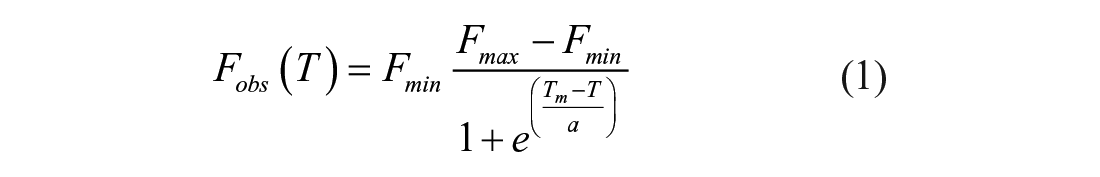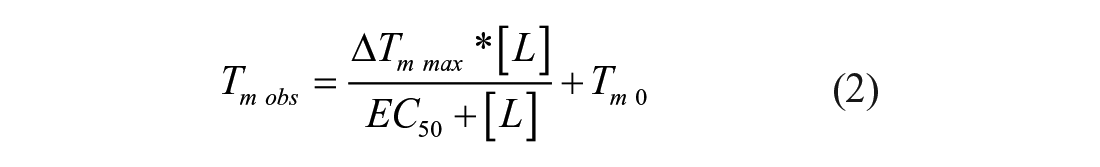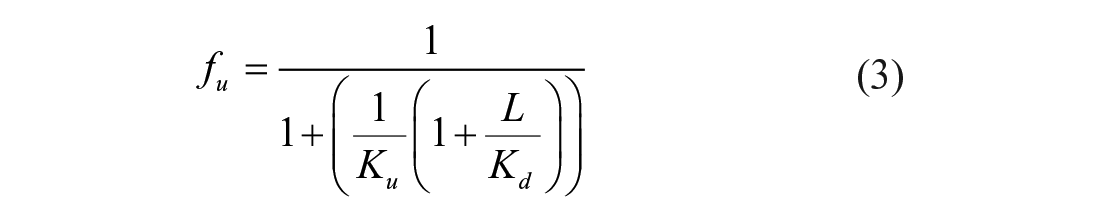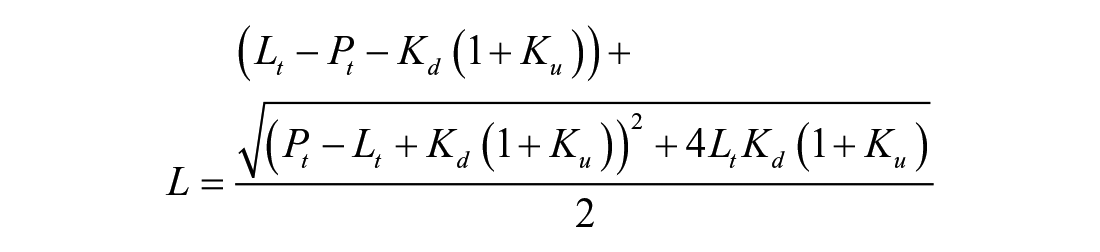Abstract
Small molecules that bind the SARS-CoV-2 nonstructural protein 3 Mac1 domain in place of ADP-ribose could be useful as molecular probes or scaffolds for COVID-19 antiviral drug discovery because Mac1 has been linked to the ability of coronaviruses to evade cellular detection. A high-throughput assay based on differential scanning fluorimetry (DSF) was therefore optimized and used to identify possible Mac1 ligands in small libraries of drugs and drug-like compounds. Numerous promising compounds included nucleotides, steroids, β-lactams, and benzimidazoles. The main drawback to this approach was that a high percentage of compounds in some libraries were found to influence the observed Mac1 melting temperature. To prioritize DSF screening hits, the shapes of the observed melting curves and initial assay fluorescence were examined, and the results were compared with virtual screens performed using AutoDock Vina. The molecular basis for alternate ligand binding was also examined by determining a structure of one of the hits, cyclic adenosine monophosphate, with atomic resolution.
