Abstract
With the advent of targeted therapy in non-small cell lung carcinoma (NSCLC), biomarker detection has become standard practice. US Food and Drug Administration (FDA) has approved a few companion diagnostics as single gene assays to find these driver mutations. We report a case of NSCLC where initial molecular tests to detect epidermal growth factor receptor (EGFR)-sensitizing mutations by an FDA-approved assay (Cobas® EGFR Mutation Test v2) failed to identify the common EGFR mutation L858R. However, the latter was picked up on subsequent next-generation sequencing (NGS). Interestingly an additional single nucleotide variant K860I was also observed in the cis configuration. The false-negative result for L858R was assumed to be caused by the abrogation of complementarity for the probe binding site on the amplicon generated. The concurrent occurrence of L858R and K860I will not be detectable by Cobas® EGFR Mutation Test v2 (FDA-approved companion diagnostic) despite its real-world intra and inter-laboratory studies demonstrating high accuracy.
Keywords
Introduction
The spectacular advances in treating non-small cell lung carcinoma (NSCLC) in the last decade have been fueled by rapid recognition of driver mutations and the development of cognate inhibitor(s). Lung cancer is the most common cause of cancer death globally, with an average of 1.7 million deaths yearly. 1 The selection of patients with an actionable biomarker and treatment with the corresponding inhibitor is the established standard of care that provides longer overall survival and lesser adverse effects. Epidermal Growth Factor Receptor (EGFR) mutations make up the bulk of druggable alterations amongst patients of NSCLCs, especially Asians. US Food and Drug Administration (FDA) has approved two polymerase chain reaction (PCR) based tests, namely, the therascreen EGFR RGQ PCR Kit (Qiagen Manchester, Ltd.) and the Cobas EGFR Mutation Test v2 (Roche Molecular Systems, Inc), along with several centralized and federated next-generation sequencing (NGS) based assays for tissue and plasma biopsy. 2
We report a histologically proven case of NSCLC, where one of the FDA-approved in vitro diagnostic (IVD) tests failed to identify EGFR exon 21 p.L858R (c.2537 T>G) mutation that could, however, be detected on another FDA-approved NGS based assay.
Case Description
All procedures performed in the study were approved by Institutional Review Board (IRB) (vide letter no: RGCIRC/IRB-BHE/19/2022) in accordance with the 1964 Helsinki declaration and its latest amendments. An informed consent was obtained from the subjects included in the study.
A 52-year-old male, a chronic smoker with a 30-year pack history, presented with complaints of hemoptysis and breathlessness. Routine laboratory investigations were within normal limits. Chest X-ray revealed a sizeable opaque mass in the lower lobe of the right lung. A whole-body computed tomography (CT) showed a consolidative mass lesion with spiculated margins and cavitation, measuring 4.0 × 2.2 cm, in the right lower lobe abutting the costal pleura adjacent to the right seventh rib with significant lymphadenopathy and bony involvement. A CT-guided biopsy sent for histopathological evaluation revealed a tumor in a vaguely arranged acinar pattern (Figure 1). The tumor expressed Thyroid Transcription Factor 1 (TTF-1) and was negative for p40. A diagnosis of NSCLC-favor adenocarcinoma was made.
The Microphotograph Shows a Tumor with a Vague Acinar Pattern and Significant Cytological Atypia. (A) The tumor cells are immunoreactive for TTF-1. (B) NGS read out depicts a doublet mutation in EGFR, L858R mutation (c.2573T>G, Green box), and K860I (c.2579A>T, blue box).
