Abstract
Introduction:
Susceptibility factors for coronavirus disease 2019 (COVID-19) include sex and medical conditions such as asthma and rhinitis. DNA methylation (DNAm) is associated with asthma, rhinitis, and several viruses. We examined associations of asthma/rhinitis with DNAm at CpGs located on coronavirus related genes, and if these associations were sex-specific.
Methods:
In total, n = 242 subjects aged 26 years from the Isle of Wight Birth Cohort were included in the study. Linear regressions were used to examine sex specific and non-specific associations of DNAm at CpGs on coronavirus related genes with asthma/rhinitis status. Associations of DNAm with gene expression in blood were assessed for functional relevance of identified CpGs.
Results:
Statistically significant interaction effects of asthma or rhinitis with sex were identified at 40 CpGs for asthma and 27 CpGs for rhinitis. At 21 CpGs, DNAm was associated with asthma, and at 45 CpGs with rhinitis, regardless of sex. Assessment of functional relevance of the identified CpGs indicated a potential of epigenetic regulatory functionality on gene activity at 14 CpGs for asthma and 17 CpGs for rhinitis, and of those 6 CpGs for asthma and 7 CpGs for rhinitis were likely to be sex-specific.
Conclusion:
Subjects with asthma/rhinitis may have altered susceptibility to COVID-19 due to changes in their DNAm associated with these conditions. Sex specificity on association of asthma/rhinitis with DNAm at certain CpGs, and on the association of DNAm at asthma/rhinitis-linked CpGs with gene expression have the potential to explain the reported sex-specificity in COVID-19 morbidity and mortality.
Introduction
The novel coronavirus disease 2019 (COVID-19) pandemic, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) continues to spread across the world. As of September 8th, 2020, the pandemic has reached 27 236 916 confirmed cases and 891 031 deaths globally. 1 Among those infected, biological sex was found to be an important determinant of COVID-19 severity and mortality. 2 Males with COVID-19 are at higher risk for worse outcomes and mortality compared to females, independent of age. 3 Existing comorbidities and medical conditions of the respiratory tract are risk factors for COVID-19 morbidity and mortality.4-7 Asthma is a common chronic condition and rhinitis is a common childhood allergic condition that continues in adulthood, 8 but their role in COVID-19 infections remains unclear. However, recently findings have shown potential associations of pre-existing asthma and rhinitis with SARS-CoV-2 susceptibility and COVID-19 prognosis.9-12
Centers for Disease Control and Prevention (CDC) states that chronic lung diseases such as moderate to severe asthma increases the likelihood of COVID-19 severity. 13 Severe asthma defined as asthma with recent use of an oral corticosteroid has been shown to be associated with higher risk of COVID-19 and its related deaths. 14 A case study of a 3-year-old child has been reported with history of allergic rhinitis suffered from COVID-19. 15 However, some studies report low to no prevalence of asthma in COVID-19 patients.16,17 Underlying reasons for the discrepancy was not fully understood, but 1 possible explanation for asthma being protective against COVID-19 is that for asthmatic subjects on inhaled corticosteroids, such treatment can potentially protect them against COVID-19.
An epigenetic mechanism, DNA methylation (DNAm) has been shown to be associated with asthma and rhinitis in several studies.18-22 Previous studies have shown association between DNAm and viruses such as human papillomavirus (HPV), Epstein-Barr virus (EBV), Human rhinovirus.23-25 These viruses modulate epigenetic dysregulation of immune-related gene expression in host cells leading to immune suppression. 26
In this study, we examined whether asthma and rhinitis status were associated with DNAm at CpGs located on coronavirus related genes, and whether such associations were sex specific. The identified CpGs have a potential to be further assessed on their association with the risk of COVID-19 and may contribute to a better understanding on the impact of underlying asthma or rhinitis on COVID-19 susceptibility and severity.
Methods
Study population
Isle of Wight whole population birth cohort (IOWBC, n = 1456) was established in 1989/1990 on the Isle of Wight (UK) to study the natural history of allergic diseases. 27 Informed consent was provided by the parents at birth and at follow-up assessments at ages 1, 2, 4, 10 years, and by participants at 18, and 26 years of age.
