Abstract
Introduction:
Rheumatoid arthritis is associated with various cancers. Many studies have investigated physical exercise interventions as health improvements to ameliorate the risk of cancer during rheumatoid arthritis diagnosis. Recently, microRNAs were used as biomarkers for health assessment and cancer prediction in rheumatoid arthritis patients.
Methods:
The effects of exercise interventions on serum microRNAs were investigated in pristane-induced arthritis (PIA) rat models. Twelve Sprague-Dawley male rats were divided into 4 groups including non-exercise without PIA (N-EX), non-exercise with PIA (N-EX + PIA), exercise without PIA (EX) and exercise with PIA (EX + PIA). Blood samples were collected at the end of the study period to analyze miRNA biomarkers and target cancer gene predictions.
Results:
Four significant Rattus norvegicus (rno-microRNAs) may purpose as tumor suppressors were identified as potential target cancer gene candidate expressions within the 4 comparative interventional exercise groups. One rno-microRNA and target cancer gene candidate was up-regulated and 3 rno-microRNAs and their target cancer genes were down-regulated.
Conclusions:
Exercise interventions affected rno-miRNAs regulated target cancer gene candidates ITPR3, SOCS6, ITGA6, and NKX2-1 as biomarkers for cancer prognosis in rheumatoid arthritis diagnosis.
Introduction
Rheumatoid arthritis (RA) is classified as an autoimmune inflammatory condition characterized by pain, swelling, and inflammation of the joints, along with stiffness which can reduce function and impair the overall quality of life. 1 Following the diagnosis of RA, it has been shown that cytokines including IL-6 can affect the prognosis of breast cancer along with inflammation and further comorbidities.1,2
A total of 138 cases of lung and prostate solid tumors were recorded within 12 months of RA diagnosis.
3
Furthermore, those patients diagnosed with RA experienced cancer of greater severity than was the case for patients who did not have RA, with the exception of gastric cancer, where RA patients actually had less severe cases than otherwise healthy people.
4
RA patients were also found to be more likely to be diagnosed with lung cancer or lymphoma than other members of the population.
5
In contrast, the incidence of colorectal and breast cancers was shown to be lower in RA patients in comparison to the wider population.
5
It was found that RA patients were 34% more likely to experience regional or distant cancer than was the case for non-RA patients (
Earlier research has examined the incidence of RA and cancer through the use of microRNA biomarkers in studies investigating whether exercise can be used effectively to treat disease.7,8 It has been argued that miRNA biomarkers might be usefully employed for prognostic evaluations of small lung cancer cachexia as well as the metabolism in cancer cachexia and inflammation in the tissue. 7 Within the context of exercise science, it is believed that exercise can serve as a tool to improve health and support rehabilitation. 9 One study applied a novel exercise science epigenetic technique in which small non-coding microRNAs with a length of around 20 nucleotides served as biomarkers for the purpose of evaluating improvements in the health of RA patients. 10 Another research study investigating the effects of exercise in RA patients revealed no significant differences when the expressions of miRNA-155 and miRNA-16 were compared between pre- and post-exercise groups and the control. That study also found that miRNA-146a differed significantly from the control group. 8
It is possible for the target gene silencing mechanisms which facilitate mRNA degradation or translation inhibition to be determined by micro-RNAs.11,12 Meanwhile, the alternatives of the miRNA-126 target EGFL 7 have been linked to cancer due to the promotion of inflammation and angiogenesis in RA patients.12,13 Other researchers have reported that cancer could be induced by tumor necrosis factor alpha (TNFα) via the stimulation of miRNA-155, while miRNA-155 expression played a role in the regulation of IL-6 in the synovial fluid of RA patients through the stimulation of neutrophils, macrophages, and T-cells which can lead to the formation of osteoclasts as well as chronic inflammatory effects.14-16
