jarrett et al., 2019). Although advances have been made, models trained solely on tabular data fail to fully capture the domain’s complexity and semantics, limiting their ability to generalise effectively (Ruiz et al., 2024).
To overcome these limitations, neuro-symbolic (NeSy) artificial intelligence (AI) has emerged as a promising approach to integrate domain knowledge into ML models. NeSy AI combines the strengths of symbolic AI – known for logical reasoning and explainability – with sub-symbolic methods such as deep learning (Garcez & Lamb, 2023; Hitzler et al., 2022; Sarker et al., 2021). In particular, structured semantic knowledge such as knowledge graphs (KGs) has emerged as a key element in bridging the gap, providing a structured way to represent relationships between entities and capture domain-specific semantics (Bhatt et al., 2020; Gaur et al., 2018; Herron et al., 2023; Yin et al., 2019). KGs have been widely used in tasks such as KG completion (Lin et al., 2015) and link prediction (Wang et al., 2021). However, their potential to enhance ML predictions on tabular data by incorporating semantic knowledge through embeddings remains underexplored.
We propose integrating KGs into ML pipelines to enhance tabular data with structured, domain-specific information. Drawing upon techniques from the Semantic Web community, our approach begins by utilising ontologies to formalise domain semantics. We then construct KGs based on these ontologies, enriching the datasets with structured knowledge specific to the medical domain. Subsequently, we employ KG embeddings to transform the KGs into numerical vector representations suitable for ML algorithms. By embedding relationships and domain knowledge from KGs into these vectors, our methodology enhances the ML pipeline by augmenting the datasets with semantic knowledge, aiming to improve predictive performance – especially in data-scarce domains. This study specifically explores binary classification tasks in both medical predictions (heart disease and chronic kidney disease) where domain-specific structure is crucial for robust prediction. Our research is guided by the following research questions:
RQ1: How can KGs be optimally infused into an ML pipeline to enhance performance in terms of accuracy and F2 score? RQ2: How does the choice of knowledge graph embedding algorithms affect the performance of machine learning models when used to augment tabular data? RQ3: How do different ML algorithms perform when KG-based information is integrated into the input data?
To address these research questions, we took an exploratory approach, systematically investigating each aspect step by step. For RQ1, we derived five sub-hypotheses to examine how KGs can be optimally integrated into ML pipelines to enhance performance metrics such as accuracy and F2 score (with the reason for selecting these metrics explained in Section 6.2). We tested these hypotheses using eight different approaches, each incorporating KGs and embeddings in various ways. For RQ2 and RQ3, we empirically evaluated the impact of different KG embedding algorithms and ML models across two medical domains – heart disease and chronic kidney disease prediction.
Building on our previous work (Llugiqi et al., 2024), we extend and formalise our methodology for integrating KG embeddings into ML pipelines. We employ two additional embedding techniques alongside those used previously to transform the KGs into numerical vector representations suitable for ML algorithms. We developed and tested different approaches based on five sub-hypotheses derived from our first research question, providing a comprehensive evaluation of their impact on model performance in heart and kidney disease prediction. Our study demonstrates the effectiveness of incorporating ontological knowledge into the ML training process, highlighting the potential for improved predictive performance in data-scarce domains and its applicability across various fields where ontologies can be developed or expanded.
The remainder of this article is organised as follows: In Section 2, we define the key concepts that we use in our work. This is followed by an overview of related work in Section 3. In Section 4, we present an overview of our proposed approach, with more detailed explanations about our approaches provided in Section 5. Our experimental analysis is discussed in Section 6, where we outline the goals and setup of our experiments. In Section 7, we present and analyse the outcomes of our experiments. Finally, we summarise our findings and outline directions for future work in Section 8.
In this section, we outline the problem we aim to address, followed by introducing the key concepts that we use throughout the article, beginning with ontologies, KGs and knowledge graph embeddings.
Problem Description
In this study, we address the challenge of predicting heart disease and chronic kidney disease using patient medical records in tabular data format. Each dataset can be represented as a table
We focus on binary classification to predict the presence or absence of these diseases. Formally, given the dataset