Clinical and demographic variables
Status of asthma and rhinitis at age 26 years were included in the study. Asthma at 26 years was defined using ISAAC questionnaire as “ever had asthma” and “wheezing or whistling in the chest in the last 12 months” and/or “current treatment for asthma.” Rhinitis at 26 years was defined as “In the past 12 months have you had a problem with sneezing or a runny or a blocked nose when you did not have a cold or the flu?.” 28 Sex information was based on the questionnaire data and hospital records collected at the time of birth. Active smoking at 26 years was recorded as “yes” if the subject was a current smoker. Body mass index (BMI) was calculated using weight in kilograms divided by height in meters-squared at 26 years. Socio-economic status (SES) was defined based on household income, number of rooms, and maternal education.
Coronavirus related genes and CpGs
Using keywords “Coronavirus” and “Coronavirus silent sweep” in GeneCards (www.genecards.org/), 29 genes potentially related to the pathogenesis of SARS-CoV-2 were extracted. A score for each gene (number of times shown to be associated with coronavirus in previous studies) was used to select genes for this study. A scree plot of the scores in descending order was applied to select genes; the minimum score for a gene to be included in our study is such that its score is at a turning point with a large decrease followed by a flat pattern of scores (Rathod, Rathod and Rahimabad, 2021). 30 Information regarding the coronavirus associated CpGs on these selected genes, chromosomes, the location relative to CpG islands, and location on genes were identified using the Illumina’s manifestation file.
DNA methylation (DNAm)
DNA was extracted from peripheral whole blood samples using the standard salting out procedure at age 26 years. 31 DNA concentration was determined by PicoGreen dsDNA quantitation (Molecular Probes, INC. OR, USA) or Qubit (Thermofisher, MA, USA). One microgram of DNA was bisulfite-treated for cytosine to thymine conversion using the EZ 96-DNA Methylation Kit (Zymo Research, Irvine, CA, USA) for each sample. The methylation level for each CpG was assessed using the Infinium MethylationEPIC BeadChips (Illumina, Inc., CA, USA). Arrays were processed using a standard protocol 32 with multiple identical control samples assigned to each bisulfite conversion batch to assess assay variability. Using Illumina BeadStation, the beadchips were scanned and the methylation level (beta value) was calculated for each queried CpG locus using the Methylation module of BeadStudio software.
Intensities of DNAm were quantile normalized using the minfi package in R. 33 The intensities were then converted to β values, defined as a ratio of methylated (M) over the sum of methylated and unmethylated (U) probes (β = M/[c + M + U]), where c is a constant to prevent zero in the denominator. Batch effects correction using the ComBat R function was applied to the β values. β values close to 0 or 1 tend to suffer from severe heteroscedasticity. Therefore, base-2 logit transformed β values (denoted as M-values) were used in the analysis. 34 In addition, CpG sites were excluded from the analyses if DNAm was missing for at least 1 subject due to low quality or missing at random, or if the minor allele frequency of a probe SNP at that site is >7% (ie, ~⩾10 out of 1456 subjects expected to have the minor allele in the cohort) and the probe SNP was within 10 base pairs of the targeted CpG site.
As whole blood is a mixture of various cell types, 35 there is a need to adjust for cell composition to remove the potential confounding effects. 36 Cell type proportions were estimated using the minfi package.37,38 The estimated cell type proportions of CD4+ T cells, natural killer cells, neutrophil, B cells, monocytes, and eosinophil cells were included in the analyses as adjusting factors.
Gene expression
Gene expression levels at 26 years from peripheral blood samples of subjects in IOWBC were measured using paired-end (2 × 75 bp) RNA sequencing with the Illumina Tru-Seq Stranded mRNA Library Preparation Kit with IDT for Illumina Unique Dual Index (UDI) barcode primers following manufacturer’s recommendations. All samples were sequenced twice using the identical protocol and for each sample the output from both runs were combined. FASTQC were run to assess the quality of the FASTQ files (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Reads were mapped against Human Genome (GRch37 version 75) using HISAT2 (v2.1.0) aligner. 39 The alignment files, produced in the Sequence Alignment Map (SAM) format, were converted into the Binary Alignment Map (BAM) format using SAMtools (v1.3.1). 40 HTseq (v0.11.1) was used to count the number of reads mapped to each gene in the same reference genome used for alignment. 41 Normalized read count FPKM (Fragments Per Kilobase of transcript per Million mapped reads) were calculated using the countToFPKM package (https://github.com/AAlhendi1707/countToFPKM) and their log transformed values were used for data analysis.
