Abstract
In hormone-receptor-positive breast cancer, the main treatment is tamoxifen. Tamoxifen is a weak antiestrogen that requires prior activation to its most active metabolite, endoxifen, which has superior antiestrogenic activity. In the search for predictive biomarkers to optimize treatment and reduce tamoxifen resistance, studies have concluded that endoxifen concentrations are associated with CYP2D6 activity and that the concentration of endoxifen and other metabolites (whose activities are also linked to the activity of other pharmacogenes) may be associated with tamoxifen efficacy. The aim of this review is to highlight the evidence concerning the contribution of pharmacogenes (CYP2D6, CYP2C9/19, CYP3A4, UGT, and SULT) to tamoxifen metabolism and the effect of the genetic variation to which they are subject on the concentration of active tamoxifen metabolites.
Introduction
Breast cancer, which develops in the mammary glands, is the most frequently diagnosed malignant tumor in women, accounting for one in every eight diagnoses, with 2.3 million new cases in both sexes combined. It is estimated that 685,000 women died of breast cancer in 2020, that is, 16% or one in six cancer deaths. 1 These high incidence and death rates make breast cancer a real health problem worldwide.
There are several treatment options, including hormone therapy, proven to reduce the recurrence rate of hormone-sensitive (hormone receptor-positive) breast neoplasms in pre- and post-menopausal patients. 2 Around 80% of newly diagnosed elderly patients have hormone receptor-positive breast cancer. 3 The hormonal treatment of first choice remains tamoxifen, a selective estrogen receptor modulator. Its use over 5–10 years improves survival and reduces the recurrence rate. This antiestrogen requires metabolic activation into two main metabolites: endoxifene and 4-hydroxy-endoxifene, and both have a 100 times greater affinity for estrogen receptors. This metabolic conversion is mainly carried out by one of the cytochrome P450 family isoenzymes, CYP2D6. 4
However, tamoxifen metabolism appears to be complex, involving other genes such as CYP2C and CYP3A, SULTs, and UGTs.
Despite the efficacy of tamoxifen, there is inter-individual variation in response to treatment. It may have several causes, such as gender, age, drug–food and drug–drug interactions, obesity, comorbidity, renal and hepatic function, pregnancy, circadian rhythm, as well as predominantly genetic factors. 2 These variations happening in DNA sequences are mostly single nucleotide polymorphisms (SNP).
A linear regression analysis to assess the independent effect of age on the variability of tamoxifen and endoxifen concentrations showed that age explained up to 5.1% of the variability, independently of CYP2D6 phenotype. 5
Due to the lack of metabolite concentration measurement in the tamoxifen test, CYP2D6 activity remains the key determinant of endoxifen concentration. According to the literature, the CYP2D6 genotype explains up to 50% of endoxifen variability and around 9% of 4 hydroxy-tamoxifen variability. 6
The aim of this review is to highlight the evidence concerning the contribution of pharmacogenes (CYP2D6, CYP2C9/19, CYP3A4, UGTs, and SULTs) to tamoxifen metabolism, and the effect of the genetic variation to which they are subject on the concentration of active tamoxifen metabolites.
Methods
A literature review was conducted in the PUBMED databases. The search focused on articles published from inception to July 2023. Keywords included breast cancer, cytochrome P450, tamoxifen, and genetic polymorphisms.
This review included studies on tamoxifen metabolism, the association of SNPs with drug response, and the impact of pharmacogenes. In addition, this review excluded studies on neoplasms other than breast cancer, treatments other than tamoxifen, or other studies not reporting sufficient data.
