Abstract
Nucleic acid aptamers that specifically bind to other molecules are mostly obtained through the systematic evolution of ligands by exponential enrichment (SELEX). Because SELEX is a time-consuming procedure, the in silico design of specific aptamers has recently become a progressive approach. HIV-1 surface glycoprotein gp120, which is involved in the early stages of HIV-1 infection, is an attractive target for RNA and DNA aptamer selection. In this study, four single-stranded DNA aptamers, referred to as HD2, HD3, HD4, and HD5, that had the ability of HIV-1 inhibition were designed in silico. In a proposed non-SELEX approach, some parts of the B40 aptamer sequence, which interacted with gp120, were isolated and considered as a separate aptamer sequence. Then, to obtain the best docking scores of the HDOCK server and Hex software, some modifications, insertions, and deletions were applied to each selected sequence. Finally, the cytotoxicity and HIV inhibition of the selected aptamers were evaluated experimentally. Results demonstrated that the selected aptamers could inhibit HIV-1 infection by up to 80%, without any cytotoxicity. Therefore, this new non-SELEX approach could be considered a simple, fast, and efficient method for aptamer selection.
Introduction
The application of anti-HIV aptamers is one of the most progressive approaches with great potential for the diagnosis and treatment of AIDS. 1 Nucleic acid aptamers are small single-stranded DNA (ssDNA) or RNA (ssRNA) molecules capable of adopting complex tertiary structures enabling the specific binding to other molecules. Aptamers can specifically recognize a wide range of targets, including metal ions, organic molecules, viruses, and whole cells. 2 Valuable advantages of aptamers such as high specificity, sensibility, accuracy, cost-efficiency, small size, and the nonimmunogenic nature make them a powerful diagnostic and therapeutic tool for human diseases. Moreover, the possibility of in vitro synthesis offers remarkable flexibility in the design and modification of aptamers. 3 Aptamers that target HIV type 1 (HIV-1) reverse transcriptase, RNaseH, Tat, Gag, integrase, nucleocapsid, and gp120 are some of the most studied examples. HIV-1 surface glycoprotein gp120 interacts with the human surface receptor CD4 and induces receptor conformational changes leading to HIV-1 entry into the host cell. Because of its importance in the viral infection process, several gp120 RNA aptamers have been selected to block gp120 and the CD4 interaction. 3,4 Aptamer B40 and its truncated derivative B40t77 were reported have high specificity to the HIV-1 R5 strain and neutralization in peripheral blood cells. 5 In addition, another shorter B40 derivative, UCLA1, is able to inhibit HIV-1 entry at the nanomolar range. B40 and UCLA1 have been shown to reduce virus infectivity of peripheral blood mononuclear cells by up to 85% and 82%, respectively. Although several RNA aptamers have been isolated to block the interaction of gp120 and CD4 and prevent HIV-1 entry, 6 the design of a DNA aptamer has been less considered, and thus, there are no reports on DNA aptamer selection to bind gp120 specifically.
Aptamers are mostly selected from a large random library of nucleic acid sequences through an iterative process known as SELEX. This method, which consists of multiple rounds of selection and amplification, is a time-consuming procedure. 7 Furthermore, selected aptamers require truncations to obtain smaller aptamers. However, as computational methods combine in vitro and in silico approaches and provide some prediction about aptamer affinity and bioconjugate stability, they could provide higher efficiency for aptamer selection. 8 In 2016, Ahirwar and coworkers 9 suggested an approach in which the criteria used for the selection of aptamer-alike sequences to target estrogen receptor alpha were the inverted repeat nature of estrogen response elements and formation of stable hairpins. Also, RNA analogs were modeled using AutoDock Vina, HADDOCK, and PatchDock. 7 In the screening method of Chushak and Stone, 8 decreasing the simple structural motifs and increasing the stability of structures were the main parameters considered in aptamer selection. They analyzed the secondary structure of more than 2.5 × 108 RNA sequences to select 100,000 sequences for achieving tertiary structure library. In the next step, computational docking limited the number of aptamers up to 5 × 103 sequences. Ultimately, a library of 103 to 104 RNA sequences were used for the experimental verification.