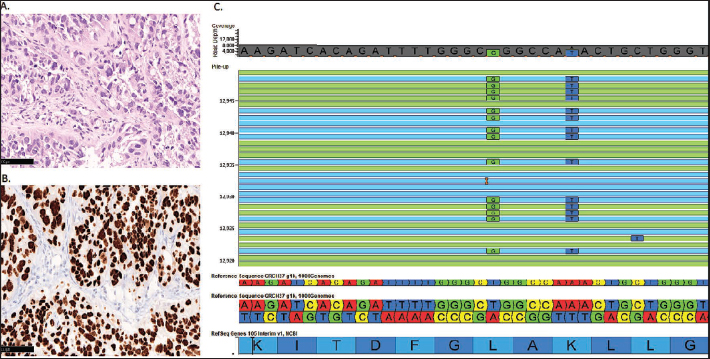
Biomarker testing returned negative for EGFR sensitizing mutation (Cobas EGFR Mutation Test v2 [Roche Molecular Systems, Inc]), ALKprotein expression by immunohistochemistry (Ventana D5F3 rabbit monoclonal antibody clone), and ROS1 protein overexpression by immunohistochemistry (D4D6, rabbit monoclonal antibody, Cell signaling Technology, Cambridge, UK) performed on Ventana benchmark XT immunostainer (Ventana Medical Systems, Tucson, AZ, USA).
The comprehensive molecular testing for additional actionable biomarkers was done with oncomine focus assay interrogating 52 significantly mutated genes for copy number variations, single nucleotide variants, and fusion rearrangements. The details of the panel can be found elsewhere. 3 Surprisingly, two EGFR exon 21 mutations, L858R and K860I, were identified in the cis configuration at a variant allele frequency of 42% (Tumor fraction: 70%), as seen below in the image from the golden helix genome browser (Figure 1). 4 The patient was started on osimertinib and is in partial response per RECIST 1.1 post six months of treatment.
Discussion
Although the most commonly used FDA-approved EGFR detection assays are capable of detecting p.L858R mutation (c.2573T>G/ c.2573_2574TG>GT, exon 21, NM_005228.5), the Cobas® EGFR Mutation Test v2 failed to detect it in the given case, despite its real-world intra and inter-laboratory studies demonstrating high accuracy. Following an absence of actionable genetic alteration in EGFR, ALK, and ROS1, comprehensive molecular testing was done employing NGS, which fortuitously revealed L858R-K860I doublet mutation. The co-occurrence of L858R-K860I has been reported previously in the literature with a frequency of 0%–4%. 5 Curiously, EGFR exon 21 p.K860I (c.2579A>T, NM_005228.5) has never been observed to occur alone. The therascreen EGFR RGQ PCR Kit (Manufacturer) and Cobas EGFR mutation test v2 are not designed to identify EGFR exon 21 p.K860I mutation. Besides, the proximity of the single nucleotide variant (c.2579A>T, NM_005228.5) to the EGFR exon21 p.L858R (c.2573T>G/c.2573_2574TG>GT, NM_005228.5) abolishes the probe binding site for the latter, leading to a false-negative result for the second most common EGFR sensitizing mutation L858R.
Onozawa et al. 5 also described a case with a false-negative result for EGFR exon21 p.L858R mutation, which was further evaluated at National Cancer Centre Japan (NCC) on a panel revealing a cis L858R-K860I doublet in a smoker patient similar to our case. EGFR mutations are often observed in never-smokers; the cis L858R-K860I doublet mutation in EGFR is described in smokers and non-smokers in the literature; however, due to the rarity of the mutation, the association with smoking history is not well established. Also, the patients harboring the EGFR exon21, p.L858R-K860I doublet mutation are noted to respond with tyrosine kinase inhibitors (TKIs) similarly to EGFR exon21 p.L858R. 5 This makes a strong case not to miss this doublet and more so because the same has been seen in as many as 4% of patients harboring EGFR mutation(s).
The present case showed no EGFR mutation in the FDA-approved companion diagnostic RealTime-PCR-based single gene assay. However, a combination of two sensitizing modifications was identified in NGS. We contend that NGS-based analysis will identify additional actionable biomarkers and the additional uncommon EGFR sensitizing mutations not detectable in the approved companion diagnostic assays, as seen in the present case.
Author contributions
Conceptualization: AM. Data Curation: HD, SM. Formal Analysis: AM, HD, SN. Investigation: AM, HD, SN, DK, MKP. Methodology: AM. Project Administration: AM. Resources: SM, DK, MPK. Supervision: AM. Visualization: HD, SN. Writing – original draft: HD. Writing – review & editing: AM
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
