Abstract
The emergence of coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) threatens the health of the global population and challenges our preparedness for pandemic threats. Previous outbreaks of coronaviruses and other viruses have suggested the importance of diagnostic technologies in fighting viral outbreaks. Nucleic acid detection techniques are the gold standard for detecting SARS-CoV-2. Viral antigen tests and serological tests that detect host antibodies have also been developed for studying the epidemiology of COVID-19 and estimating the population that may have immunity to SARS-CoV-2. Nevertheless, the availability, cost, and performance of existing viral diagnostic technologies limit their practicality, and novel approaches are required for improving our readiness for global pandemics. Here, we review the principles and limitations of major viral diagnostic technologies and highlight recent advances of molecular assays for COVID-19. In addition, we discuss emerging technologies, such as clustered regularly interspaced short palindromic repeats (CRISPR) systems, high-throughput sequencing, and single-cell and single-molecule analysis, for improving our ability to understand, trace, and contain viral outbreaks. The prospects of viral diagnostic technologies for combating future pandemic threats are presented.
Introduction
The outbreak of coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has posed a major global health crisis.1–4 Viral diagnostic technologies are essential to rapidly detect, interrogate, and respond at all stages to mitigate the COVID-19 pandemic. For example, rapid diagnostic tests that are sensitive and specific are needed to identify patients infected with SARS-CoV-2 in a timely manner. The ability to perform COVID-19 diagnostics at the point of care will dramatically improve patient management and infection control. 5 Accurate laboratory tests are also required for zoonotic, environmental, and epidemiological investigations of coronaviruses. The availability of diagnostics is key for resuming normal activities after a lockdown and preventing the next wave of coronavirus disease outbreak.
The knowledge of the virology and pathogenesis of coronaviruses provides a foundation for the diagnosis of SARS-CoV-2. Coronaviruses are enveloped viruses that consist of a positive-sense single-stranded RNA genome (27–32 kb). 6 Six species of human coronavirus have been known prior to SARS-CoV-2: 229E, OC43, NL63, HKU1, severe acute respiratory syndrome coronavirus (SARS-CoV), and Middle East respiratory syndrome coronavirus (MERS-CoV).6,7 Four of these human coronaviruses (229E, OC43, NL63, and HKU1) are most common and cause mild colds. 