Abstract
Stroke-related numbness and weakness (SRNW) are resultant symptoms of post-stroke sufferers. Existing research has supported the use of Huangqi Guizhi Wuwu Tang (HGWT) particularly for SRNW; however, their mechanisms of action have not been fully elucidated. Therefore, this study aimed to investigate the mechanisms of action of HGWT components targeting SRNW-related proteins through a computational molecular docking approach. Target proteins associated with SRNW were identified through DrugBank database and Open Targets database. Chemical compounds from each herb of HGWT were identified from the Traditional Chinese Medicine Systems Pharmacology and Analysis Platform (TCMSP). Autodock Vina was utilized and the cut-off criterion applied for protein-ligand complexes was a binding affinity score of ≤ -9.5 kcal/mol; selected protein-ligand complexes were identified using 3D and 2D structural analyses. The protein targets PDE5A and ESR1 have highlighted interactions with compounds (BS040, DZ006, DZ058, DZ118, and HQ066) which are the key molecules in the management of SRNW. PDE5A have bioactivity with the amino acid residues (Val230, Asn252, Gln133 and Thr166) throughout PDE5A-cGMP-PKG pathways which involved reduction in myofilament responsiveness. ESR1 were predicted to be critical active with site residue (Leu346, Glu419 and Leu387) and its proteoglycans pathway involving CD44v3/CD44 that activates rho-associated protein kinase 1 (ROCK1) and ankyrin increasing vascular smooth muscle. In conclusion, HGWT may provide therapeutic benefits through strong interactions between herbal compounds and target proteins of PDE5A and ESR1. Further experimental studies are needed to unequivocally support this result which can be valuable to increase the quality of life of post-stroke patients.
Introduction
Stroke-related numbness and weakness (SRNW) is a common symptom in post-stroke patients. Over 60% of post-stroke patients have experienced functional impairment of the body (such as numbness) and over 90% of them have suffered from weakness of body or extremities.1‐3 SRNW may cause pain, limited limb mobility, and affect patients’ quality of life.4‐6 Current management for SRNW includes drug and non-drug interventions.6‐10 Anti-spastic drugs are commonly used for severe numbness cases; however, they may have adverse consequences such as dry mouth, gastrointestinal disturbance, sleep disorder and dizziness.9,11,12 There is no other specific drug for SRNW. Thus, non-drug interventions such as rehabilitation programmes (eg, posture management, exercise, and physical modalities) could be used as primary treatments to alleviate SRNW.4,5,10,13 Nonetheless, due to limited therapeutic effects of these non-drug interventions for SRNW, post-stroke symptoms may become worse and worse. Furthermore, it is worth noting that long-term management of SRNW in post-stroke patients may lead to a financial burden to the patients, their families and carers, and potentially reduce work productivity.13,14 Due to the unsatisfied current management of SRNW, increasing SRNW patients consider seeking an alternative treatment option for better therapeutic outcomes and fewer adverse effects. 4
Chinese herbal medicine has been increasingly appreciated as an option for SRNW management. Huangqi Guizhi Wuwu Tang (HGWT) is one of the classical herbal formulas recorded for treating stroke-like symptoms in classical literature of Chinese medicine.15,16 This formula originated from Jin Gui Yao Lue (Synopsis of Prescriptions of the Golden Chamber) written by Zhang Zhong-Jing during the Han Dynasty of China about 1800 years ago. 17 In modern days, it has been used to treat numbness in many diseases including SRNW in clinical practice.15,18,19 However, HGWT's mechanisms of action are unknown. This study aimed to investigate the potential compounds of HGWT for the SRNW management through a herb-target molecular docking approach.
Materials & Methods
Identification and Preparation of Protein Targets
Target proteins associated with SRNW were identified through two databases, including DrugBank database (https://go.drugbank.com) 20 and Open Targets platform online database (http://www.opentargets.org). 21 Firstly, stroke or post-stroke related target proteins were identified from both databases. Then, the muscular numbness and weakness-related target proteins were identified from the Open Targets platform. Secondly, the overlapping target proteins between the two groups of candidate target proteins were chosen and the identified targets were entered into the STRING database (https://string-db.org) (platform 2020) to analyze their protein-protein interactions. Thirdly, the protein symbols of these target proteins were searched in the Uniprot database (https://www.uniprot.org/) to identify appropriate target protein sequences using the online BLAST tool. The identified sequences were then submitted to Swiss-Model (https://swissmodel.expasy.org/) to build three-dimensional (3D) structures and to repair missing regions. The models were then saved in PDB format. Furthermore, editing of each selected target protein was performed where necessary to remove their disordered regions, particularly for the potassium and calcium channels, and water molecules were removed, using Biovia Discovery Studio Visualizer 2019. Finally, the prepared target proteins in PDB format were converted to PDBQT (Protein Data Bank, Partial Charge [Q], & Atom Type [T]) format using the PyRx for molecular docking. These proteins were treated as receptors and used for molecular docking.