Baseline for ML prediction on tabular data.
For example, given a patient’s data (e.g., age 62, female, asymptomatic, resting blood pressure 140, cholesterol 268, no fasting blood sugar, max heart rate 160, downsloping slope and thalassemia), our model aims to determine the likelihood of heart disease. Similarly, a record for kidney disease might involve attributes such as age 68, blood pressure 70, specific gravity 1.01 and blood urea 54. The objective is to accurately predict disease presence.
Due to the sensitive nature of medical data, datasets in this domain are often limited or partially incomplete, impacting model performance. This scarcity of data, combined with varying data quality, presents a challenge to achieving optimal prediction accuracy, necessitating robust preprocessing and, potentially, data augmentation strategies to improve model generalisability and reliability.
Given the problem definition described in the previous subsection, our approach aims to augment these datasets by integrating semantic information to enhance predictive capabilities. To achieve this, we leverage
Ontology:
Originally a philosophical term, ontology refers to the study of existence and the nature of being. In computer science, Gruber Gruber (1993) redefined ontology as ‘explicit specifications of conceptualisations’, where a conceptualisation represents a simplified, abstract view of a domain to capture essential aspects. An ontology establishes a standardised vocabulary for knowledge sharing within a specific domain. Formally, an ontology represented as
Knowledge Graphs:
KGs expand on ontologies by capturing not only the structured relationships between concepts but also the specific instances and values within a domain. Originally popularised by Google in 2012 (Singhal, 2012) to enhance search understanding, KGs have since become integral in a range of applications, providing a structured, machine-readable format to represent knowledge. We define the KG as
KG embeddings:
While KGs provide a structured representation of entities and their relationships, they can become highly complex as the number of entities and relations grows. To enable efficient computation, learning and reasoning over KGs, knowledge graph embeddings (KGEs) are commonly used (Bordes et al., 2013; Lin et al., 2015; Wang et al., 2014). KGEs transform entities and relations from a discrete symbolic space into a continuous vector space, capturing the structure and semantics of the KG in a form that is compatible with ML algorithms. KGE algorithms can be broadly categorised into three main types based on their methodology and objectives: translational distance models, semantic matching models and random walk-based models. In the following, we briefly describe the embedding algorithms used in our experiments: Node2Vec (Grover & Leskovec, 2016) and Rdf2Vec (Ristoski & Paulheim, 2016) as random-walk-based models that leverage the graph structure, DistMult (Yang et al., 2014) as a semantic matching model and TransH (Wang et al., 2014) as a translational model.
Node2Vec uses a flexible random walk strategy to combine depth-first and breadth-first sampling, allowing it to capture various structural features of the graph whether they are labelled or unlabelled, directed or undirected. Node2Vec employs random walks, incorporating an adjustable bias parameter that allows for targeted exploration of local neighbourhoods as well as a broader global search. RDF2Vec is designed specifically for RDF (Resource Description Framework) graphs within the Semantic Web, RDF2Vec generates embeddings for entities and relations by leveraging random walks to create sequences from the graph. These sequences are then transformed into embeddings using Word2Vec, making RDF2Vec particularly effective at capturing the semantic and relational attributes present in RDF data. While both RDF2Vec and Node2Vec utilise random walks, RDF2Vec focuses more on semantic relationships within the context of the Semantic Web, whereas Node2Vec emphasises structural characteristics applicable to a wider range of graph types. DistMult is a semantic matching model that uses a bilinear scoring function to evaluate the interactions between entities and relations in a KG. In this model, each relation is represented as a diagonal matrix, simplifying the bilinear form to a weighted element-wise multiplication of entity embeddings. While this approach effectively captures pairwise relationships, it inherently assumes that all relations are symmetric, which may restrict its expressiveness for datasets containing asymmetric relations. TransH is a translational model that represents entities as vectors and relations as hyperplanes in the embedding space. Each relation is associated with a specific hyperplane and a translation vector on that hyperplane. Entities are projected onto the hyperplane of a relation before the translation operation is applied. This method allows entities to have different representations in the context of different relations, enabling the model to capture complex and diverse relationships thereby improving its ability to represent multiple types of relationships in a KG.
We review related work on (i) the categorisation of neuro-symbolic approaches, positioning our work within these categories, (ii) we discuss the use of ML models in disease prediction and (iii) enhancing ML predictions with semantic knowledge, and we conclude by discussing the novelty of our approach.
Categorisation of Neuro-Symbolic Approaches
In recent years, the field of neuro-symbolic AI has gained significant attention due to its potential to combine the strengths of both symbolic and sub-symbolic AI (Garcez & Lamb, 2023; Hitzler et al., 2022; Sarker et al., 2021). Symbolic AI excels at logical reasoning and explainability, while sub-symbolic approaches, such as deep learning, have proven effective in pattern recognition and data-driven decision-making. Combining these approaches, neuro-symbolic AI seeks to leverage the best of both worlds: the learning capability of sub-symbolic methods and the structured, interpretable reasoning of symbolic methods. Several efforts have focussed on categorising neuro-symbolic approaches. Kautz et al. Kautz (2022) classify neuro-symbolic systems into six types based on the interaction between neural networks and symbolic reasoning.
Similarly, Sheth et al. Kursuncu et al. (2019); Sheth et al. (2019) identify three levels of knowledge infusion in neural models: shallow, semi-deep and deep.
Dash et al. Dash et al. (2022) categorise methods for integrating domain-specific knowledge into deep neural networks into three main approaches: enhancing input data, modifying the loss function and adjusting the network architecture. Our research aligns with the
Van Harmelen and ten Teije Van Harmelen and Ten Teije (2019) introduced a conceptual framework known as ‘boxology’, which outlines various patterns for integrating machine learning with semantic web technologies. Breit et al. Breit et al. (2023) expanded this framework by identifying 44 distinct patterns used in hybrid learning and reasoning techniques, based on a review of around 500 papers from 2010 to 2020. Our approach falls under the
As a summary, our approach falls under the
Machine Learning Models in Disease Prediction
The application of ML in healthcare has attracted significant research interest due to its potential. Kraivsnikovic et al. Kraišniković et al. (2025) proposed an approach leveraging fine-tuned BERT models to analyse German pathology reports. Their work highlights how domain-specific adaptations can enhance the interpretability and utility of ML models in medical diagnostics by effectively capturing contextual representations. Additionally, ML algorithms have been successfully employed in predicting diseases such as heart disease (Katarya & Meena, 2021; Rani et al., 2021; Shah et al., 2020; Yadav et al., 2023) and kidney disease (Chittora et al., 2021; Rady & Anwar, 2019; Vijayarani et al., 2015; Yildirim, 2017), using various techniques such as data preprocessing, feature selection and hyperparameter tuning to enhance prediction accuracy.
Several studies have also explored combining ML methods to further improve performance. For instance, Mohan et al. Mohan et al. (2019) combined random forest and linear methods to enhance heart disease prediction, while Ali et al. Ali et al. (2019) introduced a framework for heart failure prediction using dual support vector machine (SVM) models – one for feature selection and the other for the prediction task.
Although these models demonstrate good predictive performance, they often rely on extensive preprocessing (Hassler et al., 2019), feature selection and hyperparameter tuning to achieve optimal results. Moreover, the effectiveness of ML models can be limited by insufficient or sub-optimal quality data. In this context, healthcare ontologies (Chute & Çelik, 2021; El-Sappagh et al., 2018; Ivanović & Budimac, 2014; Jovic et al., 2007; Pisanelli, 2004) offer a structured, semantically rich layer of information that can enhance the contextual understanding of ML models, which is further explored in this work.
Enhancing ML Predictions with Semantic Knowledge
Recent research has increasingly focussed on integrating semantic knowledge, such as KGs and ontologies, into ML models to enhance their performance. KGs have been widely applied in various domains, notably improving feature extraction and entity representation in natural language processing tasks. For instance, Moussallem et al. Moussallem et al. (2019) demonstrated how augmenting neural machine translation systems with KGs improved the translation quality by enhancing the semantic understanding of terminological expressions. Similarly, KG-based input enhancement has been shown to improve recommendation systems and community detection, enhancing both accuracy and explainability (Bhatt et al., 2020).
Moreover, KG-augmented neural networks have demonstrated improved performance in text classification and natural language inference tasks. Annervaz et al. Annervaz et al. (2018) showed that integrating structured knowledge from KGs not only improved model accuracy but also allowed models to perform well with less labelled data, addressing the common issue of data sparsity. Ziegler et al. Ziegler et al. (2017) adopted a similar approach by incorporating semantic knowledge through graph embeddings for credit card fraud detection, demonstrating how the injection of background knowledge – such as public holidays from DBpedia into neural models could enhance classification outcomes.
In addition to NLP and fraud detection, Szilagyi et al. Szilagyi and Wira (2018) applied semantic knowledge in smart building management by integrating taxonomies, schemas and logic rules with ML models. This hybrid system optimised building management by combining data-driven insights with rule-based reasoning, showing the potential of semantic knowledge in enhancing decision-making processes. Huang et al. Huang et al. (2023) introduced an Abductive Learning with KG approach that automatically mines logic rules from KGs and integrates them into ML models using a knowledge-forgetting mechanism to filter irrelevant information, thereby improving model performance even with limited labelled data.
In healthcare, Gazzotti et al. Gazzotti et al. (2019) demonstrated how augmenting sparse electronic medical records (EMRs) with ontological resources improved the predictive capabilities of ML algorithms, specifically in hospitalisation prediction. Ontologies such as DBPedia, Wikidata and the more domain specific ones provide structured medical knowledge, enabling a richer representation of patient data. Similarly, Ruiz et al. Ruiz et al. (2024) introduced the PLATO method, which uses a KG to regularise a multilayer perceptron for tabular datasets, showing that semantic knowledge can help ML models handle high-dimensional and low-sample-size data more effectively.
These studies highlight the growing importance of integrating semantic knowledge into ML models to address challenges such as data quality, sparsity and explainability across different domains. Table 1 provides an overview of these studies, outlining the types of semantic knowledge used, the domains or tasks covered, the ML models applied, the incorporation of KGEs and the integration methods employed. These approaches generally fall into two main categories: (i) direct integration of structured knowledge through explicit rules or ontological features and (ii) representation learning via KGEs, where entities and relations are embedded into a continuous vector space, allowing downstream ML models to leverage the semantic structure.
Summary of Related Work on Integrating Semantic Knowledge Into ML Models.

Anatomical Therapeutic Chemical Classification, National Drug File - Reference Terminology, International Primary Care Classification
Summary of Related Work on Integrating Semantic Knowledge Into ML Models.