7 SARS-CoV 8 and MERS-CoV 9 are highly pathogenic and can cause life-threatening illness. SARS-CoV-1 and SARS-CoV-2 are two different strains of SARS-CoV. The genomes of coronaviruses share a high similarity in their arrangement, encoding 16 nonstructural proteins (nsp1–nsp16) that are involved in viral RNA transcription and replication and 4 major structural proteins: spike (S), envelope (E), membrane (M), and nucleocapsid (N).6,10,11 The life cycle of coronavirus initiates from the binding of S protein to the host receptor. SARS-CoV-1 12 and SARS-CoV-213,14 bind to angiotensin-converting enzyme 2 (ACE2), and MERS-CoV 15 binds to dipeptidyl-peptidase 4 (DPP4). After the entry of the virus, the genomic RNA starts to translate the replicase proteins, followed by the replication of genomic RNA and subgenomic RNA for translation. Following the translation of the structural proteins S, E, M, and N, the viruses are assembled and released.11,16,17
In this article, we review the principles and considerations of major viral detection technologies along with their applications in COVID-19 diagnostics (
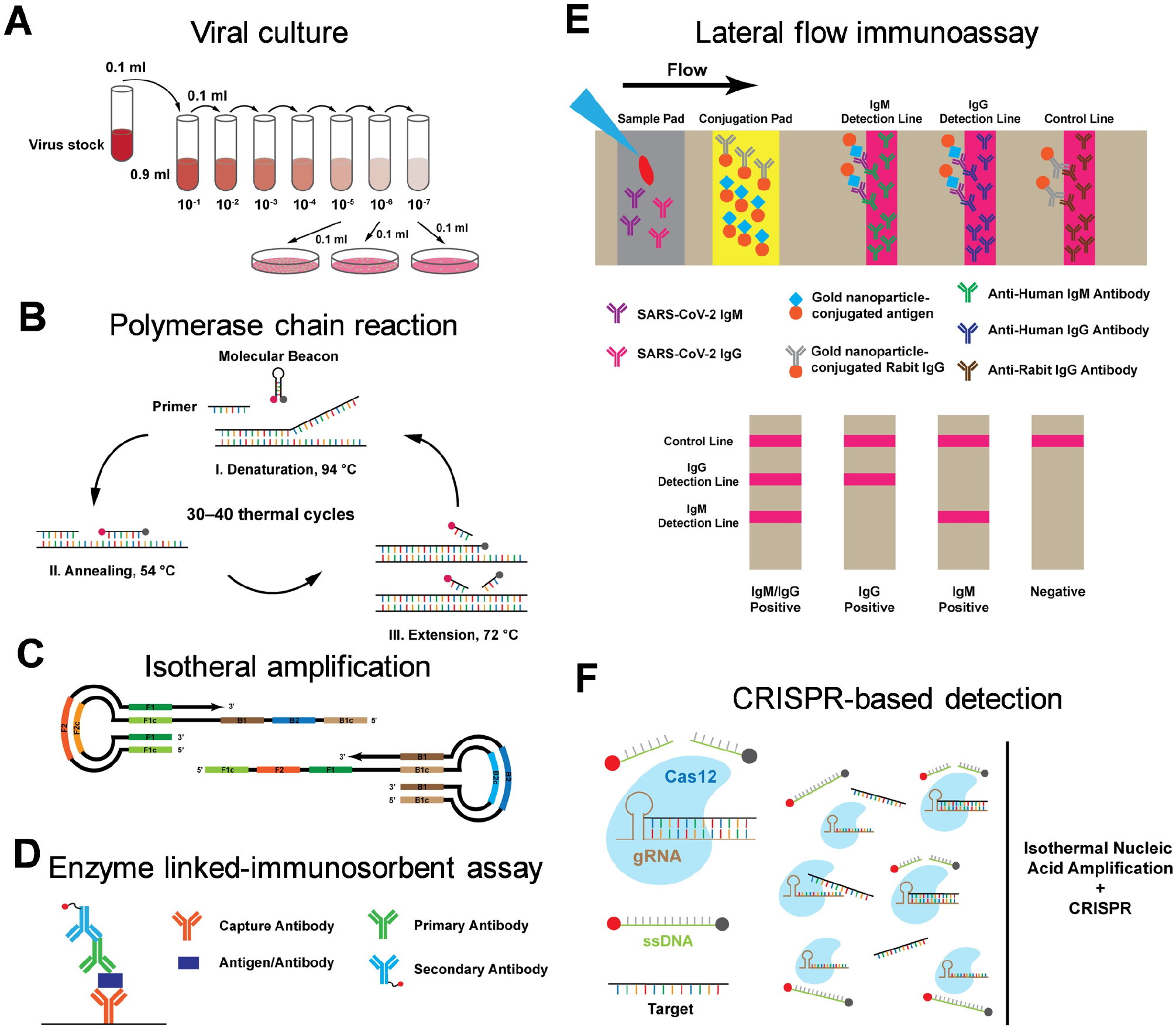
Common viral diagnostic schemes. (
Nucleic Acid Detection Based on PCR
PCR is an important biochemistry and molecular biology technique that exponentially amplifies nucleic acid in vitro via enzymatic replication. For viral detection, the viral DNA serves as the template, and a primer pair is designed to amplify a region of the DNA through thermal cycling. Conventionally, gel electrophoresis is applied to determine the presence of the target sequence and the size of the amplicon. Variants of PCR were developed to detect RNA viruses and to quantify the targets in the specimen. In reverse transcription PCR (RT-PCR), the target RNA is first reverse-transcribed to a complementary strand of DNA (cDNA) by the reverse transcriptase, and the resulting cDNA can then be amplified through PCR. Real-time PCR, also called quantitative PCR (qPCR), amplifies the target and detects the amplicon with an intercalating dye or a molecular beacon in real time for estimating the initial target concentration. Combining the two variants, real-time reverse transcription PCR (qRT-PCR or rRT-PCR) can quantify the concentration of RNA virus in a sample (
The design of the primer set is critical for the performance of a viral assay. Emery et al. tested primer sets targeting various regions of the genome of SARS-CoV-1. 20 The limit of detection (LOD) for primers targeting nucleocapsid (N) was as low as two copies per reaction. The analytical sensitivity and reproducibility were higher than with primers targeting the RNA-dependent RNA polymerase (RdRp) with a LOD of 7.5 copies per reaction. In contrast, a primer study of SARS-CoV-2 indicated a higher analytical sensitivity for primer targeting the RdRp gene than the N gene. 21 The CDC currently recommends SARS-CoV-2 qRT-PCR assays should target the N gene due to its high expression level. 22
Other important considerations of nucleic acid assays for COVID-19 diagnostics are the specimen type and the associated sample collection procedures. The specimen type can influence the sensitivity and reliability of viral assays. Using qRT-PCR, Wang et al. detected SARS-CoV-2 from several patient specimen types, including bronchoalveolar lavage, fibrobronchoscope brush biopsy, sputum, nasal swabs, pharyngeal swabs, feces, and blood. 