Abstract
Lipid droplets (LDs) are central hubs in cellular lipid handling. They serve as lipid storage organelles and are involved in neutral lipid biosynthesis and breakdown as well as in the production of phospholipids and sterols. For communication with other organelles, LDs are heavily engaged in contact sites. The molecular basis of these structures is formed by proteins or protein complexes termed tethers, which attach partner organelles to the surface of LDs. Here, we describe the structural and functional characteristics of recently identified LD tethers. Intriguingly, these LD tethers have additional features, such as the structural capacity to form tri-organellar contacts, domains specialized for interorganellar bulk lipid transfer, and connections to specific lipid metabolism enzymes, which might collectively contribute to the key role of LDs in cellular lipid flux.
Panta Rhei: Lipid Droplets and Lipid Flux
Cellular lipid flux is a key aspect of life. Lipids serve diverse crucial biological roles, for example, as building blocks of all cellular membrane systems, as highly efficient energy supply, and as intra- and intercellular signaling molecules (Athenstaedt and Daum, 2006; van Meer et al., 2008; Wymann and Schneiter, 2008). These functions critically depend on compartmentalization of lipid metabolism reactions and on directed lipid transport between different cellular compartments (Stefan et al., 2017). For example, the endoplasmic reticulum (ER) is a key player in the synthesis of membrane lipids. Lipid-based adenosine triphosphate (ATP)-production through fatty acid breakdown via β-oxidation happens in mitochondria and peroxisomes. The endolysosomal compartment is involved in lipid recycling. While lipids fulfill crucial roles, they can also have adverse lipotoxic effects (Schaffer, 2003). Particularly problematic are free saturated fatty acids, which compromise cellular functions and induce cell death at elevated levels (de Vries et al., 1997; Hardy et al., 2000; Cnop et al., 2001; Listenberger et al., 2001, 2003). Fatty acids essentially are detergents, and thus have the potential to directly damage cellular content. Furthermore, they can drive adverse metabolic alterations, such as the pronounced formation of ceramides and other lipid species that are cytotoxic at higher levels (Paumen et al., 1997; Maedler et al., 2001). To increase the cell's lipid storage capacity and prevent lipotoxicity, excess fatty acids and sterols are esterified to form neutral lipids, specifically triacylglycerols (TAGs) and sterol esters, which represent lipid storage forms (Athenstaedt and Daum, 2006). Lipid droplets (LDs) are ubiquitous organelles that house most of the cell’s neutral lipids. LDs have a unique architecture, with their whole internal compartment being made up of neutral lipids, which are covered by a phospholipid monolayer. In addition, LDs contain a set of surface proteins, many of which are lipid metabolism enzymes (Thiam et al., 2013; Walther et al., 2017; Olzmann and Carvalho, 2019).
Due to their key roles in lipid storage and metabolism, LDs are a cellular hub of lipid flux (Olzmann and Carvalho, 2019). To prevent lipotoxic effects and safeguard cellular integrity, lipid flow to and from LDs has to be performed in a controlled way. In recent years, it is emerging that material transfer between LDs and the other cellular organelles mainly depends on nonvesicular mechanisms mediated by contact sites (Goodman, 2008; Barbosa and Siniossoglou, 2017; Schuldiner and Bohnert, 2017; Olzmann and Carvalho, 2019), places of close organelle apposition (Scorrano et al., 2019). The molecular basis for the formation of these structures is provided by specialized proteins termed tethers (Eisenberg-Bord et al., 2016), which physically link the surfaces of distinct organelles and hold them at a close distance, generally a few tens of nanometers apart.
Never Let Me Go: Tether Proteins Are Key Players in Contact Sites
Identification and characterization of tether proteins is currently one of the key objectives of the contact site field, as knowledge of the identity of tethers is a prerequisite for experimental contact site manipulation. It has been found that multiple types of tether machineries exist (Eisenberg-Bord et al., 2016). Their unifying feature is the ability to associate with two distinct organelle surfaces at the same time. This is accomplished by transmembrane domains, lipid anchors, or by membrane-binding domains such as pleckstrin homology (PH), phox homology (PX), and C2 domains. Contact site tether machineries can be monomeric or oligomeric. Importantly, the term monomeric tether in this context does not imply that the tether protein would never associate with other proteins or even form a homooligomer. Instead, it means that all structural features required for attaching two organelles to each other are contained within one single polypeptide chain. Monomeric tether proteins are often anchored at one organelle via membrane integral domains and dock onto the partner organelle via membrane-binding domains. Examples for this type of tether are the extended synaptotagmins (Saheki and De Camilli, 2017) and some members of the oxysterol-binding protein-related protein (ORP) family (Tong et al., 2016). VAMP-associated protein (VAP) is an ER protein that is part of numerous heterodimeric tether complexes. It binds to two phenylalanines in an acidic tract (FFAT) as well as FFAT-like motifs present in diverse VAP-partner proteins, many of which contain membrane-binding domains specific for other organellar surfaces, such as ceramide transfer protein (CERT), StAR-related lipid transfer protein 3 (STARD3), protein tyrosine phosphatase interacting protein 51 (PTPIP51), and several ORP family members (Murphy and Levine, 2016). In addition to these VAP-containing heterodimeric complexes, structurally diverse oligomeric complexes exist that are formed by interaction of integral or surface proteins of distinct organelles, for example, the ER mitochondria encounter structure (ERMES) at ER–mitochondria interfaces (Kornmann et al., 2009), the nucleus vacuole junction (NVJ) at contacts between nuclear ER and vacuole (Pan et al., 2000), and the mitochondrial contact site and cristae organizing system (MICOS) that is involved in tethering the inner and outer mitochondrial membranes (Harner et al., 2011; Hoppins et al., 2011; von der Malsburg et al., 2011). Interestingly, most, if not all tether proteins have additional functions apart from tethering. Many tethers contain a domain with a hydrophobic cavity optimized for incorporation of lipids, which mediates lipid transfer between the tethered organelles (Wong et al., 2019). Such lipid transfer domains comprise synaptotagmin-like mitochondrial lipid-binding protein (SMP) domains, VAD1 analog of StAR-related lipid transfer (VASt) domains, ORP domains, and many more. Furthermore, some tethers, such as extended synaptotagmin 1, C2CD2L/TMEM24, and OSBP, are able to adapt their tethering activity to environmental stimuli such as changes in cytosolic calcium and membrane lipid levels (Giordano et al., 2013; Mesmin et al., 2013; Lees et al., 2017).