Anatomical Therapeutic Chemical Classification, National Drug File - Reference Terminology, International Primary Care Classification
Although this prior research has demonstrated that embedding-based methods can improve ML performance, most existing work relies on large, general-purpose KGs, such as Wikidata or DBpedia. In contrast, our recent study Llugiqi et al. (2024) introduced four approaches for augmenting tabular data with KGEs using two embedding algorithms, focussing on smaller, domain-specific ontologies. These initial methods illustrated the potential of embedding-driven enrichment exploring how semantic context could be systematically incorporated into tabular datasets.
Building on that foundation, this article proposes four additional approaches that calculate various metrics in the embedding space to enrich tabular data with additional semantic context. Furthermore, we employ two more KG embedding algorithms, extending the methodology and formalising our approaches. We also perform a more thorough evaluation of the proposed techniques, applying them to the prediction of heart disease and chronic kidney disease.
Compared to other studies shown in Table 1, our method goes beyond simply embedding entities and relations, we exploit the embedding space itself to derive meaningful metrics that further enrich tabular features. Moreover, instead of relying on extensive, generic KGs, our approach leverages small, existing domain-specific ontologies (or select subsets of existing big ontologies), which we populate with relevant tabular data to form task-specific KGs. Such domain-focussed strategies remain underexplored within medical prediction and systems. By emphasising smaller ontologies and extracting deeper semantic insights from the embedding space, our work aims to advance semantic knowledge integration for ML in medical and other specialised domains.
To improve the performance of ML models, we leverage KGs to enrich tabular datasets with semantic information. This section outlines the two core steps of our approach. First, in Section 4.1, we explain the construction of KGs using instances from the tabular data, as shown in the top part of Figure 2. This step focuses on building KGs that capture deeper relationships within the data. Next, in Section 4.2, we focus on integrating KG embeddings into the ML pipeline. This includes various augmentation strategies designed to enhance model performance by incorporating structural and relational information from the KG into the training data, as depicted in the bottom part of Figure 2.
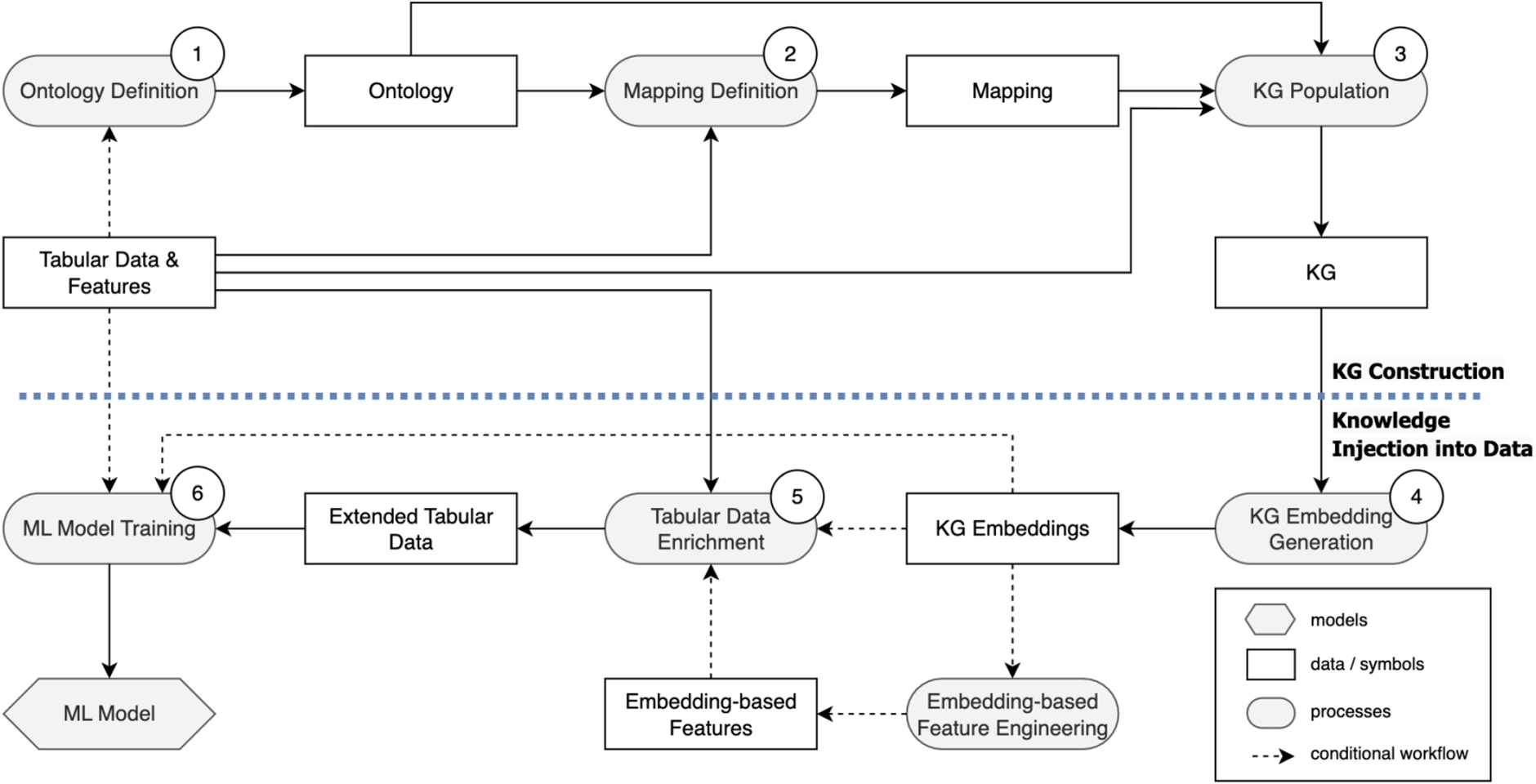
Overview of the proposed approach including (i) KG construction (top) and (ii) knowledge injection into data (bottom) (adapted from Llugiqi et al. (2024) following the boxology notation (van Bekkum et al., 2021)).
For our approach to enrich ML input data with supplementary knowledge, constructing KGs is essential. They serve as structured representations of domain knowledge, capturing the semantics of the data and allowing for the integration of ontological information into datasets. This enrichment allows ML models to leverage contextual and relational information, enhancing their predictive capabilities. The upper part of Figure 2 illustrates the methodology used for building these KGs, which represent data that was initially captured in tabular form. The following steps provide a formal description of this construction process.
Step 1: Ontology Definition
The first step in constructing the KG is defining an ontology, which is used to capture domain semantics and provide a structured framework for enriching the datasets. There are different ways to develop an ontology, represented as
Step 2: Mapping Definition
The process of mapping dataset features to the concepts in the ontology is crucial for using instances from tabular data to populate the ontology and, consequently, construct a KG. A key aspect of this mapping process is the
Step 3: Knowledge Graph Population
The knowledge graph is constructed by utilising the ontology
This preprocessing phase ensures that features from the tabular data
In Section 4.1, we outlined the construction of enriched data structures that capture deeper semantics beyond the raw data. This section will now focus on transforming these enriched structures into a vectorised format suitable for ML, and on the optimal strategies for augmenting the input data, as illustrated in the lower part of Figure 2.
Step 4: Knowledge Graph Embedding Generation
With the populated KG with enriched data structures, the subsequent step is to prepare the KG for ML model training. This requires transforming the KG into a vector space representation suitable for ML models, using KGE algorithms. Having a knowledge graph
Step 5 & 6: Tabular Data Enrichment and ML Model Training
After computing KGEs, our objective is to explore the integration of these embeddings to enhance the performance of ML models. We experimented with different approaches for augmenting the training set using KGEs. First, we established a baseline that trains ML models using only tabular data
Proposed Approaches for Tabular Data Enrichment and ML Model Training
In this section, we outline the eight distinct approaches we explored for integrating KG embeddings into the training dataset, each designed to evaluate the impact of enriched semantic information on model performance. 1
Embeddings As ML Model Inputs (EmbedOnly)
We begin by our initial objective to investigate whether training a model on the vector representations generated from these KGs, using various embedding algorithms, could reveal underlying patterns and relationships within the data. Therefore, we define our first sub-hypothesis as follows.
To explore this, we first explored the EmbedOnly approach, focussing solely on the embeddings to assess their standalone effectiveness in capturing meaningful insights as shown in Figure 3.
Embedding vectors, highlighted in yellow, serve as inputs to the ML model.