23 The mean cycle threshold of nasal swabs was 24.3, which was lower than other specimens ranging from 31.1 to 34.6, suggesting that the nasal swab is the most sensitive specimen for COVID-19 diagnostics. In agreement, most COVID-19 diagnostic kits developed at the early stage of the crisis were based on nasal swab samples. Saliva tests, however, were also developed for COVID-19. The saliva-based approach provides a convenient alternative to swab tests, which reduces the risk of exposing health professionals to the virus and eases the shortage of swabs, personal protective equipment, and other medical supplies. A qRT-PCR test using home-collected saliva samples (Rutgers Clinical Genomics Laboratory, Newark, NJ) was authorized on May 7, 2020, by the FDA. 19
Currently, qRT-PCR is the gold standard for viral detection, and various commercially available qRT-PCR platforms have been adopted for COVID-19 diagnostics.2,9,21,24,25 If the viral target sequence is identified, accurate and sensitive qRT-PCR assays can be developed quickly in response to a viral outbreak. Direct detection of viral RNA allows early diagnosis of COVID-19 regardless of the symptoms and immune response of the patient. The limitations of qRT-PCR lie in the substantial assay time, the cost and availability of the equipment, false-positive results due to contamination, and the requirement of a clinical laboratory. A qRT-PCR test for coronavirus, which involves sample collection, RNA extraction, reverse transcription, and qPCR, could take several hours. The procedures are typically conducted in well-equipped clinical laboratories. The transportation of patient samples and the logistics of the clinical laboratory procedures can further increase the turnaround time. While feasible, an integrated, point-of-care diagnostic test for coronavirus based on qRT-PCR would be relatively expensive. Hence, efforts are being devoted to developing viral nucleic acid tests that are rapid, cost-effective, and automated for point-of-care diagnostics.
Nucleic Acid Detection Based on Isothermal Amplification
Isothermal nucleic acid amplification methods have been developed to simplify the thermal cycling process and reduce the assay time.
26
For instance, loop-mediated isothermal amplification (LAMP) applies four to six primers to recognize different regions of the target sequence and repeatedly amplifies the sequence through a stem-loop structure at a single temperature (~60 °C) (
More recent studies proposed variations of RT-LAMP to overcome these limitations. Shirato et al. introduced a fluorescence quenching probe into the RT-LAMP assay that specifically detected the primer-derived signals while reducing nonspecific signals. 31 The combination of RT-LAMP and a vertical flow visualization strip (RT-LAMP-VF) was proposed by Huang et al. 32 The result was detectable by the naked eye, and the assay allowed point-of-care detection of MERS-CoV within 35 min. Cai et al. improved LAMP by using phosphorothioated DNA as primers (PS-LAMP) for enhancing the efficiency of loop formation and extension. 33 PS-LAMP lowered the operating temperature to 40 °C while maintaining comparable performance with regular LAMP.