Exercise may represent a novel means of mitigating the suffering of RA and cancer patients.17,18 A number of studies have sought to examine the application of exercise as a means of inhibiting tumorigenesis, limiting infections, and building up resistance to tumors. 19 Furthermore, the symptoms of some cancers, especially breast cancer, can be reduced through an exercise routine of moderate intensity conducted daily for up to an hour. 17 More recently, it has been found that exercise can support the effects of numerous miRNAs which play a role in the suppression of tumors. 20 In the case of bladder, ovarian, colorectal, prostate, breast, and gastric cancers, the circulation of alternative c-miRNA-133 underwent expression in solid tumors, and served to suppress those tumors in patients who maintained regular exercise. 21 Other research findings have indicated that the expression of miR-206 was positively affected by exercise and could limit cell growth in breast cancer through the inhibition of the G1/S transition, thereby achieving the suppression of cell proliferation. 22 The expression of miRNA 21, miRNA Let-7a, and miRNA16 was also enhanced by exercise, while the expression of miRNA15b was inhibited. The effects upon miRNAs duly influenced the relationship with pro-apoptotic target genes whereby a reduction in PTEN, PDCD4, RAS mRNA and anti-apoptotic resulted in an increase in Bcl-2 mRNA when tests were conducted in rats. 23 This research involving rat models examined the influence of exercise in early stage RA as a predictor of cancer in the context of miRNA expression associated with target cancer gene candidates.
Methods
Animals
Twelve 11-week-old Sprague-Dawley male rats were housed at Chulalongkorn University Laboratory Animal Center (CULAC). The rats were assigned to 4 cages with 3 animals per cage and injected with 100 µl of pristane-induced arthritis (PIA)24,25 at the dorsal side of the tail as the control group with PIA and the exercise group with PIA. The environment was maintained under a 12/12 light-dark cycle at 22°C with ad libitum food and water.
Exercise intervention
After 2 weeks of quarantine and another week of treadmill training preparation, the animals were divided into 4 groups (3 rats per group). The first group was assigned as non-exercise without PIA induction (N-EX). The second group was assigned as non-exercise with PIA induction (N-EX + PIA). The third group received high-speed treadmill exercise (20-25 m/min) for 60 minutes without PIA induction (EX), while the fourth group received high-speed treadmill exercise (20-25 m/min) for 60 minutes with PIA induction (EX + PIA). The animals were weighed every 2 days (Figure 1). Animals in the exercise groups were allowed to rest for 15 minutes when they showed signs of fatigue (stay at the end of the treadmill or be unable to run with sound or electrical stimulation). On day 45, all rats were euthanized by inhalation of CO2.

Allocation of the 4 sample groups into 4 exercise conditions: (a) exercise without PIA induction group (EX), (b) exercise with PIA induction group (EX + PIA), (c) non-exercise without PIA induction group (N-EX), (d) non-exercise with PIA induction group (N-EX + PIA), and (e) blood sample collection .
Sample collection
EDTA-blood samples were collected from the external jugular vein (Figure 1). Plasma was separated by centrifugation (8 minutes), aliquoted and stored at −80°C until assay. Total RNA was extracted using TRIzol Reagent and miRNeasy Mini Kit (Qiagen), and then quantified and qualified by Agilent 2100/2200 Bioanalyzer (Agilent Technologies, Palo Alto, CA, USA), NanoDrop (Thermo Fisher Scientific Inc.) and 1% agarose gel electrophoresis. The RNA samples were dried in a GenTegra tube under vacuum.
Small RNA library preparation
The 3’SR adaptor for Illumina was ligated to the small RNA using the 3′ligation enzyme. To prevent adaptor-dimer formation, the excess 3′SR adaptor was a hybrid with the SR RT primer. The 5′SR adaptor for Illumina was ligated to the small RNA using the 5′ligation enzyme and first-strand cDNA was synthesized using ProtoScript II reverse transcriptase. Each sample was then amplified by PCR using P5 and P7 primers, with both primers carrying sequences that annealed with flow cells to perform bridge PCR and P7 primer carrying a 6-base index allowing for multiplexing. PCR products of ~150 bp were recovered and cleaned using polyacrylamide gel electrophoresis (PAGE). The final DNA library was validated using an Agilent 2100/2200 Bioanalyzer (Agilent Technologies Palo Alto, CA, USA) and quantified by a Qubit 3.0 Fluorometer (Invitrogen, Carlsbad, CA, USA).