Introduction
Direct-acting antivirals (DAAs) are desperately needed to treat COVID-19 patients and stem the devastation caused by the current SARS-CoV-2 pandemic. DAAs are typically developed from potent inhibitors of viral enzymes or high-affinity ligands of viral proteins. For example, remdesivir inhibits the SARS-CoV-2 RNA-dependent RNA polymerase, halting SARS-CoV-2 replication. 1 Based on past experiences, any effective DAA therapy will likely require a cocktail of more than one antiviral agent because drug resistance evolves rapidly. Methods are therefore needed to rapidly identify small-molecule drug-like ligands for as many SARS-CoV-2 proteins as possible. The ~29,900-nucleotide SARS-CoV-2 genome encodes many potential DAA targets, including 16 nonstructural proteins (nsps), 4 structural proteins, 6 accessory proteins, and possibly many others. Most SARS-CoV-2 nsps are products of the rep1b open reading frame that encodes a short (aka ORF1a) and long (aka ORF1b) polyprotein because an internal RNA hairpin occasionally causes translational frameshifting. The subject of this study is the multifunctional 945-amino acid-long nsp3 protein that cleaves the three junctures separating nsp1, nsp2, and nsp3. 2 SARS-CoV-2 nsp3 is most likely tethered to the ER with two ubiquitin-like domains (Ubl1 and Ubl2), two papain-like protease domains (PLP1pro and PLP2pro), three macrodomains (Mac1, Mac2, and Mac3), a nucleic acid-binding domain, and a hypervariable region facing the cytoplasm.
In a previous study, Frick et al. 3 characterized the SARS-CoV-2 Mac1 domain (aka the X domain) and its ability to bind ADP–ribose (ADPr). Here, we report the results of pilot screens designed to find drug-like Mac1 domain ligands, which might facilitate DAA design, or which could be useful as molecular probes. A differential scanning fluorimetry (DSF; aka the thermal shift, or ThermoFluor)4–6 assay was optimized and used to screen 726 compounds in the National Institutes of Health clinical collection (NIHcc), the National Cancer Institute (NCI) mechanistic set (540 compounds), and Sigma-Aldrich’s 1280-compound Library of Pharmacologically Active Compounds (LOPAC1280). Since up to 5% of compounds in each set influenced the apparent melting temperature, compounds were prioritized using information derived from individual melting curves and virtual screens performed with AutoDock Vina. 7 A high-resolution structure of one hit compound, cyclic adenosine monophosphate (cAMP), was determined in complex with SARS-CoV-2 Mac1, revealing at atomic resolution the capacity of the binding cleft to accommodate other ligands.
Materials and Methods
Purified SARS-CoV-2 Mac1 protein was prepared as described previously. 3 DSF assays were performed in 96-well PCR plates using an Eppendorf Mastercycler ep Realplex Quantitative Realtime PCR System, with each well containing 19 µL of master mix (5 µM Mac1) and 1 µL of a compound stock (10 mM for screening) or DMSO. The master mix was prepared by adding 20 µL of 500 µM Mac1 and 2.5 µL of 5000× SPYRO Orange protein gel stain (Sigma-Aldrich cat. S5692) to 1977.5 µL of buffer (20 mM MOPS, 25 mM NaCl, pH 7). The 96-well PCR plate was then sealed by a clear adhesive film and centrifuged at 1100 rpm for 5 min. The temperature was raised from 20 to 95 °C at a rate of 2 °C/min while measuring the fluorescence in the “TAMRA” channel. Each plate included both negative (DMSO) and positive (ADPr) controls. Tm values were calculated by fitting the data to eq 1 using either GraphPad Prism (GraphPad, La Jolla, CA) or TSA-CRAFT (https://sourceforge.net/projects/tsa-craft/). 8
In eq 1, Fobs (T) is the observed fluorescence at each temperature (T), Fmin is the minimum observed fluorescence, Fmax is maximum observed fluorescence, and a is a Hill slope. Two methods were used to estimate the affinity of Mac1 from DSF. First, the observed melting temperatures were plotted versus ligand and fit to eq 2 to determine the amount of compound needed to cause a change in the melting temperature of 50% (EC50), and a nonlinear regression was used to estimate ΔTm max (the maximum change in Tm) from the melting temperature of Mac1 in the absence of ligand (Tm 0):
The dissociation constant (Kd) of Mac1 and ADPr and the equilibrium constant describing protein unfolding (Ku) were also estimated using isothermal analysis as described by Bai et al. 9 Briefly, normalized melting curves were used to calculate the fraction of protein unfolded at a particular temperature (fu) and those values fitted to the total ligand (Lt) and protein (Pt) concentrations using nonlinear regression and eq 3:
where
Computational ligand screening was performed using both the unligated (6WEY and 6VXS) and ADPr-bound (6W02) forms, using the program AutoDock Vina. 7 The protein files were downloaded directly from the Protein Data Bank (PDB) and processed as described below before submitting for screening. All solvent molecules (HETATM) were removed from the files. Polar hydrogen atoms were added and Kollman charges were included in the protein files. The converted protein and ligand file pdbqt libraries were uploaded to a parallel computing cluster and run with the following parameters: energy difference = 4; number of recorded modes = 20; and exhaustiveness was set to 12. The docking box location was configured prior to using AutoDock tools. After the docking calculation was complete, the locations, orientations, and binding affinities of the top candidates were examined using UCSF Chimera and tabulated for comparison.
