Abstract
Background
Allergic rhinitis (AR) is the most common inflammatory disorder of the upper airway caused by aberrant immune responses to allergens in genetically predisposed individuals. Recently, the long noncoding RNA (lncRNA) antisense noncoding RNA in the INK4 locus (ANRIL) has been identified as a novel genetic factor associated with increased AR risk.
Objectives
This study aimed to evaluate the potential correlation of ANRIL gene single nucleotide polymorphisms (SNPs) with AR risk in the Kurdish population of Kermanshah, Iran.
Methods
In this case–control study, 130 AR patients and 130 healthy controls were recruited to genotype for two SNPs of the ANRIL gene (rs1333048 and rs10757278) using the Tetra-primer amplification refractory mutation system polymerase chain reaction (T-ARMS-PCR) method.
Results
Our results showed no significant difference for the alleles and genotypes frequency distribution of lncRNA ANRIL SNPs (rs1333048 and rs10757278) between AR patients and healthy controls (p > 0.05). Additionally, the dominant, additive and recessive genetic models of both SNPs were not associated with altered susceptibility to AR risk (p > 0.05).
Conclusion
The results demonstrated that the ANRIL gene rs1333048 and rs10757278 polymorphisms might not be associated with susceptibility to AR in the Kurdish population of Kermanshah, Iran
Introduction
Allergic rhinitis (AR) is a chronic inflammatory disease of the nasal mucosa characterized by nasal rhinorrhea, sneezing, itching, and congestion. Epidemiological studies have reported that the prevalence of AR is 10% to 30% of adults and up to 40% of children worldwide.1,2 The characteristic inflammation in AR is induced by immunoglobulin E (IgE)-mediated reactions against the inhaled allergens, during which different inflammatory cells, including T helper (Th)2 and Th17 cells, mast cells, basophils, and eosinophils are activated. Activation of the immune cells during AR results in the production of a variety of proinflammatory cytokines and mediators, such as interleukin (IL)-13, IL-4, IL-5, IL-17, tumor necrosis factor (TNF)-α, histamine, and leukotrienes, contributing to the increased inflammation of AR.1,3 In addition to environmental exposures, genetic predisposition and epigenetic modifications might be involved in the dysregulation of the immune system in AR.4,5
Emerging evidence has placed a new emphasis on the role of long noncoding RNAs (lncRNAs) in regulating immune responses and the pathogenesis of several immune-related disorders, such as allergic diseases.6–8 lncRNAs are a class of RNAs containing more than 200 nucleotides in length with no or very low protein-coding potential. 9 Based on findings from other studies, lncRNAs can participate in epigenetic regulation of gene expression at transcriptional, posttranscriptional, translation, and posttranslational levels through interacting with DNA, mRNAs, microRNAs (miRNAs), and proteins. Furthermore, lncRNAs also contribute to a variety of biological processes, including embryonic development, cell differentiation, and metabolism.9,10
Antisense noncoding RNA in the INK4 locus (ANRIL), as one of the earliest discovered lncRNAs, plays an important role in the development and progression of inflammation-related diseases, such as systemic lupus erythematosus, cardiovascular disease (coronary artery disease [CAD]), diabetes, and cancers.11–14 The ANRIL gene is located in the chromosome 9p21 region, and it has been identified as a novel genetic factor associated with the pathogenesis of AR. 15 According to a recent study, long noncoding RNA (lncRNA) ANRIL expression in the nasal mucosa of AR patients was upregulated compared to controls. It was positively correlated with increased AR risk, severity, and inflammation. 15 More interestingly, lncRNA ANRIL expression was discovered to be positively correlated with increased TNF-α, IL-4, IL-6, IL-13, and IL-17. In contrast, it was negatively correlated with interferon (IFN)-γ and IL-10 mRNA expressions, suggesting that it was related to elevated inflammation of AR. 15
In recent years, given the critical role of lncRNA ANRIL in the pathogenesis of various inflammatory diseases, single nucleotide polymorphisms (SNPs) of the ANRIL gene have gained widespread attention.16,17 Although the role of ANRIL in regulating immune responses and expressing some AR-related factors has been acknowledged, the participation of ANRIL genetic variants in AR has not yet been assessed. Genetic variants of the ANRIL gene may act as a risk or protective factor for AR development. Regarding the indispensable contribution of lncRNA underlying inflammatory disease, this is the first study aimed to evaluate the role of ANRIL gene polymorphisms with AR susceptibility in the Kurdish population of Kermanshah, Iran.
