Abstract
RNA molecules have a variety of cellular functions that can drive disease pathologies. They are without a doubt one of the most intriguing yet controversial small-molecule drug targets. The ability to widely target RNA with small molecules could be revolutionary, once the right tools, assays, and targets are selected, thereby defining which biomolecules are targetable and what constitutes drug-like small molecules. Indeed, approaches developed over the past 5–10 years have changed the face of small molecule–RNA targeting by addressing historic concerns regarding affinity, selectivity, and structural dynamics. Presently, selective RNA–protein complex stabilizing drugs such as branaplam and risdiplam are in clinical trials for the modulation of SMN2 splicing, compounds identified from phenotypic screens with serendipitous outcomes. Fully developing RNA as a druggable target will require a target engagement-driven approach, and evolving chemical collections will be important for the industrial development of this class of target. In this review we discuss target-directed approaches that can be used to identify RNA-binding compounds and the chemical knowledge we have today of small-molecule RNA binders.
Introduction
The central dogma of molecular biology states that DNA is transcribed to RNA, which is then translated to protein. This dogma implies two notions: (1) only proteins are capable of the complex processes that enable a cell to function, and (2) nucleic acids, especially RNA, serve only as passive carriers of genetic information for protein synthesis. However, the recent Encyclopedia of DNA Elements (ENCODE) project determined that only ∼1.5% of the human genome codes for proteins, yet more than 50% is transcribed from DNA to RNA.
1
This finding suggests that RNAs have a more significant role to play in cellular function than previously believed (
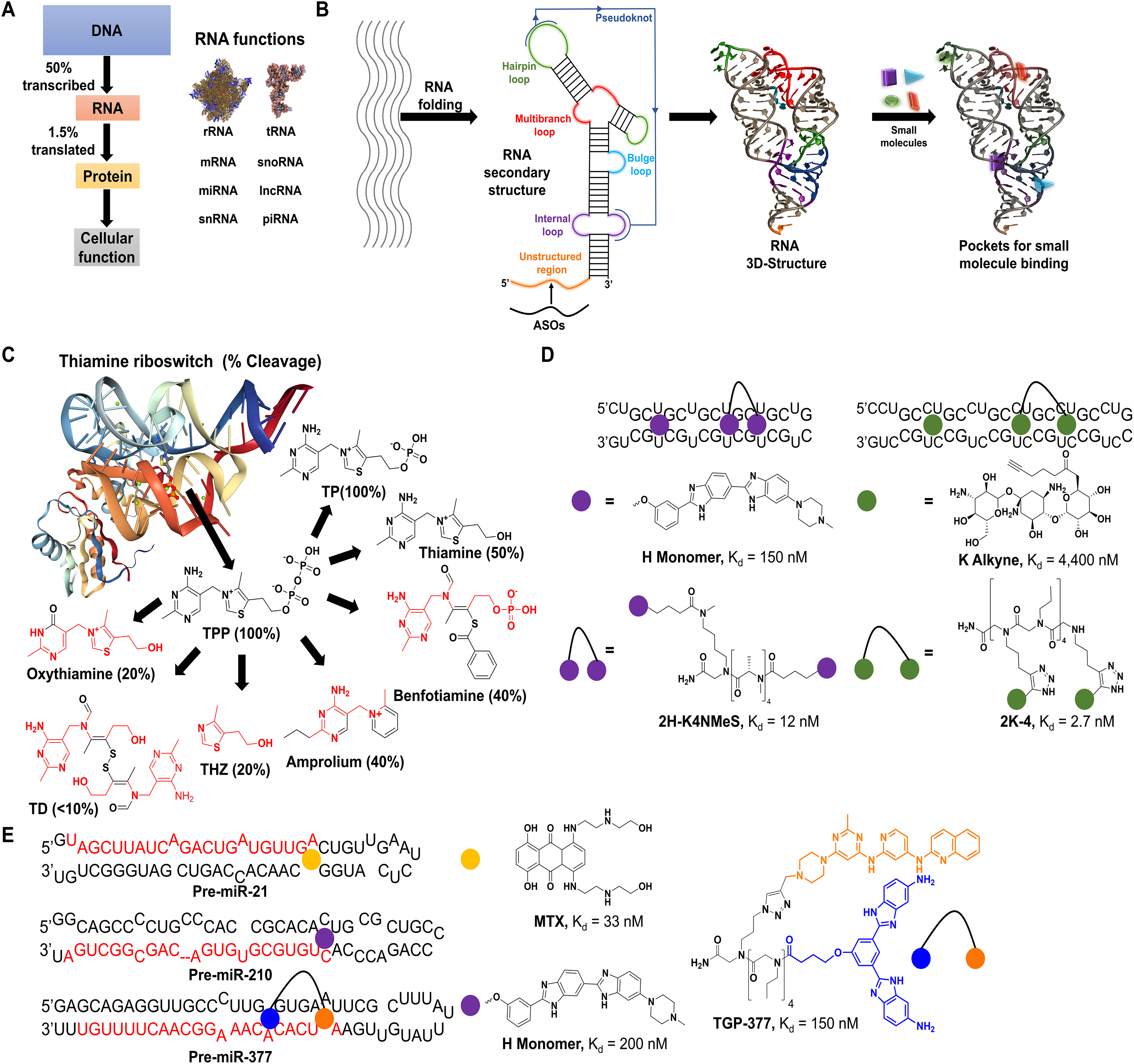
Schemes of key advances in RNA–small molecule binding. (
Initial efforts to drug RNA utilized antisense oligonucleotides (ASOs) that target RNA by Watson–Crick base pairing. This method has achieved moderate success in FDA-approved treatments
6
but contains several limitations including complications with efficacy, delivery, and off-target effects.
7
Its most notable drawback is its dependence on targeting unstructured regions of RNA, which is nevertheless intrinsically structured (
The 3D folds of RNA—including helical regions, hairpin loops, bulges, multibranched loops, and internal loops, as well as longer-range interactions like pseudoknots
12
—create potential pockets for the binding of small-molecule drugs (
Since the publication of these seminal works, more than 200 small-molecule binders to RNA have been identified
19,20
to target a wide variety of RNAs, from repeat expansions that cause myotonic dystrophy types 1 (DM1; r[CUG]) and 2 (DM2; r[CCUG]) (
Many reviews are available that focus on individual methods used to identify RNA-binding small molecules. However, a comprehensive review summarizing all of the tools available for this purpose has yet to emerge. Thus, the focus of this review is to provide a broad overview of the methodologies available to identify RNA-binding small molecules and cite examples of these methods being used to identify ligands of potential therapeutic benefit. The methods described herein are divided into three categories: (1) label-based methods, such as fluorescent intercalator displacement (FID), Förster resonance energy transfer (FRET), differential scanning fluorimetry (DSF), microscale thermophoresis (MST), and surface plasmon resonance (SPR); (2) label-free methods, including small-molecule microarrays (SMMs), mass spectrometry (MS)-based approaches, and nuclear magnetic resonance (NMR) spectroscopy; and (3) predictive approaches like virtual screening. These methods will be described in the context of their individual merits for assessing small-molecule RNA-binding capacity and additional potential for high-throughput screening (HTS) where applicable.