Statistical analysis
Linear regressions were applied to evaluate the association of DNAm at each CpG site on the coronavirus-related genes (dependent variable) with asthma status (asthma-free as reference) at age 26 years. The same analytical model was applied to the status of rhinitis (rhinitis-free as reference). As previous studies have shown sex differences in asthma prevalence42-44 as well as COVID-19 severity and mortality,3,45-47 interaction effects of asthma/rhinitis, and sex on DNAm were assessed. Subjects with asthma free and males are the reference groups.
Association between DNAm and gene expression
To assess the biological relevance of the identified CpGs showing an association between DNAm and asthma or rhinitis status, we further assessed their association with gene expression. Linear regressions were applied to evaluate the association of DNAm levels with gene expression for the CpG mapped genes as well as the nearby genes on the same chromosome. When selecting nearby genes, we considered 500 000 base pairs (bps) (250 000 bps up stream and 250 000 bps downstream of a CpG site) following suggestion in a recent study. 18 Gene expression was used as dependent variable, DNAm and sex (male = reference group) as independent variables in the model. Additionally, to assess sex-specificity of DNAm and expression association, an interaction term of DNAm × sex was also included in the model as we previously found this association to be different in both sexes (Patel, Solatikia and Zhang, 2021). 52 Statistical significance was considered at α = .05 level. All the analyses were performed using SAS 9.4.
Sensitivity analysis
Of the CpGs that were found to be associated with asthma and rhinitis (interaction and main effects, they were further assessed using the same analytical model as above with covariates BMI, active smoking, and SES at age 26 years included in the model. We would examine whether the interaction effects of asthma or rhinitis with sex were changed in terms of direction of associations after including these covariates in the model.
Results
Following the elbow method of a scree plot on gene scores of >2.8, 66 genes were selected shown to be associated with coronavirus in the literature. Additionally, 14 genes shown to be associated with coronavirus silent sweep infection were also included. Thus, in total 80 genes were selected for the purpose of this study (Supplemental Table 1). Of the 80 genes, we had data for 78 genes on autosomes. In total, 1555 CpGs mapped to 78 genes were extracted from Illumina manifestation file. Of these, 4 CpGs with “ch” prefix were excluded. After preprocessing and exclusion of CpG sites with missingness, low quality, or close to probe SNPs, DNAm at 677 CpGs were available at 26 years in IOWBC (n = 242). Among these 242 subjects, 41 (females = 26, 17.3%) had asthma and the remaining without asthma (females = 124, 82.7%), and 120 (females = 78, 52%) had rhinitis and others without rhinitis (females = 72, 48%). We also examined the difference in asthma prevalence between males and females (Supplemental Tables 4(A) and 4(B)). In the complete cohort, the difference in prevalence between males and females was marginally significant (P-value = .08) for asthma, and was statistically significant for rhinitis (P-value < .001). For both health conditions, prevalence was higher in females. For the subsamples included in the current study, the prevalence in females was also higher, but the difference between the 2 sexes was not statistically significant.
We assessed the association of asthma/rhinitis and DNAm at each of the 677 CpGs, and whether such an association was sex-specific using linear regressions. Two separate models were tested for. Model 1 assessed interaction of asthma and sex while model 2 assessed interaction of rhinitis and sex. Of the 677 CpGs, statistically significant interaction effects with sex were observed at 40 CpGs with asthma (26 genes) (Supplemental Table 2(A)) and 27 CpGs (17 genes) with rhinitis (Supplemental Table 3(A)). A potential sex-specific association of asthma/rhinitis with DNAm was shown at 36 CpGs for asthma (Figure 1a and Supplemental Table 2(A)) and 25 CpGs for rhinitis (Figure 1b and Supplemental Table 3(A)). CpG sites cg05927401 and cg25866075 were identified in both the analyses of asthma and of rhinitis, and at each of these 2 CpGs, the association of asthma/rhinitis with DNAm was sex-specific.