Identification and Preparation of HGWT-Compounds
HGWT-compounds were identified through examination of the chemical constituents of HGWT's individual ingredients. HGWT contains five herbs including Astragali Radix (Huang qi, Astragali), Paeoniae Radix Alba (Bai shao, Paeoniae), Cinnamomum Ramulus (Gui zhi, Cinnamomum), Jujubae Fructus (Da zao, Jujubae) and Zingiberis Rhizoma Recens (Sheng jiang, Zingibers). Chemical compounds from each herb were found from the Traditional Chinese Medicine Systems Pharmacology and Analysis Platform (TCMSP, https://tcmspw.com). 22 After removing the duplicated compounds, the molecular structures were obtained and downloaded from PubChem 23 as SDF files and translated into the PDB format using Biovia Discovery Studio Visualizer 2019 which provides molecular visualization as a multi-objective optimization. The molecules were treated as ligands that were used for molecular docking. Figure 1 illustrates the procedures for preparing candidate target proteins as receptors and HGWT-compounds as ligands for computational molecular docking.
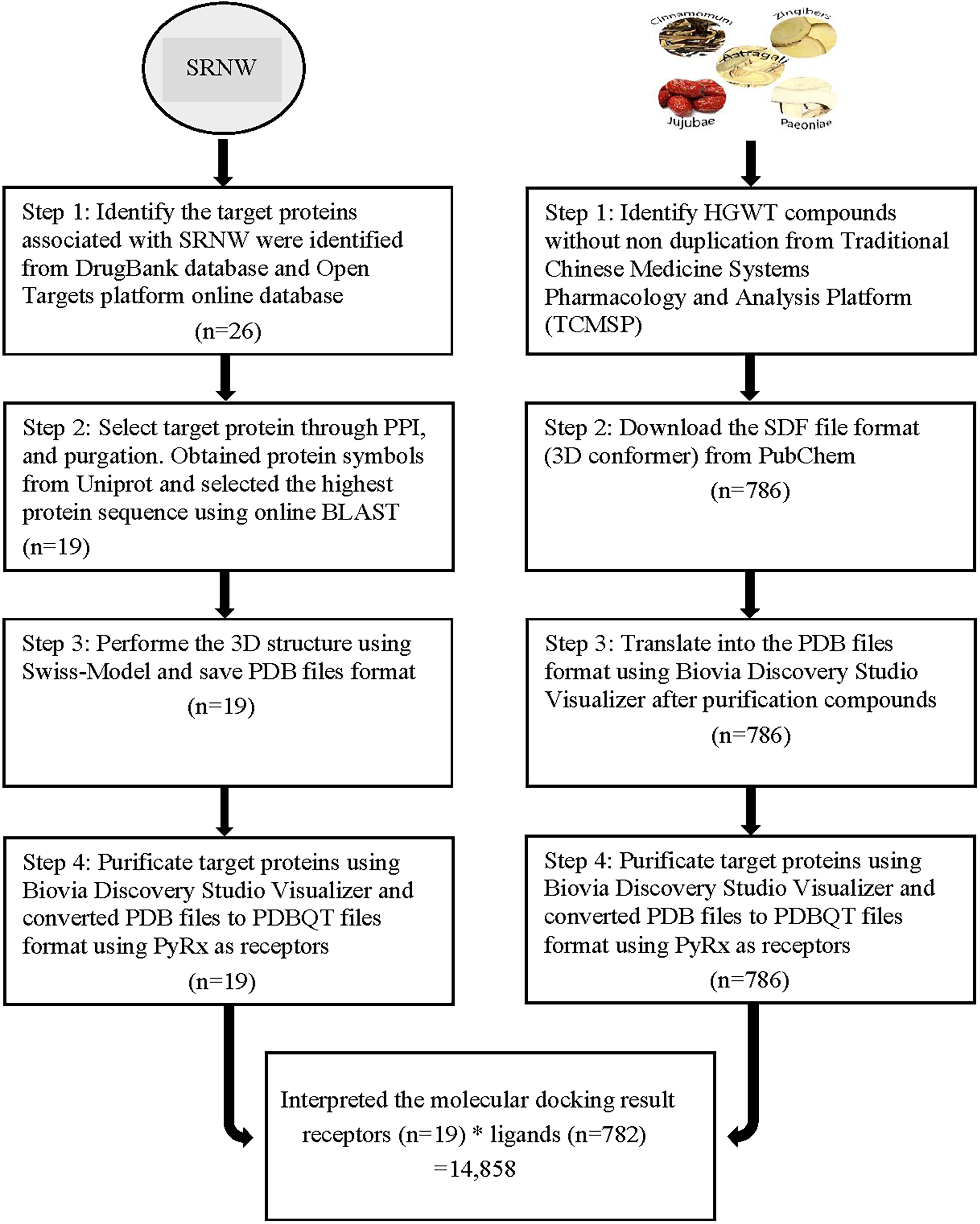
Procedures for molecular docking between target proteins and compounds. The target proteins were obtained from two databases for stroke-related numbness and weakness (SRNW), processed to PDB files format, selected for the highest protein sequence using online BLAST, and then built 3D structure from Swiss-Model. Compounds were searched from the Traditional Chinese Medicine Systems Pharmacology and Analysis Platform (TCMSP) and downloaded in SDF files format from PubChem, then changed to PDB files format through Biovia Discovery Studio Visualizer. Both target proteins and compounds translated to PDBQT files format using PyRx. Finally, molecular docking was performed using high performance computing facilities, and 14,858 results were obtained from the docking between target proteins (n = 19) and herbal compounds (n = 782), in which four compounds did not have a significant predicted binding affinity value in the molecular docking result.
Molecular Docking and Analysis
We used a similar molecular docking approach to our previous published paper. 24 Specifically, all the receptors from candidate target proteins and all the ligands from HGWT-compounds were loaded into PyRx (Virtual Screening software for Computational Drug Discovery, Autodock Vina) which elucidates the biochemical process between small molecule and protein using the model of interaction,25,26 while Autodock Vina use the Broyden-Fletcher-Goldfarb-Shanno (BFGS) method that optimizes the predicted docking poses by minimizing