Label-Based Methods for Identifying an RNA Binder
Small-molecule binding to RNA can be assessed by a number of methods, most of which involve labeling the RNA at its terminus with a fluorescent probe such as 5-carboxyfluorescein (5-FAM), cyanine dyes (Cy5, Cy3), rhodamine, and Alexa Fluor dyes (
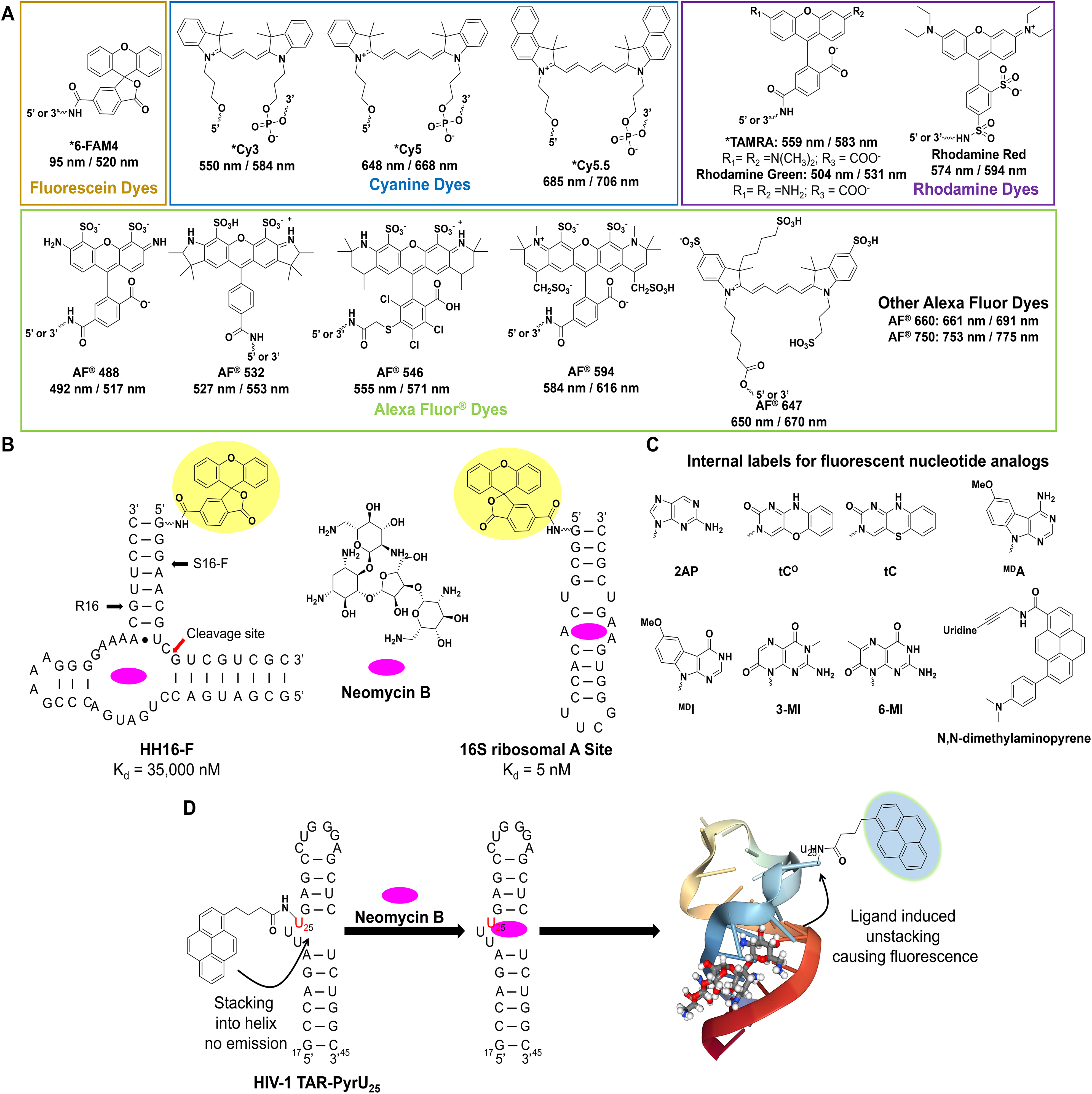
Common probes used for label-based methods to assess RNA binding. (
Direct Binding with End-Labeled RNAs
The fluorescence of labeled RNAs can be directly monitored to measure the binding interaction of small molecules to the RNA. This is due to differential stacking of the label with the helix, which can be sensitive to perturbations in the RNA’s structure that occur upon ligand binding. Numerous examples have demonstrated the application of this method to assess a small molecule’s RNA binding, notably the work of Llano-Sotelo and Chow. This group conjugated the S16 component of the hammerhead (HH) ribozyme with fluorescein on the 5′ terminus and, upon annealing with the R16 component, generated the HH16-F construct that could detect binding of neomycin B with a Kd of 35 µM (
Fluorescent Nucleotide Analogs
Internally labeled nucleobase analogs can serve to combat the limitations outlined above, as a number of strategies have been employed to increase their fluorescence properties. These methods include using isosteres, such as 2-aminopurine (2AP), that are intrinsically fluorescent; synthesizing base modified variants where the π system is extended; or using extended nucleobase analogs, where known fluorophores are conjugated onto the parent base via a rigid or flexible linker (
Generally speaking, the methods referenced above are applied to the study of RNA conformational dynamics, but in theory they can be applied to the study of small-molecule binding to RNA as well. This is exemplified by Blount and Tor’s work with identifying binders to HIV-1 TAR using pyrene U-labeled TAR RNA (
Fluorescence Polarization