Other isothermal nucleic acid amplification methods have also been developed for viral detection. Nucleic acid sequence-based amplification (NASBA) uses two primers and three enzymes, namely reverse transcriptase, RNase H, and T7 RNA polymerase, to amplify the nucleic acid target isothermally, and was demonstrated for SARS-CoV-1 detection.34–36 The recombinase polymerase amplification (RPA) assay exponentially amplifies the nucleic acid with a strand-displacement strategy driven by recombinase and primers, 37 and the assay time was only 3 to 7 min for MERS-CoV detection. 38 Rolling circle amplification (RCA) proposed by Fire et al. generates self-propagating strand displacement initiated by ligated circular DNA from the target template, amplifying the nucleic acid signal to 109-fold within 90 min.39,40 Due to the involvement of multiple primer sets and enzymes, isothermal amplification strategies are relatively complex and often require substantial optimization and validation. Nevertheless, isothermal amplification provides rapid and sensitive testing without bulky instruments, which is particularly attractive for viral diagnosis in remote settings. For instance, a portable isothermal amplification platform (Abbott, Chicago, IL) received the FDA’s EUA for COVID-19 detection on March 27, 2020. 19 The system detects positive samples in as fast as 5 min and negative samples in 13 min.
Immunological Tests Based on Viral Antigen
Viral antigen tests represent another strategy for detecting SARS-CoV and for investigating the pathogenesis of coronaviruses. The virion structure of SARS-CoV-2 consists of the viral RNA genome bound with the N proteins in the viral envelope anchoring the M, E, and S structural proteins.7,13,16,41,42 The nucleocapsid protein is the most abundant protein in coronavirus and can be used as a diagnostic marker for SARS-CoV detection.43–46 SARS-CoV-1 nucleocapsid proteins were shown to be detectable in nasopharyngeal aspirate, urinary, and fecal specimens. 44 With specific antibodies against the nucleocapsid protein, viral antigen tests can distinguish SARS-CoV from other human coronaviruses, such as OC43 and 229E.
Virus antigens can be detected by common protein detection techniques, such as immunohistochemistry (IHC) and enzyme-linked immunosorbent assay (ELISA). IHC labels target antigens in fixed cells or tissues with primary and secondary antibodies to locate the proteins of interest in the sample. For instance, Shieh et al. characterized the distribution of SARS-CoV-1 in the lung tissue of a patient by IHC and revealed coronavirus particles in pneumocytes.
47
In contrast, ELISA immobilizes the target antigen on a substrate, followed by the binding of an antibody that yields an absorbance signal (
Viral antigen tests provide an economical and convenient diagnostic method when a qRT-PCR system is not available. Due to the low false-positive rate, viral antigen tests have a high clinical specificity. Antigen detection is limited by the availability of specific antibodies, however, and the sensitivity of antigen tests is relatively low due to the high false-negative rate. Lau et al. conducted a comprehensive study on detecting SARS-CoV-1 antigens and compared the detectable time of the antigen in different specimens. 44 The nucleocapsid protein was detectable in nasopharyngeal aspirate, urine, and fecal samples from days 6 to 24, 11 to 31, and 8 to 32 after the onset of the illness, respectively. This time delay after the onset of illness increases the false-negative rate and reduces the clinical sensitivity of the assay, limiting the potential of viral antigen tests for early diagnostics. A COVID-19 antigen test (Quidel Corporation, San Diego, CA) based on fluorescence immunoanalysis received the FDA’s EUA on May 8, 2020. 19
Immunological Tests Based on Host Antibody
Serological tests, or antibody tests, typically detect IgM and IgG in blood. IgM and IgG are antibodies generated against a viral infection in the adaptive immune response, reflecting that a person was infected recently or in the past. 49 Spike protein and nucleocapsid protein are the most common immunogens for coronavirus serological tests.50,51 The antibody response of SARS-CoV-1 suggests that IgM is produced prior to IgG and can be detected in the early stage, while IgG has a higher expression level and is detectable for 3 years after the initial exposure. 52 The antibody response of SARS-CoV-2 is under intensive investigation. Zhao et al. reported the antibody response profile to SARS-CoV-2, showing that the median seroconversion time of IgM and IgG were, respectively, 12 and 14 days after the onset of illness. 53
Similar to viral antigen testing, ELISA, immunofluorescence assay, and other protein assays can be applied for SARS-CoV-1, MERS-CoV, and SARS-CoV-2 antibody testing.51,54–56 These techniques, however, are relatively costly for large-scale studies, require supporting equipment, and are difficult to implement in remote settings. In contrast, a lateral flow immunoassay, or a test strip, is cost-effective, standalone, and easy to operate. A typical lateral flow strip consists of a sample pad, conjugation pad, detection line, and control line.