Small RNA sequencing
Libraries with different indices were multiplexed and loaded on an HiSeq (Illumina, San Diego, CA, USA). Sequencing was carried out using a 2 × 150 paired-end (PE) configuration and image analysis and base calling were conducted by HiSeq Control Software (HCS) + OLB + GAPipeline-1.6 (Illumina) using HiSeq instrument image analysis.
Data analysis
The FASTQ sequences were examined for quality control using FastQC software (v0.10.1). The raw sequencing reads were adaptor trimmed and quality filtered (<
Data interpretation
All rno-miRNAs and target cancer gene results were interpreted using our computerized analysis, with target cancer gene functions reviewed from genecards.org, nature.com, cancerindex.org, http://ncbi.gov and genome.jp/keg. 31
Results
Mi-RNA expressions
The volcano plots shown in Figure 2 represent differential expressions of rno-miRNA in a rat model. The 213 rno-miRNAs were significant, with 51 known rno-miRNAs and 162 novel rno-RNAs. The 67 rno-miRNAs showed up-regulation, while 145 rno-miRNAs showed down-regulation. Twenty significant rno-miRNAs were up-regulated (log2) and 31 rno-miRNAs were down-regulated (−log2). All known rno-miRNAs are presented in Table 1.

(a) EX versus EX + PIA. A total of 82 rno-miRNAs had significant expressions in the EX group compared to the EX + PIA group. Twenty rno-miRNAs showed up-regulation (red) log2 and 64 rno-miRNAs showed down-regulation (blue) −log2. (b) N-EX + PIA versus EX + PIA. A total of 46 rno-miRNAs had significant expression in the N-EX + PIA group compared to the EX + PIA group. Ten rno-miRNAs showed up-regulation (red) log 2 and 39 rno-miRNAs showed down-regulation (blue) −log2. (c) N-EX versus EX. A total of 66 rno-miRNAs had significant expression in the N-EX group compared to the EX group. Thirty-five rno-miRNAs showed up-regulation (red) log 2 and 31 rno-miRNAs showed down-regulation (blue) −log2. (d) N-EX versus N-EX + PIA. A total of 16 rno-miRNAs had significant expression in the N-EX group compared to the N-EX + PIA group. Three rno-miRNAs showed up-regulation (red) log2 and 13 rno-miRNAs showed down-regulation (blue) −log2.
Comparative exercise intervention groups for known rno-miRNA sequencing: Exercise versus Exercise + PIA.

The table shows the regulation of 26 rno-miRNAs in the Exercise group compared to the Exercise + PIA group as 10 up-regulation (log2) and 16 down-regulation (−log 2).
Non-exercise + PIA versus Exercise + PIA.

The table shows the regulation of 12 rno-miRNAs in the Non-exercise + PIA group compared to the Exercise + PIA group as 4 up-regulation (log2) and 8 down-regulation (−log 2).
Non-exercise versus Exercise.

The table shows the regulation of 10 rno-miRNAs in the Non-exercise group compared to the Exercise group as 4 up-regulation (log2) and 6 down-regulation (−log 2).
Non-exercise versus Non-exercise + PIA.