FP is the selective excitation of a fluorophore using plane polarized light. The selection arises from the orientation of the molecules and is such that only a fraction of them will have their dipole in a parallel orientation relative to the excitation polarization. Depending on the species that the fluorophore is conjugated to—that is, a small molecule, nucleic acid, or protein—the species will tumble at different rates, a result of Brownian motion relative to the lifetime of the fluorophore, randomizing its orientation and with small species tumbling faster than larger species. Thus, when a photon is emitted, it will be unpolarized, and the slower-tumbling larger species will retain some of the original polarization (
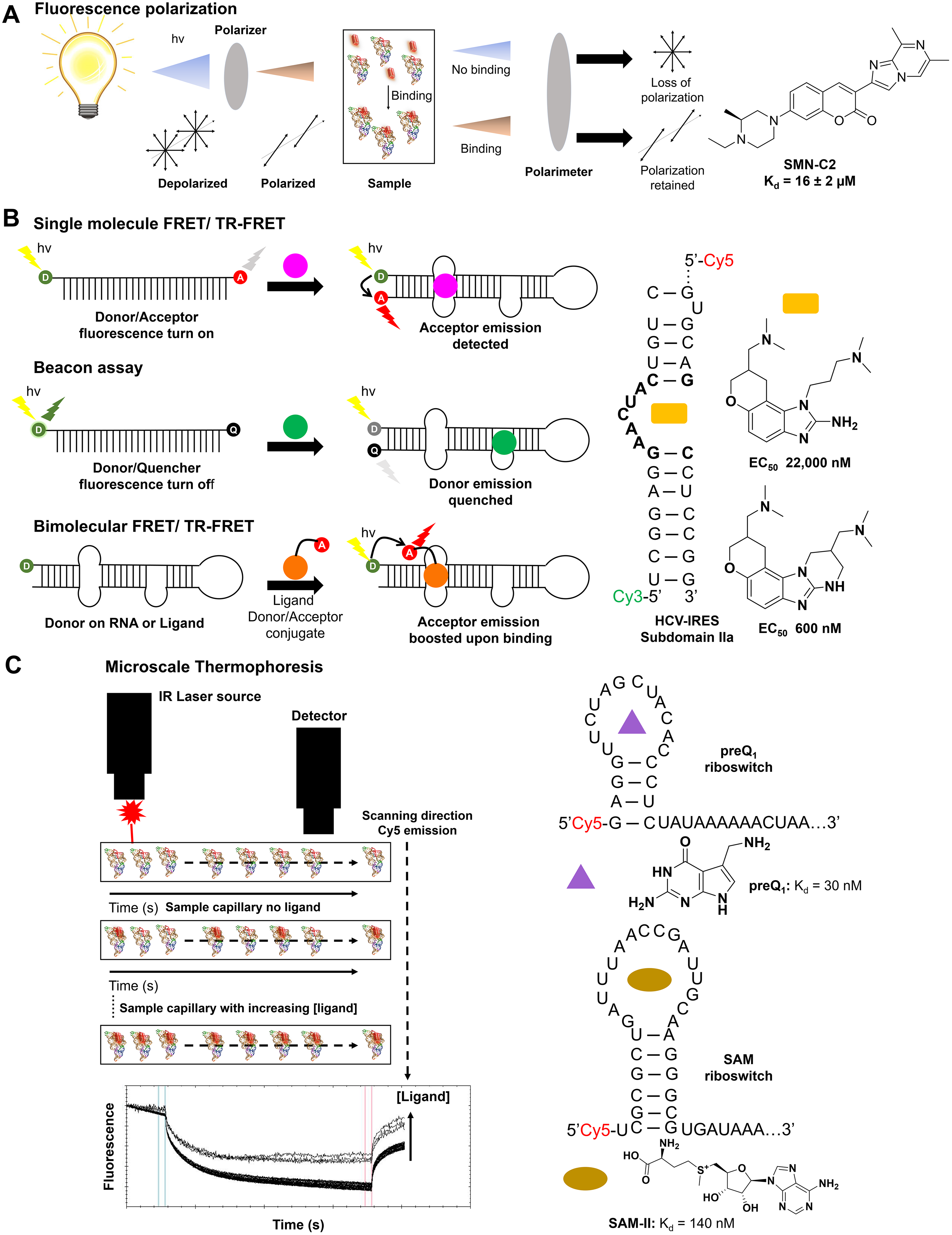
Schematics of label-based assays with examples. (
The FP assay has seen widespread use in the study of protein–nucleic acid and nucleic acid–small molecule binding analyses. Here, we focus on the latter application. In a pioneering study, PTC Therapeutics and F. Hoffmann-La Roche identified three novel splicing modifiers, SMN-C1, SMN-C2, and SMN-C3, that correct improper splicing of survival of motor neuron 2 (SMN2) in spinal muscular atrophy (SMA) patients.
49
This prompted a study of the molecules’ mode of action, initiated by J. Wang and colleagues, who demonstrated by the synthesis of an SMN-C2-Phe-coumaine (SMN-C2-Phe-Cu) analog that SMN-C2 binds an AGGAAG stretch in exon 7 of SMN2 with a Kd of 16 ± 2 µM, which was confirmed by cleavage analysis in vitro (
FRET-Based Approaches for RNA Binding
FRET and beacon assays are widely used methods for studying small-molecule binding to RNA in vitro and in vivo. This method uses a donor fluorochrome and an acceptor fluorochrome to form a donor–acceptor pair that is then conjugated to the biomolecule(s) of interest. The donor’s emission spectrum must sufficiently overlap with the excitation of the acceptor and be in sufficient proximity (1–10 nm) to allow FRET.
