Abstract
Objective
Head and neck cancers are mostly composed of head and neck squamous cell carcinoma (HNSCC). The incidence and mortality of HNSCC are higher in countries with emerging health care systems, particularly Africa. Given that they are more genetically diverse, characterization of polymorphism in African HNSCC may result in the identification of distinct molecular targets as compared with the known HNSCC candidate genes. This study objective is to review the current evidence of genetic data on HNSCC among African populations as well as to demonstrate any distinctions as compared with known candidate genes and to appraise any research gaps.
Data Sources
Publications that interrogated susceptible gene polymorphisms to African-based populations with cancer were reviewed for this study.
Review Methods
Our search methodology was modeled after the Cochrane systematic review protocol, which included MeSH terms and keywords related to cancer, polymorphisms, and African countries.
Results
Seven articles studying 2 HNSCC cancer types in 3 of 54 African countries met the inclusion criteria. Thirteen polymorphisms from 10 genes were screened (NOS3, CYP1A1, CYP2D6, NAT1, NAT2, NQO1, IL-10, IL-12, IL-8, COX2). All articles were screened for polymorphisms based on a polymerase chain reaction–based technique. All polymorphs suggested association to HNSCC, with 10 of 13 polymorphs demonstrating a statistically significant association.
Conclusion
Studies on known HNSCC candidate genes should be undertaken in Africa, particularly among sub-Saharan Africans. Importantly, these studies should be large scale with multiple HNC sites and with use of high-throughput methods.
Head and neck cancers are the seventh-leading cancer by incidence worldwide. 1 They are mostly composed of head and neck squamous cell carcinoma (HNSCC), which develops in the mucous membrane of several locations, such as the mouth, throat, larynx, nose, and sinuses. Despite recent therapeutic advances, the 5-year overall survival is still at ~50%. 2 In 2012, the worldwide burden of HNSCC was ~600,000, and the recorded death was ~300,000.1,2 This incidence and mortality of HNSCC are higher in countries with emerging health care systems, due to ineffective prevention strategy, late screening, and suboptimal diagnostic and therapeutic tools. 3 According to the World Bank, 71% (41/98) of low- and middle-income countries are in Africa (https://data.worldbank.org/income-level/low-and-middle-income). There have been reports of dissimilarities in the genetic etiology of cancer based on ethnic differences in mutations. 4 This is due mainly to locus heterogeneity among ethnic groups, immune-mediated response, and corroboration of Eurasian population reports.4,5 These Eurasian populations contribute a subset to the human genome and do not fully characterize susceptible variants, particularly those of Africans, who exhibit higher genetic diversity and demonstrate less gene linkage disequilibrium. 5 Several genetic disease studies have provided ample evidence of overlap in disease-causal alleles between ancestry groups and diverse communities.6,7 This overlap of the underlying causal allele at several loci across ancestry improves the power to detect disease-susceptible loci and improve causal variant mapping. 8 Our group has published the most comprehensive catalog of susceptible genes and polymorphisms in African population–based cancers with limited reports on HNSCC candidate alleles. 9 While there are several reports on HNSCC candidate genes,10,11 there remains no catalog of candidate genes for African population–based HNSCC. Reviewing genetic polymorphisms in HNSCC in African populations will appraise the current evidence and gene distinction from the known HNSCC candidate genes.
Etiology of HNSCC
Tobacco use, alcohol consumption, and human papillomavirus (HPV) are identified as the main causative factors of initiation and progression of HNSCC. While the correlation of E6/E7 mRNA gene expression to the genotoxic activity of HPV 16/18 in oropharyngeal cancer has been established,12,13 there is yet to be an understanding of the molecular correlation between oral cancer and nicotine subunits/tobacco alkaloids that give rise to tobacco-specific N-nitrosamines.14,15 Except for high-risk HPV status, quite a number of these molecular risk factors have limited clinical use.
Genetics of HNSCC
The major cellular function often altered during carcinogenesis is the cell cycle regulation. Cell cycle–regulating genes targeted in HNSCC by carcinogens or HPV oncogenes are mainly those encoding p53 proteins and retinoblastoma pathways. 13 Several studies have investigated the genomic landscape of HNSCC with high-throughput methods and have reported significant single-nucleotide polymorphisms and copy number variations in candidate genes as well as a distinct genetic landscape associated with HPV status.10,11,16 These genes comprise TP53 (40%-60%), CDKN2A (12%-16%), NOTCH1 (10%-12%), PIK3CA (8%), FAT1 (5%-8%), HRAS (5%-8%), CASP8 (5%-8%), and PTEN (3%-7%).10,16
This study aimed to catalog the current evidence of the knowledge and genetic data (cohort, cross-sectional, and case-control studies) on HNSCC among African populations, as well as appraise any distinction between known candidate genes and research gaps for future studies.