In this study, we proposed a novel non-SELEX screening approach for the selection of aptamers. We used HEX and HDOCK docking tools to analyze the affinity of 1000 ssDNA sequences to gp120. Low docking scores and binding of the aptamer to conserved regions and nonglycosylated residues of Gp120 were the main parameters considered in the aptamer screening. The aim of this study was to find a simple and fast method for aptamer selection without the need for facilities and high levels of bioinformatics knowledge.
Materials and Methods
The practical steps of the suggested approach can be summarized as several rounds of modification and docking to obtain lower docking scores. Thus, the first criterion was lower docking scores. After that, binding of the aptamer to conserved regions and nonglycosylated residues of Gp120 were considered the main parameters for aptamer selection. Details of each step of the aptamer-screening approach are described in the following sections.
Selection of gp120 Aptamer and Prediction of the Secondary and Tertiary Structures
Three strains of HIV-1 (accession numbers AB703608.1, AB703613.1, and AB703614.1) and the RefSeq (NP_579894.2) Gp120 sequences were obtained from NCBI (https://www.ncbi.nlm.nih.gov/). To start, the interaction of the B40 aptamer with the gp120 sequences was studied using two docking tools: Hex software and HDOCK online web server (http://hdock.phys.hust.edu.cn/). HDOCK finds homologous templates of the given sequences and then builds the structures from the monomer or complex templates for docking. Hex is a docking and superposition program that uses spherical polar Fourier correlations to accelerate the calculations. Moreover, YASARA software was used to visualize the docking output data. Then, B40 parts with more interacted residues were separated for use as an aptamer sequence. To obtain a better interaction between the aptamer and gp120, which resulted in lower docking scores, some modifications and deletions were performed, and DNA sequences with a nucleotide length of 30 to 50 were generated. In addition, secondary and tertiary structures were predicted to perform further studies about the interaction conditions between aptamer and Gp120.
Secondary structures of the ssDNA aptamers were generated by the Mfold web server (http://unafold.rna.albany.edu/?q=mfold). The Mfold web server consists of some closely related software applications used for the prediction of nucleic acid hybridization, folding, and melting temperatures and it has been used in many recent, important scientific reports. The thermodynamic details of each aptamer structure were also predicted using the mentioned server. 10 Avogadro (https://sourceforge.net/projects/avogadro/files/latest/download), a powerful software for the prediction of the tertiary structure of macromolecules, was used for three-dimensional (3D) structural modeling of the aptamer. 11 Similarly, SWISS-MODEL Server (https://swissmodel.expasy.org/) was used for modeling the 3-D structure of Gp120. 12 Finally, the output data in PDB format were used for docking studies.
The obtained PDB files of Gp120 and aptamer molecules were used as the input receptor and ligand molecule, respectively, in Hex and HDOCK. The obtained docking scores of the two methods were compared. In addition, the interacted residues of both the ligand and receptor molecules were identified by YASARA software. Because B40 is a long (117nt) ssRNA aptamer, some modifications were made to obtain lower docking scores (higher negative scores) in both docking tools to achieve more affinity between the truncated ssDNA aptamer and gp120. This procedure was done using the genetic algorithm. In summary, the aim of our attempt was to maintain the favorite modifications (which lead to lower docking scores) and remove the others. Also, aptamers that bind to variable regions and glycosylated sites of Gp120 were excluded in the next step. As Gp120 is a highly glycosylated protein, aptamers that interact with glycosylated residues were not considered. On the basis of the mentioned criteria, four aptamer sequences were prepared for further in vitro studies.
In Vitro Study of the Anti-HIV Effect of the Aptamer
Blood Sampling and Lymphocyte Culture
Three blood samples with a volume of 8 mL were taken from volunteers who were seronegative for HIV. The collected blood was immediately transferred into sterilized tubes containing heparin to prevent coagulation. Centrifugation of the blood samples in Ficoll-Hypaque (1800 rpm for 20 min) separated the lymphocyte from other blood compartments. Lymphocytes of the intermediate layer were cultured in DMEM medium containing antibiotics and 10% fetal bovine serum, transferred to a 24-well plate, and incubated at 37 °C for 1 h. To avoid microbial contamination, all of these steps were carried out in sterile conditions in a laminar flow hood. DMEM culture medium and Ficoll-Hypaque were purchased from Gibco-BRL (Grand Island, NY).