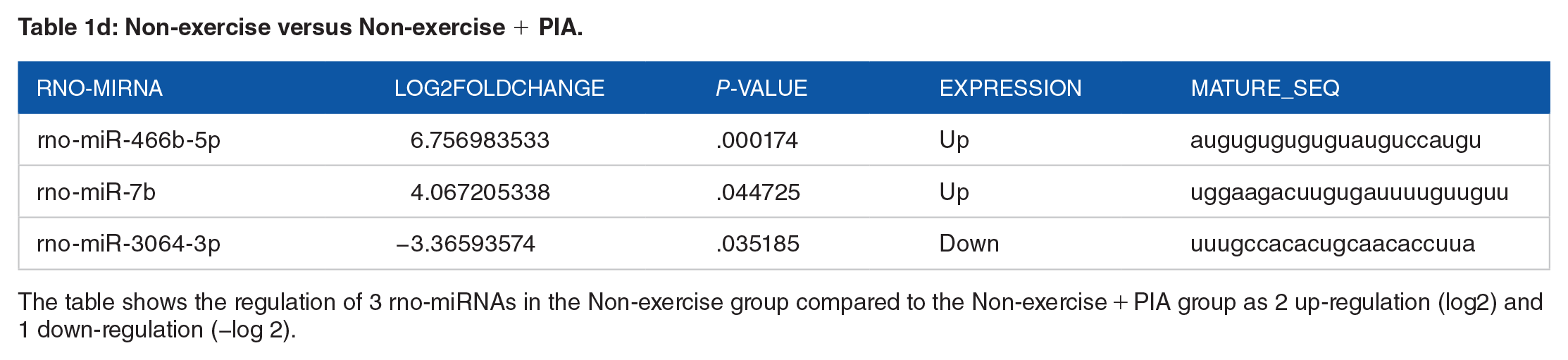
The table shows the regulation of 3 rno-miRNAs in the Non-exercise group compared to the Non-exercise + PIA group as 2 up-regulation (log2) and 1 down-regulation (−log 2).
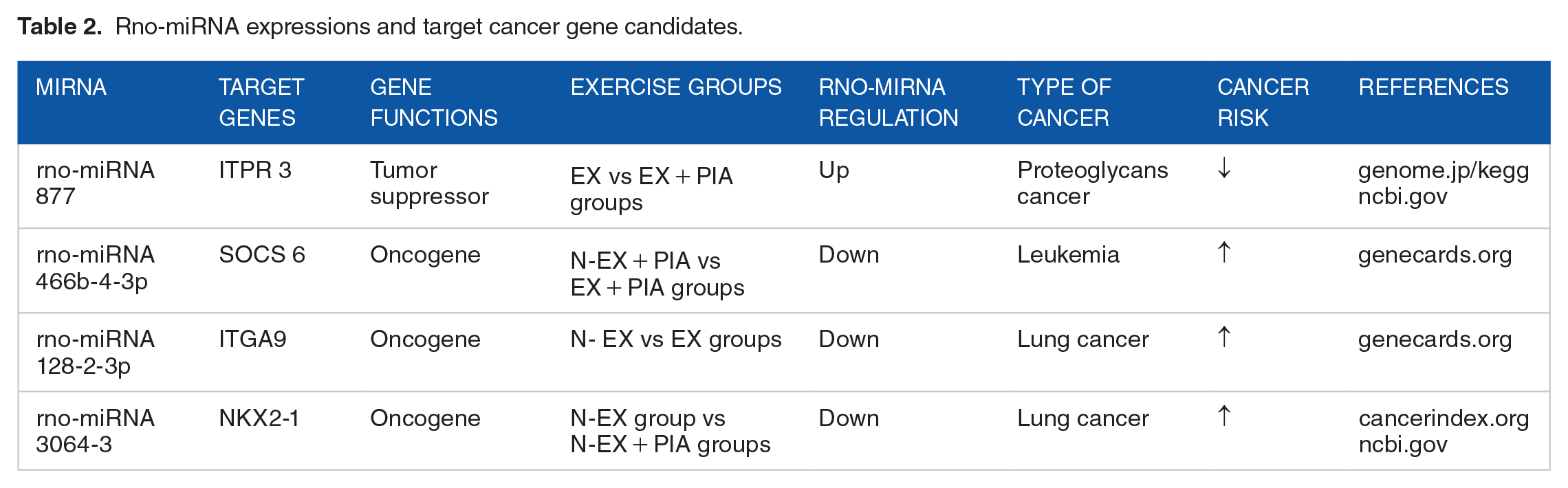
The 4 rno-miRNAs target cancer gene candidates in the comparative interventional exercise groups are listed in Table 2, while the same 4 specific rno-miRNAs with varied other target genes from our computational predictions are shown in Figure 3.