Materials and Methods
Our search methodology was modeled after the Cochrane systematic review protocol. MeSH terms and keywords comprising and related to cancer, HNSCC and subtypes, genetic polymorphism, single-nucleotide polymorphisms, Africa, and African regions and countries were used for the initial search (see
Table 1
). Publications were selected that interrogated susceptible gene polymorphisms to African-based populations with cancer, comprising North Africans and sub-Saharan Africans, including black South Africans. Those whose combined ancestral origins were not African were excluded (eg, white Africans, African Americans, and African Europeans). Publications were selected with the search tools PubMed (1950-2019), Ovid Medline (1946-2019), and Web of Science (1900-2019). References from these selected publications were searched and appropriately included. The English-language title, abstract, and full text of selected publications were read to identify publications suitable for this review. Genetic studies considered were genetic (candidate gene) association studies and sequencing studies for germline and somatic mutations. As such, purely clinical or pathologic articles were excluded. A modified Quality Assessment Tool–Strengthening the Reporting of Genetic Association (QAT-STREGA) was used to assess the selected articles for study quality.
17
In summary, this QAT-STREGA tool assessed the presence of the following: power calculation, case population characteristics, cancer diagnosis, case and control screening, measure of association with odds ratio, genotyping error, Hardy-Weinberg equilibrium, and rs identification number (per the National Center for Biotechnology Information’s variation resource single-nucleotide polymorphism database). The QAT-STREGA score evaluated 9 items, and each had a score of 0 or 1, which can add to a maximum score of 9; a score
Search Strategy to Identify Publications on African Head and Neck Squamous Cell Carcinoma.