Mac1 was prepared for crystallization as described before 3 with the following modifications. First, the plasmid was modified to express one additional N-terminal residue (E206) and C-terminus was shortened by three residues such that it encoded residues 206–374 of the SARS-Cov-12 nsp3. After purification and TEV protease cleavage, the tag-free protein was concentrated to 20 mg/mL in 20 mM HEPES, pH 7.5, 150 mM NaCl, and 1 mM TCEP. Crystallization was accomplished as described for the Mac1·AMP complex 10 (1 µL of concentrated Mac1 was mixed with 1 µL of 30% PEG 4000 and 0.1 M MES, pH 6.5). Plate-shaped crystals grew in 3–7 days at 22 °C.
The Mac1·cAMP complex was prepared by soaking the crystal in a solution containing 35% PEG 4000 and 20 mM cAMP for 30 min. Cryoprotection was accomplished by briefly soaking the crystal in 35% PEG 4000, 20 mM cAMP, and 20% glycerol before plunging it into liquid nitrogen. Diffraction data were collected on Life Sciences Collaborative Access Team (LS-CAT) beamline 21-ID-F at the Advanced Photon Source of the Argonne National Laboratory, which is fitted with a fixed-wavelength beam at 0.97872 Å and a MarMosaic M300 detector. The data were collected with an oscillation width of 0.5° per image for a total oscillation of 180°. The data were processed using HKL2000;
11
data collection statistics are provided in
The structure was determined by molecular replacement in PHASER
12
using PDB ID 6WEY,
3
with solvent molecules and B-factor information removed, as the search model. The model underwent iterative rounds of (re-)building in COOT
13
and refinement in PHENIX.refine.14,15 Translation–libration–screw (TLS) refinement provided a more realistic treatment of the atomic displacement parameters; TLS groups were identified by phenix.find_tls_groups. Model refinement and validation statistics are provided in
Results
Optimized DSF Assay for SARS-CoV-2 Mac1
DSF has been used previously to study ligand binding to viral macrodomains.16,17 In DSF experiments using SARS-CoV-2 Mac1, the presence of ADPr raised the Mac1 melting temperature in a concentration-dependent manner (
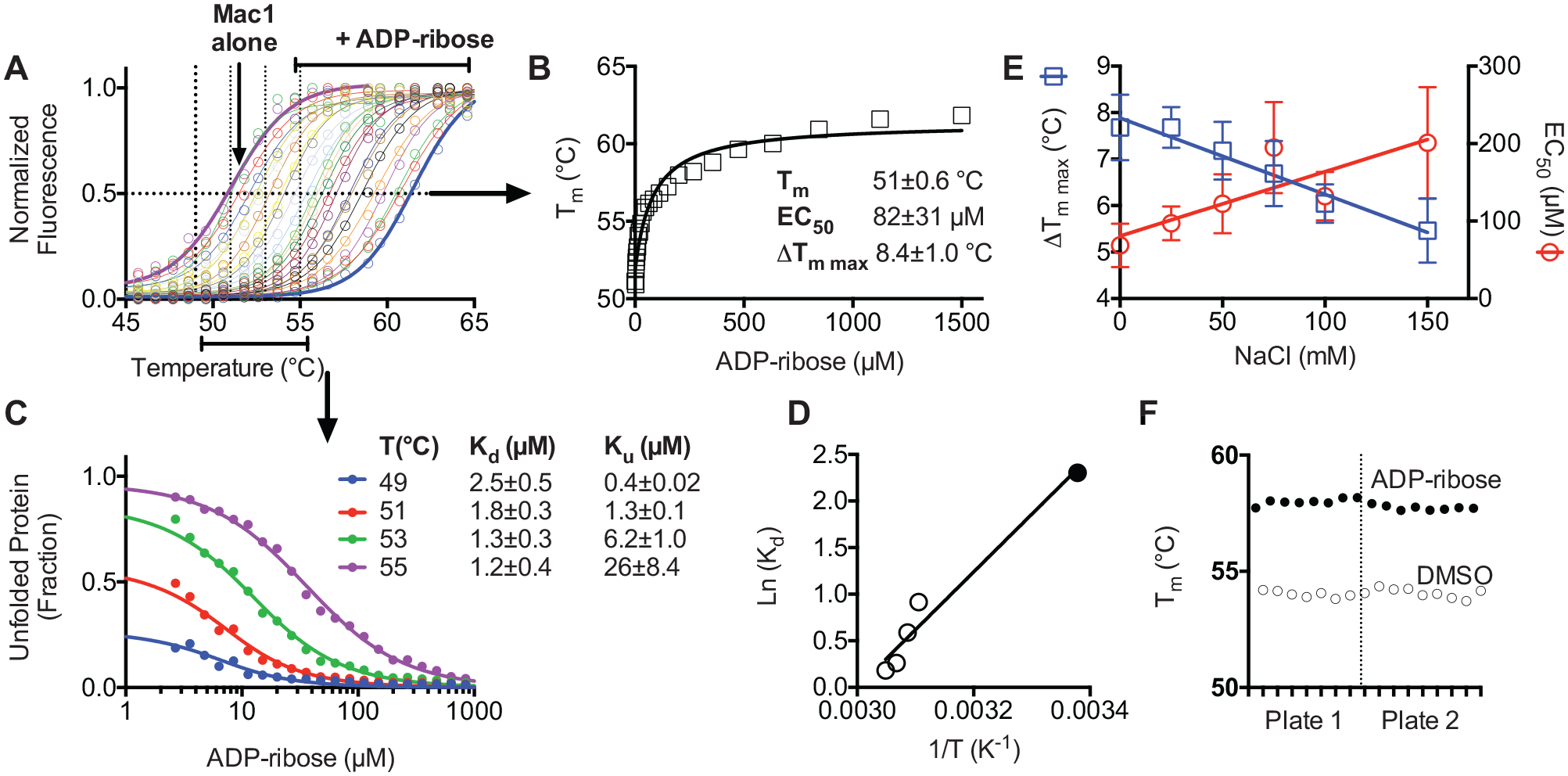
DSF assay optimization. (