55
This process is the nonradiative transfer of energy between two fluorochromes in close proximity due to the coupling of their dipole moments, the efficiency of which is affected by many factors, the most important being the selection of the FRET pair. The following factors should also be considered for optimal assay performance: (1) the fluorescence lifetimes of the donor and the acceptor, (2) photobleaching of the fluorochromes, (3) the quantum yields of the donor and acceptor, and (4) bleed-through of donor emission into the acceptor emission channel. Emphasis is placed on points 1, 3, and 4 as these predominantly affect the assay’s sensitivity. FRET assays can be used to study both intra- and intermolecular binding events, where the donor–acceptor pair is conjugated to one species in the former case or two different species in the latter case (
Single-Molecule FRET
smFRET studies the changes of a biomolecule that is simultaneously labeled with both the donor and acceptor fluorochromes and has seen extensive use in identifying RNA-binding small molecules. Hermann and colleagues used a Cy3/Cy5 FRET pair conjugated to a mimic of the hepatitis C virus internal ribosome entry site (HCV-IRES) subdomain IIa riboswitch, aiming to identify novel benzimidazole modulators of the riboswitch’s conformation to inhibit viral translation. In doing so, two structurally similar derivatives were identified, exhibiting EC50 values of 22,000
35,56
and 600 nM,
57
respectively, in FRET binding assays (
Time-Resolved FRET
To overcome the shortcomings of bulk FRET measurements, TR-FRET can be employed. TR-FRET combines time-resolved fluorescence with FRET and hinges on the use of a short-lifetime donor with a long-lifetime acceptor, coupled with a delay in measurement of acceptor emission. This delay ensures sufficient decay of donor emission and background fluorescence, maximizing signal-to-noise ratios and assay sensitivity. Utilizing this approach, the intermolecular interaction of r(CUG) repeats that bind to muscleblind-like protein 1 (MBNL1) was studied using a biotinylated RNA with a streptavidin-XL665 tag and an MBNL1-His 6 /Anti-His-Tb as the donor–acceptor pair. 60 This RNA protein complex is responsible for DM1-associated splicing defects, and it is believed that disruption via small-molecule binding to the r(CUG)exp can provide lead therapeutics. Parkesh et al. identified 17 compounds from a computationally optimized library of 40 small molecules to inhibit MBNL1 binding by >85%, with IC50 values ranging from 50 to 1000 µM. 60,61 Rzuczek et al. further examined this model, identifying 28 r(CUG)exp RNA binders from a library of 320 small molecules (IC50 = 30–130 µM), with 5 of 28 being bioactive in cellulis. 62 Using multivalent approaches to rationally design RNA-binding ligands, bioactivity and potency were improved 1000-fold to low nanomolar levels. 21
Beacon Assays
Beacon assays also use a FRET-based sensor to study the presence of a specific nucleic acid in a PCR, or the binding of ligands to a specific sequence. Initially discovered by Tyagi and Kramer in their effort to study the origin and movement of mRNAs 63 in cells, the beacon probe is usually 25–35 nucleotides long and conjugated with a donor fluorophore on the 5′ end and a quencher fluorophore on the 3′ end, forming a donor–quencher (D/Q) pair. The first and last 5–7 nucleotides are complementary, forming a hairpin that brings the D/Q pair into close proximity and quenches the donor’s fluorescence. The single-stranded apical loop then targets the DNA or RNA of interest via Watson–Crick base pairs. Upon hybridization with the target DNA/RNA, the D/Q pair is separated, restoring the donor’s fluorescence. Factors to consider when designing a beacon probe include the size and sequence of the targeting loop for efficient hybridization to the target, the size of the stem for hairpin formation, and the selection of an adequate D/Q pair to maximize the signal-to-noise ratio. The interplay of these properties on assay design is summarized in greater detail by Zheng et al. 64
Beacon assays have been applied successfully to identify RNA-binding small molecules to a variety of targets like miRs and viruses. miRs are 22- to 23-nucleotide short noncoding RNAs that bind to the 3′ UTR of a target mRNA via Watson–Crick base pairing, downregulating its translation. 65 Bose et al. employed a DNA beacon (5′-FAM/3′-BHQ1) targeting the guide strand of miR-27a. The group identified six aminoglycosides that inhibited pre-miR-27a processing in vitro and further demonstrated that these aminoglycosides were bioactive in MCF-7 cells, displaying an IC50 of ≈20 µM. 66 Unlike the two-component beacon used by Bose et al., Bell et al. used a single-molecule beacon to study compounds capable of inhibiting binding of the Gag protein to stem loop 3 (SL3) in the human immunodeficiency virus 1 (HIV-1) Ψ packaging domain. This afforded NSC260594 from the Library of Pharmacologically Active Compounds (LOPAC) collection of 1280 compounds, establishing it as the best hit out of 78 primary hits with an IC50 of 4.5 µM. 67 Interestingly, further study of this compound by selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE) demonstrated that NSC260594 induced global changes in the packaging domain that limit incorporation of the viral guide RNA into virions. 68
The Arenz group also employed a single-molecule beacon to study the inhibition of Dicer processing of pre-let-7 by dodecapeptides. As a proof of concept, the aminoglycoside kanamycin was shown to bind to and inhibit processing by Dicer, as indicated by a 40% reduction in emission from the D/Q probe at 100 µM. Analysis of the three dodecapeptides S117 (Ac-NH-SSIYALEPDQKG-CONH2), S417 (Ac-NH-RYNIKKEFNEFG-CONH2), and S186 (Ac-NH-AKPYSQRRKTSG-CONH2) demonstrated 10%–20% inhibition for S117 and S417 and 85% for S186, with dose-dependent reductions in fluorescence observed for all RNA constructs tested. 69 The Duca group used a similar beacon approach to identify inhibitors of miR-372. Using neomycin–nucleobase and unnatural nucleobase conjugates, they showed that the conjugates designated as 3e and 3f were able to achieve ∼100% inhibition of in vitro Dicer processing and bound to the pre-miR with Kd values of 16 ± 8.8 and 14 ± 7.6 µM. 70 Staedel and colleagues used this approach to screen a library of 640 polyamines identifying a spermine–amidine conjugate that bound with a Kd of 150 nM and inhibited Dicer processing with an IC50 of 1.1 µM. The bioactivity of this ligand, measured by cell growth inhibition, was further confirmed in gastric cancer cells. 71
Microscale Thermophoresis
MST is the measure of molecular movement in a microscopic temperature gradient created by heating the sample via infrared radiation (
Recent studies have applied MST to study the binding of small molecules to RNA. One of the first to apply this method to RNA was Gaffarogullari et al. in 2013, studying the catalytic activity of the Diels–Alderase ribozyme that catalyzes Diels–Alder cycloaddition reactions. Using this method, the group not only obtained the Kd of each substrate to the ribozyme but also determined a cooperativity effect of 2.6-fold between the diene and dienophile for binding to the active site. They also found that entropy drove the energy penalty observed for dissociation of the diene.
75
In addition, Moon et al. showed that MST could be applied to the binding of Rev to Rev response element (RRE) RNA (Kd = 4.8 nM), as well as to the inhibition of that interaction by competing off REV with neomycin (Ki = 2300 nM). The group also determined the binding constants of the prequeuosine1 riboswitch from two different organisms to cognate ligands PreQ1 (Kd = 26 nM) and SAMII (Kd = 140 nM), on par with affinities previously reported in the literature (
Differential Scanning Fluorimetry
DSF has seen widespread use in the identification of ligands binding to proteins. 80,81 Recently, this method has been adapted to include RNA–ligand binding interactions, initially utilized by Silvers et al. studying RNA stability in the presence of Mg+ and F– for the fluoride riboswitch. 30 The method uses an RNA-binding dye, like RiboGreen or SYBR Green, that discriminates between structured and unstructured RNA (single- vs double-stranded) by binding and emitting fluorescence. As the temperature of the sample increases, the RNA begins to unfold, causing a decrease in fluorescence along with a signal reduction due to thermally induced dye diffusion. This experiment affords a Tm, the temperature at which 50% of the RNA is unfolded and is a direct measure of the RNA’s thermal stability under these conditions. The ligand can then be added, after which the Tm is again measured to determine the gain or loss in RNA stability as a result of binding. This method has many advantages over normal fluorescence and calorimetric approaches, such as its insensitivity to pipetting errors, its small sample volumes, and its amenability to HTS campaigns as a result of the previous two points.