This structural and functional diversity of tethers has created the need for a clear definition of what exactly a tether is. It was therefore suggested that in order to be termed a tether, a protein/protein complex needs to fulfill three criteria that can be experimentally tested: (a) Tethers must be enriched or exclusively located at contact sites. (b) They must have the structural capacity to bind to two organelles at the same time. (c) And tethers must exert a tethering force (Eisenberg-Bord et al., 2016).
While tethers are a key topic in contact site research, the molecular basis of tethering of LDs to other organelles has been largely enigmatic until recently. Frequent observation of lipidic structures linking LDs to the ER, and occasionally to other organelles, has highlighted a potential contribution of lipidic tethers to LD contact site formation (Schuldiner and Bohnert, 2017). Organization and stability of such lipidic bridges at least partially depend on atypical proteinaceous contact site factors, such as the lipodystrophy factor seipin, a protein localized to LD–ER contacts that forms homooligomeric rings (Binns et al., 2010; Sui et al., 2018; Yan et al., 2018), which possibly reside within the lipidic bridge and which are required for their integrity (Grippa et al., 2015; Salo et al., 2016; Wang et al., 2016; Salo et al., 2019). Within the last years, it has become apparent that LD contact sites furthermore house proteins that fulfill the established criteria for typical LD tether proteins. These LD tether proteins will be described in detail in the following chapters.
Tethers at the LD–ER Interface
LD de novo biogenesis is a complex process that takes place at the ER membrane. According to the prevailing hypothesis, neutral lipids are synthesized by ER localized enzymes and initially released between the two leaflets of the ER membrane, where they form lipid lenses that ultimately bud toward the cytoplasm (Walther et al., 2017; Nettebrock and Bohnert, 2019). The resulting surface phospholipid monolayer of mature LDs often stays connected with the outer ER membrane leaflet via lipidic bridges (Ohsaki et al., 2008; Jacquier et al., 2011; Wilfling et al., 2013; Grippa et al., 2015; Wang et al., 2016).
Due to the key role of the ER in LD biogenesis, and the tight functional interplay of these two organelles in lipid metabolism, it is not surprising that a large number of proteins have been detected at LD–ER interfaces (Hugenroth and Bohnert, 2019). These comprise factors directly involved in the LD biogenesis process, factors required for maintenance and regulation of the unique lipidic continuity of the two organelles, and classical contact site proteins found in most types of contacts, such as tethers and lipid transfer proteins. Importantly, many LD–ER contact site proteins are multifunctional, for example, seipin is required for efficient LD biogenesis (Cartwright et al., 2015; Salo et al., 2016; Wang et al., 2016) but also involved in contact site maintenance (Fei et al., 2011; Wolinski et al., 2011; Grippa et al., 2015; Han et al., 2015; Wolinski et al., 2015; Salo et al., 2016; Salo et al., 2019), and Snx14 (see later) acts both as an LD biogenesis factor and as an LD–ER tether (Bohnert, 2019; Datta et al., 2019). A complete overview of all known residents of the LD–ER interface as well as a classification of these factors can be found in a recent review (Hugenroth and Bohnert, 2019). Here, we will primarily focus on those proteins that display the characteristics of canonical tether proteins described earlier and in Eisenberg-Bord et al. (2016).
Monomeric Tethers for Spatial Organization of Organelles and Enzymes
The yeast protein Mdm1 has originally been identified as a component of the NVJ, a large contact site between the nucleus and the vacuole (yeast lysosome-like organelle; Henne et al., 2015). Intriguingly, Mdm1 is not uniformly distributed within the organelle contact area marked by the NVJ tether complex Nvj1-Vac8, but instead enriched on its outer rim (Hariri et al., 2018). Mdm1 has two N-terminal transmembrane domains that anchor the protein in the ER membrane. A Phox homology associated (PXA), a regulator of G-protein signaling (RGS), a Phox homology (PX), and a C-terminal Nexin domain are exposed to the cytosol (Bohnert, 2019; Figure 1a). The PX domain interacts with phosphatidylinositol 3-phosphate, which is enriched in the vacuolar membrane, thus mediating nucleus–vacuole tethering (Henne et al., 2015). Interestingly, the NVJ acts as an LD biogenesis hot spot under nutrient stress conditions (Hariri et al., 2018). Pronounced LD biogenesis at the NVJ coincides with other nutrient stress induced processes, including an expansion of the NVJ (Roberts et al., 2003; Hariri et al., 2018) and recruitment of several facultative NVJ resident proteins, specifically the putative lipid transfer proteins Vps13 and Nvj2 and the phosphatidic acid phosphatase Pah1 (Toulmay and Prinz, 2012; Barbosa et al., 2015; Lang et al., 2015). Mdm1 co-localizes with LDs at the rim of the NVJ (Hariri et al., 2018), and artificial relocalization of the protein to the cortical ER results in aberrant LD biogenesis in this ER region (Hariri et al., 2019), indicating that Mdm1 has a role in LD biogenesis. Consistently, lack of Mdm1 results in inefficient incorporation of fatty acids into TAG, delayed LD de novo biogenesis, and increased sensitivity to lipotoxic fatty acids (Hariri et al., 2019). Hariri et al. (2019) generated a set of Mdm1 mutants and found that the PXA domain has multiple roles in Mdm1 function. First, the PXA domain contains a hydrophobic helix that localizes to LDs when expressed separately and mediates together with the N-terminal membrane integral domain association with LDs. This indicates that Mdm1 does not only bridge two organelles, but instead mediates formation of organellar three-way junctions consisting of ER, vacuole, and LDs. Second, the PXA domain binds to free fatty acids in vitro, which together with the observed defect in TAG biogenesis of mdm1 mutants points toward a link to conversion of fatty acids into neutral lipids. And third, it interacts with the fatty acyl-CoA synthetase Faa1 (Figure 1a). Mdm1 thus combines tri-organellar tethering with the spatial organization of an enzymatic activity (Hariri et al., 2019).