the value and the derivative of the scoring function.27,28 Docking calculations enable the prediction of the binding positions of small-molecule ligands to protein targets, and also produce a quantitative estimation of binding affinity values, both of which help to elucidate the mechanisms of action of a library of compounds at the molecular level. All of the receptors and ligands were converted to PDBQT format using PyRx. Molecular docking between target proteins and chemical compounds was conducted using the ‘Gadi’ high performance computing facilities hosted by the National Computational Infrastructure (NCI) in Canberra, Australia.
Further analyses were performed on selected ligand-receptor complexes which were predicted to be important for HGWT's mechanisms of action. These complexes were chosen if they satisfy both of the following criteria. The docking-predicted receptor-ligand complexes must include i) one of the top three receptors with the highest average binding affinity values with respect to the ligands; and ii) one of the top three ligands with the highest average binding affinity at the receptors. Complexes that satisfy the above were selected for further specific analyses of ligand-residue interactions.
The binding site and ligand interactions between the identified targets and HGWT's compounds were then analyzed using Biovia Discovery Studio Visualizer 2019. The strength of molecular-level interactions was determined by ligand-receptor binding affinity. The more negative the binding affinity has the higher the binding energy between compounds and target proteins. 29 We hypothesized that ligand-receptor complexes with higher binding affinity scores may show more effectiveness in treating SRNW. 30 Additionally, we performed 3D and two-dimensional (2D) analyses to visualize the specific residue-level interactions between ligands and the highest potential inhibition from the targets and thus to further investigate the active sites in the binding pockets for selected receptors and ligands.
Results
Selected Target Proteins and HGWT Compounds
Twenty-six target proteins were identified associated with SRNW. Seven of them were removed for the following reasons: six target proteins (CACNA1A, CACNA1S, KCNH2, IKBKB, SELP and HMGCR) contained structures that were too large for bind docking to be performed, and one target protein (CA1) was removed as it did not have a protein-protein interaction (PPI), and the PPI analysis was provided in Figure 2. As a result, a total of 19 target proteins were selected as candidate target proteins for molecular docking, as detailed in Table 1.