(a) Effects of asthma on 40 CpGs of coronavirus genes, stratified by sex-specific effects. At 36 CpGs, opposite effects of asthma on DNAm were observed in males and females. (The Y-axis is for regression coefficients of asthma status. Negative values indicate that on average DNAm was lower in asthmatic subjects compared to subjects without asthma. The X-axis lists the CpG with their mapped gene names.) (b) Effects of rhinitis on 27 CpGs of coronavirus genes, stratified by sex-specific effects. At 25 CpGs, opposite effects of rhinitis on DNAm were observed in males and females. (The Y-axis is for regression coefficients of rhinitis status. Negative values indicate that on average DNAm was lower in rhinitis subjects compared to subjects without rhinitis. The X-axis lists the CpG with their mapped gene names.)
For CpGs that we did not observe significant interaction effects between sex and asthma/rhinitis status, we examined the main effects of asthma/rhinitis on DNAm at these CpGs. In total, 637 CpGs for asthma and 650 CpGs for rhinitis were included in the analyses. Statistically significant associations of DNAm were observed at 21 CpGs with asthma (17 genes) (Supplemental Table 2(B)) and at 45 CpGs with rhinitis (31 genes) (Supplemental Table 3(B)). One CpG site, cg16628205, was common between the 21 and 45 identified CpGs.
Results from sensitivity analyses indicated factors such as BMI, smoke exposure, and SES did not substantially affect the results. Of the 40 identified CpGs showing asthma and sex interaction effects, statistical significance remained at 31 CpGs (~80%) with the same direction of associations as in the model without these covariates (Supplemental Table 5(A)). Of the 21 CpGs with statistically significant main effects of asthma, after including the covariates, at 18 CpGs (~90%), statistical significance was intact, and the directions of association were the same as those when covariates were not included (Supplemental Table 5(B)). For rhinitis, similar results were obtained; interaction effects at 21 (80%) of the 27 identified CpGs were intact (Supplemental Table 6(A)), and main effects at 38 (~84%) of the 45 CpGs were not affected (Supplemental Table 6(B)).
To assess the biological relevance of 61 (=40 with sex-interactions +21 with main effects only) and 72 (=27 with sex-interactions +45 with main effects only) CpGs that showed association with asthma and rhinitis respectively, we evaluated the association of DNAm at these CpGs with the expression of genes that the CpGs were mapped to as well as their nearby genes by considering a window of 500 000 base pairs (bps) of the CpG site (250 000 bps up and down stream). Additionally, we also assessed whether such associations were sex-specific. We did not have gene expression data for gene CRP (from the study on asthma and rhinitis) and TMPRSS2 (from the study on rhinitis). For asthma, statistically significant interaction effects were observed at 6 of the 61 CpG sites on their association with expression of 5 genes (Table 1), and at 8 CpG sites, DNAm was associated with expression of 8 genes regardless of sex (Table 2). For rhinitis, statistically significant interaction effects were observed at 6 CpG sites on their association with expression of 7 genes (Table 3), and at 11 CpG sites, DNAm was associated with expression of 8 genes regardless of sex (Table 4). In particular, an increase in DNAm was associated with downregulation of 5 genes in males and 4 genes in females for asthma, indicated by negative regression coefficients for each sex. While for rhinitis, an increase in DNAm was associated with downregulation of 4 genes in males and 5 genes in females. For the sex non-specific associations, an increase in DNAm was associated with downregulation of 7 genes for asthma and 6 genes for rhinitis. For the remaining genes, an upregulating relationship was identified. The flowchart of the study is shown in figure 2.
Asthma: sex-specific association of DNAm at 6 CpGs with expression of their mapped genes as well as nearby genes (male = reference group).

Common genes between the findings related to asthma and those on rhinitis (Table 1 and 3) are in bold font.
Abbreviations: Chr., Chromosome; Est., main estimate; Int., interaction coefficient of the asthma with the sex.
P-values for the interaction effects of CpG × sex.
Asthma: sex non-specific association of DNAm at 8 CpGs with expression of their mapped genes as well as nearby genes (male = reference group).