Rno-miRNA expressions and target cancer gene candidates.

Varied target genes and cancer gene candidates of the 4 rno-miRNAs. (a) EX versus EX + PIA groups. All the target genes of rno-miR-877 (red) and target cancer genes associated with proteoglycans in cancer metastasis (blue). (b) N-EX + PIA versus EX + PIA groups. All the target genes of rno-miR-466b-4-3p (red) and target cancer genes associated with leukemia (blue). (c) N- EX versus EX groups. All the target genes of rno-miR-128-2-3p (red) and target cancer genes associated with lung cancer (blue). (d) N-EX versus N-EX + PIA groups. All the target genes of rno-miR-3064-3p (red) and target cancer genes associated with lung cancer (blue).
The computation results examined all significant rno-miRNAs from each comparative interventional exercise group and selected significant rno-miRNAs from each comparative group that had only the cancer gene target candidates. We identified gene functions by looking at single gene target functions from all study group results by following references (1) genome.jp/keg.ncbi.gov, (2) genecards.org, and (3) cancerindex.org.ncbi.gov. Once the cancer gene candidates were identified, we chose those genes and their own target rno-miRNAs as our results.
Table 2 shows that the interventional physical exercise groups reduced the risk of cancer. Group 3 (EX) showed the lowest risk of cancer compared to the other study groups. Group 4 (EX + PIA) showed lower risk of leukemia than Group 2 (N-EX + PIA) and lower risk of lung cancer compared to Group 1 (N-EX).
Discussion
Fifty-one rno-miRNAs were expressed with varied target genes. Twenty rno-miRNAs were up-regulated and 31 were down-regulated. Four significant rno-miRNAs targeted 4 cancer gene candidates in the interventional exercise groups. Interestingly, our results also found that expressions of significant rno-miRNAs were different from the hypothesis in 3 out of the 4 interventional exercise groups.
Our results indicated that rno-miRNA-877 up-regulations in the EX group target ITPR 3 expressions associated with proteoglycans in cancer compared to the EX + PIA group. The up-regulation of rno-miRNA 877 target ITPR 3 in the EX group is associated with proteoglycans in cancer. Results from our computation predictions data and KEGG.jp show that rno-miRNA 877 regulated target specific genes involved in the activation of Teff immune cells and other immune systems. 32 Moreover, rno-miRNA 877 is an activator of CD 28 and also a signal of the IL 2 related proliferator, especially for T-cells. 33 Expression of rno-miRNA 877 and CD 28 also play a role in controlling Treg cell mediation (multi T-cell form) to suppress other antigens including pre-cancer cells from carcinoma and prevent autoimmunity. 34 Without PIA induction, the T-cell and Fas ligand normally control the immune system and tumor necrosis factor (TNF) but with PIA condition, down-regulation of rno-miRNA 877 cannot normally regulate the immune system such as CD 28, T-cells and Fas ligand. Cancer then occurs when cancer genes are expressed after chronic rheumatoid arthritis. 32 Furthermore, some studies suggested that proteoglycans in protein can induce signal transduction via toll-like receptors, interleukins, interferons and chemokines. Exercise may stimulate signaling of proteoglycans to cytokines, and increase the immunity to suppress cancer cells. For example, decorin, as a type of proteoglycans in protein can improve signaling by suppressing cytokine and transforming growth factor (TGF)-ß to impact tumor cell death if exercise occurs. 16
Rno-miRNA 466b-4-3p that target SOCS 6 expressions associated with leukemia were down-regulated in the N-EX + PIA group compared to the EX + PIA group. Down-regulation in miRNA 466b-4-3p and expression of SOCS 6 related to inside cellular environment change, caused different expression levels in both miRNA and SOCS 6 within the 2 exercise groups. Moreover, the expression of the SOCS family, including SOCS 6, was also altered in rheumatoid arthritis patients and resulted in a change in the cellular status to induce leukemia. 35 Our study did not measure the SOCS 6 expression levels but focused on the possibility of SOCS 6 mechanisms relating to leukemia with exercise. The SOCS 6 regulates the tyrosine kinase receptor associated with immunity, inflammation and leukemia with various mechanisms. 36 Some researchers found that the Flt3 tyrosine kinase receptor was regulated by SOCS 6 mutation that transformed to acute myeloid leukemia (AML), while others suggested that Flt3 regulated SOCS 6. 37 However, both regulation and mechanism results were inconclusive.