Protein-protein interaction for network functional enrichment analysis. 19 potential target proteins are indicated in three different colours which indicates that they are clustered to three groups according to a specified markov cluster algorithm (MCL) inflation parameter using string database. Eleven potential target proteins (ADRB2, CASP3, FGF2, INS, MAPK3, PDE5A, PPARA, REN, SOD1, T TR, and VEGFA) are coloured in red, indicating that these proteins are partially biologically connected. Six proteins (AR, ESR1, HDAC2, HDAC4, PCNA, and SOST) were interacted in a cluster and they are represented in green. Another two proteins (APP and HPR) were interacted in different clusters and they are shown in blue. In analysis of these protein networking: number of nodes: 19; number of edges: 66; average node degree: 6.95; average local clustering coefficient: 0.81; expected number of edges: 26; PPI enrichment p-value: 3.65e-11. All nodes have been coloured, indicating the query proteins and first shell of interactors, and the edges represent protein-protein associations that predict interactions with different colour connections (green: gene neighbourhood; red: gene fusions; blue: gene co-occurrence; yellow: text mining; black: co-expression; and grey: protein homology).
19 Potential Target Proteins.

Note: 1: according to Uniprot database; 2: David database (KEGG pathways).
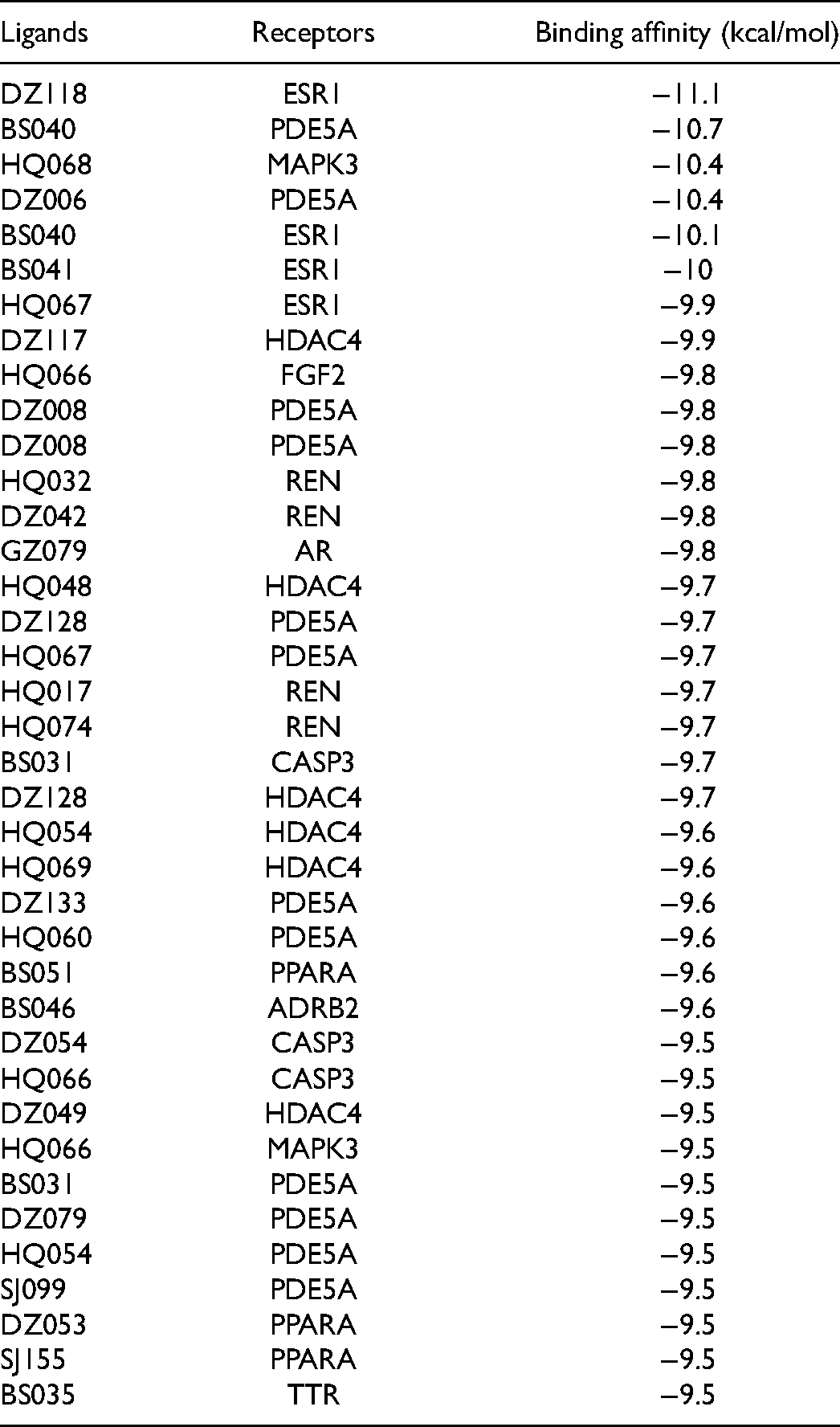
A total of 786 compounds were identified from HGWT's ingredients, including 85 from Astragali Radix (Huang qi), 83 from Paeoniae, 220 from Cinnamomum, 133 from Jujubae, and 265 from Zingiberis. Among the 786 compounds, four compounds did not have a significant predicted binding affinity value in the molecular docking result (DZ006, DZ009, DZ051, and SJ041). The molecular docking between 19 targets and 782 compounds yielded 14,858 results, as shown by the top 20 ranked higher binding affinity scores in Supplementary file 1. The binding scores between all the compounds and target proteins ranged from −0.8 to −11.1 kcal/mol and their average binding score was −6.0 kcal/mol. The selected complexes with binding affinity scores ≤-9.5 kcal/mol are listed in Table 2. The molecular docking results indicated that each herb in HGWT has been predicted to comprise at least one compound targeting a distinct protein target. It is hypothesized that compounds with more negative binding energy (ie, highest docking energies) are predicted to be more activated between compounds and target proteins.29,31 The compound with the highest binding affinity energy was DZ118 (a compound from Jujubae) against Estrogen receptor alpha (ESR1) with a binding affinity score of −11.1 kcal/mol; the