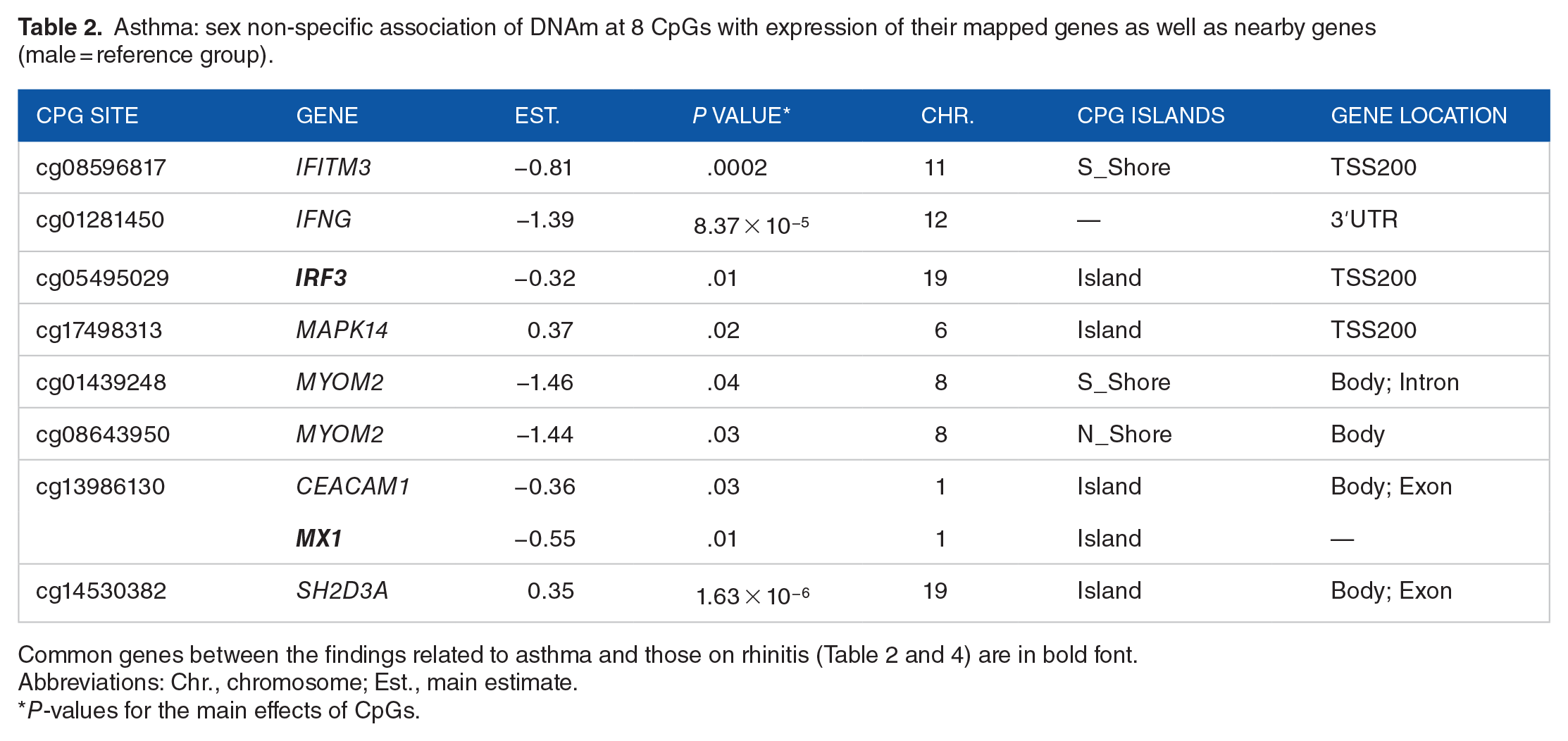
Common genes between the findings related to asthma and those on rhinitis (Table 2 and 4) are in bold font.
Abbreviations: Chr., chromosome; Est., main estimate.
P-values for the main effects of CpGs.
Rhinitis: sex-specific association of DNAm at 6 CpGs with expression of their mapped genes as well as nearby genes (male = reference group).

Common genes between the findings related to asthma and those on rhinitis (Table 1 and 3) are in bold font.Abbreviations: Chr., chromosome; Est., main estimate; Int., interaction coefficient of the rhinitis with the sex.
P-values for the interaction effects of CpG × sex.
Rhinitis: sex non-specific association of DNAm at 11 CpGs with expression of their mapped genes as well as nearby genes (male = reference group).

Common genes between the findings related to asthma and those on rhinitis (Table 2 and 4) are in bold font. Abbreviations: Chr., chromosome; Est., main estimate.
P-values for the main effects of CpGs.