One example of this approach is the study of HIV trans-activating response element (TAR)-binding ligands like acepromazine and its analogs by Sztuba-Solinska et al. 82 Initially identified from SMMs, DSF studies identified two compounds that exhibited 4–8 °C increases in Tm, with a thienopyridine being most active. Direct binding by 2AP assays (see McGovern-Gooch and Baird for more detail 40 ) afforded a Kd of 2.4 ± 1.1 µM. More recently, Baird and colleagues utilized the differential scanning FRET method to study the effect of Mg2+ ions on the conformation landscape of the c-di-GMP aptamer, identifying that maximal binding-induced conformation changes occur at near-physiological Mg2+ concentrations. 83 Demonstrating the high-throughput nature of this assay, Matarlo et al. identified natural product inhibitors of pre-miR-21 biogenesis. By screening a library of >3800 pure natural products, 10 compounds were identified as hits to bind to pre-miR-21 and stabilize its structure, with minimal cytotoxicity in HCT-116 cells. The compound bPGN was identified as having a Kd of 400 nM and a demonstrated knockdown of miR-21, as well as an upregulation of programmed cell death protein 4 (PDCD4) and phosphatase and tensin homolog (PTEN), both downstream targets of miR-21. 84 This method is relatively nascent in its application to the study of RNA-binding small molecules but has significant potential to provide high-throughput analysis of ligand effects on RNA stability, otherwise unattainable by conventional UV melting.
Fluorescent Intercalator Displacement
FID assays are a commonly used method for identifying novel RNA-binding matter. Initially used to identify ligands that bound DNA using intercalators such as ethidium bromide (EtBr) 85,86 and thiazole orange (TO), 87,88 the method has since been applied to the study of RNA–ligand interactions, employing a wider array of indicators. The assay functions by using a fluorescent indicator of known affinity to the RNA of interest and then precomplexing them together, enhancing or quenching the indicator’s emission. A library of small molecules can be subsequently screened against the complex to identify potential RNA-binding ligands, because as unknown ligands bind to the RNA, the indicator’s emission is altered (quenched/enhanced). This change can assess the binding capacity of each ligand to the RNA and then rank them accordingly. Common indicators include EtBr, TO 89 and its derivatives TO-Pro-1 90,91 and TO-Pro-3, X2S, 92 2AP, 93,94 and SYBR dyes. 95
FID assays using small molecules have been employed with great success in the identification of novel RNA binders. Zhang et al. identified mitoxantrone and sanguinarine as binders to the RRE of HIV-1 RNA, using X2S as their indicator with a hit rate of 0.2%. 92 Similarly, Tran and Disney identified eight RNA binders (hit rate, 19%) from a library of small molecules focused on RNA-binding chemotypes such as benzimidazoles and pentamidines, with Kd values ranging from 4 to 160 µM. 91 Incorporating a more high-throughput orientation, Haniff et al. used TO-Pro-1 bound to r(AAUU), r(AUAU), r(GGCC), and r(GCGC) base pairs to study whether small molecules can selectively bind to these base pairs, as well as their ability to discriminate between the orientation of the pairs in tandem (i.e., AU/AU or alternating AU/UA). This work identified 28 novel RNA-binding ligands from a library of 3200 compounds (hit rate, 0.8%). This library yielded AU and GC pair-selective base pair binders with Kd values ranging from 0.04 to 3 µM. 79
An alternative to RNA-binding small-molecule indicators is the use of RNA-binding protein indicators. RNA-binding proteins such as the trans-activator of transcription (Tat) and Rev protein of HIV-1 have been extensively studied for their capacity to bind TAR and RRE RNA, respectively. 96 To increase the generalizability of this approach, Gobel, Crothers, and colleagues identified that the highly basic and positively charged regions of the Tat protein, specifically residues 49–57 (RKKRRQRRR), 97 -99 contributed to its RNA-binding properties. This was successfully applied as a minimized fragment to identify RNA–small molecule interactions by Matsumoto et al., 100 Patwardhan et al., 101,102 and Weeks and Crothers. 98 Similarly, Luedtke and Tor 85,103 found that residues 34–50 (TRQARRNRRRRQRERQR), which contribute to Rev’s RNA-binding capacity, could be used in HTS to identify selective disrupters of the RRE–Rev interaction. 85,103 In these and other studies, peptide fragments were labeled with 5-FAM and tetramethylrhodamine (TAMRA) on the N- and C- terminus, respectively, for Tat 100 to create a FRET pair as the readout. In the case of Rev, the peptide was labeled with 5-FAM only, with subsequent binding being measured by FP. Such a strategy was also applied by Wang et al. in their study of the binding of aminoglycosides to 16S ribosomal RNA. 104 These two methods provide unique advantages and disadvantages to the study of RNA–small molecule interactions and are discussed in depth by Wicks et al. 105
Surface Plasmon Resonance
SPR, like many of the methods described herein, has seen widespread application in the study of protein binding. SPR occurs when polarized light hits a metal surface at the interface of materials with different refractive indices. At the appropriate angle of incidence, surface electrons of the metal become excited and oscillate, absorbing some light while reflecting the rest. If one of the surface compositions is changed, its refractive index also changes, thereby shifting the resonance angle required to absorb light. 106
Win and colleagues identified codeine binding aptamers by affinity selection using Sepharose beads conjugated with codeine. Lead aptamers were then assessed for their binding affinity to codeine using SPR. This was completed by chemical modification of a CM5 sensor chip to display codeine, affording two aptamers with Kd values of 2 and 4 µM, respectively. 107 Conversely, Hendrix et al. used biotinylated RNAs to bind to the surface of a CM5 sensor chip, revealing that neomycin B bound to three sites in domain II of RRE with submicromolar affinities. 108 The advantage of using the latter biotinylated method over the former chemical modification is found in the simplicity and ease by which the surface can be conjugated.