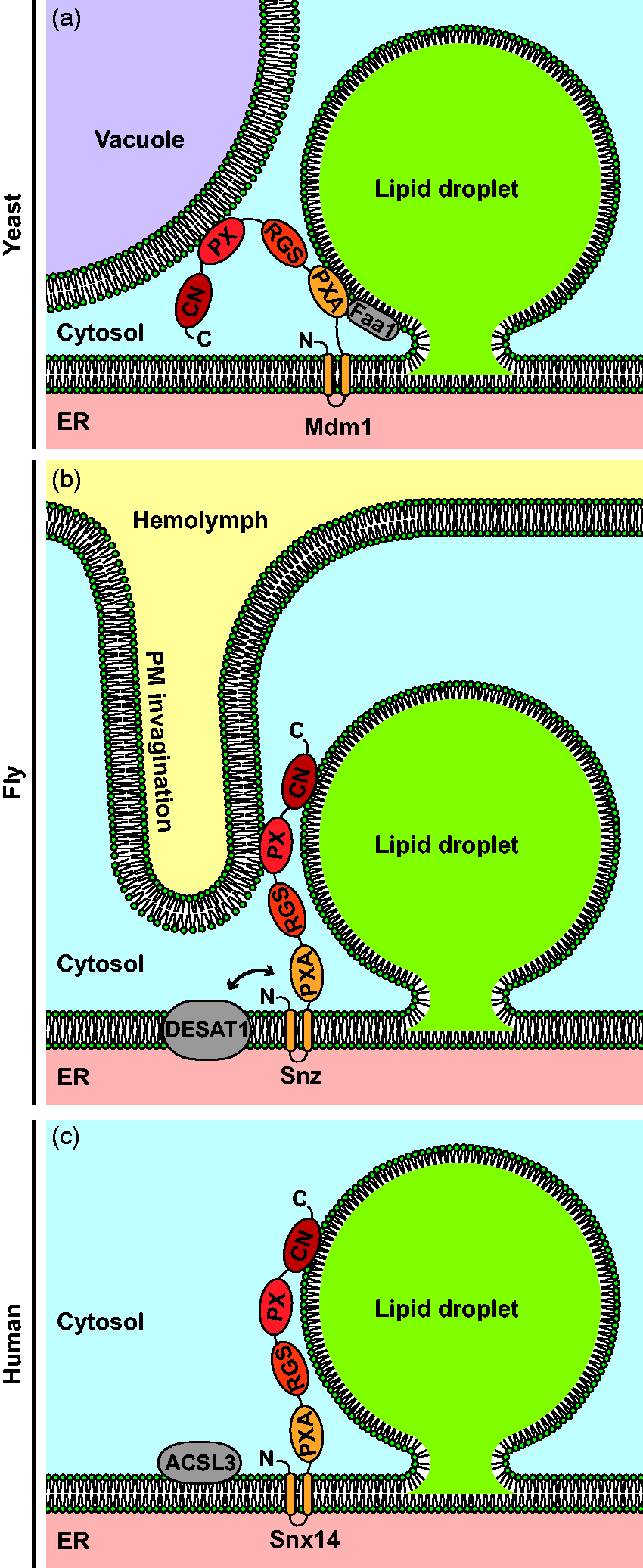
Sorting Nexin Proteins Mediate Spatial Organization Beyond LD–ER Contact Formation. Panel A: Yeast Mdm1 forms tri-organellar lipid droplet (LD)–ER–vacuole contacts and interacts with the fatty acyl-CoA synthetase Faa1. Panel B: Drosophila melanogaster Snazarus (Snz) mediates LD–ER–plasma membrane (PM) contacts and interacts with the stearoyl-CoA desaturase DESAT1. Panel C: In humans, sorting nexin 14 (Snx14) colocalizes with the fatty acyl-CoA synthetase ACSL3 at LD–ER contact sites. All three sorting nexin proteins share the same domain structure, consisting of N-terminal transmembrane segments, followed by phox homology associated (PXA), regulator of G-protein signaling (RGS), phox homology (PX), and C-terminal nexin (CN) domains. ER = endoplasmic reticulum.
Snazarus is an Mdm1 homolog in Drosophila melanogaster and the two proteins also share key functional aspects, albeit in different subcellular contexts. Like Mdm1, Snazarus is an integral ER protein, and the two proteins have the same domain architecture (Suh et al., 2008; Henne et al., 2015; Figure 1b). Similar to Mdm1, Snazarus binds to LDs, although via a different domain than Mdm1, namely, the C-terminal Nexin domain (Hariri et al., 2019; Ugrankar et al., 2019). Intriguingly, unlike most PX domains, the PX domain of Snazarus does not bind to phosphatidylinositol 3-phosphate, the signature lipid of the endolysosomal compartment. Instead, the isolated Snazarus PX domain binds to liposomes containing the plasma membrane phospholipids phosphatidylinositol 3,4-bisphosphate and phosphatidylinositol 4,5-bisphosphate, as well as other polyphosphorylated phosphatidylinositol variants and posphatidylserine (Ugrankar et al., 2019). Consistently, the Snazarus PX domain lacks the conserved canonical phosphatidylinositol 3-phosphate-binding motif and instead comprises a noncanonical putative binding site for polyphosphorylated phosphatidylinositol variants (Chandra and Collins, 2018; Chandra et al., 2019; Ugrankar et al., 2019). In vivo, efficient formation of Snazarus foci at the plasma membrane depends on the phosphatidylinositol 4,5-bisphosphate synthesizing enzyme dPIP5K, indicating that Snazarus binds to plasma membrane phosphatidylinositol variants via its PX domain. In summary, Snazarus might form a tri-organelle contact composed of ER, LD, and plasma membrane, similarly to the role of Mdm1 at ER–LD–vacuole interfaces (Ugrankar et al., 2019; Figure 1a and b).
Snazarus is highly expressed in cells of the fat body, which is the dedicated fat storage tissue in fly (Suh et al., 2008; Ugrankar et al., 2019). These cells contain two structurally and functionally distinct subpopulations of LDs: large LDs located in the cell’s center, and tiny peripheral LDs sitting right under the plasma membrane (PM). Snazarus foci colocalize with plasma membrane invaginations that tightly wrap around the small peripheral LDs (Ugrankar et al., 2019; Figure 1b). Intriguingly, these peripheral LDs appear to be generated mainly from lipids derived from lipophorin particles, which transport lipids from gut enterocytes to different organs (Canavoso et al., 2001), while biogenesis of the large central LDs depends predominantly on de novo lipogenesis mediated by the fatty acid synthase FASN1 (Ugrankar et al., 2019). Snazarus deletion results in disorganization of the peripheral LD pool. Nonetheless, Snazarus does not appear to be crucial for LD–PM interaction, as wrapping of PM invaginations around LDs still occurs in its absence (Ugrankar et al., 2019). Snazarus overexpression results in elevated TAG levels and in an enlargement of the fat body peripheral LDs. Interestingly, Snazarus is physically linked to several fatty acid handling enzymes, among them the stearoyl-CoA desaturase DESAT1 (Figure 1b). In the absence of DESAT1, increased TAG formation in response to Snazarus overexpression does not occur (Ugrankar et al., 2019). Thus, similarly to Mdm1, Snazarus links tri-organellar tethering to positioning of a lipid metabolism enzyme, a functional combination that could be important for incorporation of extracellular lipids into peripheral LDs (Hariri et al., 2019; Ugrankar et al., 2019).