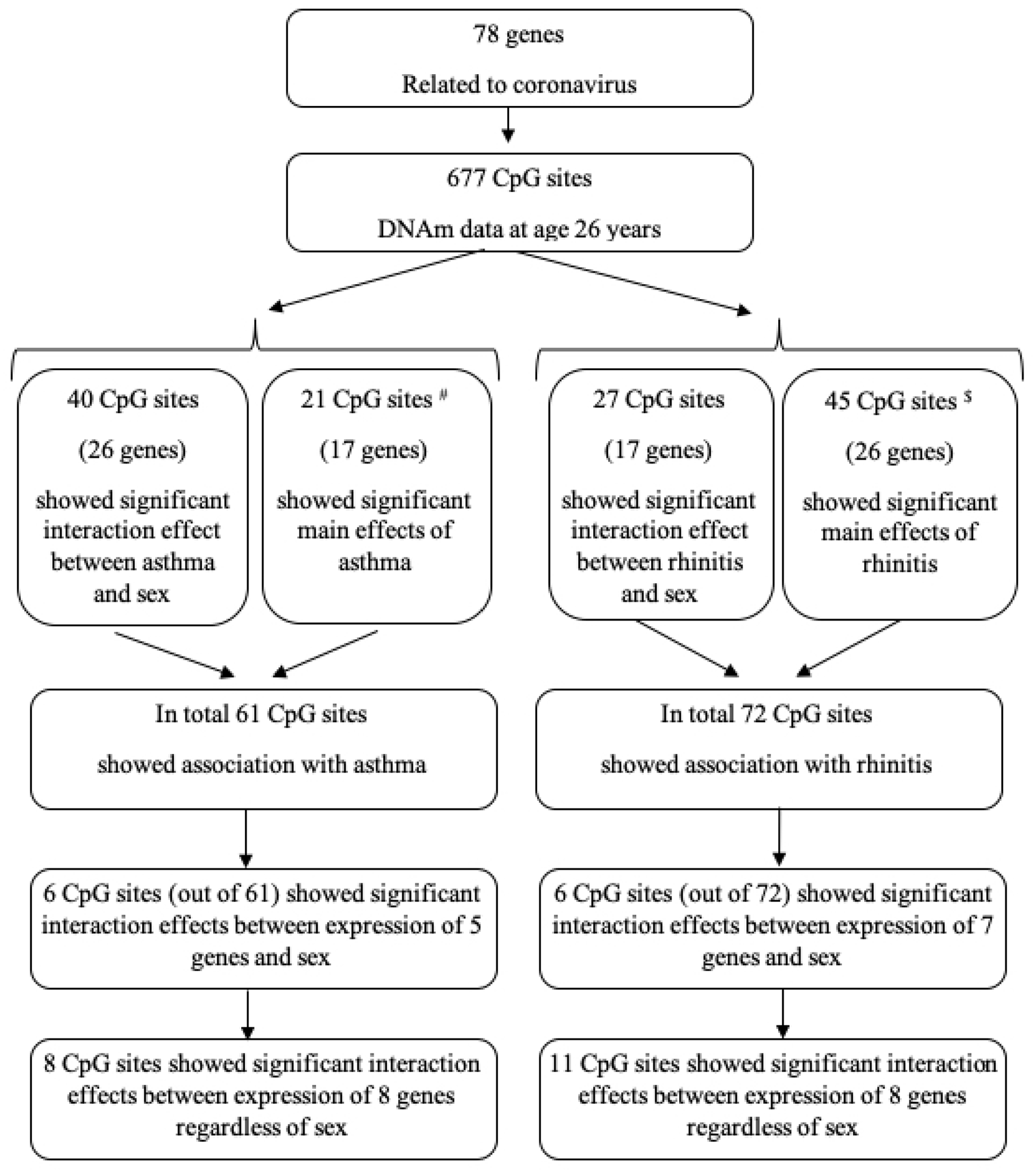
Flowchart of the study.
Discussion
In this study, we assessed the association of asthma/rhinitis with DNAm at CpGs on genes associated with coronavirus in a sex-specific and non-specific manner. We identified statistically significant interaction effects between sex and asthma/rhinitis at 40 CpGs (26 genes) for asthma and 27 CpGs (17 genes) for rhinitis. In addition, at 21 CpGs (17 genes) with asthma and at 45 CpGs (31 genes) with rhinitis, asthma/rhinitis was associated with DNAm regardless of sex. Our sensitivity analysis indicated that the associations of DNAm at these CpGs with asthma/rhinitis were not substantially impacted by potential confounding factors such as BMI, smoking exposure, or SES. Assessment of the biological relevance on the identified CpGs indicated a potential of epigenetic regulatory functionality on gene activity at some of the CpGs and such epigenetic functionality was likely to be sex specific. In addition, for most genes showing association between DNAm and gene expression, the association was stronger in males (indicated by larger effect sizes for males).