57
The fluid motion is driven by the capillary force on loading of the sample. The target antibody binds the gold nanoparticle–conjugated antigen, which is then captured by the detection antibody on the detection line, and the remaining gold nanoparticle–conjugated antigen is captured on the control line (
The main limitation of serological tests lies in the long seroconversion time. Zhao et al. applied ELISA to test its sensitivity for COVID-19. 53 The result shows that the sensitivity was less than 40% for IgM and IgG detection in the first week after the onset. The value increased to 94.3% for IgM and 79.8% for IgG 15 days after the onset, and the overall sensitivity is 82.7% and 64.7% for IgM and IgG. Therefore, the sensitivity of serological tests for early-stage coronavirus detection is relatively low. Another limitation of antibody tests is the cross-reactivity from conserved antigens of other viruses, leading to false-positive results. 50 Additional efforts are required for selecting proper immunogens to ensure the specificity. Serological tests can, however, provide important data for understanding viral epidemiology and guiding public policy. The cost-effectiveness and point-of-care nature of serological tests make them promising candidates for population-level investigations. The FDA issued an EUA for a SARS-COV-2 IgG/IgM test (Cellex, Research Triangle Park, NC) on April 1, 2020. 19
Emerging Technologies
CRISPR technology represents an emerging approach for nucleic acid detection. CRISPR was first developed as an RNA-guided DNA endonuclease for gene editing.59,60 Taking advantage of CRISPR nuclease activity on RNA recognition, Zhang et al. developed a CRISPR-Cas13a (CRISPR-associated protein 13a)–based technology for nucleic acid detection, termed Specific High-Sensitivity Enzymatic Reporter UnLOCKing (SHERLOCK).61–63 Engineered Cas13a–CRISPR-RNA (crRNA) was designed to bind with the target RNA sequence, which was amplified by isothermal amplification (RPA) and transcribed to RNA. On Cas13a-crRNA binding with target RNA, Cas13a was activated and collaterally cleaved the nearby RNAs, which release a signal due to the cleavage of a reporter RNA.
61
Broughton et al. reported the first CRISPR-Cas12-based detection of SARS-CoV-2 (
High-throughput sequencing, or next-generation sequencing, contributes to various areas in the battle against COVID-19 outbreaks, such as assay development, genetic vaccine design, and outbreak analysis. 66 High-throughput sequencing refers to the integration of advanced amplification, sequencing, and data analysis strategies to achieve high-throughput and genome-wide sequencing.67–69 Multiple platforms, such as HiSeq (Illumina, San Diego, CA),70–72 SOLiD (ThermoFisher Scientific, Waltham, MA), 73 BGISEQ (BGI, Shenzhen, China), 74 and MinION (Oxford Nanopore Technologies, Oxford, United Kingdom),71,75 have been developed for high-throughput sequencing. Researchers have been applying high-throughput sequencing for viral discovery and investigation.76–78 Since the outbreak of COVID-19, extensive research uses high-throughput sequencing, such as metagenomic sequencing, to detect and analyze SARS-CoV-2.79–81 For instance, the development of COVID-19 nucleic acid tests was facilitated by the sequence of SARS-CoV-2. The key advantage of metagenomic sequencing is an unbiased sampling that does not require hypothesis-based primer design. Metagenomics detects the entire genome of the virus, identifies a wide range of viruses, and discovers new or unexpected viruses. 82 Nevertheless, current high-throughput sequencing technologies are slow and expensive, and require library preparation and substantial computational analysis. These features limit high-throughput sequencing technology in current clinical practice. High-throughput sequencing, however, enables researchers to track the mutation and evolution of viruses, providing insights into the origin and propagation of viruses.