Snx14 is a human Mdm1/Snazarus homolog. It shares the basic domain structure of both Mdm1 and Snazarus with N-terminal transmembrane domains, followed by a PXA, RGS, PX, and a C-terminal Nexin domain (Bohnert, 2019; Figure 1c). Snx14 displays all characteristic features of an LD–ER tether protein (Datta et al., 2019) as defined in (Eisenberg-Bord et al., 2016; and earlier). (a) Defined location at the contact site: Snx14 localizes to the LD–ER interface, although not under all metabolic conditions. In the absence of exogenous fatty acids in the growth medium, Snx14 localizes dispersed throughout the ER membrane, while it forms foci at LD interfaces in the presence of fatty acids (Datta et al., 2019). (b) Structural capacity for tethering two organelles: Using truncation constructs, it was shown that Snx14 has the structural features required to mediate LD–ER tethering. ER binding was mapped to the N-terminal hydrophobic domains, while LD binding depends on an amphipathic helix located within the C-terminal Nexin domain (Datta et al., 2019). (c) Tethering activity: Finally, loss of Snx14 results in reduced LD–ER contacts, while overexpression leads to enhanced contacts, showing that the protein has tethering activity (Datta et al., 2019). Unlike for Mdm1 and Snazarus, there are, however, no indications that Snx14 mediates a tri-organellar contact site. This is consistent with previous findings that Snx14 has an altered PX domain with a closed phosphatidylinositol 3-phosphate-binding pocket that does not show membrane-binding in vitro (Mas et al., 2014; Bryant et al., 2018), indicating that this protein lacks a third membrane-binding domain.
Snx14 overexpression promotes fatty acid-dependent LD biogenesis, while it has no visible effects on LDs in the absence of fatty acids. Unlike for Mdm1 and Snazarus, a physical interaction of Snx14 with lipid metabolism enzymes has to date not been reported, but Snx14 is functionally linked to ACSL3, a fatty acid-CoA synthetase (Figure 1c). ACSL3 is an early marker for growing LDs (Kassan et al., 2013), and during fatty acid induced LD biogenesis, Snx14 gets recruited to these ACSL3 foci. In the absence of ACSL3, however, Snx14 stays dispersed in the ER even in the presence of fatty acids, indicating that Snx14 affects LD formation in an ACSL3-dependent manner (Datta et al., 2019).
In summary, this group of sorting nexin proteins is involved in LD tethering to the ER and further partner organelles and combines this with mediating spatially confined enzymatic activities.
VAP-Dependent LD–ER Tethers
VAP proteins are integral ER proteins that act as receptors for many different proteins on the cytosolic ER surface (Murphy and Levine, 2016). Among other functions, VAPs have important roles in ER contact sites.
VPS13A and VPS13C are VAP-interactors that localize at contacts between the ER and LDs. Both proteins are also found at other organelle interfaces: VPS13A at ER–mitochondria contacts (Kumar et al., 2018; Yeshaw et al., 2019) and VPS13C at ER-lysosome/late endosome contacts (Kumar et al., 2018; Figure 2). Mutations in VPS13A are linked to chorea acanthocytosis (Rampoldi et al., 2001; Ueno et al., 2001), while VPS13C is connected to early-onset Parkinson’s disease (Lesage et al., 2016). Both VPS13A and C are large multidomain proteins, consisting of an N-terminal Vps13α domain, followed by putative WD40, Dbl homology (DH)-like and PH domains (Kumar et al., 2018; Figure 2). The Vps13α domain of VPS13C is more than 500 amino acids larger than the one in VPS13A due to a putative internal sequence duplication. Both Vps13α domains contain a FFAT motif, which is crucial for VAP-mediated binding to the ER (Kumar et al., 2018; Yeshaw et al., 2019).

VPS13 Proteins Localize to Multiple Contact Sites. VPS13A (left) localizes to lipid droplet (LD)–ER and mitochondria–ER contacts, while VPS13C (right) resides in LD–ER and lysosome–ER contact sites. Both VPS13 proteins consist of an N-terminal Vps13α domain, followed by a WD40, DH-like and PH domain. The Vps13α domain of VPS13C is approximately 500 amino acids larger than the one in VPS13A. Both VPS13 proteins are bound to the ER via interaction of their FFAT motifs with the ER anchor VAP. ER = endoplasmic reticulum; PH = pleckstrin homology; DH = Dbl homology; FFAT = two phenylalanines in an acidic tract.
VPS13A and VPS13C function as organelle tethers and very likely also as interorganellar lipid transfer proteins. VPS13A depletion results in a reduction of ER–mitochondria contacts as well as in mitochondrial and LD abnormalities, particularly mitochondrial fragmentation and an increase in LD numbers and motility (Yeshaw et al., 2019). VPS13A overexpression leads to reduced LD motility (Yeshaw et al., 2019) and enlarged ER–mitochondria contacts (Kumar et al., 2018), while co-overexpression of VPS13C and VAP results in endosomes that are almost completely wrapped in ER membranes (Kumar et al., 2018). Thus, VAP-VPS13 complexes appear to have the ability to induce organelle contacts, a feature characteristic for tethers. In the absence of VAP or upon mutation of the FFAT motif, VPS13A gets lost from the ER and decorates the surface of mitochondria and LDs uniformly. Vps13α domains expressed alone localize to the ER, while the same proteins with mutated FFAT motive are dispersed in the cytosol. The WD40-like domain of VPS13A was detected in the cytosol, while the respective VPS13C domain binds to lysosomes and endosomes (Kumar et al., 2018). LD as well as mitochondrial binding of VPS13A has been mapped to the C-terminal DH-like and putative PH domain (Kumar et al., 2018; Yeshaw et al., 2019). An amphipathic helix within this region localizes to LDs when expressed on its own. LD surface binding via amphipathic helices has also been detected for some sorting nexin LD tethers (see earlier) and for many LD surface proteins (Rowe et al., 2016; Prévost et al., 2018).