Preparation of Aptamer DNA Sequences and Single-Cycle Replication HIV
Aptamer sequences were purchased from Gene Fanavaran (Tehran, Iran). Single-cycle replication (SCR) HIV was obtained from former experimental studies of Behbahani and coworkers. 13
Cytotoxicity Assay
A portion of the cultured lymphocyte cells was transferred to 96-well plates. Subsequently, two concentrations of aptamer, 25 and 40 μM, were added to different wells in triplicate and incubated at 37 °C overnight. The negative control group was treated with medium only. Then, 12 µL MTT was added to each well and incubated for 24 h. The optical density (OD) of the samples was measured at 450 nm using a microplate reader, and cell viability was determined based on the following equation: cell viability (%) = (OD treated/OD control) × 100,
where the OD control was recorded in the absence of DNA aptamer and OD treated was in the presence of aptamer.
HIV Inhibition Assay
An Ag/Ab HIV kit (bioMérieux, Craponne, France) was used to test the inhibitory activity of selected aptamers on gp120-CD4 binding. In summary, 15 µL of SCR HIV (with titers up to 6 × 104 infectious units/mL) was preincubated for 30 min in the presence of four different concentrations of aptamer (10, 25, 40, and 55 μM) and one random sequence aptamer with no cytotoxicity and HIV inhibition effect as control, which were then added to the cultured lymphocytes. Negative control samples were treated with only sterile distilled water. Moreover, different concentrations of azidothymidine (AZT), 1, 0.1 and 0.01 μM, were used to compare the results of the two treatment groups. The plate was incubated at 37 °C overnight. All other steps were accomplished according to the manufacturer’s procedure. Then, the optical density of each well was measured at 450 nm by a microplate reader to determine the inhibitory efficacy of the aptamer to HIV entry. Finally, the following equation was used to calculate the inhibitory effect of aptamer: inhibition percentage (%) = [1 – (OD treated/OD control)] × 100,
where OD control and OD treated were measured in the presence of random and designed aptamer sequences, respectively.
Statistical Analysis
The inhibition percentage and cytotoxicity means ± standard deviations (SDs) was estimated for three independent experiments, and each was carried out in duplicate. Microsoft Excel 2010 was used for drawing graphs. The differences between treated and control groups were compared by Wilcoxon rank-sum test using SPSS software.
Results
Design of gp120 Aptamer
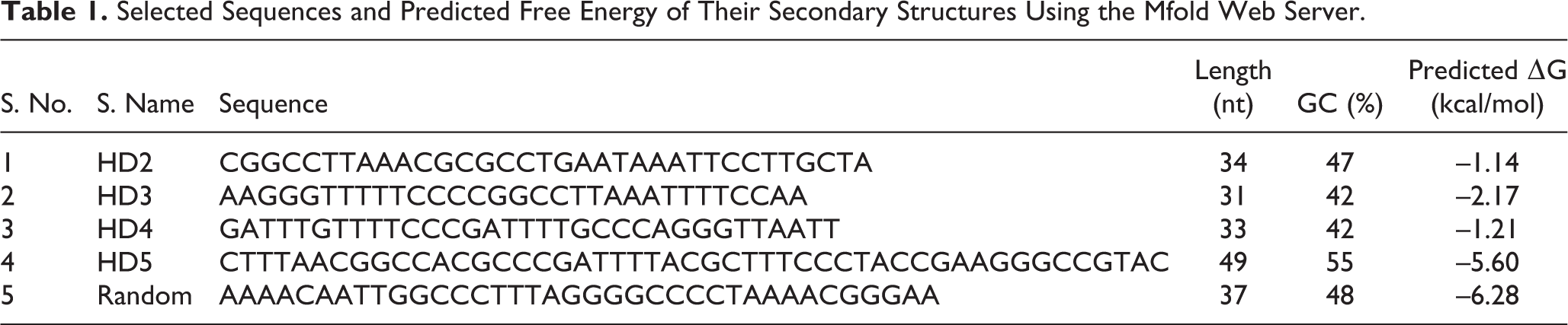
The characteristics of the four selected aptamers and a random sequence are shown in Table 1 . Among the selected sequences, the aptamer HD5 had the longest sequence length (49 nucleotides), highest GC mass (55%), and greatest secondary structure stability (–5.6). In addition, the secondary structures of these sequences predicted by the Mfold web server are illustrated in Figure 1 . As shown in this figure, in comparison with the other aptamers, sequence HD5 adopted the most complicated structure with several loops and stems.