For each instance
Building upon EmbedOnly approach, we define our second sub-hypothesis as follows.
This led us to design approaches that integrate both embeddings and tabular features, aiming to see if the KG information could complement and enrich the existing dataset. Thus, we investigated EmbedAugTab and other subsequent approaches that leverage embeddings for data augmentation based on this intuition.
EmbedAugTab approach involves training ML algorithms on datasets that integrate the original tabular data with additional columns derived from embeddings, as illustrated in Figure 4 and presented in Algorithm 2. For each instance Tabular dataset enrichment with embedding vectors, highlighted in yellow, used as inputs for the ML model.

Utilising embedding vectors directly to augment the tabular data may introduce noise. Thus, our sub-hypothesis is defined as follows.
This led us to introduce the DistAugTab and ClustAugTab approaches, which aim to selectively extract meaningful information from the embeddings to improve the learning process.
In DistAugTab approach, we enhance the tabular dataset
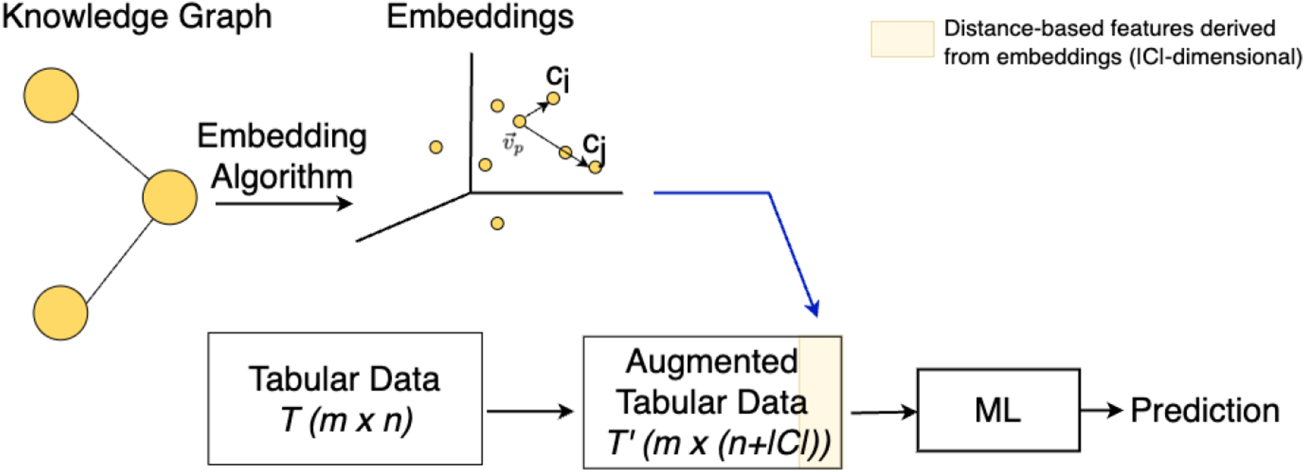
Tabular dataset enrichment with distance measures from the KG (highlighted in yellow), used as inputs for the ML model.

The new columns are calculated by determining the Euclidean distance between the embedding vector
By including these distance features, we aim to capture how closely each instance’s embedding aligns with the class centroids, thereby potentially improving the model’s ability to differentiate between target classes. For example, in the healthcare domain, the target classes could represent the presence or absence of a disease
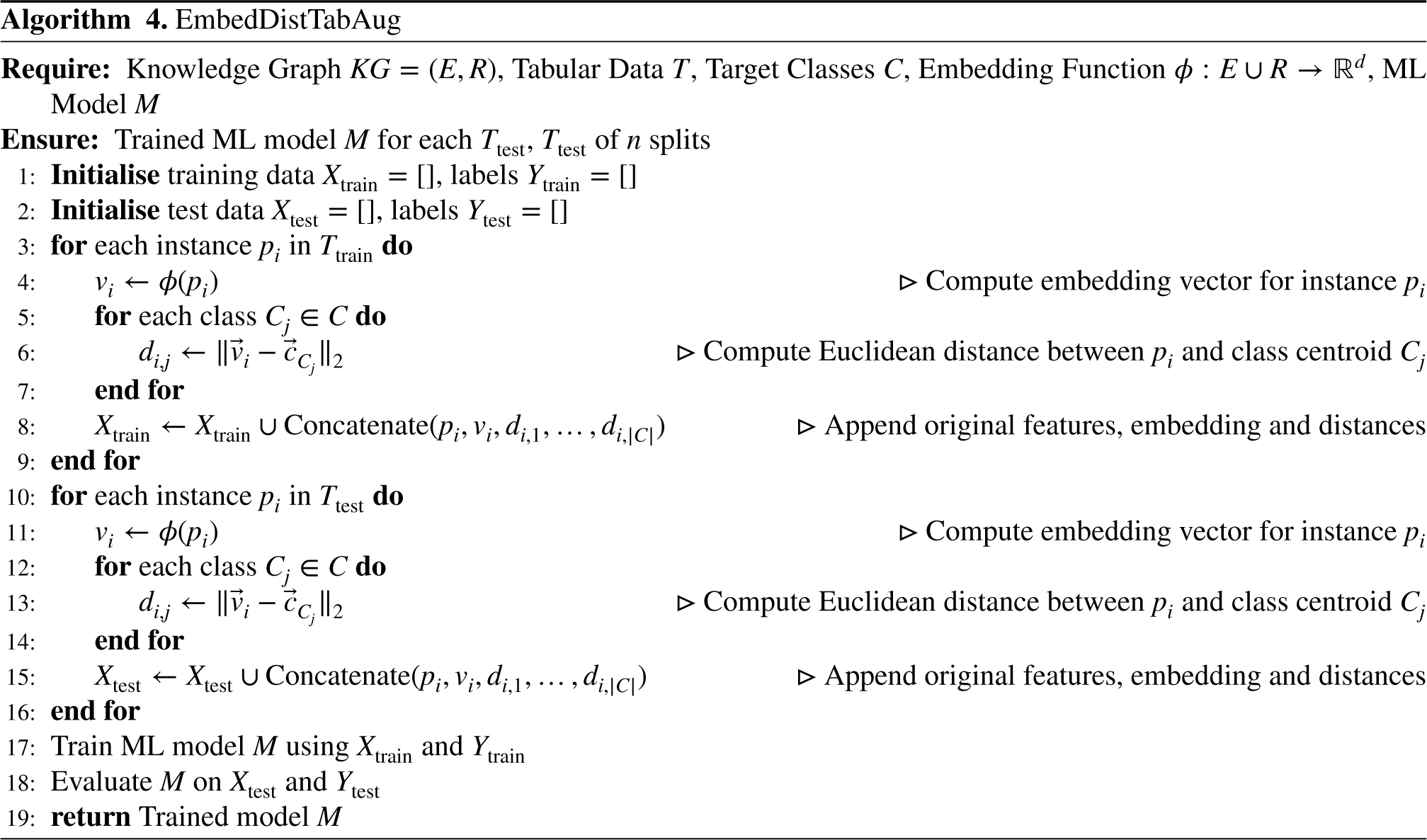
This approach augments the tabular dataset by incorporating both embedding vectors and distance-based features, as depicted in Figure 6 and presented in Algorithm 4. For each instance Tabular dataset enrichment with distance measures from the KG and vector embeddings, highlighted in yellow, used as inputs for the ML model.