There are manifold indications that VPS13 proteins transport lipids at contact sites. Yeast Vps13 has not been detected at LD contact sites so far, and its partner proteins differ from those of the human proteins. Nonetheless, it is also a contact site protein and localizes to contacts between the vacuole and mitochondria as well as the nuclear ER (Lang et al., 2015). The Vps13α domain of yeast Vps13 has a rod-like shape according to negative stain electron microscopy (Kumar et al., 2018). This domain has been found to simultaneously bind to approximately 10 glycerophospholipid molecules, with a preference for phosphatidylcholine. A Vps13α domain fused to a synthetic membrane tethering module was found to transport glycerophospholipids between tethered liposomes in vitro. The crystal structure of an N-terminal fragment of this domain from Chaetomium thermophilum has been solved. This fragment forms a novel type of fold with a hydrophobic cavity with larger dimensions than typical lipid transfer domains. According to structure predictions, Vps13α likely contains several such lipid binding/transfer modules or possibly consists of one very long lipid transfer tube (Kumar et al., 2018). The latter hypothesis is consistent with the described rod-shape of yeast Vps13α. VPS13 thus might act as a lipid transfer protein for a special type of bulk lipid transfer, similar to the autophagy protein ATG2, which has structural similarities to VPS13 and mediates lipid transport from the ER to the growing autophagosome (Valverde et al., 2019).
A further protein that could potentially form a VAP-dependent LD–ER bridge in humans is the oxysterol-binding protein-related protein 2 (ORP2). Numerous functions have been proposed for this protein, for example, roles in lipid metabolism, lipid transfer, steroid hormone biogenesis, cytoskeleton organization, and signaling (Olkkonen et al., 2019). Likewise, ORP2 has been detected in various subcellular locations. In addition to its cytosolic pool, ORP2 has been detected on LDs (Hynynen et al., 2009), at LD–ER contact sites (Kentala et al., 2015; Weber-Boyvat et al., 2015), at late endosomes (Koponen et al., 2019), and at the cell cortex (Hynynen et al., 2009; Kentala et al., 2015, 2018; Wang et al., 2019).
While some ORPs are multidomain proteins with several membrane-binding modules, ORP2 has a rather simple architecture, consisting of a FFAT motive at the very N-terminus, followed by an oxysterol-binding domain. The structures of several oxysterol-binding domains have been solved (Im et al., 2005; Tong et al., 2013), including the one of ORP2 (Wang et al., 2019). This domain contains a hydrophobic cavity that can bind to lipids. Binding to different types of phosphoinositides and sterols has been experimentally demonstrated (Xu et al., 2001; Hynynen et al., 2005; Suchanek et al., 2007; Hynynen et al., 2009; Koponen et al., 2019; Wang et al., 2019). In general, ORPs are bispecific lipid transfer proteins that mediate counter-transport of sterols and phosphoinositides. In a recent study, it was shown that ORP2 delivers cholesterol to the plasma membrane and removes phosphatidylinositol 4,5-bisphosphate from this membrane in a process that is independent from the FFAT motif. Localization to the plasma membrane depends on binding to phosphatidylinositol 4,5-bisphosphate, likely via positively charged residues on the surface of the protein (Wang et al., 2019). In the presence of a high-affinity oxysterol variant, ORP2 recruitment to the plasma membrane is strongly enhanced (Hynynen et al., 2009; Wang et al., 2019), probably because oxysterol binding abolishes lipid transport, resulting in elevated phosphatidylinositol 4,5-bisphosphate levels at the plasma membrane. Under the same condition, wild-type ORP2, but not an ORP2 mutant defective in oxysterol binding, is lost from LDs (Hynynen et al., 2009).
As ORP2 can bind to the LD surface and its interactor VAP is an ER protein, ORP2-VAP complexes theoretically could have the structural capacity to tether LDs to the ER. Nonetheless, there is currently no clear evidence for such a function. Loss of ORP2 has in fact been shown to lead to an increase rather than a decrease in LD–ER contact. On the other hand, contact expansion in response to exposure to fatty acids was found to be abolished in the absence of ORP2 (Kentala et al., 2018). This points toward a regulatory rather than a tethering function of ORP2 at the LD–ER contact site. The exact role of ORP2-VAP in LD–ER interaction remains to be determined.
Oligomeric Tethers at LD–ER Interfaces
There are further examples of protein complexes that can bridge the cleft between the ER and adjacent LDs. Diacylglycerol acyltransferase 2 (DGAT2) located on the LD surface has been found to physically interact with the fatty acyl-CoA synthetase FATP1 on the ER. Interaction of these two lipid metabolism enzymes supports LD growth, by mediating localized TAG synthesis at the LD–ER interface (Xu et al., 2012).
The small GTPase Rab18 has been detected both on LDs (Martin et al., 2005; Ozeki et al., 2005; Liu et al., 2007; Pulido et al., 2011; Li et al., 2017; Bersuker et al., 2018; Xu et al., 2018) and on the ER (Gerondopoulos et al., 2014). Rab18 depletion leads to a reduced number and abnormal expansion of LDs in adipocytes and Leydig cells (Xu et al., 2018), and Rab18 overexpression induces pronounced LD–ER contacts (Ozeki et al., 2005). In its GTP-bound form, Rab18 interacts with the C-terminus of ZW10, a component of the NRZ (NAG, RINT1, ZW10) multisubunit tethering complex (Gillingham et al., 2014; Xu et al., 2018). Crystal structures of a homologous complex from yeast indicate that the NRZ complex forms an elongated structure consisting of numerous short α-helices (Ren et al., 2009). LD–ER tethering is mediated by interaction of the NRZ complex with the ER-localized SNAREs Syntaxin18, Use1, and BNIP1 (Xu et al., 2018). Functionally, Rab18 is required for efficient LD formation in some (Xu et al., 2018; Salo et al., 2019), but not in all cell types (Jayson et al., 2018). A further factor that is physically connected to Rab18 and that affects LD–ER contacts is the protein double FYVE-containing protein 1 (DFCP1; Gao et al., 2019; Li et al., 2019). LD–ER contacts respond to changes in DFCP1 levels, with loss of DFCP1 leading to reduced, and overexpression resulting in increased contacts (Li et al., 2019). Loss of DFCP1 also leads to further LD alterations, namely, an increased LD number and a decreased average size (Gao et al., 2019; Li et al., 2019). In the presence of oleic acid, DFCP1 localizes to LD precursors associated with the ER, which fuse to form expanding LDs. The effects of DFCP1 and Rab18 on LDs depend on each other. Efficient targeting of Rab18 to LDs requires the presence of DFCP1 and vice versa. Similarly, efficient expansion of LD–ER contacts in response to overexpression of either component depends on the presence of the other protein (Li et al., 2019). In summary, Rab18, NRZ, SNAREs, and DFCP1 might cooperate to mediate LD–ER contacts in a cell type-specific manner.