Single-cell and single-molecule analysis represent important tools for virology. Viral culture techniques, such as the plaque formation assay and the endpoint dilution assay, are traditional methods for quantitative detection of viruses (
Summary
Rapid, reliable, and economical detection of SARS-CoV-2 is of great importance for diagnosis of COVID-19. Existing diagnostic techniques have distinctive characteristics ( Table 1 ). Nucleic acid tests, such as qRT-PCR and isothermal amplification, are robust and sensitive methods for early-stage detection of viral infections. With the development of lateral flow immunoassays and other microfluidic approaches, serological tests and antigen tests detect markers in minutes, instead of hours or days, for population-scale screening of viral infection. Emerging molecular biology techniques and engineering platforms (e.g., CRISPR and high-throughput sequencing) have great potential to enable novel diagnostic platforms. As evidenced by the COVID-19 crisis, the availability, speed, and accuracy of current viral diagnostic technologies remain limited for rapid response to global pandemics. Advances in diagnostic technologies will be required to enhance our ability to combat future viral outbreaks.
Comparison of COVID-19 Diagnostic Tests.
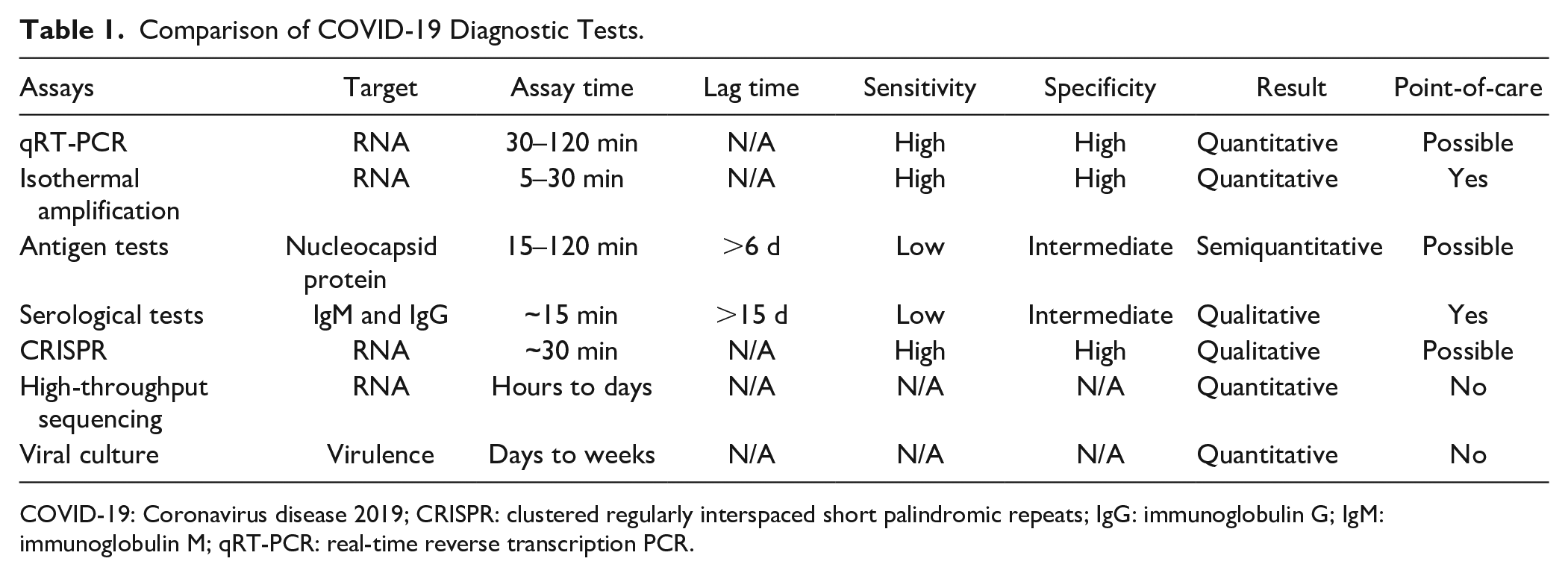
COVID-19: Coronavirus disease 2019; CRISPR: clustered regularly interspaced short palindromic repeats; IgG: immunoglobulin G; IgM: immunoglobulin M; qRT-PCR: real-time reverse transcription PCR.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received the following financial support for the research, authorship, and/or publication of this article: This work is supported by the Penn State COVID-19 research initiatives.