Results
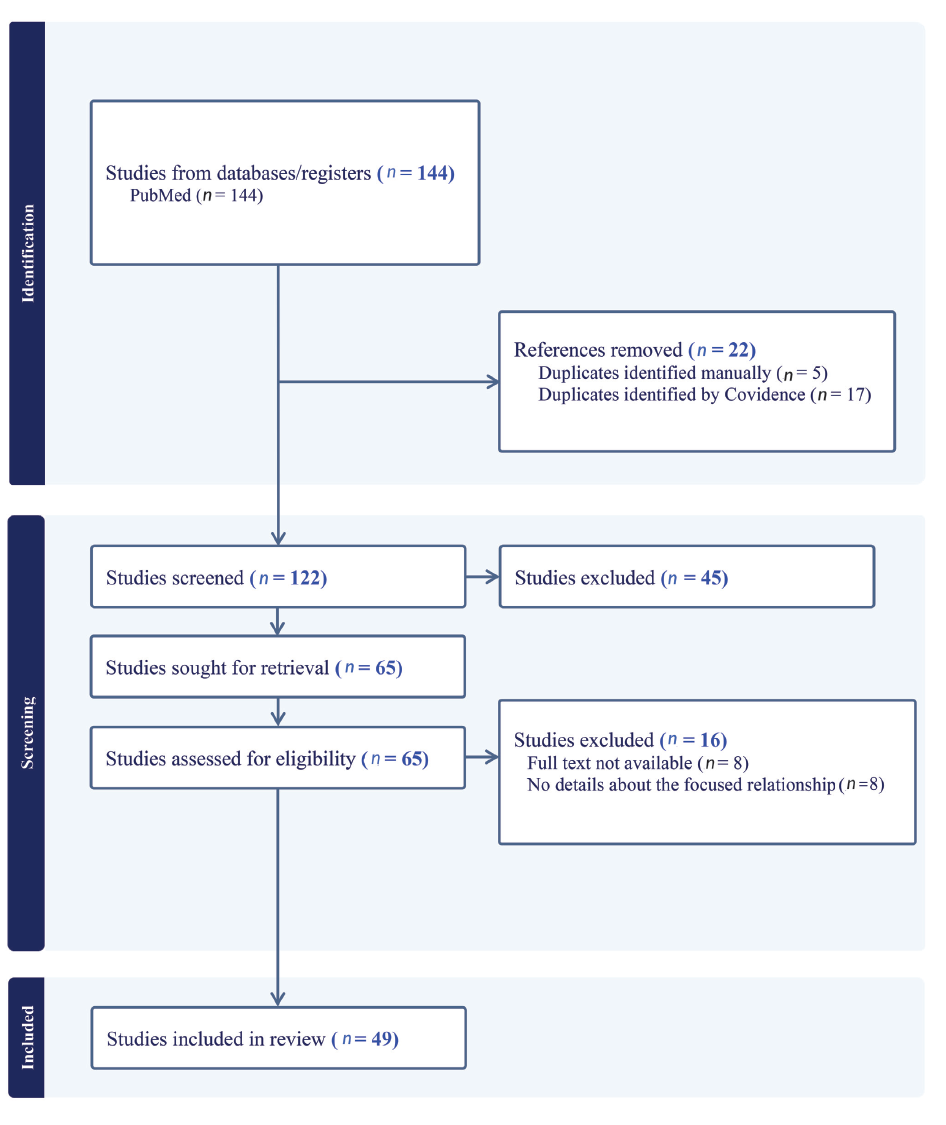
Using the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) technique or flow diagram, a systematic search on PubMed identified 144 studies with the keywords: breast cancer, cytochrome P450, tamoxifen, and genetic polymorphisms. Among these, 22 duplicates were detected and excluded. The titles and abstracts of the remaining 122 studies were examined, leading to the exclusion of 45 articles deemed irrelevant. This left 65 studies for further evaluation. In the assessment for eligibility (n = 65), 16 studies were excluded. Specifically, eight studies were removed due to the absence of the complete text, and another eight were excluded for lacking pertinent information on the relationship between genetic polymorphisms and clinical outcomes.
Consequently, 49 studies were included in the review, each meeting the defined eligibility criteria and providing comprehensive information on the focused relationship. The contents of these studies were subsequently analyzed and discussed in this academic paper. The flow diagram is shown in Figure 1.
Flow Diagram (PRISMA).
In the early 1980s, landmark studies conducted by the Nolvadex Adjuvant Trial Organization (NATO) 7 and the National Surgical Adjuvant Breast and Bowel Project (NSABP) 8 provided the first evidence of clinical benefit for early-stage breast cancer patients receiving adjuvant tamoxifen. 9
Similarly, a meta-analysis of patients from 20 trials, comparing around 5 years of tamoxifen treatment versus no adjuvant tamoxifen, reported a safe reduction in the risk of breast cancer recurrence and death over 15 years. Slightly ER-positive patients fared better than others. 10
In another meta-analysis, the same group reported a 31% reduction in annual mortality in ER-positive patients after 5 years of tamoxifen administration, largely independent of chemotherapy. 11
Tamoxifen Metabolism
Tamoxifen is a prodrug, which is converted into several derivatives by phase I enzymes. Of these, 4-hydroxytamoxifen (4 OH-TAM) and endoxifen have the highest affinity for the estrogen receptor.