LD Tethering to Lysosomes
In yeast, LD–organelle interactions show a growth phase-dependent dynamic behavior that includes extensive association with the lysosome-like vacuole. LDs are dispersed across the ER surface in logarithmic growth phase, but then cluster adjacent to the NVJ in the beginning of stationary phase (Wang et al., 2014; Barbosa et al., 2015; Eisenberg-Bord et al., 2018; Hariri et al., 2018). The tri-organellar LD–ER–vacuole tether Mdm1 (see earlier) mediates a burst of spatially confined LD biogenesis at the nucleus–vacuole interface (Hariri et al., 2018, 2019). Mdm1 binds to LDs via its N-terminal transmembrane and PXA domains (Hariri et al., 2019) and to the vacuole via its PX domain (Henne et al., 2015), thus acting as an LD–vacuole tether (Hariri et al., 2019; Figure 1a). A further factor involved in LD clustering at the NVJ is the lipid droplet organization (LDO) machinery consisting of Ldo16 and Ldo45. In the absence of LDO, LD clustering at the NVJ is compromised (Eisenberg-Bord et al., 2018). The exact functional role of the LDO machinery remains to be determined, but it has been found to interact with the LD biogenesis factor seipin (Eisenberg-Bord et al., 2018; Teixeira et al., 2018). Promethin/LDAF1 is a human protein with structural similarity to LDO that also interacts with seipin (Castro et al., 2019; Chung et al., 2019). Promethin/LDAF1 is directly involved in the process of LD biogenesis (Chung et al., 2019), pointing toward a potential role of LDO in spatially confined LD biogenesis at the NVJ. During progression into late stationary phase, LDs do not stay clustered at the NVJ, but instead again change their localization. They move away from the NVJ and onto the surface of the vacuole (van Zutphen et al., 2014; Wang et al., 2014). LDs form tight contact sites with the vacuolar membrane and remain attached until they ultimately are taken up into the vacuole by microlipophagy (van Zutphen et al., 2014; Wang et al., 2014). The tether proteins that mediate this tight LD–vacuole attachment are currently unknown. Intriguingly, the vacuolar membrane segregates into sterol-rich and sterol-poor domains around the same time (Toulmay and Prinz, 2013; Wang et al., 2014), with the LDs being bound to the sterol-rich membrane domains (Wang et al., 2014). Intriguingly, loss of the LDO machinery results in impaired membrane segregation as well as a defect in lipophagy (Teixeira et al., 2018).
In mammalian cells, LD–lysosome proximity has also been observed (Kaushik and Cuervo, 2015; Schroeder et al., 2015; Valm et al., 2017), but their molecular basis is currently poorly understood. Functionally, LD–lysosome association has been implicated in LD consumption. The multifunctional small GTPase Rab7 is required for efficient nutrient stress-induced LD–lysosome association and lipophagy (Schroeder et al., 2015). Perilipins 2 and 3 have been identified as substrates of chaperone-mediated autophagy (CMA; Kaushik and Cuervo, 2015), a process in which proteins containing a special pentapeptide motif are recognized by the heat shock protein Hsc70 and transferred to the lysosome-associated membrane protein 2A (LAMP-2A) for translocation into the lysosomal lumen (Kaushik and Cuervo, 2018). A block in CMA-dependent perilipin degradation results in reduced LD binding of adipose triglyceride lipase and of autophagy factors and thus in impaired LD consumption both by lipolysis and by lipophagy. Mutation of the perilipin 2 CMA motif was found to lead to decreased LD–lysosome contacts, indicating that the proteins involved in CMA could possibly form a complex to physically bridge the surfaces of the two organelles, but the molecular details underlying this observation remain to be determined (Kaushik and Cuervo, 2015).
LD Tethering to Mitochondria and Peroxisomes
Contact sites between fatty acid storing LDs and fatty acid oxidizing mitochondria as well as peroxisomes might form the basis for highly efficient energy conversion units. Recent studies are starting to uncover the molecular bases and the functions of these contacts.
Perilipins and LD–Mitochondria Contacts
LDs and mitochondria are important players in energy metabolism. Both organelles presumably utilize contact sites rather than vesicle trafficking as main route of interorganellar communication (Eisenberg-Bord and Schuldiner, 2017; Schuldiner and Bohnert, 2017). Contact site-based LD–mitochondria interactions are extensive and appear to serve different metabolic functions. LD–mitochondria contacts are particularly abundant in heart as well as skeletal muscle cells, hepatocytes, and in brown adipose tissue, cell types with a high energy demand (Vock et al., 1996; Tarnopolsky et al., 2007; Shaw et al., 2008; Shiozaki et al., 2011; Wang et al., 2013; Boutant et al., 2017). These contacts have been found to expand under starvation conditions (Herms et al., 2015; Rambold et al., 2015; Nguyen et al., 2017; Valm et al., 2017). In skeletal muscle, LD–mitochondria contacts grow in response to endurance training (Tarnopolsky et al., 2007; Shaw et al., 2008). Mitochondria mediate the breakdown of fatty acids by β-oxidation, TCA cycle, and oxidative phosphorylation to harvest their energy. LDs store fatty acids in the form of neutral lipids and indications for direct handover of fatty acids from LDs to mitochondria under starvation conditions have been reported. This process has been found to depend on lipolysis, on LD refueling by autophagy, and on functional mitochondrial dynamics, specifically on the fusion machinery (Rambold et al., 2015). It is conceivable that fatty acid channeling from LDs to mitochondria might be an efficient and safe way to drive lipid-dependent ATP production while preventing lipotoxic effects. On the other hand, LD–mitochondria contacts have also been connected to LD expansion (Nguyen et al., 2017; Benador et al., 2018). Under starvation conditions, autophagy-derived fatty acids were found to be channeled into newly formed LDs in close contact to mitochondria, a process that protected mitochondria from lipotoxic effects (Nguyen et al., 2017). In brown adipose tissue, it was found that the subpopulation of mitochondria in contact with LDs displayed fundamental metabolic differences to other mitochondria: mitochondria tethered to LDs produced ATP via oxidation of pyruvate, but displayed low fatty acid oxidation. In contrast, mitochondria not physically connected to LDs had higher β-oxidation activity. Furthermore, association with mitochondria resulted in LD expansion, while cold exposure-induced thermogenesis via β-oxidation co-occurred with loss of LD–mitochondria contact. Collectively, this implies that physical contact with mitochondria can support LD growth, potentially by contributing ATP required for fatty acid activation into fatty acyl-CoA for TAG synthesis (Benador et al., 2018). Consistent with these indications for roles of LD–mitochondria contacts beyond fatty acid channeling, yeast mitochondria, which do not perform β-oxidation, also form contact sites with LDs (Shai et al., 2018). Future work will be required to untangle these different observations on the biological roles of LD–mitochondria interactions.