second strongest binding affinity was between cGMP-specific 3′,5′-cyclic phosphodiesterase (PDE5A) and BS040 (a compound from Paeoniae) with a binding affinity score of −10.7 kcal/mol; the third highest binding affinity (-10.4 kcal/mol) was predicted for two ligand-protein complexes, including HQ068 (a compound from Astragali) against Mitogen-activated protein kinase 3 (MAPK3) as well as DZ006 (a compound from Jujubae) against PDE5A.
Binding Affinity between Ligands and Receptors with Scores ≤ −9.5 kcal/mol.
Top Three Receptors with Highest Average Binding Affinity Against Ligands
The top three receptors with the highest average binding affinity against ligands are PDE5A (average binding score: −6.7 kcal/mol), ESR1 (average binding score: −6.6 kcal/mol), and Beta-2 Adrenergic Receptor (ADRB2, average binding score: −6.6 kcal/mol) (Table 3).
Top Three Receptors with Highest Average Binding Affinity Results.

The binding affinity scores of PDE5A with its top three ligands ranged from −10.7 to −9.8 kcal/mol, ESR1 ranged from −11.1 to −10.0 kcal/mol and ADRB2 ranged from −9.5 to −9.0 kcal/mol. Thus, ESR1 has a substantially higher binding affinity than PDE5A and ADRB2. PDE5A has predicted strong binding with BS404, DZ006, DZ117 and HQ066, for which DZ117 and HQ066 have the same binding affinity score as −9.8 kcal/mol while ESR1 has high predicted binding affinity values with DZ058, DZ006, and DZ118. PDE5A and ESR1 both interact with DZ006 compounds which are predicted to have active interactions. ADRB2 interacts with BS040, DZ057 and DZ123 as an average high binding affinity receptor. PDE5A, ESR1 and ADRB2 are the proteins that exhibit the highest average binding affinity with the ligands of HGWT in this study.
Top Three Ligands from Each Ingredient of HGWT
The top three ligands from each HGWT ingredient with the highest average binding affinity against all the receptors are the significant compound of each herb and are provided in Table 4. Among the compounds of Paeoniae, BS040 was predicted to interact strongly with two receptors, PDE5A (binding affinity −10.7 kcal/mol) and ESR1 (binding affinity −9.9 kcal/mol). The top three ligands from Jujubae interacted with ESR1 and PDE5A, with DZ118 showing the highest binding affinity. Cinnamomum had the highest interaction with HDAC4 (binding affinity energy −9.7 kcal/mol) with GZ079. HQ068 had the highest binding affinity energy (-10.4 kcal/mol) with MAPK3 in Astragali compounds. Finally, Zingiberis had the highest interaction with PDE5A and PPARA. Their binding affinity energy was −9.5 kcal/mol.
Huangqi Guizhi Wuwu Tang's Herbal Ingredients with Their Significant Ligands Predicted from top Three Binding Affinity with Receptors.