Selected Sequences and Predicted Free Energy of Their Secondary Structures Using the Mfold Web Server.
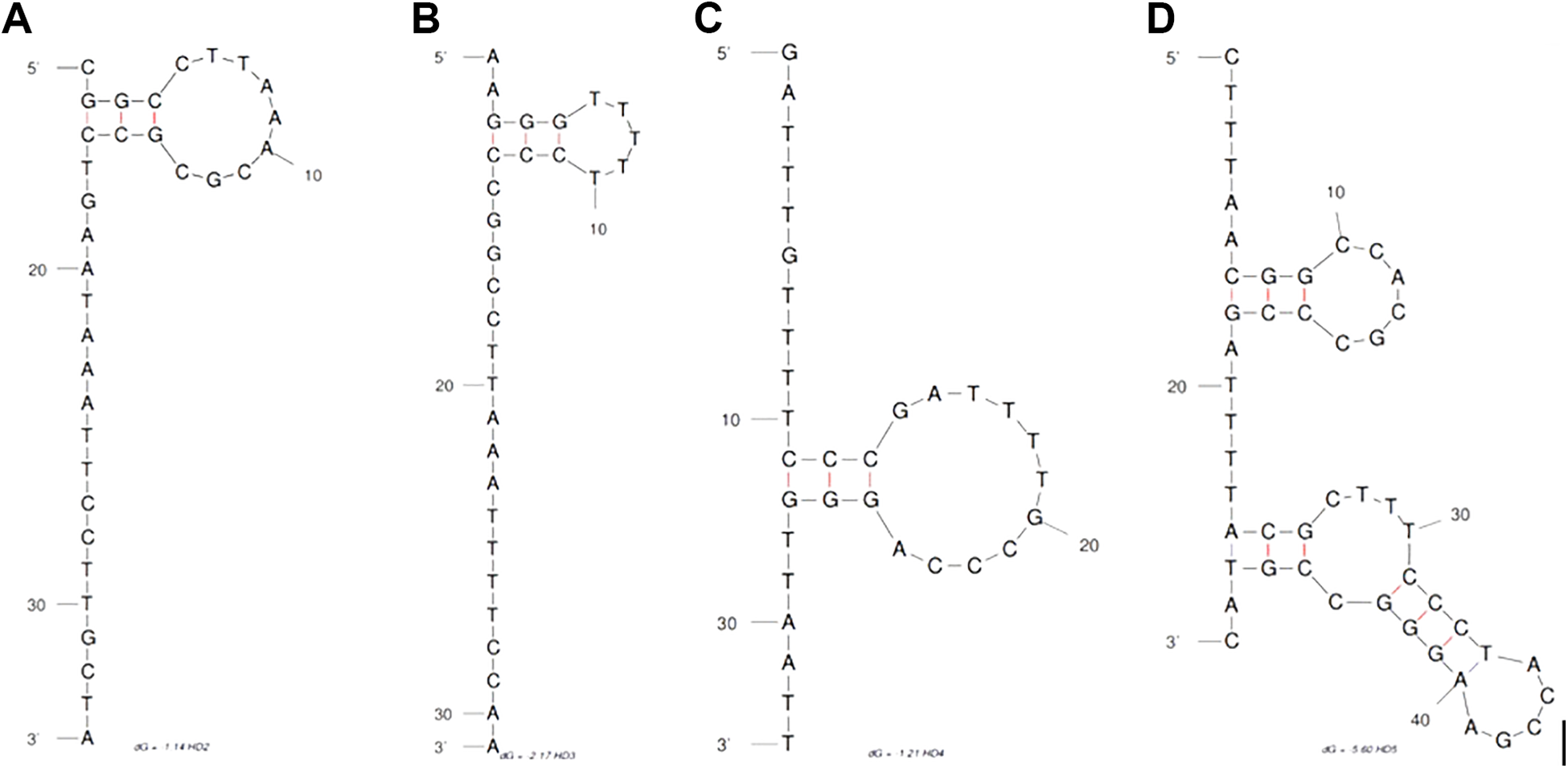
Secondary structures of four selected sequences predicted by the Mfold web server. (
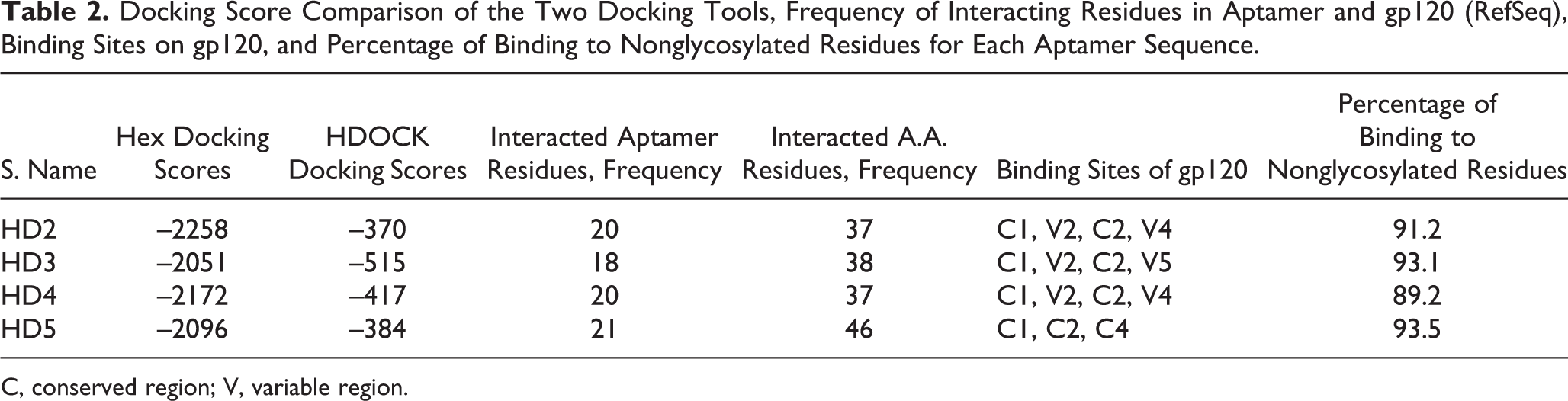
In Table 2 , the docking scores obtained from the two docking tools are compared. Four sequences with the best docking scores from both docking tools were selected. HD2 and HD4 showed the best Hex docking scores, whereas HD3 and HD4 achieved the best HDOCK scores, respectively. In fact, the HD4 sequence obtained one of the best docking scores from both docking tools.
Docking Score Comparison of the Two Docking Tools, Frequency of Interacting Residues in Aptamer and gp120 (RefSeq), Binding Sites on gp120, and Percentage of Binding to Nonglycosylated Residues for Each Aptamer Sequence.
C, conserved region; V, variable region.
Moreover, the predicted binding sites of aptamer and Gp120 as well as the binding residues of each molecule were obtained by the HDOCK web server and YASARA software, respectively. HD5, with the highest sequence length, had the highest frequency of interacted aptamer and gp120 residues. Based on the results of the docking tools, all aptamers interacted with conserved (C1 and C2) and variable regions (V2, V4, and V5), but HD5 interacted with only gp120 conserved regions (C1, C2, and C4). Thus, it appears that this aptamer can recognize various strains of this protein and bind them with high affinity ( Table 2 ). Furthermore, the maximum interaction with nonglycosylated gp120 residues was observed in HD3 and HD5 aptamers.