Materials and Methods
Study Area and Population
The study population consisted of 130 AR patients and 130 age- and gender-matched healthy controls. All samples were chosen from the Kurdish population of Kermanshah province, located in the west of Iran. The Kurds, the most ancient indigenous people, lived mainly in the Zagros Mountains for thousands of years and had their own sociocultural values. To note, only the Kurdish population was included in this study, and participants with other ethnicities were excluded. An allergist diagnosed all patients with AR according to diagnostic criteria described by the Practical Guideline for managing AR. 18 Healthy controls were collected from blood donors with no history of inflammatory, allergic, or autoimmune disorders. The study protocol was approved by the ethics committee of Kermanshah University of Medical Sciences (IR.KUMS.REC.1399.1008). After being informed of the study and obtaining written consent from all study participants, blood sampling was conducted.
DNA Extraction and SNP Genotyping
The genomic DNA samples from peripheral blood of the patients and control subjects were extracted by the salting-out method. The purity and concentration of the extracted DNA samples were determined by the NanoDrop 2000-UV-Vis spectrophotometer (Thermo Scientific, USA).
Genotype determination for selected SNPs in the ANRIL gene (rs1333048 and rs10757278) was performed by the tetra-primer amplification refractory mutation system polymerase chain reaction (T-ARMS-PCR) method. The specific primers for both SNPs were designed by the PRIMER1 online tool (http://primer1.soton.ac.uk/primer1.html). The location of the SNPs and primers sequence are shown in Table 1.
The sequence of primers and product size for T-ARMS-PCR.
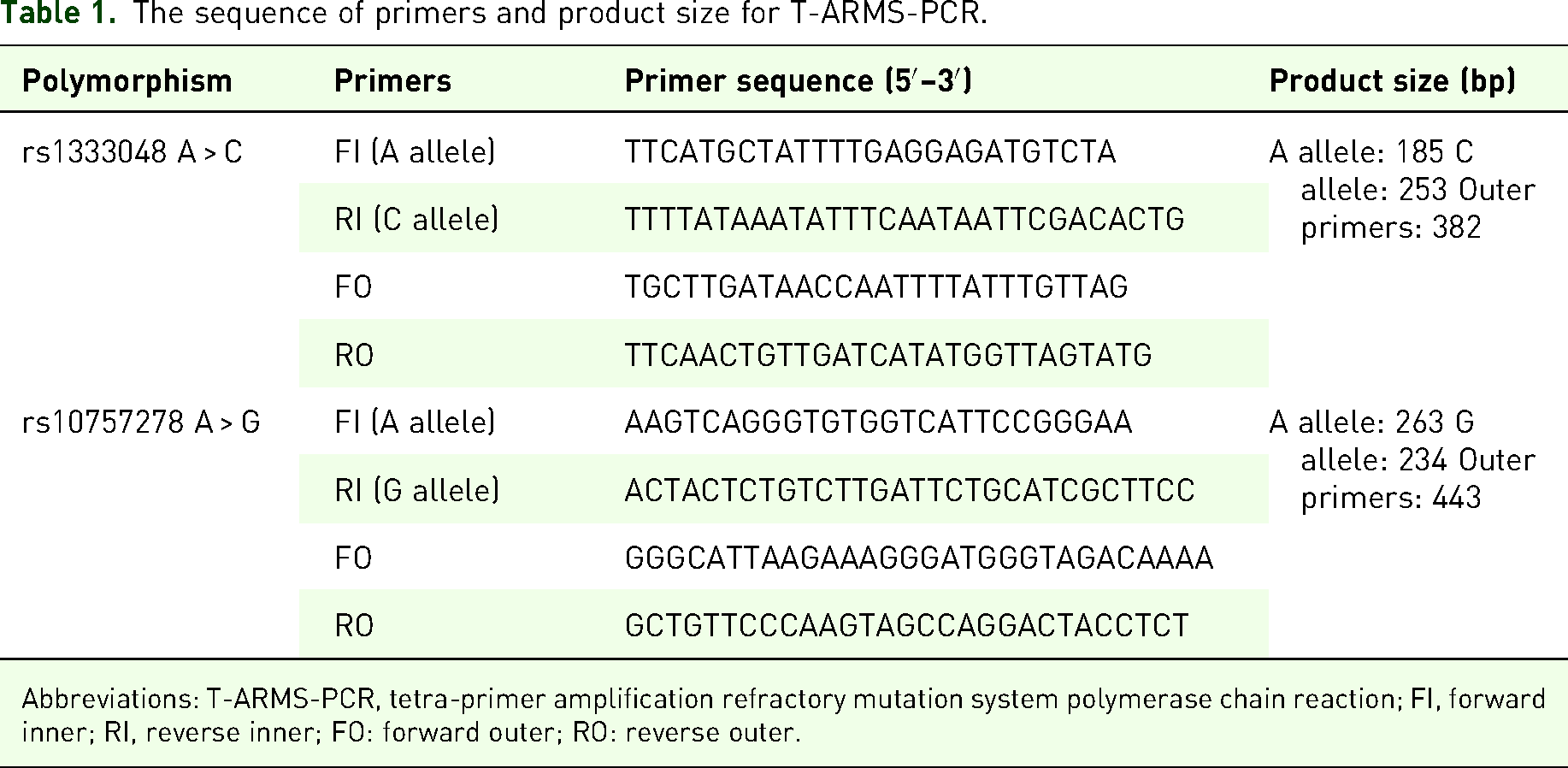

Abbreviations: T-ARMS-PCR, tetra-primer amplification refractory mutation system polymerase chain reaction; FI, forward inner; RI, reverse inner; FO: forward outer; RO: reverse outer.