The research and ethics review board of the Lagos State University College of Medicine required no ethics approval for this study.
Results
The study search protocol identified 27 publications from PubMed (n = 16), Ovid Medline (n = 8), and Web of Science (n = 3; see Figure 1 ). Based on the search of the titles and review of abstracts, 21 publications were removed for not being related to HNSCC or genetic polymorphisms or variants. The remaining 6 publications had their full text reviewed and were adjudged to have met the inclusion criteria. One publication from the references was added to make of total of 7 eligible publications. Two HNSCC types were studied in these selected articles: nasopharyngeal and laryngeal carcinomas.
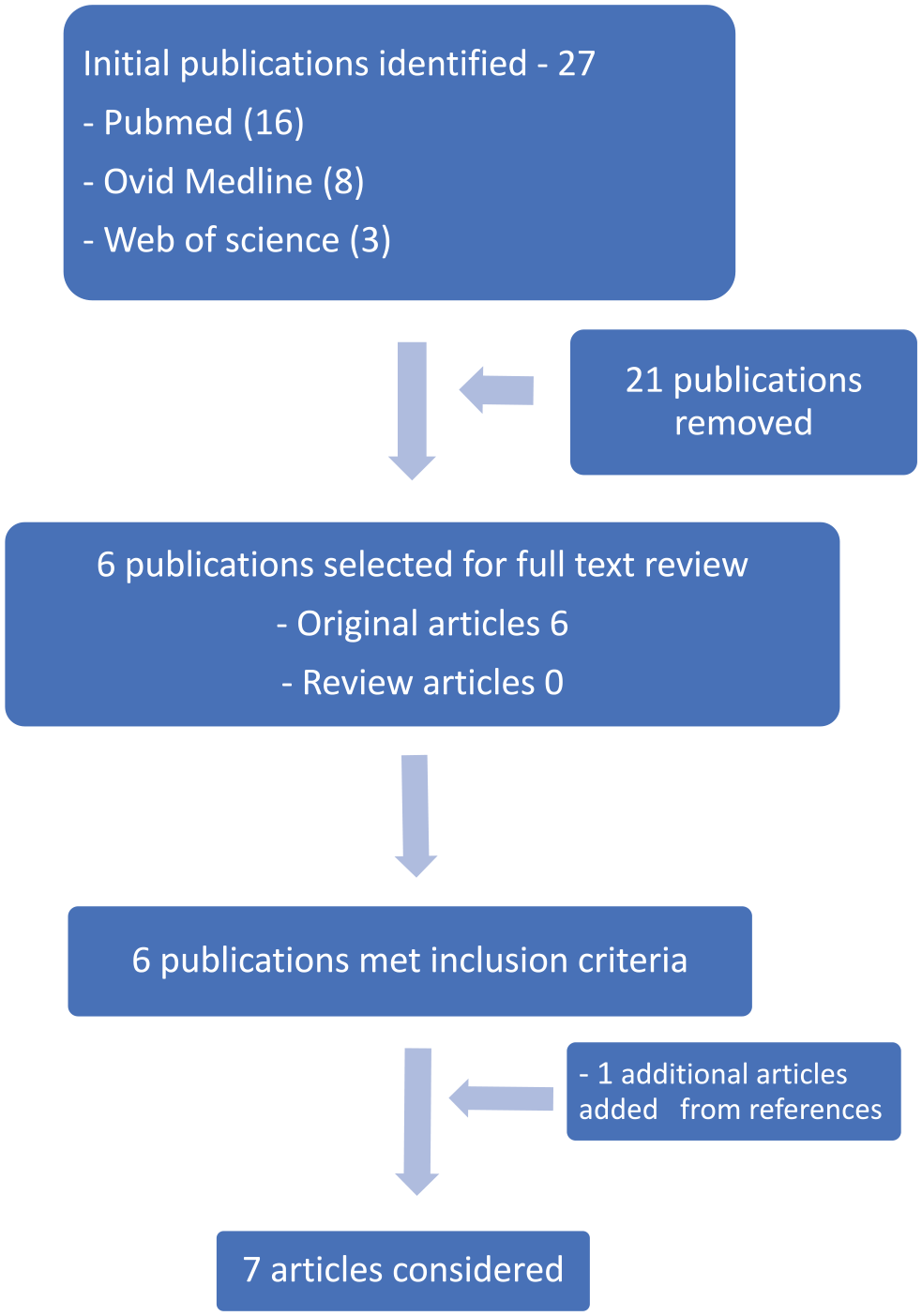

Search strategy flowchart.
A total of 6 genetic studies on nasopharyngeal cancer and 1 study on nasopharyngeal and laryngeal cancers were undertaken in 3 of 54 (6%) African countries: Tunisia, Algeria, and Morocco.18-24 All studies were case-control studies and published between 2007 and 2019 (see Tables 2 and 3 ). Sample sizes ranged from 137 to 392 (median, 180) for cases and 168 to 365 (median, 261) for controls. Cancer diagnosis for all cases was by histology. Tissues for DNA extraction were cancer tissue for cases and matching peripheral blood for controls. All studies screened for variants with polymerase chain reaction–based techniques. Thirteen polymorphisms from 10 genes were screened: NOS3, CYP1A1, CYP2D6, NAT1, NAT2, NQO1, IL-10, IL-12, IL-8, and COX2. All studies reported risk ratios; 6 of 7 studies tested the Hardy-Weinberg equilibrium; and 4 of 7 studies reported the rs numbers per the National Center for Biotechnology Information. All studies had a QAT-STREGA score >5, and 10 of 13 variants showed a statistically significant association to target gene polymorphism. Additional details of these studies are shown in Table 2 and Table 3 .
Quality Assessment of Genetic Studies.