In Vitro Study of Aptamer Anti-HIV Effect
The result of the anti-HIV activity of aptamers is shown in Figure 2 . The results demonstrated that all selected aptamers have an anti-HIV effect at different concentrations (10, 25, 40, and 55 μM). However, the most effective dose to reduce virus titer was achieved by 10 μM of each aptamer, resulting in a 49% to 80% inhibition. Aptamer HD4 at a concentration of 10 μM inhibited 80.14% viral replication. It was followed by HD5, HD3, and HD2, which caused 78.4%, 74.48%, and 49.23% HIV inhibition, respectively. Also, it appears that experimental results were more consistent with the HDOCK docking predictions.
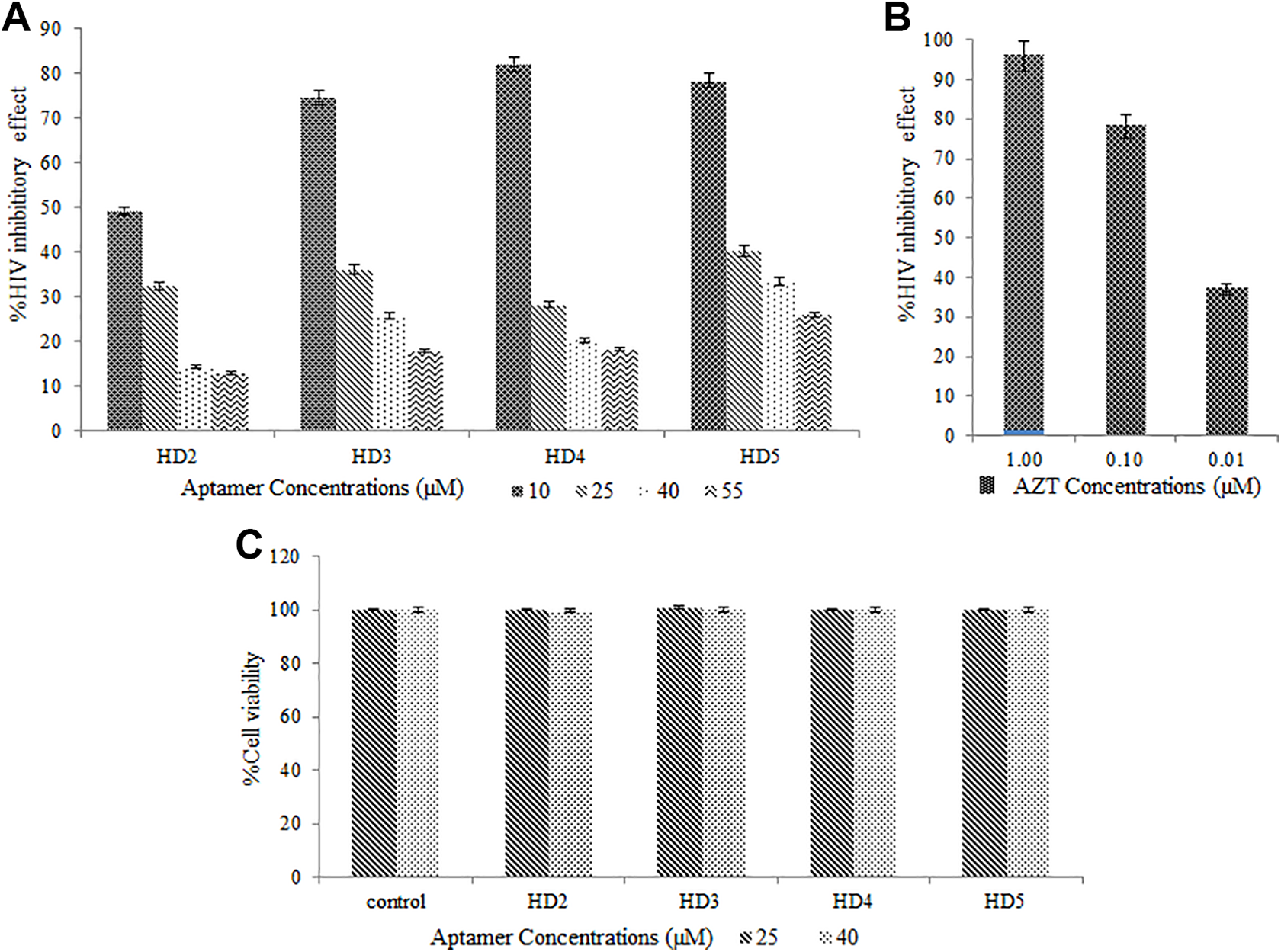
Inhibition percentage of HIV-1 replication caused by (
Cytotoxicity Assay
In the application of the aptamer in the body, the potential for adverse effects on cell function should be studied. Thus, lymphocyte cell viability was compared in aptamer treated and control samples after MTT addition. The measurement of cell viability showed that synthetic aptamers have no considerable cytotoxicity on cultured lymphocytes ( Figure 2 ).