DMSO did not change the melting curve even at concentrations as high as 10% (v/v), and similar results were also obtained when titrations with ADPr were repeated in various buffers, with the pH ranging from 6.5 to 8.0, or in the presence of various concentrations of divalent metal cations (Mg2+ or Mn2+). In contrast, the ionic strengths of the assay buffers influenced the results, with the largest ΔTm values and lowest EC50 values being obtained at the lowest ionic strengths (
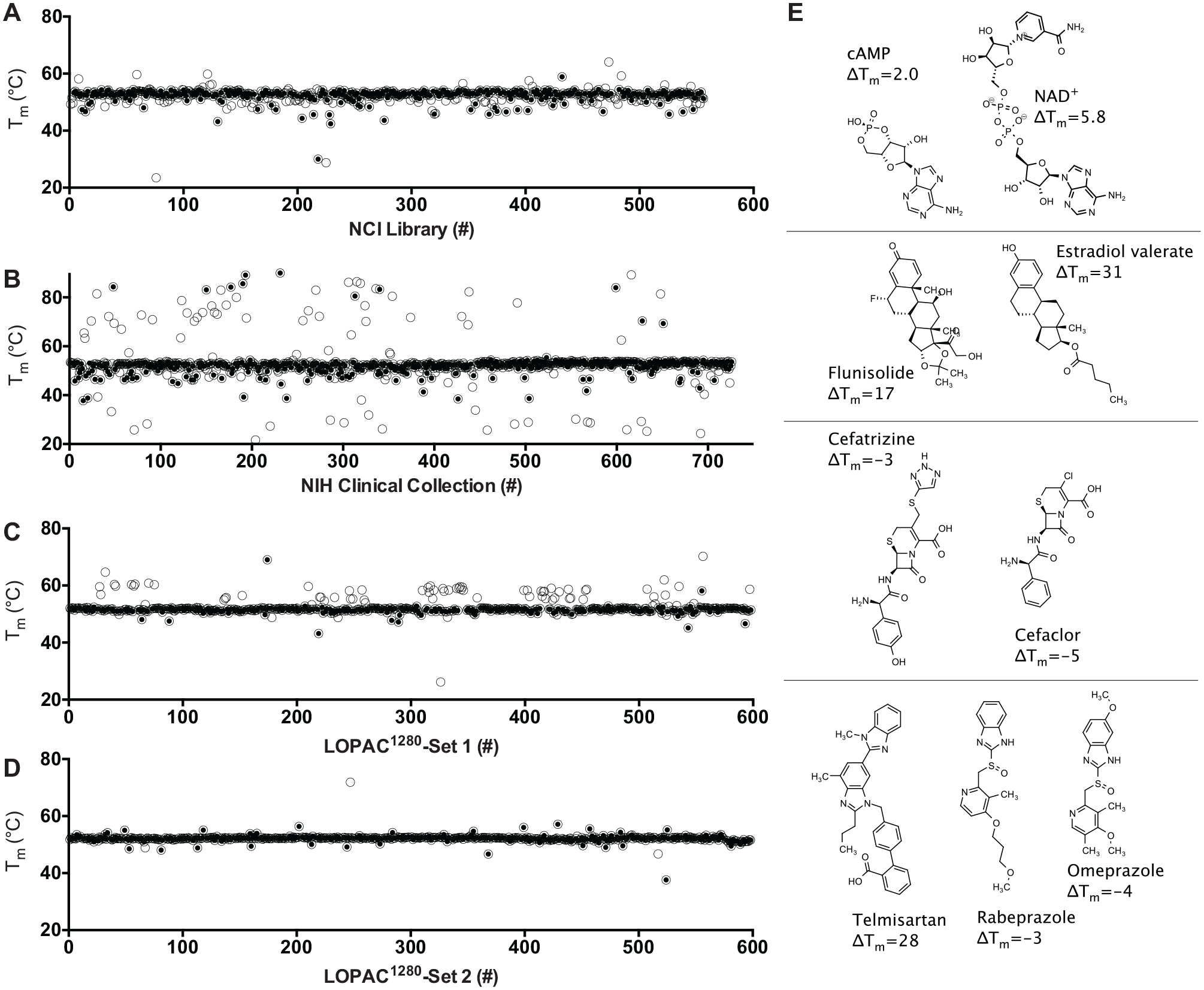
DSF screens of FDA-approved drugs and drug-like compounds for SARS-CoV-2 Mac1 ligands. Tm values calculated by fitting melting curves to eq 1 (open circles) obtained for Mac1 in the presence of each compound in (
Prioritizing Hits in DSF Screens
Screening results reveal that the major limitation of this approach is that a high percentage of compounds in some libraries influence the observed protein melting temperature, which was particularly evident with the NIHcc. Many of these compounds either quench fluorescence, fluoresce themselves, or interact with the reporter dye. To exclude such compounds in follow-up experiments, the TSA-CRAFT software package was used to identify what it defines as “typical” curves (