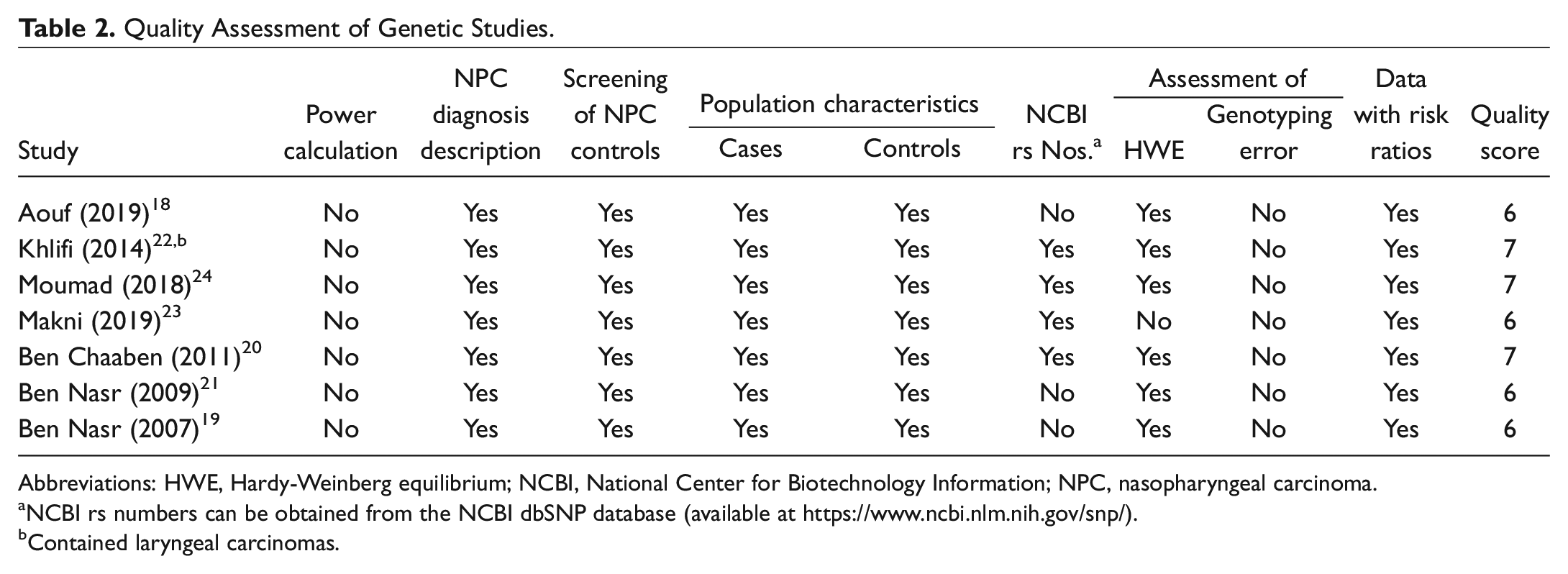
Abbreviations: HWE, Hardy-Weinberg equilibrium; NCBI, National Center for Biotechnology Information; NPC, nasopharyngeal carcinoma.
NCBI rs numbers can be obtained from the NCBI dbSNP database (available at https://www.ncbi.nlm.nih.gov/snp/).
Contained laryngeal carcinomas.
Gene Polymorphisms and Methodologies of Genetic Studies.
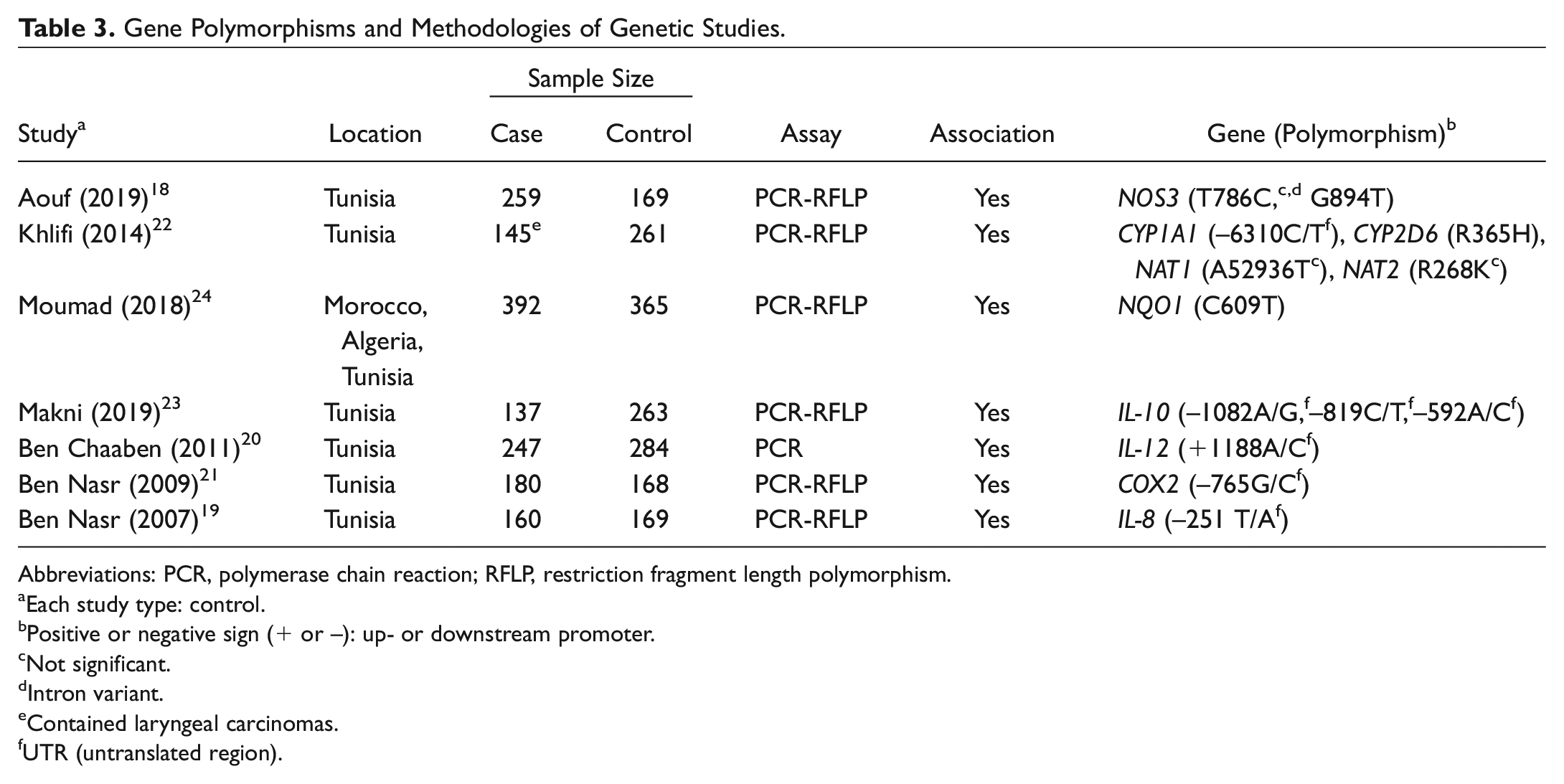