The PCRs were carried out in microtubes with a total volume of 15 μl, containing 0.5 μl of genomic DNA as a template, 0.7 μl of each inner primer, 0.3 μl of each outer primer, 7 μl of multiplex PCR master mix, and 5.5 μl of DNAase-free distilled water. PCR amplification was performed with an initial denaturation at 95 °C for 5 min, followed by 35 cycles of 95 °C for 45 s, annealing for 30 s at 52 °C for rs1333048, 30 s at 60 °C for rs10757278, and extension for 55 s at 72 °C, with a final extension of 72 °C for 5 min. The amplified products were electrophoresed on 2% agarose and stained with DNA-safe stain. According to the agarose gel electrophoresis for rs10757278 A > G, the AA genotype was represented if two fragments with 443 and 262 bp were generated, the AG genotype was detected in three fragments with 443, 262, and 236 bp, and the GG genotype was identified if two fragments with 443 and 236 bp were generated. Based on the agarose gel electrophoresis for rs1333048 A > C, the AA genotype produced 2 fragments of 381 and 184 bp, the AC genotype yielded 3 fragments of 381, 252, and 184 bp, and the CC genotype had 2 fragments of 381 and 252 bp. A schematic representation of the study protocol is shown in Figure 1.
A schematic representation of the study protocol.
Statistical Analysis
A comparison of genotype and allele distribution between cases and controls was analyzed by the chi-square test. Using logistic regression analyses, the relation of genotypes with AR risk was estimated by calculating odds ratios (ORs) and their 95% confidence intervals (CIs). The web-based SHEsis software (version 4.2) was used to analyze pairwise linkage disequilibrium and haplotype structure. The accordance of genotype frequencies with Hardy–Weinberg equilibrium (HWE) was performed using the chi-square test. Differences with a P-value <0.05 were considered statistically significant. Statistical analyses were performed using the SPSS software version 23 (SPSS, Chicago, IL, USA).
Results
Demographic Characteristic
The study population consisted of 130 AR patients, involving 60 males (46.15%) and 70 females (53.85%) and 130 control subjects, containing 60 males (46.15%) and 70 females (53.85%). The mean age of AR patients and the control group was 35.81 ± 10.75 and 35.92 ± 10.45, respectively. There were no statistically significant differences between the 2 groups of AR patients and healthy controls for age and gender (P > 0.05), indicating that the study groups were matched regarding age and gender.