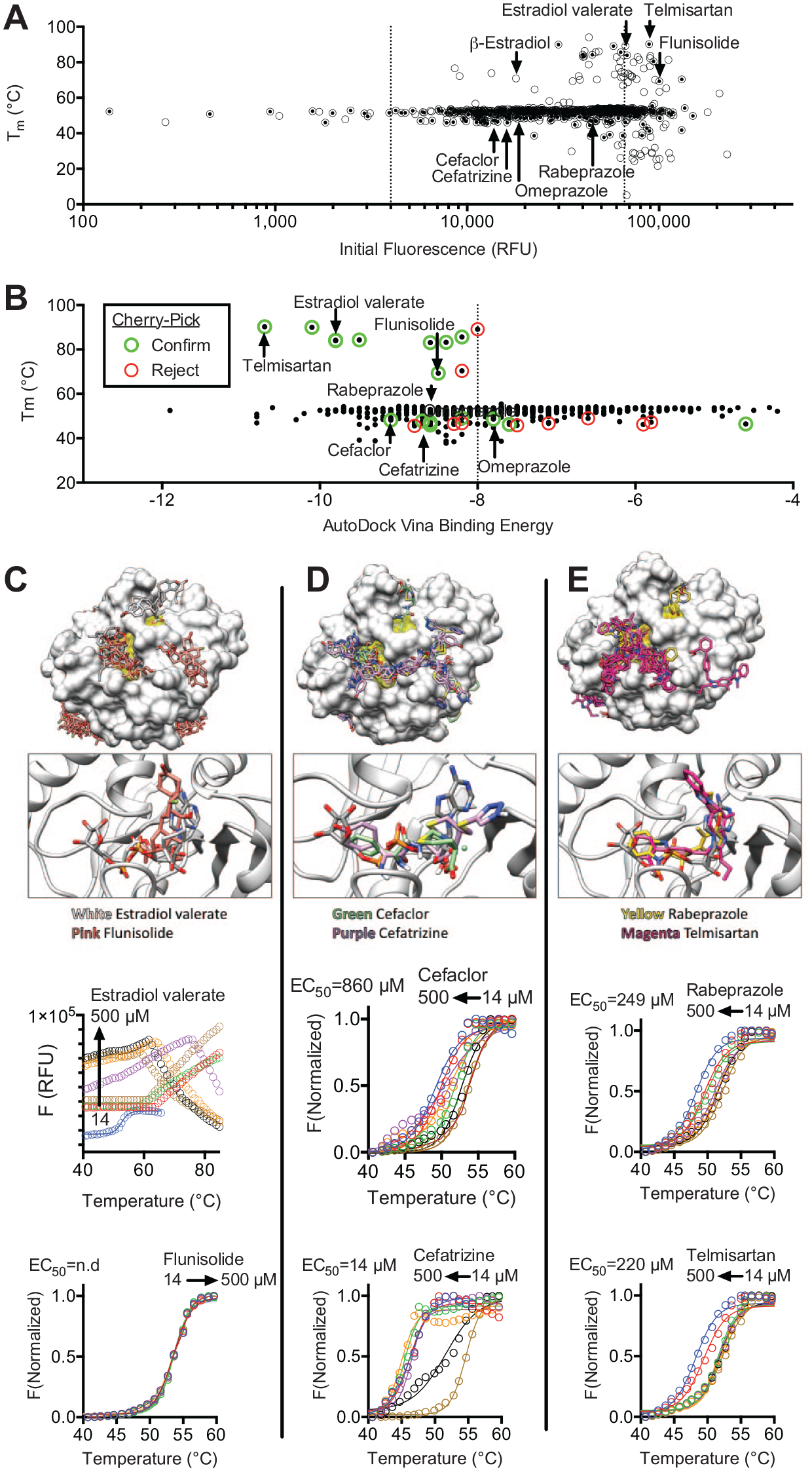
Methods to prioritize hits from DSF screens. (
As another method for hit prioritization, “virtual” screens were performed with SARS-CoV-2 Mac1 crystal structures (PDB files 6WEY, 6W02, and 6VXS) as targets in the program AutoDock Vina (
By combining these prioritization methods, compounds that more likely bind Mac1 could be differentiated from those that likely do not. For example, all nucleotides that were hits yielded results similar to those seen with ADPr. However, the steroids either yielded atypical curves (β-estradiol) or increased the DSF assay fluorescence (estradiol valerate and flunisolide). When new aliquots of selected compounds were purchased, abnormal melting curves were observed with estradiol valerate and no effects were observed with flunisolide (
Structure of cAMP Bound to SARS-CoV-2 Mac1
The protein construct used to determine the structure of the apoenzyme (PDB ID 6WEY; nsp3 residues 207–377) crystallized with such tight packing that it proved impossible to obtain structures of the ligand-bound Mac1 protein. Thus, we were in the