Expressions of SOCSs and SOCS 6 associated with rheumatoid arthritis involve several cytokines such as IL-6, IL-10, IL-12 and IL-15 that interfere with Janus kinases (JAKs) signal transducers and activator of transcription (STAT). 35 In chronic rheumatoid arthritis inflammation, SOCSs can alter the expression and regulate JAK/STAT as mutation induces leukemia. 38 By contrast, with exercise JAK/STAT phosphorylation may increase, while satellite cell proliferation differentiated with transcription of the activating cell cycle. 39 Thus, the EX + PIA group could have less SOCS 6 expression than the N-EX + PIA group. Exercise conditions could induce a healthy cell differentiation cycle, influence the cell microenvironment and suppress the development of leukemia. However, limited information exists about how SOCS 6 alters the expression of leukemia prognosis in rheumatoid arthritis under exercise.
Down-regulation of rno-miRNA 128-2-3p in the N-EX group is associated with target lung cancer cell ITGA 9 expressions (interacts with beta 1 subunit generating α9β1) compared to the EX group. The expression of ITGA 9 occurs in many types of carcinoma cancer tumors. 40 In small cell lung cancer, ITGA 9 can be altered on CpG island by hypermethylation or hypomethylation when lifestyle changes. 40 ITGA 9 is also associated with many types of mechanisms and pathways. Exercise could affect ITGA 9 expression or down-regulation in some miRNAs. Our results in the N-EX group showed greater expression of ITGA 9 than the EX group, possibly related to lung extracellular-matrix cell communications that interacted as critically important in various cell functions. 41 Under exercise, lung functions and cells are changed by body temperature, airway diameter, lung connective tissue, and smooth muscle activation. These factors changed the lung cell environment and may be associated with immunoreactivity in the α 9 subunit in epithelial lung tissue. 42 Our study results indicated that the N-EX group might have more ITGA9 expressions than the EX group but results were unclear. Further investigations are required to confirm these assumptions.
In the final study group, down-regulation of rno-miRNAs 3064-3 in the N-EX group target NKX2-1 expressions compared to the N-EX + PIA group. The NKX2-1 gene encodes homeobox protein or NKX2-1 protein specific to the brain, lungs and thyroid gland, which is involved in the regulation of morphogenesis during early embryonic development. 43 The NKX2-1 gene also encodes the thyroid transcription factor 1 (TTF-1) which explicitly regulates the thyroid gland, lungs, and diencephalon (a part of the brain) as it is a tissue-specific nuclear transcription factor. The protein binds to the thyroglobulin promoter and regulates the expression of thyroid-specific genes. The thyroid gland helps to produce and regulate hormones, particularly hormones involved in brain growth and development. This protein can be used as a marker to determine lung and thyroid gland tumors. The NKX2-1 was induced by activin A via SMAD2 signaling in a human embryonic stem cell differentiation model. 44 Gene mutations and deletions are associated with many genetic diseases, such as benign hereditary chorea, choreoathetosis, congenital hypothyroidism and neonatal respiratory distress (also known as a brain-lung-thyroid syndrome), and probably also associated with thyroid cancer and lung cancer.