In this approach, referred to as ClusterAugTab, we augment the tabular dataset by first computing embeddings for the data Tabular dataset enrichment with embedding clusters’ membership (highlighted in yellow), used as ML model inputs.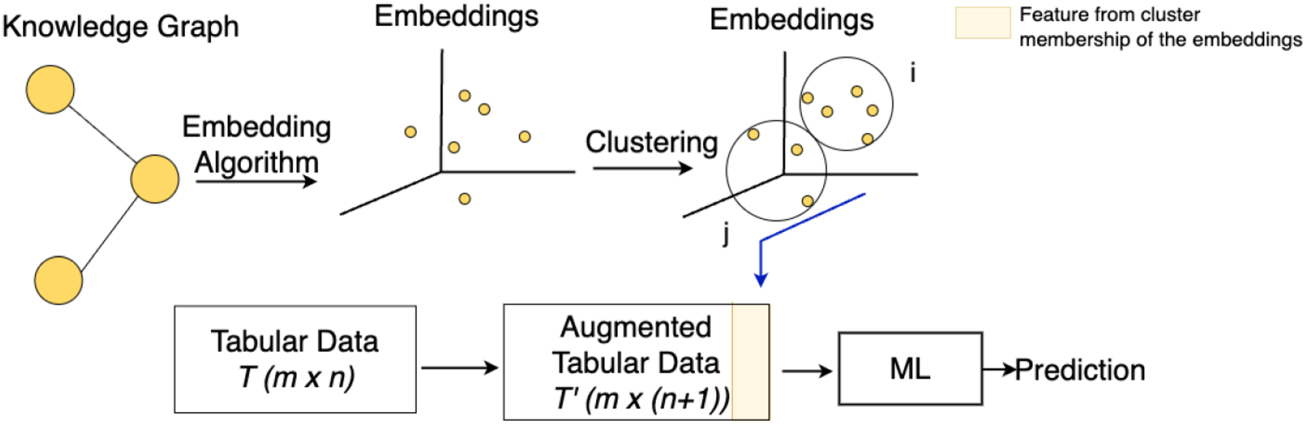

This approach, the tabular dataset is augmented by integrating both embedding vectors and cluster memberships, as shown in Figure 8 and detailed in Algorithm 6. For each instance Tabular dataset enrichment with embedding clusters’ membership and vector embeddings, highlighted in yellow, used as ML model inputs.

To further optimise the integration of KG information, we hypothesised that interactions between embeddings and existing features could reveal complex patterns. We define the sub-hypothesis as follows.
By developing approaches that compute these interaction terms, we aimed to enrich the feature space, enabling the model to capture dependencies arising from the integration of KG-derived and tabular features. This approach, implemented in the InteraAugTab approach, offers a multi-dimensional perspective that aims to improve accuracy and F2 score.
InteraAugTab approach augments the tabular dataset by incorporating interaction terms derived from the original features, as illustrated in Figure 9 and presented in Algorithm 7. For each instance Tabular dataset enrichment with feature interaction (highlighted in yellow), used as ML model inputs.

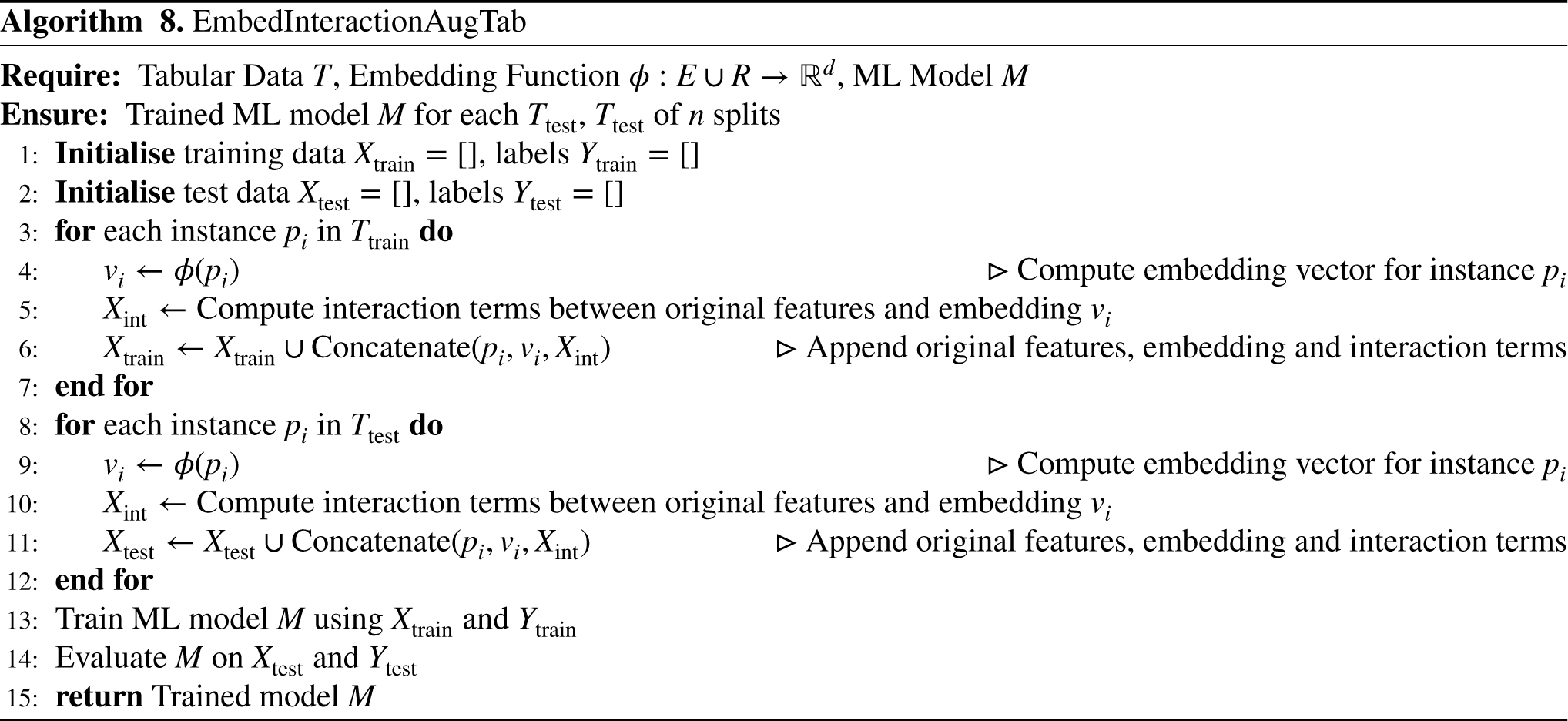
In this approach, referred to as EmbedInteractionAugTab, we augment the tabular dataset by incorporating both the embedding vectors and the interaction terms between the original features and the embedding vectors, as shown in Figure 10 and presented in Algorithm 8. Similar to InteraAugTab approach, the embeddings are computed and the interaction terms. The augmented dataset Tabular dataset enrichment with feature interaction and vector embeddings, highlighted in yellow, used as ML model inputs.

To address the risk of high dimensionality, which can adversely affect the performance of certain models, we implemented a dimensionality reduction step using the PCA algorithm (Abdi & Williams, 2010). This reduction was specifically applied to approaches integrating embeddings, namely EmbedOnlyRed, EmbedAugTabRed and EmbedDistAugTabRed. We define our sub-hypothesis as follows:
In this section, we discuss the experimental goals that guide our investigation in Section 6.1 and in Section 6.2 we discuss the experimental setup and materials used to achieve these goals.
Experimental Goals
The goal of our experimental evaluation is to investigate the use of KGs through KGEs to enhance the predictive performance of ML methods. We leverage the semantic structure of the ontologies, to represent the instances with more semantics and then through our proposed approaches use these to augment the tabular dataset for a better ML performance. The specific goals of our experiments are as follows:
Experiment Setup
Heart disease dataset consists of 303 instances, with 14 features capturing various patient health indicators relevant to diagnosing heart disease such as heart rate and cholesterol. Chronic kidney disease contains 400 instances and 25 features, capturing various health metrics related to chronic kidney disease such as blood pressure and albumin levels.
Both datasets contain a mix of categorical and numerical attributes, making them suitable for testing the integration of KGE with tabular data. Additional details about the datasets’ features can be found in Appendix 8.
The The The
For chronic kidney disease, we only used the third approach, extracting a sub-ontology from SNOMED-CT, due to the lack of ontologies specific to this domain. An overview of the ontologies used for both domains, including the count of classes and properties, is presented in Table 2.
Details of the Ontologies for Heart and Kidney Disease Domain.