Allele and Genotype Frequencies of SNPs
The genotype distribution and allele frequencies of the 2 investigated SNPs, rs1333048 and rs10757278 in the ANRIL gene in healthy controls and AR patients are summarized in Table 2. The distribution of genotype frequencies of rs1333048 (but not rs10757278) was in accordance with HWE in both patient and control groups.
Distribution of Allele and Genotype Frequencies for ANRIL Gene Polymorphisms in Patients With AR and Controls.

Abbreviations: ANRIL, antisense noncoding RNA in the INK4 locus; AR, allergic rhinitis; SNP, single nucleotide polymorphism; OR, odds ratio; CI, confidence interval.
The 2 SNPs were successfully genotyped for AR patients and healthy controls. Select T-ARMS-PCR gels for rs10757278 and rs1333048 polymorphisms are shown in Figures 2 and 3, respectively.

Products of lncRNA ANRIL rs10757278 A > G polymorphism on 2% agarose gel. Homozygotes wild AA genotype (443 and 262 bp); heterozygous AG genotype (443, 262, and 236 bp); mutant GG genotype (443 and 236 bp); NG (negative control).

Products of lncRNA ANRIL rs1333048 A > C polymorphism on 2% agarose gel. Homozygotes wild AA genotype (381 and 184 bp); heterozygous AC genotype (381, 252, and 184 bp); mutant CC genotype (381 and 184 bp); NG (negative control).
Our results revealed no significant differences in the allele and genotype frequencies of ANRIL SNPs (rs1333048 and rs10757278) between AR patients and healthy controls (P > 0.05). In addition, the genetic models of dominant, additive, and recessive as well as haplotype frequencies showed no significant association with AR risk (P > 0.05; Tables 2 and 3).
Distribution of Haplotypes for ANRIL Gene Polymorphisms in AR Patients and Control Group.

Abbreviations: ANRIL, antisense noncoding RNA in the INK4 locus; AR, allergic rhinitis; OR, odds ratio; CI, confidence interval.
Discussion
In the current study, we investigated the association between the ANRIL gene polymorphisms, including rs1333048 and rs10757278, with susceptibility to AR in the Kurdish population of Kermanshah, Iran. Our results showed no significant association between these 2 polymorphisms and AR risk.
From an etiopathological point of view, AR is considered a complex multifactorial inflammatory disease in which genetic factors are critically involved in the activation of inflammatory responses. 19 Although many studies have identified a large number of genetic factors involved in the pathogenesis of AR, the underlying mechanisms of gene dysregulation have not yet been identified. Recently, noncoding RNAs (ncRNAs), such as miRNAs and lncRNAs, as crucial regulators of gene expression, have been suggested to be involved in the regulation of immune responses and the pathogenesis of several immune-related disorders.8,20–23
Regarding the nature of AR as Th2-mediated responses and the role of lncRNAs and miRNAs in the regulation of Th2 responses, no wonder that altered expression of these ncRNAs might contribute to the pathogenesis of AR. Intriguingly, the importance of miRNAs was determined in AR pathogenesis. For example, upregulation of miR-126-5p, miR-19a-5p, and miR-26a-5p was reported in the nasal mucosa of AR patients compared to healthy control. 23 Recently, a study indicated that lncRNA ANRIL expression in the nasal mucosa of AR patients was remarkably increased. It was positively associated with the upregulation of proinflammatory cytokines (IL- 4, IL-6, IL-13, and IL-17) as well as disease severity, such as itching and congestion. Hence, these observations suggest that the lncRNA ANRIL might be involved in the pathogenesis of AR. 15 Although it is not clear how lncRNA ANRIL might play a role in regulating AR-related cytokines and miRNAs, it appears that the sponging of antiinflammatory miRNAs, such as miR-Let7e and miR-18b by lncRNA ANRIL might be an explanation.
Interestingly, the binding of ANRIL to an ANRIL binding transcriptional factor (Yin Yang 1) may be considered another mechanism, leading to upregulating IL-6 and IL-8 expression in human endothelial cells.15,24–26 It is worth emphasizing that signal transducer and activator of transcription 1 (STAT1) signaling contributed to ANRIL expression, which participates in the regulation of immune response through induction of the proinflammatory cytokine IFN-γ. 27 Given the association between the nuclear factor (NF)-κB pathway and the crucial role of ANRIL in the expression of several proinflammatory genes regulated by NF-κB,28,29 ANRIL might be a missing puzzle piece in the pathogenesis of AR. However, this speculation should be further investigated.