Current findings acknowledge the sex difference in COVID-19 susceptibility.3,45-47 The prevalence of asthma and rhinitis was different by sex in our complete cohort, which was consistently observed in the study samples (although the difference was statistically insignificant), with females having a higher prevalence. This has been shown in earlier studies (Patel, Solatikia and Zhang, 2021).48-52 The disproportionality in asthma and rhinitis between males and females possibly contributed to the observed interaction effects of such health conditions with sex on DNAm of genes linked to coronavirus in the present study. Although further investigations are certainly needed to help us better understand such associations, with a focus on CpGs located on genes shown to be associated with coronavirus, our results of sex-specificity on the association of asthma/rhinitis with DNAm and on the association of DNAm with gene expression support sex-specificity in COVID-19 susceptibility and severity. 3 Our findings have a potential to help explain the disproportionality of COVID-19 between males and females. In patients with asthma and rhinitis, the DNAm might have been altered and thereby influence the risk of coronavirus susceptibility or severity of manifestations. This suggests that gene expression regulated by epigenetics, for example, DNAm, may play a critical role in a host’s susceptibility of coronavirus. 26 Further epidemiological and experimental investigations are warranted to examine the epigenetic functionality of the identified CpGs and assess their role in the occurrence of diseases linked to coronavirus.
At 3 genes, HLA-B, IRF3, and MX1, for both asthma and rhinitis, DNA methylation was associated with expression of genes. All these 3 genes have been shown to play a role in pathogenesis of SARS-CoV-2 infection. First, HLA-B was shown to be associated with the severity of SARS-CoV-2 infection. 53 Alleles of HLA-B gene such as HLA-B*07:03, HLA-B*46:01 have been shown to be associated with SARS-CoV-1 susceptibility. 54 As HLA-B*46:01 is expected to have few possible binding peptides for SARS-CoV-2, individuals with this allele may be vulnerable to COVID-19. While HLA-B*15:03 allele has highly conserved SARS-CoV-2 peptides making individuals with this allele to have cross-protective T-cell dependent immunity. 53 Other studies have shown HLA-B alleles HLA-B*07:03, 55 HLA-B*46:01, 56 and HLA-B*52:01:01:02 57 to be significantly associated with severity of COVID-19. A significant correlation of HLA-B*27:07 was observed in a group of 99 Italian patients affected by severe or extremely severe course of COVID-19. 58 Interestingly, DNAm has a quite strong potential of sex-specific regulatory functionality on HLA-B gene (indicated by rather small P-value 6.1 × 10−6), which deserves further investigation regarding its potential to explain infection severity differentiation between sex. Second, IRF3 inactivates interferon signaling which aids viral replication of SARS-CoV-2 infection.59-61 Gene expression of IRF3 has been significantly shown to be upregulated in COVID-19 patients by approximately 250-fold compared to uninfected patients. 62 Another study showed SARS-CoV-2 inhibits activation of IRF3 thereby slowing IFNβ production upon infection. 63 Third, the gene MX1, an interferon responsive gene target, was found to be highly upregulated in SARS-CoV-2 infected normal human bronchial epithelial cells suggesting an activation of antiviral interferon innate response. 64 MX1 antiviral gene expression was significantly higher in COVID-19 patients when compared to non-COVID-19 patients, has been shown to be triggered by SARS-CoV-2, thereby playing a crucial role in inhibiting viral replication. 65 Minor alleles of 5 SNPs near MX1 were found to be correlated with decreased risk to develop severe COVID-19 and high level of MX1 expression in blood. 66 MX1 has been suggested as a potential candidate for drug targets in COVID-19 treatment.65-67 All these 3 genes have been previously demonstrated to be associated with SARS-CoV-2 pathogenesis. Our study from a different angle examined these genes in terms of their role in the connection between asthma or rhinitis and the susceptibility of COVID-19. We showed that DNAm at CpGs on these genes was associated with asthma or rhinitis, which was further associated with expression of genes. Our findings indicated that asthma or rhinitis might have resulted in DNAm changes on these genes and the potential regulatory functionality (shown by their association with gene expressions) of these changes in methylation levels at CpGs of these genes may influence gene expression, thereby influencing the COVID-19 susceptibility and severity among people with asthma and/or rhinitis. We hope our epigenetic findings will help us better understand the pathogenesis of COVID-19 from the epigenetic perspective.