One alternative to using chip-based SPR methods is biolayer interferometry (BLI), functioning on the same principles as SPR, but instead of measuring the shift in resonance angle of resonating plasmons, it measures the interference patterns of reflected light. Generally, a BLI sensor consists of two reflective surfaces, one inside the sensor that does not interact with the analyte, and one outside the sensor that does. The internal surface serves as a reference, while the external surface measures the change in refractive index as a result of analyte binding. This method has seen many applications in the study of nucleic acid binding, including recent work by Disney et al. that has applied this method extensively to the study of r(CGG)exp, r(CUG)exp, and r(G4C2)exp expansions. The group identified high-affinity binders such as 2HE-5NME (Kd = 50 nM),
109
3K-4 (Kd = 5 nM),
22
and a hydroxy ellipticine derivative dubbed
Label-Free Methods for Identifying RNA Binders
Small-Molecule Microarrays
SMMs offer unique advantages over plate-based assays for the identification of RNA-binding ligands. SMMs are highly miniaturized, therefore using less material; they have an excellent signal-to-noise ratio due to their small sample volumes; and they can exponentially increase the throughput with which RNA-binding ligands can be identified. This method involves the printing of small molecules onto a glass surface, either covalently
111
or noncovalently,
112
using a robotic printer. Once adhered, a labeled RNA is hybridized to the glass surface and then washed to remove unbound species. Hits are identified by accumulation of the label onto the points at which compounds were printed (
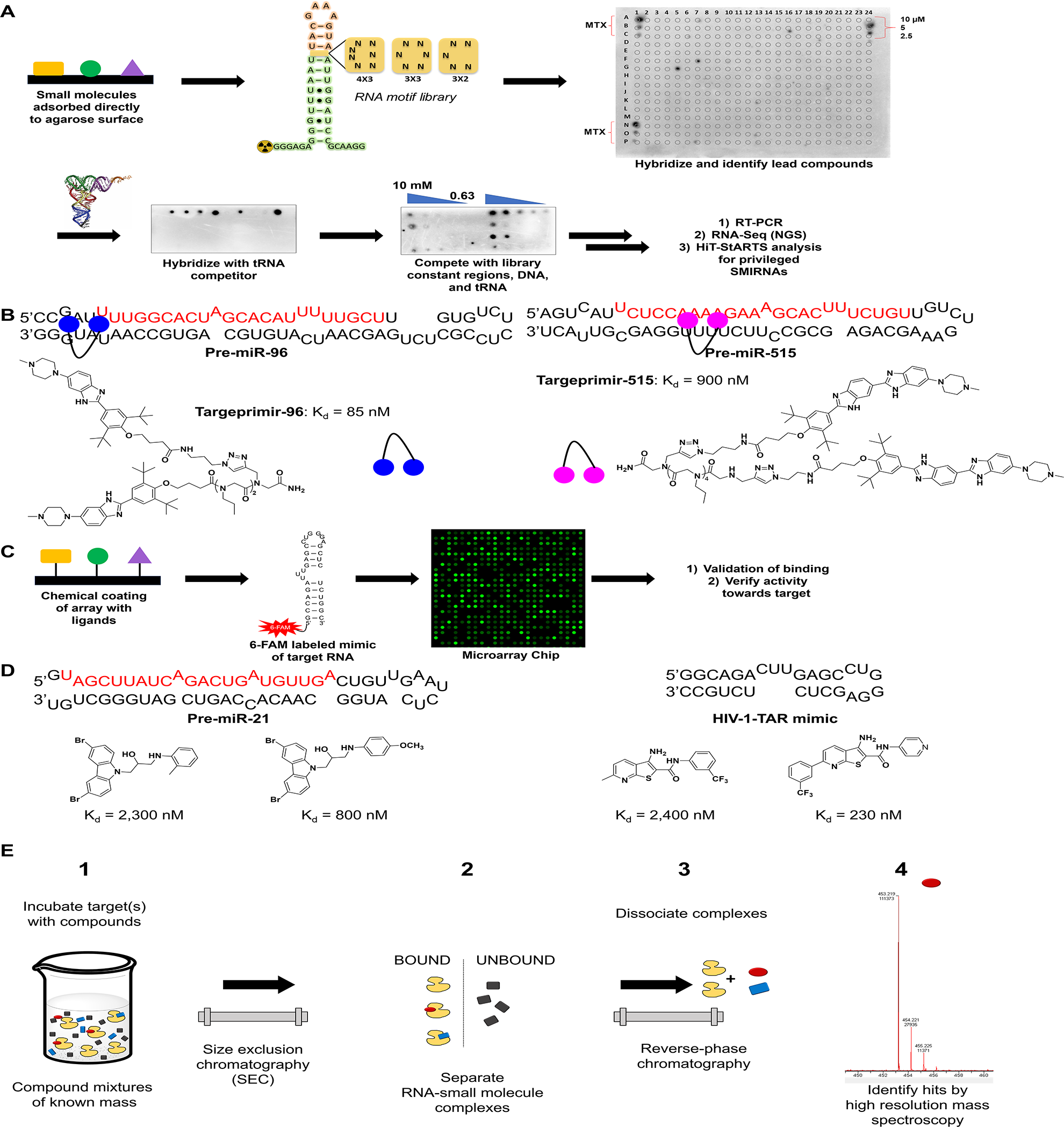
Schematic of SMM methodologies to identify RNA binders. (
Disney et al. and Schneekloth et al. have both done extensive work in the use of SMMs to identify small molecules binding to RNA. Two-dimensional combinatorial screening (2DCS), a method created by the Disney lab, utilizes glass slides coated with an agarose matrix used to bind small molecules to the surface. A γ-32P-labeled RNA library is then hybridized with the array in the presence of excess competitor oligonucleotides and washed, followed by excision of bound RNA and subsequent sequencing (
In contrast to the Disney group’s 2DCS library versus library screening, other increasingly targeted approaches using SMMs have also been employed. Such methods as those developed by Schneekloth et al.
118
use dye-labeled mimics of the target RNA to scan their arrays for novel binders. These arrays are constructed by covalently displaying the ligands on the surface of the array, which differs from the current 2DCS methods, which require no such linkage (
MS-Based Approaches
Historic MS-based approaches for identifying small molecules that bind to nucleic acid focused on direct MS detection of nucleic acid–small molecule complexes.
120
-126
These include approaches such as MASS (multitarget affinity/specificity screening)
127
and DOLCE-MS (detection of oligonucleotide ligand complexes by ESI-MS).
128
Limitations due to buffer restraints (MS interferers, i.e., K+ and Mg+), RNA size, and the need for high mass resolution and accurate mass detection hinder the use of these approaches for general large-scale screening for small molecules targeting RNA. Indirect affinity selection–mass spectrometry (AS-MS) methods have been developed for screening large multiplexed compound sets under a variety of conditions.
126,129
-131
Recently, Merck (Kenilworth, NJ) applied their AS-MS method, referred to as the Automated Ligand Detection System (ALIS), to screen compounds against numerous RNA targets, an approach validated with ncRNA riboswitches, their ligands, and a large unbiased small-molecule library.