The molecular machineries underlying contact sites between LDs and mitochondria are only partially understood. Depletion of the SNARE protein SNAP23 results in decreased LD–mitochondria contact and defective β-oxidation (Jägerström et al., 2009). Furthermore, the long chain acyl-CoA synthetase ACSL1 located on the mitochondrial surface was found in close proximity to SNAP23 (Young et al., 2018), but the biological role of this interplay is unclear.
In brown adipose tissue, a potential tethering complex has been reported that consists of perilipin 1 and mitofusin 2 (MFN2; Boutant et al., 2017; Figure 3a). Perilipin 1 is an LD surface protein, while MFN2 is mainly located to the mitochondrial outer membrane. Together with its homolog MFN1, MFN2 mediates mitochondrial fusion (Mattie et al., 2019). MFN2 has a cytosolic N-terminal GTPase domain followed by a heptad repeat, a transmembrane domain that anchors the protein in the outer mitochondrial membrane, and another heptad repeat domain at the C-terminus (Mattie et al., 2018). In addition to its role in mitochondrial fusion, MFN2 is also involved in mitochondrial contact sites. An ER localized MFN2 pool mediates ER–mitochondria tethering via interaction with mitochondrial mitofusins (de Brito and Scorrano, 2008). A peroxisomal pool of the yeast mitofusin Fzo1 is involved in the contact site between mitochondria and peroxisomes (Shai et al., 2018). Boutant et al. (2017) found that in brown adipose tissue, MFN2 co-precipitates perilipin 1, particularly under lipolytic conditions. Furthermore, in the absence of MFN2, LD–mitochondria contacts were found to be reduced, consistent with an LD–mitochondria tethering function (Boutant et al., 2017). However, due to the broad impact of MFN2 on mitochondrial morphology, it remains possible that these effects on LD contacts are indirect.

LD Tethering to Mitochondria and Peroxisomes. Perilipin (PLIN) 1 (A) and 5 (B) are lipid droplet (LD) surface proteins that affect communication with mitochondria and have been implicated in LD–mitochondria contact sites. Perilipin 1 has been suggested to interact with mitochondrial MFN2 in brown adipocytes. The mitochondrial binding partner of perilipin 5 is unknown. Panel C: MIGA2 mediates LD–mitochondria contacts in white adipocytes. MIGA2 is anchored in the mitochondrial outer membrane by two transmembrane segments, and binds to the LD surface via a C-terminal amphipathic helix (red). Its middle domain (MD) contains a FFAT motif that can interact with the ER protein VAP (not shown). Panel D: Spastin and ABCD1 link LDs to peroxisomes. Spastin has been detected on the LD surface. It is bound to LDs via an N-terminal hydrophobic domain. Its cytoplasmic domain consists of a microtubule interacting and trafficking (MIT) domain, a peroxisome-interacting (PXI) region, and a C-terminal AAA-ATPase domain. Spastin mediates LD–peroxisome bridging via binding of the PXI region to the peroxisomal ABC transporter ABCD1.
A further member of the perilipin family, perilipin 5, has an established key role in the interplay between LDs and mitochondria. High levels of perilipin 5 induce LD–mitochondria contacts, and this effect depends on the proteins C-terminal 20 amino acids (Wang et al., 2011; Benador et al., 2018). Perilipin 5 has been detected at LD–mitochondria interfaces by superresolution microscopy, further supporting a potential tethering role (Gemmink et al., 2018). Perilipin 5 is predominantly expressed in oxidative tissues (Wolins et al., 2005; Yamaguchi et al., 2006; Dalen et al., 2007). In muscle, it is upregulated in response to training (Peters et al., 2012; Louche et al., 2013; Vigelso et al., 2016). While Perilipin 5 is clearly somehow involved in LD–mitochondria association, it is currently unclear how this protein may be attached to the mitochondrial outer membrane (Figure 3b).
MIGA2, a Novel LD–Mitochondria Tether
Perilipin 5, likely the most heavily studied protein in the interplay between LDs and mitochondria, is mostly absent from white adipocytes (Wolins et al., 2006), but nonetheless, abundant LD–mitochondria contact sites have been observed also in this cell type (Novikoff, 1980; Cohen et al., 2004). Recently, mitoguardian 2 (MIGA2), a protein previously connected to mitochondrial fusion (Zhang et al., 2016), has been identified as a novel player in LD–mitochondria contacts in white adipocytes (Freyre et al., 2019). MIGA2 had been connected to fat storage before, as MIGA2 knockout mice show a strong reduction in body fat mass (Bassett et al., 2012; Podrini et al., 2015; Zhang et al., 2016). Interestingly, MIGA2 mRNA and protein levels were found upregulated during the process of adipocyte differentiation (Freyre et al., 2019). Artificial overexpression of MIGA2 in adipocytes led to pronounced contact site formation between mitochondria and LDs, as well as between mitochondria and the ER. Extensive LD–mitochondria interaction was also observed when MIGA2 was expressed in COS7 cells, which normally display little LD–mitochondria contacts and usually also do not express MIGA2, indicating that MIGA2 might be able to mediate contact formation independent of other factors (Freyre et al., 2019). Indeed, a structure-function analysis of MIGA2 revealed that it has the molecular features of a monomeric LD–mitochondria tether (Figure 3c). MIGA2 consists of two N-terminal transmembrane segments, a middle domain, and a C-terminal amphipathic domain. The N-terminal transmembrane segments anchor the protein in the mitochondrial outer membrane. LD binding depends on the amphipathic domain at the protein’s C-terminus (Freyre et al., 2019). Amphipathic helices are typical LD surface-binding motifs (Rowe et al., 2016; Prévost et al., 2018) that also mediate LD binding of several LD tethers (Kumar et al., 2018; Datta et al., 2019). Intriguingly, MIGA2 constructs lacking the transmembrane segments required for association with mitochondria were found to be unable to bind to LDs despite the presence of the amphipathic LD-binding domain (Freyre et al., 2019). In contrast, a synthetic construct consisting of the cytoplasmic domain of MIGA2 fused to the C-terminus of the mitochondrial outer membrane protein TOM20 efficiently induced LD–mitochondria association in a manner dependent on the C-terminal LD-binding domain, similar to native MIGA2. Furthermore, a fusion protein consisting of the ER membrane protein calnexin and the LD-binding domain of MIGA2 induced LD–ER association. Taken together, this suggests that LD binding of the amphipathic helix of MIGA2 depends on association of MIGA2 with a partner organelle (Freyre et al., 2019). Interestingly, the middle domain of MIGA2 contains a FFAT motif (Murphy and Levine, 2016), which mediates interaction with the ER protein VAP (Freyre et al., 2019). MIGA2 thus acts as a contact site tether and coordinates three organelles, mitochondria, LDs, and the ER. Functionally, preadipocytes lacking MIGA2 showed a defect in TAG synthesis, LD expansion, and in differentiation into mature adipocytes. Importantly, feeding of differentiating preadipocytes with radiolabeled glucose revealed that MIGA2 knockout cells were unable to convert glucose into TAG. MIGA2-dependent organelle coupling might thus be involved in coordinating de novo lipogenesis (Freyre et al., 2019). MIGA2 is present in diverse tissues (Bassett et al., 2012), and its role in organelle communication in cell types other than white adipocytes will be an interesting topic for the future.