unusual position of trying to get looser packing and poorer resolution. Adding a single N-terminal residue to the protein (nsp3 residues 206–374) was enough to change the packing from orthorhombic (P212121) to monoclinic (P21). To verify binding, each of the compounds in
Figure 2
was both co-crystallized with this new Mac1 construct and soaked into crystals of the apoprotein. Despite considerable effort, we were only able to obtain a complex structure with cAMP. This model contains one molecule of Mac1 in the asymmetric unit, comprising 166 amino acids (residues 208–373), 135 water molecules, and 1 molecule of cAMP. The cAMP binds in the cleft between the β2-α2 loop (residues K248–V253) and the β5-α5 loop (residues L331–D339); the bottom of this cleft is formed by strand β2 (
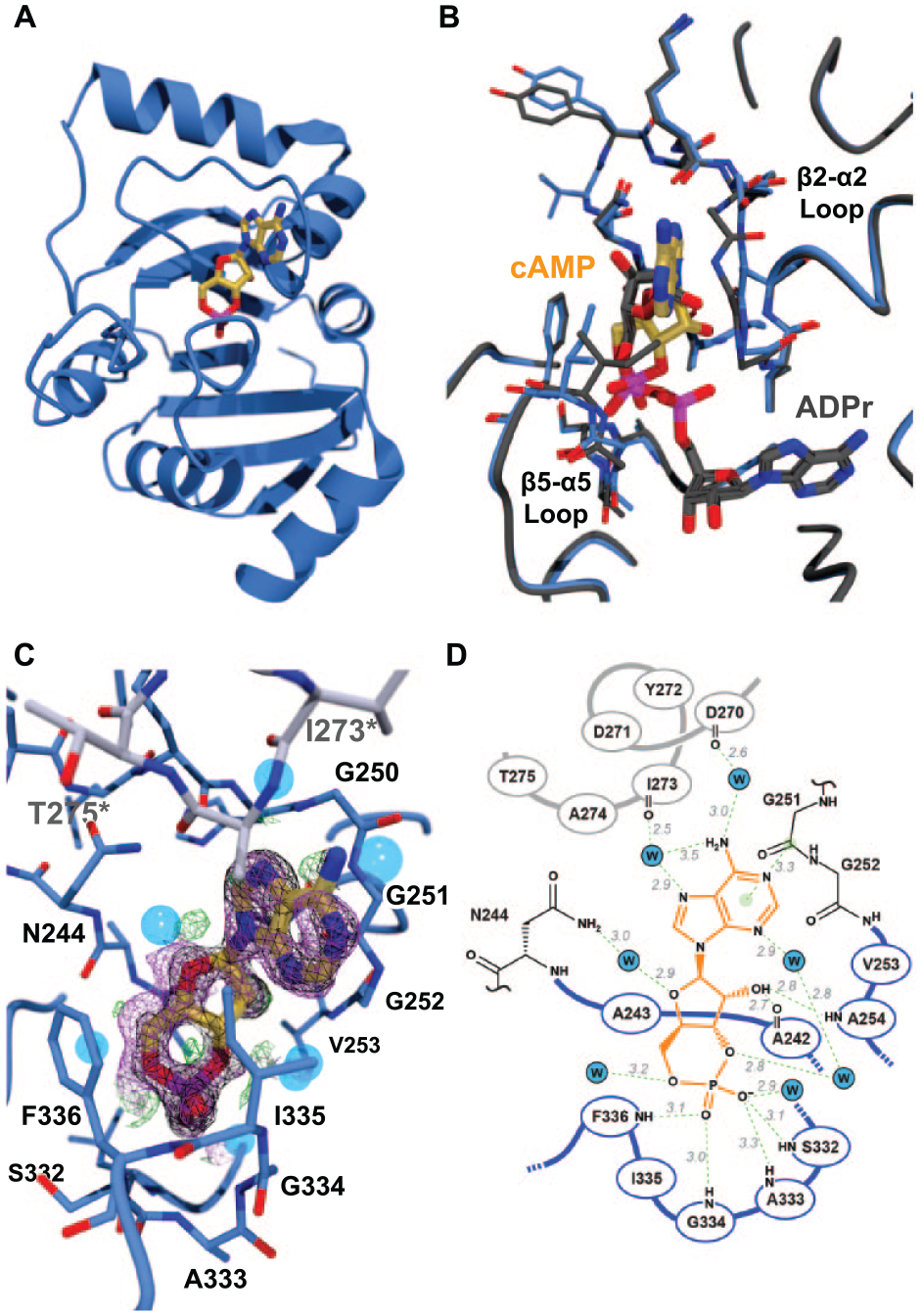
The Mac1 protein binds cAMP in an unexpected way. (
The binding mode of cAMP was compared with those of ADPr (PDB ID 6W02;
10
Discussion
The idea that SARS-CoV Mac1 functions as an enzyme in the cell to remove ADPr from antiviral proteins suggests that Mac1 might be an important new drug target for COVID-19.