132
-134
Here, RNA is incubated with mass-encoded small-molecule mixtures, small molecule–RNA complexes are purified by size exclusion chromatography (SEC), and the complexes dissociate through reverse-phase chromatography. Bound ligands are detected by high-resolution MS (
Dynamic Combinatorial Chemistry
An inherent limitation of the methods described above is the use of “fixed” ligand libraries generated by traditional combinatorial chemistry approaches. This culminates into a screen where the target is unable to influence the ligand pool and select the most optimal binder. One solution to this problem is the use of dynamic combinatorial libraries (DCLs) generated through dynamic combinatorial chemistry (DCC). This concept hinges on the use of “soluble” ligand binding fragments that can undergo reversible chemistry to form more complex binders and whose reaction equilibrium shifts in response to the target’s interaction with those complex species. In other words, upon binding, the ligand drives the reaction equilibrium to make more of the high-affinity species and depletes low-affinity species from the pool. 135,136
Since its first mention in the late 1990s, this methodology has been successfully applied to identify RNA-binding ligands by the Marchán and Miller groups. For example, the Marchán group, using the stem loop at the tau exon-10-intron 10 junction, used a DCL consisting of acridine, neamine, and azaquinolone derivatives with varying spacer lengths for the sulfide reactive group and two aromatic tripeptides (H-Cys-Tyr-Arg-NH2 = TyrP and [H-Cys-Trp-Arg-NH2][S-S] = TrpP). Through use of biotinylated RNA, the selection was carried out, isolating bound ligands by streptavidin pulldown. Analysis of the ligand pools found amplification of the Acr1-Nea and Acr2-Nea2 ligands, which displayed EC50 values of 5.9 and 2.1 µM, respectively, and was confirmed by optical melting analysis to stabilize the tau hairpin. 137
The Miller group has conducted extensive analyses of RNA binders deploying DCC to generate RNA-directed DCLs. For example, due to the biases of DCLs to produce dimers as a result of the entropic penalty for forming trimeric or higher-order species, the Miller group developed resin-bound dynamic combinatorial chemistry (RBDCC), whereby phase separation of the individual units enables display of the full library complexity, or the use of bifunctional resin partners that can form ternary resin-bound dynamic combinatorial libraries (RBDCLs). 138,139 Gareiss and colleagues applied these methodologies to identify lead molecules capable of displacing MBNL1 from r(CUG) repeats. To do this, an RBDCL with 11,325 members was constructed from 150 unique resin-bound cysteine-containing building blocks. This library was incubated with Cy3-labeled r(CUG)10 and bound beads were picked by fluorescence and deconvoluted to identify the ligand, which resulted in four unique ligands that bound with Kd values ranging from 41 to 1.9 µM. 140 Ofori and colleagues then used ligand 4 from Gareiss et al. and derivatized it with benzo[g]quinoline to produce an eight-compound library of ligands that were screened for binding to r(CUG) repeats by SPR and fluorescence titration. Two stereoisomeric derivatives exhibited Kd values of 40 and 70 nM, respectively, and were further tested in cellulo and in vivo, resulting in modest correction of MBNL1-mediated splicing defects in Clcn1 and Atp2a1 in HASLR mice. 141
Rolling Circle Amplification
Rolling circle amplification (RCA) has seen extensive use in biosensing applications of DNA and RNA. The method uses a circular DNA template that has a hybridization site for the analyte of interest. Upon hybridization, isothermal primer extension from the analyte is carried out, creating a concatemer, or repeated DNA sequences in series, that amplifies the detectable signal of the analyte for readout. While this method is highly sensitive, it has had few uses to study the ability of small-molecule ligands to inhibit RNA biology. The Arenz group utilized this methodology to study the inhibition of the RNA-induced silencing complex cleavage of miR-122 in vitro by aminoglycosides such as kanamycin A/B and neomycin. Here, using Sybr Gold, they measured the amount of concatemer produced upon cleavage by recombinant “minimal RISC,” kanamycin A/B, and neomycin B and found that concatemer production was reduced by ∼20%, ∼40%, and ∼70%, respectively, indicating an inhibition of pre-miR-122 cleavage by the minimal RISC. 142
Fragment-Based Approaches by NMR Spectroscopy and X-Ray Crystallography
The development of HTS technologies has led to many drug leads. 143 However, screening of large compound libraries has yielded either few hits or false positives against challenging targets. 143 An alternative strategy, fragment-based drug discovery (FBDD), consists of screening smaller sets of fragments for chemical starting points. 143 Fragments, defined as having less than 20 nonhydrogen atoms, allow for more efficient sampling of the chemical space than drug-like molecules. 143 Small fragments may bind to a greater number of sites than larger, more complicated molecules, allowing them to bind to targets such as RNAs that lack well-defined binding pockets. 143 -145 While fragments are typically weak binders, lead compounds may be developed by growing, merging, or linking fragment hits together. 144,146
NMR is a sensitive technique suitable for finding weak binders such as those with single-digit millimolar Kd values. 146,147 NMR methods for identifying fragments consist of both target- and ligand-based screens. Target-based screens monitor changes in the spectra of an unlabeled or isotopically labeled target upon addition of a ligand. 144,148,149 These methods may identify binding sites and determine binding affinities. 144,150 However, assignment of the target resonances may be complicated by spectral overlap. 144,149 Titration measurements are also slower than other biophysical methods or biochemical assays, and target-based methods require greater amounts of biomolecule target than ligand-based methods. 149
Ligand-based methods monitor changes in the spectra of a ligand upon addition of a target. 144,148,151 Screens can be carried out with cocktails of fragments and do not require assignment of the target resonances, nor large amounts of the target. 152 Methods used for ligand-based screens include saturation transfer difference (STD), water–ligand observation with gradient spectroscopy (WaterLOGSY), and T2 relaxation-edited techniques. 144,147,148,151
In an STD experiment, resonances of the target are selectively irradiated by a radiofrequency field. 148 The saturation is transferred to bound ligands, and then to free ligands in fast exchange with bound ligands. This is where signals are detected. 148,151 A separate spectrum is then taken with the irradiation frequency applied away from the target or ligand spectral range, and the spectra are subtracted. 148,153 Only resonances of ligands that bind to the protein or other targets remain. Ligand sites that are not close to the target and give consequently weaker signals may be identified by STD and used as fragment growing or linking sites. 144 Stronger STD signals are displayed for protons in close proximity to the target experiment. 144,147,148,151 In a WaterLOGSY experiment, the water signal is irradiated, and magnetization is transferred from bound water to bound ligand. 148,153 Bound ligands display signals of the same sign as target resonances. 144,147,148,154
T2 relaxation-edited experiments rely on the difference in T2 relaxation rates between small-molecule ligands and macromolecule targets. 148,150,155 -157 Larger molecules have longer rotational correlation times (or slower tumbling) due to enhanced spin–spin interactions, leading to shorter T2 relaxation times and broader lines. 150,155,157 In a T2-edited spectrum, binding of a ligand to a macromolecule leads to broadening of the ligand resonances because the ligand adopts the shorter relaxation time of the macromolecule. 150,157 The T2 relaxation time may be measured using a Carr–Purcell–Meiboom–Gill (CPMG) pulse to follow the signal decay as a function of the relaxation delay. 157 These methods are used to identify fragments that bind to a target, but with certain limitations.