More broadly, it is now clear that LD–mitochondria contact sites can serve diverse functions. The type of LD–mitochondria interplay appears to differ between distinct cell types and may also vary dependent on the exact metabolic conditions. It will be interesting to assess how LD–mitochondria tethering is regulated during cellular differentiation processes, such as adipocyte browning (Bartelt and Heeren, 2014).
Spastin Mediates LD–Peroxisome Contact Sites
A further organelle that mediates fatty acid oxidation is the peroxisome. Similar to mitochondria, contact sites to LDs that could mediate direct fatty acid handover have been frequently observed (Novikoff et al., 1973; Novikoff and Novikoff, 1973; Novikoff, 1980; Hayashi et al., 2001; Schrader, 2001; Binns et al., 2006; Thazar-Poulot et al., 2015), but the tether machineries underlying their formation have long been enigmatic. Chang et al. (2019) have recently described a tether complex consisting of LD-localized M1-spastin and the peroxisomal membrane fatty acid transporter ATP-binding cassette subfamily D member 1 (ABCD1; Figure 3d).
Spastin is a disease component frequently mutated in autosomal-dominant hereditary spastic paraplegia. The Spastin protein has been detected in numerous subcellular locations (Reid et al., 2005; Connell et al., 2009; Park et al., 2010), among them the LD surface (Papadopoulos et al., 2015). Functionally, Spastin has been mainly connected to microtubule dynamics and membrane remodeling (Lumb et al., 2012). M1-Spastin is a multidomain protein, consisting of an N-terminal hydrophobic hairpin motif, followed by a microtubule interacting and trafficking (MIT) domain, a peroxisome-interacting (PXI) region, and a AAA-ATPase domain (Chang et al., 2019). The hydrophobic hairpin can directly insert into the LD phospholipid monolayer, while M87-Spastin, a variant derived from an alternative start codon that lacks the hairpin (Claudiani et al., 2005), does not localize to LDs. The PXI domain targets to peroxisomes when expressed alone. Overexpression of M1-, but not M87-Spastin, results in the formation of extensive LD–peroxisome contact sites in a microtubule-independent way, while Spastin knockdown results in loss of contact. Interestingly, ATPase deficient Spastin mutants still localize to LDs but are unable to induce LD–peroxisome tethering (Chang et al., 2019).
ABCD1 is a peroxisomal ABC transporter. Mutations in ABCD1 are connected to X-linked adrenoleukodystrophy (Morita and Imanaka, 2012). Upon loss of ABCD1, expansion of LD–peroxisome contacts by Spastin overexpression as well as in response to experimental lipid peroxidation is abolished. ABCD1 coprecipitates together with Spastin and also with its isolated PXI domain, while ATPase mutants of Spastin coisolate lower amounts of ABCD1 (Chang et al., 2019). Interestingly, a pathogenic Spastin K388R point mutant (Blackstone, 2018) exerts a dominant negative effect on LD–peroxisome interaction and results in increased LD peroxidation. Pulse-chase experiments with a fluorescent fatty acid analog indicate that fatty acid transfer from LDs to peroxisomes is affected upon knockdown of both ABCD1 and Spastin, while Spastin overexpression results in enhanced transfer (Chang et al., 2019). The Spastin MIT domain binds to two different ESCRT III proteins IST1 and CHMP1B (Reid et al., 2005; Yang et al., 2008; Agromayor et al., 2009; Renvoisé et al., 2010). Recruitment of IST1 and CHMP1B to LDs is dispensable for Spastin-dependent contact formation, but fatty acid transfer is affected, indicating that ESCRT III proteins facilitate fatty acid mobilization, possibly by local induction of curvature at the LD surface (Chang et al., 2019).
Outlook
While LDs have long been largely disregarded by the contact site field, their importance in organization of the interaction-based organelle communication network is now starting to be appreciated. The very recent years have yielded identification of the first LD tether proteins, which display interesting functional features. It has become clear that similar to tethers between bilayer bounded organelles, LD tethers exist that additionally mediate (bulk) lipid transfer. Some LD tethers alternatively localize to other cellular contact sites, indicating that LD contact formation and function is tightly coordinated with other contact sites. Interestingly, there are examples of LD tethers that mediate spatial organization beyond formation of bi-organellar contacts, through coordination of three different types of organelles into tri-organellar structures, and by positioning of lipid metabolism enzymes next to LD contacts. Many further LD tethers with unexpected molecular features likely await their discovery. Identification and functional characterization of LD tether proteins will in the future enable us to unravel the exact biological roles of each LD contact site and to ultimately understand cellular lipid flux in health and disease.
Footnotes
Acknowledgments
I would like to thank all members of my lab for helpful discussions. I would like to apologize to those colleagues whose work could not be cited due to space limitations.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Deutsche Forschungsgemeinschaft, Cells-in-Motion Cluster of Excellence (EXC 1003—CiM), University of Münster, Germany.