One of the SNPs within the ANRIL gene has been reported to change the phenotypic traits and function of the genes, such as changes in the binding site for proteins involved in signaling pathways.27,30 For instance, the polymorphism rs10757278 of the ANRIL gene has been demonstrated to disturb the binding of STAT1. 27 In recent years, different studies have demonstrated the association of SNPs of the ANRIL gene with susceptibility to different diseases.21,30,31
Taheri et al 30 revealed that the GG genotype of rs10757278 and the AA genotype of rs1333048 A > C were associated with prostate cancer and benign prostate hyperplasia risk in an Iranian population. 30 Besides, rs10757278 was significantly associated with CAD risk in patients from Iraq, Iran, and Poland.31–33 Additionally, ANRIL gene rs10757278 and rs1333048 SNPs were shown to confer a risk for psoriasis development in an Iranian population. 23 In accordance with our study, there was no significant association between ANRIL gene rs1333048 and rs10757278 SNPs with multiple sclerosis (MS) and breast cancer.21,34 However, haplotype analysis of ANRIL gene polymorphisms (rs1333045, 1333048, rs4977574, and rs10757278) indicated a protective effect of CCGG and TAAA haplotypes in MS, while TAGG and CCGA haplotypes were significantly associated with an increased risk of MS. 21 Moreover, haplotype analysis (with an order of rs1333045, 1333048, rs4977574, and rs10757278 SNPs) demonstrated that TCGA haplotype was associated with breast cancer risk. 34 The discrepancy in the reports concerning the contribution of ANRIL polymorphisms to different diseases might stem from population-specific genetic stratification, sample size, different genotyping methods, and the involvement of other genetic factors.
In addition, other studies have revealed a genetic predisposition to type 2 inflammatory disorders. A systematic review has shown a significant association between major SNPs with chronic rhinosinusitis and the specific pathways involved. 35 Furthermore, a meta-analysis supported that IL-13 SNP rs20541 was associated with increased AR risk, particularly in Asians, while no significant association was detected in Caucasians. 36 These findings suggest that due to different sequencing and sampling methods, more studies with a wider scope are needed to identify important SNPs.
Several potential limitations in our study should be pointed out. First, we did not investigate the lncRNA ANRIL expression in the study groups and could not determine a link between the SNPs and transcription of lncRNA ANRIL. Second, the small sample size and inclusion of only 1 Iranian population can decrease the statistical power of our study. Thus, we recommended replication studies with a larger sample size to assess the involvement of ANRIL polymorphisms in AR pathogenesis. Third, the present study did not cover all of the genetic polymorphisms on ANRIL, and thus, further studies are required to identify potential causative variants. Fourth, the distribution of genotypes of rs10757278 was not by HWE in the study groups, probably due to inbreeding or small population sizes. Therefore, the obtained results should be interpreted with caution.
Conclusion
In conclusion, our results represented no significant difference between ANRIL gene rs10757278 and rs1333048 polymorphisms with AR susceptibility. Further studies are needed to indicate the possible contribution of ANRIL gene SNPs as a genetic risk factor in the pathogenesis of AR.
Footnotes
Ethics Approval
This study was approved by the ethics committee of Kermanshah University of Medical Sciences (IR.KUMS.REC.1399.1008).
Consent to Participate
All patients gave written informed consent to participate in the study extension.
Consent for Publication
Participants consented to publication.
Author Contributions
AR was involved in the concept and design of the study. SF drafted the manuscript. All authors were involved in data collection, analysis, and interpretation and approved the final manuscript.
Author Contribution(s)
Acknowledgments
The authors thank the Vice Chancellor for Research and Technology of Kermanshah University of Medical Sciences for financial support. This article was preprinted on bioRxiv preprint server37 (https://doi.org/10.21203/rs.3.rs-1401694/v1).
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Research Administration Department of Kermanshah University of Medical Sciences, Kermanshah, Iran (grant no. 990972).
Competing interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of Data and Material
All data related to particular cases are available with individual authors/clinicians that were in charge of the patients, and further data (if relevant to the report) are available on request.