The strength of this study is the availability of DNAm and expression data of genes, shown in the literature to be associated with coronavirus, enabling us to examine the association of DNAm at CpGs on those genes with asthma/rhinitis among adults, and how such association was sex-specific. To our knowledge this is the first study to examine SARS-CoV-2 related genes from an epigenetic perspective.
We aimed to provide a comprehensive epigenetic insight on genes related to coronavirus. This study has some limitations worth mentioning. First, in our data, DNA was extracted from peripheral blood cells. Coronaviruses affect different cell types, primarily, cells of the respiratory tract. 68 In our analysis, we extracted candidate genes potentially related to SARS-CoV-2 infection, and did not have information on which tissue or cells these genes were identified in. It has been shown that DNAm of blood cells has concordance with that of the respiratory system cells, 69 although some differences exist between the 2 and tissues specifically related to the respiratory system are ideal. For CpGs with DNAm associated with gene expressions, our findings supported their biological relevance and indicated that those CpGs at least had a potential to serve as markers. However, for genes not showing any association with expression of genes, studies focus on different tissues are desired to examine epigenetic perspectives of asthma and rhinitis on coronavirus related genes. Second, the candidate genes included in our study were not randomly selected or untargeted (eg, as in genome-wide studies). In such situations, forcing multiple testing would potentially increase type II error and is not recommended. 70 However, we believe it is important to further test the identified CpGs in independent cohorts to assess their reproducibility and generalizability. Third, we were not able to assess the impact of the identified CpGs on the risk of COVID-19 incidence due to lack of COVID-19 data in those subjects. Nevertheless, our findings showed that subjects with asthma or rhinitis had different DNAm compared to people without the diseases on certain genes related to coronavirus, which suggests a possibility that subjects with asthma or rhinitis may be at an altered risk of susceptibility to the SARS-CoV-2 infection due to changes in their DNAm. An assessment on the direction of risk, increased/decreased/intact, requires information on infection status, and certainly deserves further investigation.
Supplemental Material
sj-docx-1-gae-10.1177_25168657211039224 – Supplemental material for Association of Asthma and Rhinitis with Epigenetics of Coronavirus Related Genes
Supplemental material, sj-docx-1-gae-10.1177_25168657211039224 for Association of Asthma and Rhinitis with Epigenetics of Coronavirus Related Genes by Aniruddha Rathod, Rutu Rathod, Hongmei Zhang, Parnian Kheirkhah Rahimabad, Wilfried Karmaus and Hasan Arshad in Epigenetics Insights
Footnotes
Acknowledgements
The authors are thankful to the nurses and staff at the David Hide Asthma and Research Allergy Centre, Isle of Wight, UK, for their help in recruitment and sample collections, and are thankful to all the cohort participants.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Institutes of Health (NIH) research fund R01AI121226 (MPI: H. Zhang and J.W. Holloway) and by NIH research fund R01HL132321 and R01AI091905 (PI: W. Karmaus). The Isle of Wight Birth Cohort assessments were supported by NIH research fund R01HL082925 (PI: H. Arshad), Asthma UK (Grant no. 364. S.H. Arshad), and the David Hide Asthma and Allergy Research Trust.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
HZ, WK, and AR conceived the study. HZ, RR, and AR designed the analytical plan. SHA provided data. PR, WK, and HZ searched the genes related to coronavirus. AR and RR did data management and performed the statistical analyses. AR, RR, and HZ drafted the manuscript. All authors reviewed and approved the final manuscript.
Availability of Data and Materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Ethics Approval and Consent to Participate
The IoW birth cohort study was approved by the Isle of Wight, Portsmouth and Hampshire Local Research Ethics Committee (now known as the National Research Ethics Service, NRES Committee South Central—Hampshire A) (06/Q1701/34) DNA methylation related studies have been approved by the University of Memphis Institutional Review Board in Memphis, U.S. (FWA00006815).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