Abbreviations: PCR, polymerase chain reaction; RFLP, restriction fragment length polymorphism.
Each study type: control.
Positive or negative sign (+ or –): up- or downstream promoter.
Not significant.
Intron variant.
Contained laryngeal carcinomas.
UTR (untranslated region).
As none of the identified genes were similar to the HNSCC candidate genes, a review search was conducted on these identified genes and polymorphs with TCGA-HNSC project data ( Table 4 , Figure 2 ).
Gene Polymorphisms From TCGA-HNSC Project Data.
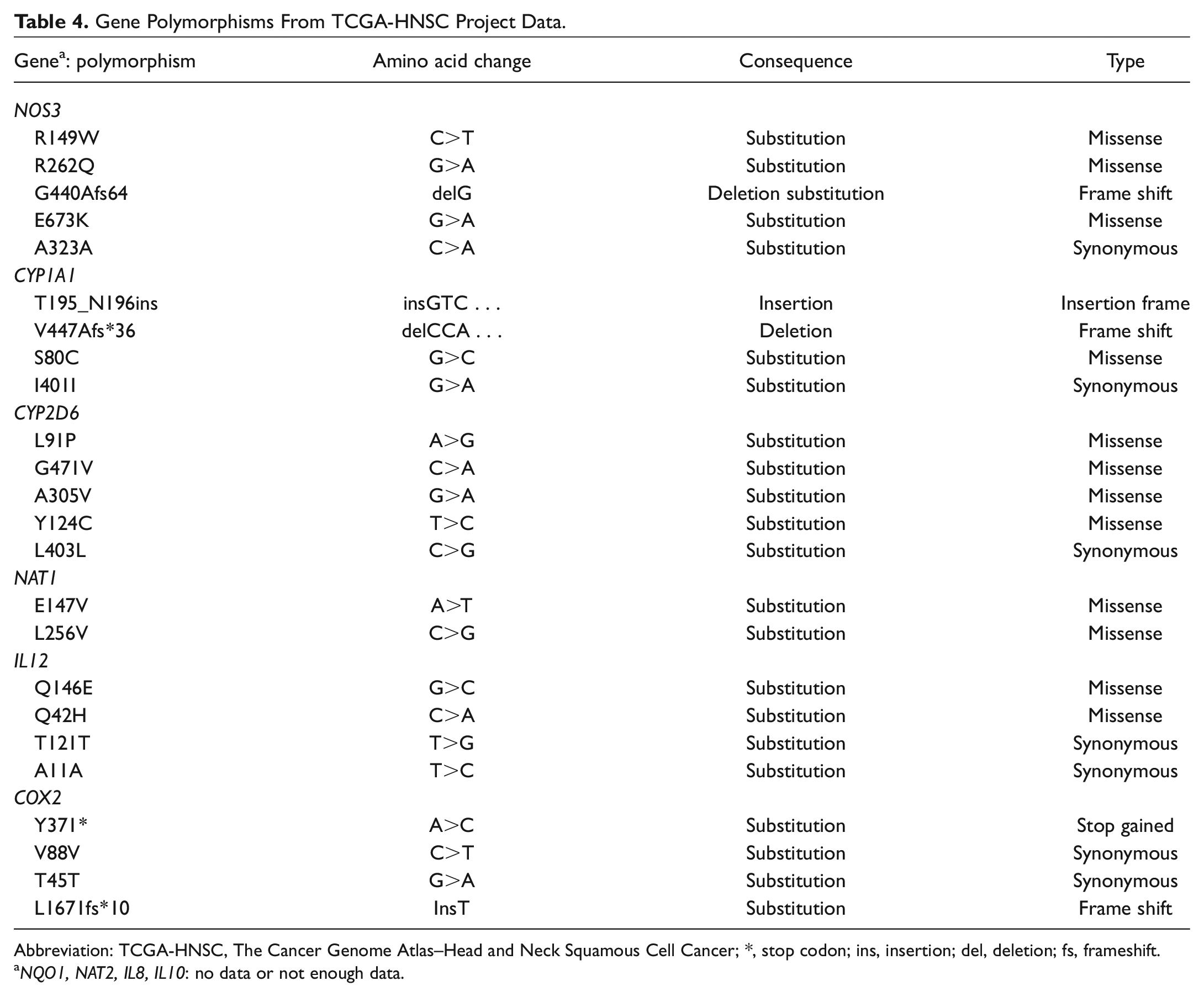
Abbreviation: TCGA-HNSC, The Cancer Genome Atlas–Head and Neck Squamous Cell Cancer; *, stop codon; ins, insertion; del, deletion; fs, frameshift.
NQO1, NAT2, IL8, IL10: no data or not enough data.
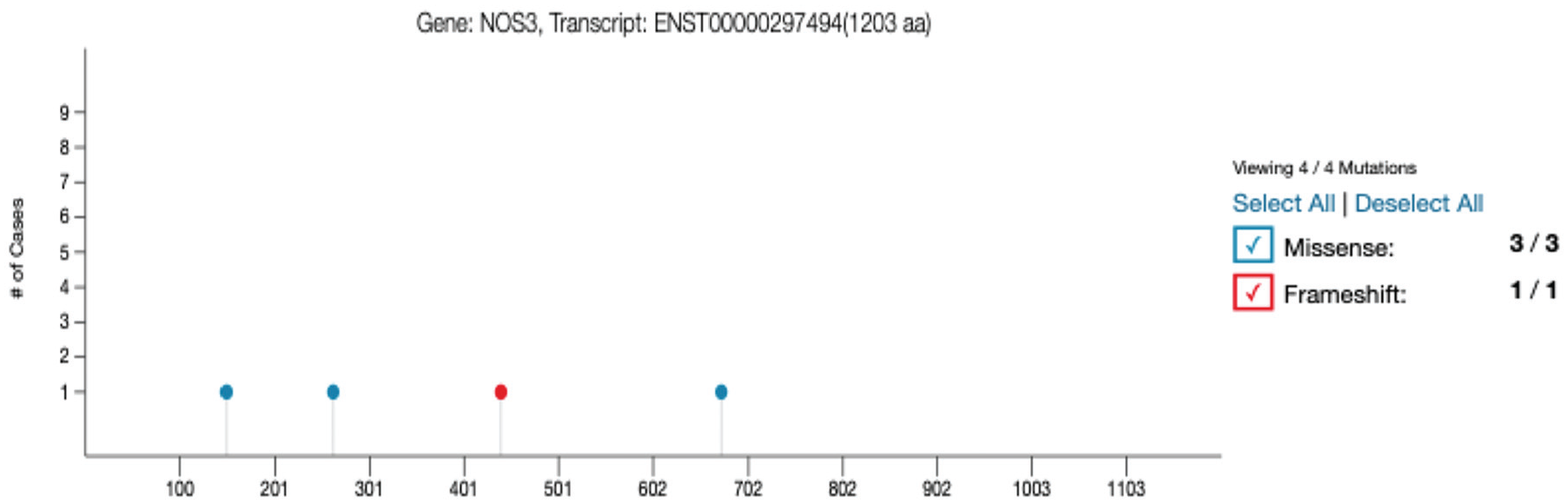
NOS3 showing mutation positions: TCGA-HNSC project database (The Cancer Genome Atlas–Head and Neck Squamous Cell Cancer).