Table 4 shows that the top three ligands from each herb interacted mostly with PDE5A (n = 10), which in turn also interacted with the following compounds: BS040, DZ006, GZ081, GG210, HQ048, HQ066, HQ067, SJ099, SJ155, and SJ051. The receptor with the second-highest number of interactions was Renin (REN, n = 4) with compounds: HQ017, HQ032, HQ074 and SJ051 and the receptor with the third-highest number of interactions was ESR1 (n = 3) with compounds: BS040, DZ058, and DZ118. However, ESR1 had higher binding affinity scores with the compounds than REN. Jujubae exerted the greatest number of ligands with the highest binding affinity in its compounds (DZ006, DZ057, DZ058, DZ117, DZ118 and DZ123) which suggests that Jujubae may contain the most substantial number of biologically active molecules against the top three proteins. PDE5A and ESR1, which interacted with these herbal compounds, are also predicted to have the highest average binding affinity as calculated over the values obtained for all herbal ligands, indicating their potential importance in the mechanisms of action of HGWT. However, one of the top three receptors with the highest average binding affinity, ADRB2, was not included in the complexes involving the top three ligands from each ingredient of HGWT. Thus, ADRB2 was excluded in the 3D and 2D analyses.
Interaction Analysis Using 3D and 2D
Receptor-ligand complexes were chosen if they contain one of the top three receptors with the highest average binding affinity against ligands as well as one of the top three ligands from each ingredient of HGWT. The receptors (PDE5A and ESR1) and ligands (BS040, DZ006, DZ058 DZ118 and HQ066) were chosen and analyzed using 3D ribbon diagrams and 2D ligand-residue interaction diagrams. The 3D interaction diagram indicates that PDE5A, two ligands (BS040 and DZ006) were predicted to bind in a similar pocket, while one ligand (HQ066) was predicted to bind in a distinct location. DZ058 and DZ118 were bound at a common specific location on the surface of ESR1, as shown in Figure 3.

3d analyses showing ESR1 and PDE5A interactions with five ligands. (A) ESR1 interactions with 2 ligands (DZ058 and DZ118); (B) PDE5A interact with 3 ligands (BS040, DZ006 and HQ066).
In terms of 2D analysis, there were two receptors (PDE5A and ESR1) and five ligands (BS040, DZ006, DZ058, DZ118 and HQ066) involved. Their interactions are shown in Figure 4. The green lines in the figures indicate hydrogen bonds; the pink lines show alkyl interactions or π-alkyl interactions; and the light-yellow lines represent π-sulfur interactions. Table 5 shows the form of hydrogen bond and other bonding interactions from each binding ligand that could be used to characterize the binding sites. Figure 5 indicates that each amino residue interaction with PDE5A and ESR1 in 3D. In the interactions between PDE5A and three ligands (BS040, DZ006 and HQ066), the amino acid residues (Val230, Asn252, Gln133, and Thr166) were predicted to form hydrogen bonds. Six amino acid residues had π-alkyl (Tyr261 and Phe251) or alkyl interactions (Ile266, Ile285, Ile229 and Val255) and one has an unfavorable interaction (Asn485). Compounds identified from Astragali, Jujubae, and Paeoniae were predicted to form more than one hydrogen bond with amino acid residues within the binding pockets, and these ligands may be especially effective for SRNW via targeting of PDE5A, since there are more hydrogen bonds found in the binding pocket than other ligands throughout the 2D analysis

2d analyses showing ESR1 and PDE5A interactions with five ligands (BS040, DZ006, DZ058, DZ118 and HQ066). (A) ESR1 and DZ058; (B) ESR1 and DZ118; (C) PDE5A and BS040; (D) PDE5A and DZ006; (E) PDE5A and HQ066. The green lines in the figures indicate hydrogen bonds; the pink lines show alkyl interactions or π-alkyl interactions; the light-yellow lines show π-sulfur interactions and red lines indicate unfavourable acceptor-acceptor.

3d analyses of each amino acid residue interactions with PDE5A or ESR1. (A) Leu346 interacted with hydrogen bond between ESR1 and DZ058; (B) Glu419 and Leu387 interacted with hydrogen bond between ESR1 and DZ118; (C) Val230 interacted with hydrogen bond between PDE5A and BS040; (D) Asn252 interacted with hydrogen bond between PDE5A and DZ006; (E) Glu133 and Thr166 interacted with hydrogen bond between PDE5A and HQ066. Note: Blue colour: ligands; green: Amin acids interacted with ligands; yellow colour: amino acid residues interacted with ligands connected hydrogen bond; green dot lines: hydrogen bond.
Details of the 2D Analyses of PDE5A and ESR1 with the Residues of Ligands.