Discussion
Although the classical method of aptamer selection, SELEX, allows researchers to isolate many specific aptamers for a wide variety of targets, it has some inevitable limitations. 14 To reduce the number of cycles and consequently provide the possibility of artifact generation, the SELEX procedure was coupled with high-throughput (HT) sequencing technologies. However, the data obtained by a single HT-SELEX experiment require some computational tools to identify aptamers with greater binding properties. 15 Therefore, the development of a new approach that provides more efficiency in the selection of specific aptamers seems necessary. 8 In the present study, an in silico screening approach was described for the selection of a new ssDNA aptamer to HIV-1 Gp120. Because of the key role of Gp120 in the initial phases of HIV infection, it was attended as the target of various RNA aptamers to block its activity. 5,6,16 The sequence of the gp120 gene is highly variable in different isolates, but it consists of a succession of five variable (V1 to V5) and five conserved (C1 to C5) regions. Because the CD4-binding site of gp120 is composed of some discontinuous sequences with major conserved regions, 17 one of the criteria mentioned in aptamer selection was binding to conserved sequences. Moreover, aptamers capable of binding to conserved regions can recognize various strains of HIV. In addition, because of the heavy glycosylation of gp120, 18 aptamers with less affinity to glycosylated regions were selected for the next screening step. In summary, the main criteria considered in the aptamer screening were low docking scores and binding of aptamer to conserved regions and nonglycosylated residues of Gp120.
On the other hand, appropriate features of ssDNA aptamers in comparison with RNA-based aptamers such as stability as well as reasonable biocompatibility suggest the use of a high potential protective and curative treatment for clinical use. Recently, some DNA aptamer sequences were isolated to CD4 by the SELEX method, but there is no report of the selection of ssDNA aptamer for gp120. The aptamers selected for CD4 caused 20% to 50% inhibition of binding HIV to Gp120. 19 In this work, we proposed a new non-SELEX method for aptamer screening that could achieve four aptamers (HD2, HD3, HD4, and HD5) with 49% to 80% inhibition of HIV-1. The inhibitory effect of the selected aptamers was comparable with some commercial drugs such as AZT. However, high concentrations of the aptamers may lead to aggregation and lower efficacy. Obtained results make this concept a method of choice for the non-SELEX approach of aptamer design, aptamer binding improvement, and aptamer truncation. In this study, we used HEX and HDOCK docking tools and then YASARA for visualization of bioconjugate structure and binding sites. Comparison of the obtained docking results of the two docking tools showed that the results of docking by HDOCK relative to HEX were more relevant to experimental data. In vitro results showed that HDOCK, which is based on a hybrid algorithm of template-based modeling and ab initio free docking, is an efficient web server for protein–protein and protein–DNA/RNA docking with an acceptable accuracy in the prediction of bioconjugate free energy. In vitro studies demonstrated that aptamers with lower docking scores show better efficiency in binding to gp120. In conclusion, finding a new, simple, fast, and accessible method for aptamer selection will help to overcome the present challenges in the development of aptamers. It appears that the suggested non-SELEX method may help to address this requirement. Moreover, selected ssDNA aptamers may be applied to the prevention of HIV-1 transmission. However, the binding affinity of these aptamers may improve in combination with drug delivery or multimeric systems, as described in former studies. 19 -21
Footnotes
Acknowledgments
Support for this study provided by the University of Isfahan is highly acknowledged.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: funding was provided by the University of Isfahan, Hezar Jareeb St., Isfahan 81746-73441, Iran.