Discussion
HNSCC can occur de novo or transform from a premalignant lesion, with the commonest being oral epithelial dysplasias (OEDs)—specifically, oral leukoplakia. Califano et al 25 in 1996 postulated the first genetic multistep characterization of morphologic changes in the oral squamous epithelium that give rise to oral squamous cell carcinoma. They suggested that loss of heterozygosity at chromosomes 3p, 9p, and 17p occurred in OEDs, reflecting early carcinogenesis, while alterations at chromosomes 11q, 4q, and 8q in oral squamous cell carcinoma indicated late-phase carcinogenesis, which may be independent of a previously existent OED. This group’s research formed the basis for researchers the world over to demonstrate HNSCC-susceptible genes categorized as candidate and established HNSCC genes.10,11
Our study reviewed all the molecular genetic research undertaken in Africa to suggest any variant distinction as compared with other established reports. It was significant to note that all 7 studies were undertaken in North African populations, indicating that sub-Saharan Africa is worst hit by this burden of a lack of research and data when compared with the North African countries. This in itself suggests that any findings might be an inaccurate reflection of the susceptible genes for African HNSCC, as North Africa shares a majority of its ancestry with Eurasia as compared with sub-Saharan Africa. 26 North Africa is surrounded by the Sahara Desert in the south and the Mediterranean Sea in the north and is bounded by Asia and Europe, with the first 2 landmarks acting as prehistoric gene flow barriers and the latter 2 heavily influencing the Maghreb gene pool. 27 More significant, the Sahara Desert acts as a gene flow barrier between the North Africa and sub-Saharan Africa.
This review reported the interrogation of several gene target variants by using a polymerase chain reaction–based molecular technique—restriction fragment length polymorphism—which locates a DNA template within a sequence. Considering the low strength of this technique, it is possible that some mutations were missed since restriction fragment length polymorphism is known to be of limited utility, with a base change sensitivity <80%. 28 While some of the techniques used may have been contemporary for their time, high throughput–based study may be necessary to interrogate for variants in the entire genome.
Surprisingly, none of the known HNSCC candidate genes were interrogated in these African studies, and significantly, the justification for interrogating some of these genes and polymorphisms showed very little correlation to the TCGA-HNSC project database. While some of these identified genes (NOS3, CYP1A1, CYP2D6, NAT1, NAT2, NQO1, IL8, IL10, IL12, COX2) are susceptible to carcinogenesis, their review search with the TCGA-HNSC project database revealed no HNSCC correlation. Perhaps a larger-scale study from multiple head and neck cancer sites and the use of high-throughput methods for variant screening may have provided more genetic data.
In addition, no study in this review undertook replication and validation assays for its genetic polymorphisms. This made it difficult to demonstrate any genetic correlation of these polymorphs to African-based populations HNSCC, thus rendering the resolution to mainly suggestive rather than statistically significant. In addition, the majority of these studies did not report a power calculation or a genotyping error, and this may have resulted in these studies being underpowered with an increased genotyping error. Furthermore, the underpowered nature of the study would not support the statistically significant association of the polymorphisms reported. An effective sample size for a threshold
Several candidate genes (TP53, CDKN2A, NOTCH1, PIK3CA, FAT1, HRAS, CASP8, PTEN) and their functional genetic polymorphisms have been identified as playing a role in HNSCC causation.10,16 According to the present study, none of these known candidate genes have been researched in Africa, highlighting a research gap in the molecular etiology of African population HNSCC.
In conclusion, studies on HNSCC candidate genes needs to be undertaken in Africa, with more of these studies in sub-Saharan Africa. Importantly, these studies should be large scale with multiple head and neck cancer sites and should employ high-throughput methods for improved causal mapping.
Footnotes
Acknowledgements
Members of the head and neck oncology research group, Medical Research Centre, Lagos State University College of Medicine.