NKX2-1, a key molecule in lung development, is highly expressed in non-small cell lung cancer (NSCLC). Two microRNAs (miRNAs) were associated with NKX2-1 as miR-365 which targets NKX2-1 and miR-33a that targets downstream of NKX2-1. 45 Our results showed that the N-EX group down-regulated rno-miRNA3064-3 compared to the N-EX + PIA group. The rno-miRNA3064-3 targeted the NKX2-1 gene, causing up-regulation of NKX2-1 protein in the N-EX group and enhancing the risk of lung cancer. In N-EX group, NKX2-1 gene induces many gene mechanisms especially on the target genes. Normally, NKX2-1 plays roles in single nucleotide polymorphism (SNP), tumor suppressor or tumor promotors which can bind to varied locus on each gene. 46 NKX2-1can influence some anti-tumorigenic genes to make them become mutation and become lung cancer without lifestyle change. 47 Binding of NKX2-1 to varied gene locus are usually in downstream, proximal promotor (upstream) and intron which also induces lung adenocarcinomas. NKX2-1 can influence EGFR gene become mutation and stimulate lung tumorigenesis. On the other hand, NKX2-1 gene can also suppress KRAS gene to inhibit lung tumorigenesis. 48 In the N-EX group, it could be possible that NKX2-1 gene can play automatically roles to be cancer suppresser to inhibit other carcinoma genes. Thus, expression of NKX2-1 gene is dependent on many factors like gene locus, gene types and base pairs etc. 46
Furthermore, the N-EX + PIA group was associated with inflammation and body immune system as a possible of PIA body introduction. Stimulation of PIA activated T helper cells (Th17) to regulate the immune system as hemostasis could suppress NKX2-1 gene expression and prevent lung carcinoma. 49 Moreover, PIA injection induced CD4+T cells to play a role as a host immune response to cancer and chronic inflammation. 50 PIA also induced CD4+T cells to develop monocytes/macrophages in response to the pathogens by enhancing CD8+T interaction with B cells. 51
The CD4+ T cells also modulate immune homeostasis to suppress pro-inflammation and prevent lung cancer formation by suppressing NKX2-1 expressions. 51 However, how the NKX2-1 genes and rno-miRNA 3064-3 relate to exercise requires further investigation.
The strengths of this study were the preliminary integration of candidate target cancer genes and exercise intervention to ameliorate the onset of chronic rheumatoid arthritis. We used our research results in varied rno-miRNAs expressions, together with the possibility of target cancer gene candidates as the mechanism for a prospective study.
Conclusions
Our results indicated that tumor suppresser rno-miRNAs in exercise activity up-regulated target cancer gene expression compared to non-exercise activity in both rheumatoid arthritis and non-rheumatoid arthritis conditions. Due to funding restrictions, our study only investigated the rno-miRNA expressions and used computerized predictions on target cancer gene candidates. Further studies could include (1) measurement of gene expression levels, (2) measurement levels of joint inflammations, and (3) analysis of cancer cell proliferation. Results offer a new modification for epigenetics and exercise science fields to investigate cancer prevention among rheumatoid arthritis patients.
Footnotes
Acknowledgements
The authors wish to acknowledge Chulalongkorn University Laboratory Animal Center (CULAC) and thank the laboratory assistants for helping with rodent care during this research.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: My personal thanks to TINARATHPATRA Co., Ltd. for funding (46/158S1) this project.
Author Contributions
VT, SP, WS, and TP organized the research ideas. PC was responsible for rno-miRNA isolation, preparation and sequencing. VS, VT, WS, and SP coordinated the data analysis and statistical tests, including writing up the results. Under the writing obligation, VT was also responsible for the manuscript with consultation from SP, VS, TP, WS, and PC. All authors have read and approved the final version of this manuscript.
Ethical Approval
This project was approved by the Chulalongkorn University Animal Care and Use Committee (Approval protocol number 2073010).