ESR1 interacted with 16 amino acid residues which formed close contacts with the ligands. Among them, three had hydrogen bonding interactions with the amino acid residues (Glu419, Leu387 and Leu346); Glu419 and Leu387 interacted with DZ0118, and Leu346 interacted with DZ058. Additionally, five amino acid residues (Ala350, Leu346, Met421, Leu525, and Ile424) involved π-alkyl interactions with the ligands; Leu346 interacted with DZ058 via π-alkyl interactions and DZ118 via hydrogen; and four amino acid residues (Leu524, Leu525, Leu540, and Leu536) had alkyl bonding with the ligands. Furthermore, Phe404 had a π-π interaction in a T-shaped configuration with DZ118; Leu525 was involved in both bonding via π-alkyl with DZ118 and alkyl interactions with DZ058; Met343 and Met421 exhibited π-sulfur bonding. Hydrogen bonding with the backbone of Leu346, Glu419 and Leu387 in the binding pocket may be a key role of receptor binding for SRNW.
The formation of multiple hydrogen bonds connected to the aromatic ring structure provides stability as predicted by the 2D analysis of molecular docking. It is proposed that the selected compounds (BS040, DZ006, DZ058, DZ118, and HQ066) may have significant bio-active effects upon PDE5A and ESR1 receptor binding, and these particular compounds are worthy of further investigation
Discussion
HGWT has been used for the management of SRNW since many post-stroke patients experience those symptoms. SRNW are not severe or life-threatening; however, they do decrease patients’ quality of life. Even though there are many clinical studies, including randomized clinical trials (RCTs), providing evidence of the effectiveness of HGWT in related post-stroke symptoms such as shoulder-hand syndrome, hemiplegia, and paralysis of the body,32‐34 the mechanisms of action of HGWT are not completely known. This study thoroughly analyzed HGWT using a computational docking method to find HGWT's mechanisms. Hence, this study is critical to provide further understanding of the mechanism of HGWT's effects on SRNW.
The present study identified target proteins using DrugBank and Open Targets databases as proteins from these databases are approved medicines and currently used for post-stroke. TCMSP provides comprehensive compounds of herbal medicines. Autodock Vina is a molecular docking programme which predicted binding energy scores between macromolecule (receptor) and herbal compound (ligand). These methods enabled us to select target proteins and compounds for docking as well as identify strong interactions efftively and efficiently.
In our study, two target proteins including ESR1 and PDE5A were shown to have a high predicted binding affinity with HGWT-compounds. In particular, five compounds (BS040, DZ006, DZ058, DZ118, and HQ066) were identified to exhibit strong interactions with target proteins and play a key role in bioactive processes involved in SRNW. The interactions with the receptors potentially play a key role in their mechanisms of action in SRNW and identifying the type of bonding with different residues of the ligands provides insight into the potential biological activity of these ligands, such as the dominant types of forces that govern their binding. In particular, the hydrogen bonds between the binding site of residues and receptors may be especially important for the activities of ligands in their activity against SRNW, as hydrogen bonds are among the strongest type of non-covalent interactions. In the current docking study, amino acid residues including Val230, Asn252, Gln133 and Thr166 of PDE5A; and Leu346, Glu419 and Leu387 of ESR1 were predicted to be critical active site residues that bind to the compound molecules in PDE5A and ESR1 via hydrogen bonding interactions.
According to a pharmacophore model study for PDE5A, promising results were obtained which predicted that the presence of a hydrogen bond and aromatic ring provided stability for ligand binding in the pocket of this protein, and ligands which these characteristics may be effective in generating vascular muscular relaxation. 35 PDE5A-cGMP-PKG pathways involve the nitric oxide stimulation of guanylate cyclase and reduction in myofilament responsiveness to Ca2+ that leads to the reduction of muscular numbness.36,37 In addition, this pathway was found to be associated with metabolic energy deficiency and decreased mitochondrial antioxidant activity in myocardial tissue in an animal experimental study. 38 Another protein, ESR1 plays a key role in facilitating the transcriptional effects of hormone functions in protein kinase in the target tissues and expressing in endothelial cells, vascular smooth muscle cells, and macrophage.39,40 According to KEGG pathways, there are nine signalling pathways involving ESR1 (endocrine resistance, chemical carcinogenesis-receptor activation, breast cancer, estrogen signalling pathway, prolactin signalling pathway, pathways in cancer, thyroid hormone signalling pathway, endocrine and other factor-regulated calcium reabsorption, and proteoglycans in cancer). 41 Among the mentioned pathways, proteoglycans in the cancer pathway has shown cytoskeleton activation, cell growth and proliferation, and survival to be similar to SRNW. This pathway involves CD44v3 which activates rho-associated protein kinase 1 (ROCK1), increases serine/threonine-protein phosphatase PP1 catalytic subunit, then acts on vascular smooth muscle causing contraction and regulation of actin cytoskeleton; CD44 also activates ankyrin which leads to cytoskeleton activation.41,42 It can be suggested that ROCK1 and ankyrin may play an important role in muscular contraction and relaxation. Also, the molecular docking results revealed that the highest binding affinity compound is Furmarine (DZ118), an alkaloid that possesses anti-cholinergic effectiveness that helps treat abnormal involuntary muscle movement. Another compound, Spiradine A (DZ006) is an alkaloid with high interactions with ESR1 and PDE5A. Spiradine A could inhibit platelet aggregation by atisine-type diterpene alkaloids (ADP) and hence significantly inhibit platelet-activating factor which is potentially induced by aspirin and regulates blood to improve the weakness of the body. 43
Furthermore, ESR1 decreases the inflammatory response through a reduction of pro-inflammatory cytokines and a dampening of inflammatory gene expression in smooth muscle cells interacted with NF-κB inhibits in KEGG pathway. 41 Therefore, we suggest ESR1 and its three amino acid residues (Leu346, Leu387 and Glu419) are important components to be targeted by ligands in the design of new treatments of SRNW.