As the plasma concentration of endoxifen is 6–12 times higher than that of 4 hydroxy-tamoxifen, and endoxifen has the lowest emergency IC50, endoxifen is considered the main active metabolite of tamoxifen. 12
The metabolic activity of cytochrome P450 enzymes can be predicted from their genotype, allowing patients to be classified as poor (PM), intermediate (IM), extensive (EM), or ultra-fast (UM) metabolizers. 13
CYP2D6
CYP2D6 plays a vital role in the synthesis of active metabolites from tamoxifen. Around 50% of the variability in endoxifen concentration can be explained by CYP2D6 genotype activity, with an association between CYP2D6 gene variation and endoxifen concentration.6, 14
Studies report a reduction in endoxifen concentration in homozygous patients for non-functional alleles compared with wild-type by 30% and 74%, 15 as well as a low endoxifen concentration in slow metabolizers compared with nominal metabolizers. 16
Nearly 30 studies have investigated the association between endoxifen and the CYP2D6 genotype, showing that the CYP2D6 genotype is now an established predictor of endoxifen concentration efficacy. 17 Studies such as Lim et al. have reported an association between CYP2D6 and lower steady-state plasma concentrations of active tamoxifen metabolites. In addition, a significantly higher rate of Z-4-hydroxy-tamoxifen formation was observed with two CYP2D6 functional alleles. 18
However, concerning the association between endoxifen concentration and drug response, the risk of recurrence and overall survival is unclear. 19 Therefore, other variables contributing to the variability of endoxifen concentration need to be considered.
Other enzymes contributing to endoxifen concentration variability include the following:
CYP2C
CYP2C contributes less than 3% to endoxifen synthesis. CYP2C9 contributes to the formation of primary tamoxifen metabolites: N-desmethyl tamoxifen and 4-hydroxy tamoxifen. 18 CYP2C9 enzymes can exhibit different metabolic activities depending on their polymorphic state. Concerning the *1A allele, CYP2C9 metabolic activity can be normal. However for alleles *3, *5, *8, *11A, and *13, it is diminished or entirely absent for *6. 4 The most common variants, CYP2C9*2 and CYP2C9*3, are the most extensively studied, with variable frequencies reported in different populations or ethnic groups. 20 Clearance of 4-hydroxy tamoxifen was found to be respectively 48% and 49% lower in CYP2C9*2 and CYP2C9*3 variants than in wild-type homozygotes. 18 On the contrary, other studies such as Teft et al. reported no association between metabolite levels in homozygotes and the two variants. 21
CYP2C19 is a highly polymorphic enzyme located on chromosome 10. CYP2C19 is involved in the metabolism of several important therapeutic drugs, and the influence of its activity on tamoxifen metabolism has attracted considerable interest. 22
According to a study by Gjerde et al., a higher formation of 4-hydroxy-tamoxifen was observed in CYP2C19*17 carriers; 23 the same allele was associated with better clinical outcomes due to the high enzymatic activity of CYP2C19*17. 23 In contrast to two other studies, there was no association between CYP2C19 genotypes and concentrations of endoxifen, 4-hydroxy-tamoxifen, and N-desmethyl tamoxifen.24, 25
CYP3A4
CYP3A is mainly involved in the conversion of tamoxifen to N-desmethyl tamoxifen and 4-hydroxy tamoxifen to endoxifen. 26
For CYP3A4, carriers of the CYP3A4*22 variant, in addition to being four times more numerous than wild-type carriers, had higher concentrations of tamoxifen, N-desmethyl tamoxifen, and 4-hydroxy tamoxifen.27, 28 This is unexpected because it is a polymorphism of reduced activity. Other studies, however, reported no association between CYP3A4 variation and plasma concentrations of endoxifen or tamoxifen, but with some metabolic ratios (endoxifen/4-OH-Tam). 29
UGTs
UGTs (UDP-glucuronosyl-transferases) are phase II metabolic enzymes that, with the addition of a glucuronide group, prepare compounds for elimination. There are two types of glucuronidation: N-glucuronidation and O-glucuronidation. N-glucuronidation occurs for both TAM and 4-OH-TAM, while O-glucuronidation occurs for 4-OH-TAM and endoxifen.30, 31
There are several UGTs, the most studied of which are UGT1A4, UGT2B7, and UGT2B15. 32 Several studies have highlighted the role of each of these in tamoxifen metabolism. 33 Nevertheless, some of these results require further validation, as in the case of UGT2B15, where patients with the UGT2B15 Lys526Thr deletion have higher concentrations of some glucuronide metabolites.