In Table 3, we illustrate the parameters used for the embedding methods, tailored to the specific characteristics of the KGs. The embedding dimensions ([64, 128, 100]) were selected to provide a range of vector sizes that are large enough to capture meaningful patterns but small enough to maintain computational efficiency. For Node2Vec and RDF2Vec, the walk length and the number of walks per node were adapted to the size and complexity of each ontology. For smaller ontologies, shorter walks and fewer iterations, while larger or more complex ontologies required slightly longer walks. We averaged performance across three embedding dimensions to provide a more robust evaluation of each method and also computed the standard deviation to capture variability across runs.
Parameters for Different KGE Methods for Different KGs.

To ensure robust evaluation, we used stratified 5-fold cross-validation, maintaining the same class distribution in each fold. For reproducibility, a fixed random seed was applied throughout the experiments. We initially experimented with a wide range of hyperparameters and, to reduce computational cost, we narrowed the range to focus on the best-performing configurations, as shown in Table 4. Results were averaged to ensure consistency across different configurations.
Parameter Grid for ML Methods.

In this section are shown the experiment results based on the experiment setup that we discussed in Section 6.2, starting with heart disease prediction, followed by kidney disease prediction. We show the concluding results for each research question.
Heart Disease Prediction
Table 5 shows the average accuracy and F2 scores, along with the standard deviation across different vector sizes of the embeddings, for four different ML models (KNN, NN, SVM, XGB). The results include the models’ baseline performance on tabular data alone, compared with their performance when the data is augmented using embeddings generated by four KG embedding algorithms (Node2Vec, RDF2Vec, DistMult, TransH). Additional results, including average recall with standard deviation across vector sizes, evaluated using different KGs, models, approaches and embedding methods, are provided in Table 11 in Appendix 8. In the following, the results are analysed based on the research questions.
Average Accuracy and F2 Scores (With Standard Deviation Across Different Vector Sizes), Across Different KGs, for Various Models, Approaches and Embedding Methods in Heart Disease Prediction.

Average Accuracy and F2 Scores (With Standard Deviation Across Different Vector Sizes), Across Different KGs, for Various Models, Approaches and Embedding Methods in Heart Disease Prediction.

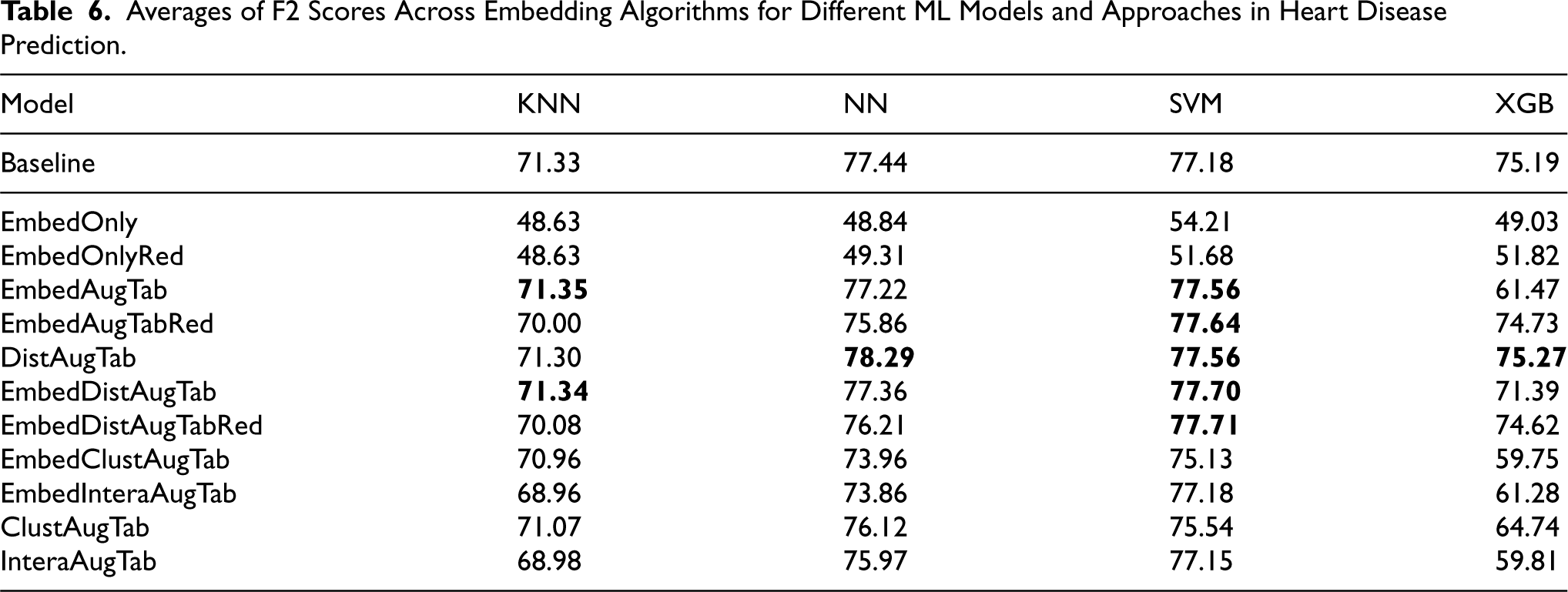
Averages of F2 Scores Across Embedding Algorithms for Different ML Models and Approaches in Heart Disease Prediction.
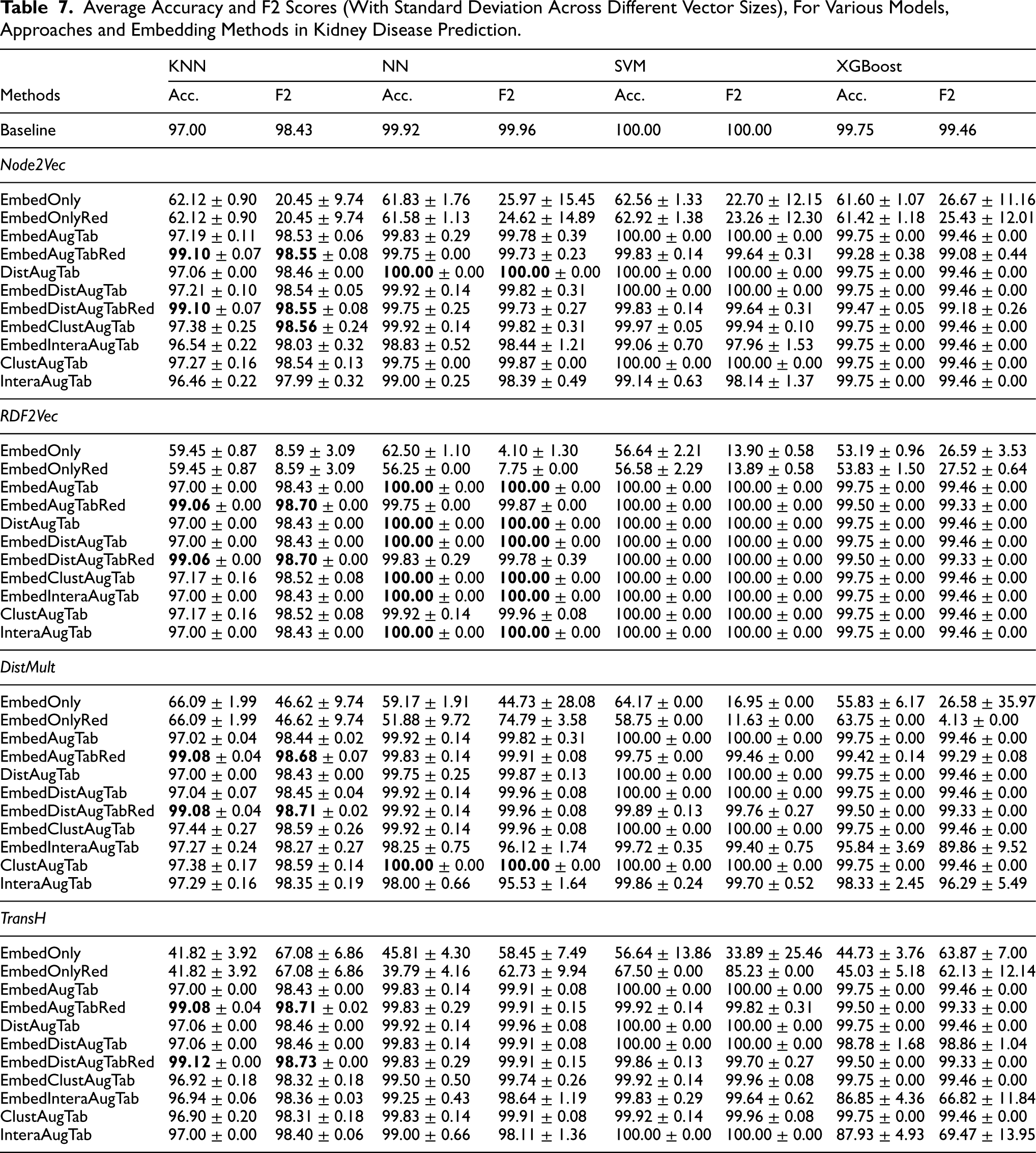
Average Accuracy and F2 Scores (With Standard Deviation Across Different Vector Sizes), For Various Models, Approaches and Embedding Methods in Kidney Disease Prediction.
Averages of F2 Scores Across Embedding Algorithms for Different ML Models and Approaches in Kidney Disease Prediction.