23
A thermal shift binding assay was therefore developed to facilitate those efforts. The main drawback with the DSF assay was the high percentage of hits for some libraries. Various methods to successfully prioritize hits are described above, with the simplest being an examination of the initial fluorescence values in DSF assays (
The most intriguing hits in DSF screens were the steroids and telmisartan (
The idea that SARS-CoV-2 ADPr-binding cleft can accommodate other ligands is supported by the x-ray structure of Mac1-bound cAMP. Surprisingly, the adenine base of cAMP does not bind in the adenine-binding cleft identified in the structures of Mac1 bound to ADPr or AMP. 10 Instead, cAMP binds in the site occupied by the β-phosphate/terminal ribose unit of ADPr. This result underscores the importance of computational modeling and experimental structure determination in assessing the hits from high-throughput screening campaigns. Based on this crystal structure, it is likely that scaffolds containing a central phosphate (or diphosphate) or sulfate group could be expected to bind to this same ADPr-binding cleft. The interaction between Mac1 and cAMP, which binds Mac1 with a similar affinity as ADPr, 3 points to other possible biological roles for Mac1 and hints that cyclic mono- or dinucleotide second messengers might allosterically modulate other nsp3 activities.
The next step in this project will be to examine the effect of promising antiviral compounds on cells harboring SAR-CoV-2 or surrogate reporter viruses. Some of the compounds above might inhibit SARS-CoV-2 replication based on the fact that Shimizu et al. showed that small molecules found in virtual screens targeting the homologous nsp3 macrodomain from Chikungunya virus inhibit replication of Chikungunya replicons. 25 However, further chemical optimization would most likely be necessary for these probes to be useful in cellular studies. Fortunately, many hits reported here are already Food and Drug Administration (FDA)-approved drugs with hundreds of analogs available to facilitate such work. Alternatively, this optimized DSF assay could be used to screen larger, more diverse libraries for more attractive probe candidates.
Supplemental Material
Supplemental_Information – Supplemental material for Discovery of Drug-Like Ligands for the Mac1 Domain of SARS-CoV-2 Nsp3
Supplemental material, Supplemental_Information for Discovery of Drug-Like Ligands for the Mac1 Domain of SARS-CoV-2 Nsp3 by Rajdeep S. Virdi, Robert V. Bavisotto, Nicholas C. Hopper, Nemanja Vuksanovic, Trevor R. Melkonian, Nicholas R. Silvaggi and David N. Frick in SLAS Discovery
Footnotes
Acknowledgements
The authors are grateful to Wilfred Tysoe and Alexander (Leggy) Arnold, for helpful discussions and technical assistance, and to Daad Saffarini, for allowing this work to proceed during the COVID-19 pandemic lockdown.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by The University of Wisconsin–Milwaukee’s generous F&A cost return policy (National Institutes of Health grant no. R01 AI088001) and is based upon work supported by the National Science Foundation under grant nos. CHE-1855199 and CHE-1903899 (N.V. and N.R.S.). This research used resources of the Advanced Photon Source, a U.S. Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by the Argonne National Laboratory under contract no. DE-AC02-06CH11357. Use of the LS-CAT Sector 21 was supported by the Michigan Economic Development Corporation and the Michigan Technology Tri-Corridor (grant no. 085P1000817).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.