This is not quite the case for the UGT1A4 Leu48Val variant, which is exclusively responsible for the glucuronidation of tamoxifen and 4 OH-tamoxifen; 31 several studies have reported an association between the concentration of glucuronide metabolites and this variant. 12
SULTs
SULTs or sulfotransferases are hepatic phase II enzymes that increase water solubility for better excretion. 17 SULT1A1 is involved in the inactivation of NDM-tamoxifen, after several consecutive reactions, to the metabolite sulfate.26, 34
With SULT1A1 and SUL1TA2 being the most expressed isoforms, studies have reported elevated concentrations of endoxifen and 4-hydroxy endoxifen in patients each carrying the most frequent variants of these two isoenzymes.12, 17 Several studies have described the genetic variation of SULT1A1 and its influence on tamoxifen metabolite concentrations and metabolic ratios. Gjerde and colleagues reported an association between the SULT1A1 genotype and NDM-tamoxifen/tamoxifen metabolic ratios. 35
A decrease in SULT1A1 enzymatic activity has been described in carriers of both variants, rs6839 and rs1042157, with an increase in endoxifen concentration and even high concentrations of metabolic ratios of tamoxifen and its 34 metabolites of 0.4%–1.3%, 36 whereas studies by Jin 37 and Fernandez-Santander 38 showed no association between SULT1A1 genotypes and tamoxifen and metabolite concentrations.39, 40
On the contrary, there is no evidence of an association between endoxifen concentration and relapse-free survival, which suggests that an association between SULT1A1 and relapse-free survival is unlikely.36, 41
Discussion
This study investigated the influence of inter-individual variability in tamoxifen metabolite enzymes on response to treatment. Although other treatments, such as aromatase inhibitors, can be used as alternatives, tamoxifen remains the first choice given the data reporting the clinical results of tamoxifen, as well as the impact of variability in the enzymes that produce tamoxifen’s active metabolites, known as cytochrome P450. Furthermore, given that several studies have shown that SNPs can influence the enzymatic activity of CYPs and other enzymes involved in tamoxifen metabolism, 42 we were interested to see whether these SNPs could affect or be associated with the clinical results of tamoxifen.
Nevertheless, despite the controversy surrounding the results observed, which we did not omit to mention in our review, we believe that direct measurement of tamoxifen metabolites could be an avenue to be exploited for tamoxifen dose adjustment even if we cannot conclude with certainty that there is indeed a significant association between variations in tamoxifen metabolites and outcomes.
Studies suggesting a significant association reported a correlation between variations in tamoxifen metabolite concentrations and the genotypes of genes responsible for metabolite synthesis, elimination, and/or survival.43–45 This contradicts other studies that found no association between the gene genotypes and clinical outcomes. In some cases, as in the study by Thota et al., the association was not significant for some genes but was significant for others, all involved in tamoxifen metabolism. 46
Moreover, it has long been established that other factors, as mentioned above, can influence inter-individual variability. A linear regression analysis to assess the independent effect of age on the variability of tamoxifen and endoxifen concentrations showed that age explained up to 5.1% of the variability, independently of CYP2D6 phenotype. 2 Similarly, the concentrations of specific tamoxifen metabolites or metabolite ratios are influenced by other factors such as ethnicity or BMI. This was the case for the concentration of N-desmethyl tamoxifen and endoxifen in the study by Saladores et al. 45
Another critical point to note is that, as, for CYP2D6, there is no lack of evidence concerning the effect of its genetic variation on endoxifen concentration, 47 CYP2D6 is the most significant contributor to tamoxifen bioactivation. 29 Indeed, multivariate regression analyses on all cohorts showed that combined genetic (CYP2C9, CYP2C19, and CYP3A5) and non-genetic (age and BMI) factors contributed to only 2.8% of the DM-Tam/endoxifen ratio compared with 53% for CYP2D6. 45
This has contributed to its use as a biomarker to guide tamoxifen treatment. Researchers were able to normalize endoxifen levels using tamoxifen dosing guided by pharmacogenetics, 48 except in the case of slow metabolizers. 49
Our results highlight the involvement of several pharmacogenes in tamoxifen metabolism, according to a multigene mode of action, except for the formation of N-desmethyl 4-hydroxy tamoxifen from N-desmethyl tamoxifen, which is mediated only by CYP2D6.
Conclusion
Evidence concerning the contribution of other pharmacogenes besides CYP2D6 in estimating the concentration of active tamoxifen metabolites seems controversial and merits further study. This is because the effect of their genetic variation on tamoxifen and metabolite concentrations may be useful to patients if the relationship between endoxifen, or a combination of its active metabolites, and tamoxifen efficacy or treatment resistance is validated. 27
This is undoubtedly a source of hope, but it is also an avenue that needs to be further exploited, and not just for CYP2D6, to establish personalized medicine.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Informed Consent
Not applicable as this article does not directly deal with subjects.
Funding
The author received no financial support for the research, authorship and/or publication of this article.