Investigating the impact of various methods for data augmentation through KGE
The different methods of augmenting tabular data with KG embeddings yield mixed results across models. Approaches such as EmbedAugTab, DistAugTab and EmbedDistAugTab often provide the most performance improvements, especially for models such as XGBoost and NN. For example, DistAugTab when Node2Vec is being used to generate the embeddings significantly improved the F2 score of XGB from 75.19 (baseline) to 90.85, highlighting the ability of XGB to effectively use the additional distance features from KG embeddings.
Conversely, SVM and KNN tend to struggle with complex augmentation methods, showing lower gains and even losses in some cases, as they are less suited to high-dimensional data and the resulting feature complexity.
In the following we consider the effectiveness of different approaches based on the sub-hypotheses H1.1 to H1.5.
The ClustAugTab approach, on the other hand also showed improvements, particularly with KNN. For example, using RDF2Vec with ClustAugTab led to better clustering of instances, resulting in improved accuracy for KNN and SVM from 81.02% to 81.18% and from 79.75% to 80.16%, respectively. KNN benefited from this approach as it relies on distance metrics to identify neighbours, and having meaningful clusters aligned with its decision-making process. Similarly, SVM showed better results using RDF2Vec for embedding generations and ClustAugTab approach for data augmentation, possibly because the cluster memberships served as a valuable feature that helped define clearer support vectors for class separation.
For instance, while EmbedAugTabRed and EmbedDistAugTabRed helped reduce overfitting for XGBoost when using DistMult to generate the embeddings by eliminating redundant features, it did not outperform the EmbedAugTab and EmbedDistAugTab methods for KNN. This suggests that, while PCA can be useful for some models it might remove valuable information that more sophisticated models can use, highlighting a trade-off between feature simplification and richness.
Investigating the impact of embedding algorithm
The choice of KG embedding algorithm has a significant impact on model performance across the various approaches. Each embedding method captures different aspects of the KG structure, influencing how well the derived embeddings integrate with the original tabular data and the model’s ability to leverage this information.
From the results shown in Table 5 and the F2 score differences illustrated in Figure 11 7 , it is evident that Node2Vec and RDF2Vec generally lead to more consistent performance improvements compared to DistMult and TransH, particularly when combined with approaches such as EmbedAugTab and DistAugTab. For example, Node2Vec embeddings with the EmbedDistAugTab approach provided the most notable gains across models, including SVM, and XGBoost. This improvement suggests that Node2Vec’s random walk-based approach is effective in preserving local neighbourhood information and graph structure, which seems to translate well into the feature space used by the models. The relational patterns it captures may align better with the tabular features, providing additional context that aids in classification.

F2 score differences relative to baseline across models and embedding methods, showing gain/loss for each approach for heart disease prediction
RDF2Vec also showed good performance, particularly with EmbedAugTab and ClustAugTab approaches. Its ability to leverage RDF graph structures and preserve semantic relationships appears to be beneficial, especially for models such as NN and SVM.
In contrast, DistMult and TransH showed more variable results. While these methods performed well in specific scenarios – such as DistAugTab with DistMult or TransH, particularly for SVM and XGBoost – they were less consistent across different approaches. For example, while DistMult’s tensor factorisation approach allows it to capture specific types of relational patterns, this does not always translate into performance gains when used for approaches such as ClustAugTab, IntraAugTab or EmbedAugTab.
Moreover, the figures show that XGBoost’s performance is particularly sensitive to the choice of embedding algorithm. While XGBoost generally excelled for DistAugTab or EmbedDistAugTab approach using Node2Vec, it underperformed with simpler methods such as EmbedAugTab when combined with TransH or DistMult. This suggests that XGBoost requires embeddings that add clear, structured relational information rather than purely dense vector representations. Thus, Node2Vec and RDF2Vec’s ability to provide richer, more interpretable representations likely aligns better with XGBoost’s learning mechanism.
In conclusion, the choice of the embedding algorithm plays a crucial role in determining the success of different data augmentation approaches. RDF2Vec consistently provides more valuable representations for enhancing model performance across a range of methods, likely due to their strength in capturing both local and global graph structures. DistMult and TransH, while potentially effective in capturing specific relational patterns, exhibit more variability and require carefully chosen augmentation methods to translate their structural information into improved model performance. These findings emphasise that selecting the right embedding algorithm is critical, as it can significantly influence how well the additional relational data is integrated into the learning process.
Investigating the impact of KGs choice
Figure 12 shows the average accuracy and F2 score across all evaluated approaches implemented with each ontology. It shows that the choice of ontology (Small, Extended, or Snomed) slightly affects model performance. Using Snomed ontology generally provides the highest accuracy and F2 scores, due to its clinically structured information from medical experts, highlighting its ability to enrich predictions. The Small KG yields the poorest results among the three ontologies, arguably due to its handcrafted nature by non-medical experts, which limits its depth and relevance to complex medical relationships.