Although this study has provided a prediction of the interactions between compounds of HGWT and target proteins of current post-stroke medications and disease target proteins, there are some limitations in this study. From the results of the binding affinity predictions, only the receptor-ligand complexes satisfy the criteria of containing one of the top three receptors and also containing one of the top three ligands with the highest average binding affinity from each ingredient of HGWT were selected for further detailed analysis. Even though these ligands enabled an initial prediction of potentially important residue interactions and enabled us to put forward a number of molecules of potential interest in the context of therapy development, some of the remaining ligands which may also have relatively high predicted binding affinity results were not included, and future work could be performed to characterize the biological activities of other compounds obtained from HGWT.
Conclusions
In summary, this preliminary study investigated the compounds of HGWT, one of the most commonly used herbal formulas for the management of post-stroke symptoms.15,16 This study predicted the binding mechanisms between the pharmaceutical target proteins from the commonly used medicine in post-stroke in DrugBank and open target protein platform, with HGWT-compounds, using a computational molecular docking method. A total of 19 receptors and 782 ligands were studied, obtaining a total of 14,858 docking results. Among these results, the ESR1-DZ118 interaction exhibited the highest binding affinity, followed by the PDE5A-BS040 complex, MAPK3-HQ068 and PDE5A-DZ006 complexes. The compounds, BS040, DZ006, DZ058, DZ118 and HQ066 exhibited the most significant interactions with target proteins PDE5A and ESR1. The ligands BS040, DZ006 and HQ066, forming hydrogen bonding interactions with Val230, Asn252, Gln133 and Thr166 with PDE5A; while ESR1 only has one predicted hydrogen bonding interaction, Leu346 with DZ058. Our study therefore predicts that five compounds (BS040, DZ006, DZ058, DZ118, and HQ066) in HGWT may have key involvement in the mechanisms of action for SRNW. These findings need to be further validated in experimental studies to guide new drug development for SRNW management in the future.
Supplemental Material
sj-docx-1-chp-10.1177_2515690X221082989 - Supplemental material for Mechanisms of Action of a Herbal Formula Huangqi Guizhi Wuwu Tang for the Management of Post-Stroke Related Numbness and Weakness: A Computational Molecular Docking Study
Supplemental material, sj-docx-1-chp-10.1177_2515690X221082989 for Mechanisms of Action of a Herbal Formula Huangqi Guizhi Wuwu Tang for the Management of Post-Stroke Related Numbness and Weakness: A Computational Molecular Docking Study by Sanghyun Lee, Andrew Hung, Hong Li and Angela Wei Hong Yang in Journal of Evidence-Based Integrative Medicine
Footnotes
Acknowledgments
We thank the National Computational Infrastructure (NCI) and the Pawsey Supercomputing Center (Australia) for the access to high-performance computing facilities. The NCI and Pawsey Centre are funded by the Australia Government. We also thank Dr Hanife Mehmet for proofreading this manuscript.
Authors’ Contributions
Sanghyun Lee performed the experiments, analyzed the data, prepared figures and/or tables, authored and reviewed drafts of the paper, and approved the final draft; Andrew Hung performed the experiments, analyzed the data, prepared figures and tables, authored and reviewed drafts of the paper, and approved the final draft; Hong Li analyzed the data, prepared figures and tables, authored and reviewed drafts of the paper, and approved the final draft; Angela Wei Hong Yang conceived and designed the experiments, analyzed the data, authored and reviewed drafts of the paper, and approved the final draft.
Ethical Approval
No ethics application is needed for this research project.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