Comparison of accuracy (left) and F2 (right) scores for different ML models across various KGs.
Investigating the performance of different ML models across various approaches and embedding algorithms
Across the evaluated models, XGBoost and NN showed the most significant improvements when incorporating various KG augmentation methods. Specifically, XGBoost’s performance saw the largest gains using the DistAugTab and EmbedDistAugTab approaches. For example, with Node2Vec embeddings combined using EmbedDistAugTab approach, XGBoost’s F2 score increased from a baseline of 75.19% to 89.27%. This can be attributed to XGBoost’s ability to effectively handle high-dimensional feature spaces, allowing it to extract valuable patterns from the distance-based features derived from the embeddings. However, XGBoost showed a lot of underperformance when the datasets were augmented with various approaches, especially when embeddings were generated with TransH and DistMult. This reduced performance may be due to the relational complexity in TransH and DistMult embeddings, which introduces interdependent features that XGBoost struggles to interpret independently.
On the other hand, KNN showed only slight performance gains when augmented with embeddings but maintained stable results across different approaches and embedding algorithms.
Looking at the average F2 scores in Table 6 when different embedding algorithms are used to generate the embeddings, we can observe that for four approaches SVM gained slightly better performance compared to the baseline, making it in general more suitable model that gains performance when additional data from KGs is being added, especially when computing the distances to the target classes, or when the vectors are added as such to augment the tabular data.
Table 7 shows the average accuracy and F2 scores, along with the standard deviation across different vector sizes of the embeddings, for different models, approaches and embedding methods in kidney disease prediction. Additional results, including average recall with standard deviation across vector sizes are provided in Table 12 in Appendix 8. In the following paragraphs, we will discuss the results based on the research questions, considering also the sub-hypothesis H1.1 - H1.5 from Section 5.
Investigating the impact of various methods for data augmentation through KGE
From Table 7, we observe that adding distance-based features to tabular data improves ML model performance, especially for KNN and NN. Although the baseline is already high, enhancements such as distance-to-class, cluster membership features and embedding vectors still boost performance. For example, KNN accuracy increases from 97% to 99.08% and 99.12% when the data is augmented with vector embeddings (EmbedAugTabRed) and with embeddings plus Euclidean distances to classes (EmbedDistAugTabRed), using TransH to generate embeddings. In the following we will discuss the hypothesis H1.1 to H1.5 based on the results.
Conversely, the performance of NN, SVM and XGBoost generally worsened after the dimensionality reduction step. This could be attributed to the already high baseline accuracy (ranging from 99.75% to 100%); further reducing the dimensionality might eliminate features that, while not highly significant, still contribute to the model’s performance. Additionally, these models can inherently manage high-dimensional data and may not benefit as much from PCA as KNN does. As a result, the reduced feature set may lack the nuanced information that these more complex models require for optimal performance.
Investigating the impact of embedding algorithm
Figure 13 shows the F2 score differences relative to the baseline across models and embedding methods, highlighting gains and losses for each approach in kidney disease prediction. The approaches EmbedOnly, EmbedOnlyRed and EmbedInteraAugTab, as well as InteraAugTab, were excluded from analysis due to their skewed performance, particularly when using DistMult and TransH to generate the embeddings, which demonstrated low performance.

F2 score differences relative to baseline across models and embedding methods, showing gain/loss for each approach for kidney disease prediction.

Different ways of infusing the KG as input into ML pipeline.
From the figure we see that RDF2VEC was shown to be the best suited algorithm among the four embedding algorithms for our approaches. It generates effective embeddings particularly for KNN where the F2 score is increased for some of the approaches and stayed the same for the others. Moreover for the SVM model it maintained a perfect F2 score of 100%, in comparison to other embedding algorithms where the performance dropped. This could indicate that RDF2VEC caputes relevant features that enhance the SVM’s ability to establish clear decision boundaries, ultimately resulting in higher predictive performance.
In contrast, Node2Vec generated effective embeddings for the KNN model, where it slightly improved performance. However, its utility decreased for SVM, XGBoost and NN, often leading to slight performance drops. This suggests that while Node2Vec captures local structural information well, it may introduce noise or irrelevant features for models such as SVM and XGBoost.
Using DistMult to generate the embeddings achieved performance gains with KNN but resulted in decreased performance for SVM. This inconsistency indicates that while DistMult enhances KNN’s ability to capture relationships, it introduces noise for SVM’s decision-making process.
Similarly, TransH performed best with KNN, especially in the EmbedAugTabRed and EmbedDistAugTabRed approaches, yet showed weaker results for other models, particularly XGBoost. This discrepancy highlights that TransH may capture specific relational aspects beneficial for KNN but lacks the broader applicability needed for more complex models such as XGBoost.
Overall, our findings suggest that RDF2Vec algorithm is the optimal choice for embedding generation for augmenting data across various models due to its ability to enhance relevant feature representation. Conversely, Node2Vec is particularly advantageous for KNN, emphasising the need to carefully select embedding algorithms based on the specific model and approach used to ensure the most effective performance enhancement.
Investigating the performance of different ML models across various approaches
Table 8 presents the average F2 scores for various ML models in kidney disease prediction, showing the impact of different approaches for combining tabular data with embeddings, averaged across multiple embedding methods. KNN showed the most notable improvements when augmented with KG embeddings across different approaches, likely due to its weaker baseline performance compared to the rest and the suitability of distance-based metrics and dimensionality reduction for this model. Excluding cases where only embeddings were used for training, NN generally maintained its performance with only slight drops in some approaches, suggesting that NN’s ability to learn complex patterns is somewhat robust to variations in feature augmentation. SVM, which achieved a perfect F2 score (100%) with only tabular data, retained this performance in EmbedAugTab, DistAugTab and EmbedDistAugTab. Similarly, XGBoost preserved its performance with the four embedding-augmented configurations, though it experienced slight declines in the remaining cases.
In this article, we proposed several innovative approaches to augment tabular data with semantic information by leveraging ontologies to capture domain semantics as shown in Figure 14. We utilised these ontologies to construct KGs, thereby enriching the datasets with structured ontological information. To make the KGs suitable for ML models, we employed KGEs to transform the graphs into a vector space representation. This process enhances the data used to train ML models by integrating domain-specific semantics, allowing the models to leverage contextual and relational information. Based on our experiment setup, we conducted experiments for heart and kidney disease prediction.
For RQ1, our experiments demonstrated that incorporating KG embeddings, particularly by augmenting tabular data with distance-based features to target classes, improves model performance in most of the cases. This enhancement is particularly evident in challenging domains such as chronic kidney disease, where accuracy and F2 scores improved despite limited room for improvement, underscoring the value of KG information for refining ML predictions, especially in data-sparse environments.
For RQ2, our findings indicate that RDF2Vec is the most effective embedding algorithm across models for both heart and kidney disease prediction, given its ability to capture relevant feature representations without performance drops. Node2Vec proved particularly beneficial for KNN in kidney disease prediction, while in heart disease prediction, Node2Vec enhanced XGBoost the most. However, XGBoost exhibited instability across approaches and embedding algorithms in both cases, suggesting the need for careful pairing of embedding methods and models.
For RQ3, in one hand for heart disease prediction overall on average SVM showed the most F2 score improvement across multiple approaches. Whereas on the other hand for kidney disease prediction, KNN showed the largest performance gains when enhanced with KG embeddings across various approaches, likely due to its weaker baseline performance and the suitability of distance-based metrics and dimensionality reduction, which complement KNN’s neighbour-based approach.
Future work will explore the effectiveness of KGs across diverse domains, particularly those with limited data, by augmenting sparse datasets to address the data dependency issues in ML models. Additionally, we plan to assess the scalability of our methods based on data size and structure and experiment with more complex ML models to further optimize the integration of KG embeddings. Furthermore, we aim to explore alternative embedding models and investigate methods for mapping literals into the embedding space to evaluate their impact on model performance.
Footnotes
Acknowledgements
This work was supported by the FFG SENSE (894802) and FAIR-AI (904624) projects, as well as by the Austrian Science Fund (FWF) BILAI 10.55776/COE12 and HOnEst (V 745-N) projects. For open access purposes, the author has applied a CC BY public copyright license to any author accepted manuscript version arising from this submission.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