Ligand-based methods may be used with mixtures of ligands and do not provide information about ligand binding sites. 144,147 STD and WaterLOGSY methods require small amounts of unlabeled biomolecule target in the micromolar range, 147,151,153 but they are often run with excess ligand over the target, a condition that may lead to a pH change in solution if the ligands contain acidic or basic moieties. 149 Furthermore, the high concentrations of compounds necessary for these methods may make it difficult to distinguish site-specific binding from nonspecific binding or aggregation. 144,149 Compound-induced partial unfolding or precipitation of the target may also give the appearance of binding or nonbinding, respectively. 144,149 Finally, ligand-based methods become challenging with high-affinity binders (Kd < 0.1 μM), which give weaker signals. 144,148
Structures of target–fragment complexes such as those mentioned previously may be further elucidated by NMR or x-ray crystallography. 144 NMR has been traditionally used to determine structures and dynamics of biomolecules in solution. 144 However, defining full NMR solution structures is a time-intensive process limited to small- to medium-sized biomolecules and requires a complete assignment of backbone resonances. 144 X-ray crystallography is commonly used to rapidly generate high-resolution structures of ligand–target complexes, but it does not provide information about affinity, and crystal contacts may block ligand binding sites. 144,149 Additionally, x-ray crystallography usually provides a single structure, without the dynamics observed for ligand–target complexes in solution. Further, overly flexible biomolecules may not be amendable to crystallization. 144,149
Lee et al. employed a target-based NMR screen to identify compounds that bound to the influenza A virus (IAV) promoter (
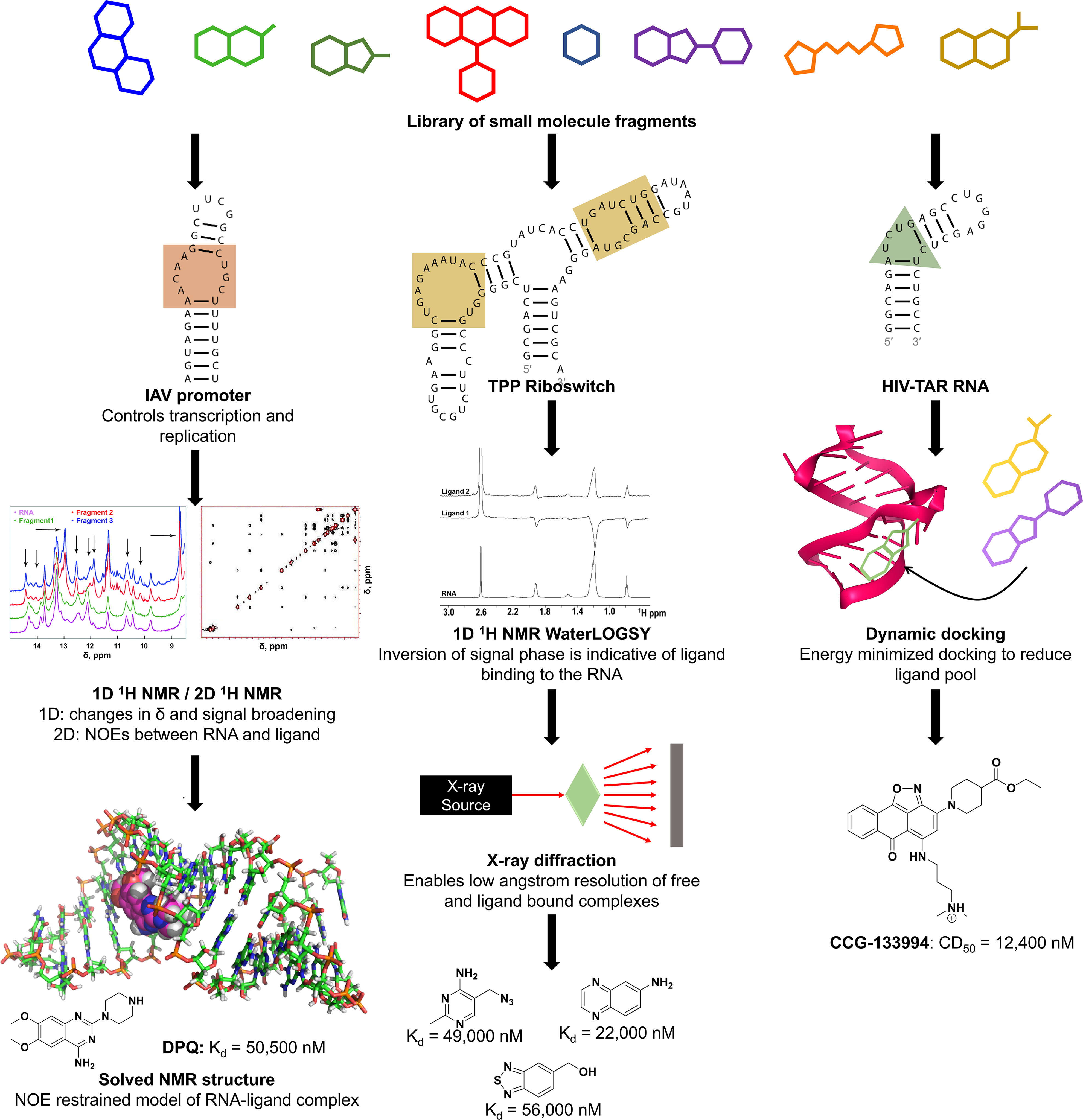
NMR, x-ray, and computational methods to screen for compounds that bind RNA. Left: Lee et al. used NMR spectroscopy to screen for ligands that bind to influenza A RNA promoter and determine the structure of one of the hits, DPQ, in complex with the RNA. 158 Center: Cressina et al. screened a library of 1300 compounds for those that bind to the E. coli TPP riboswitch and used WaterLOGSY and T2 relaxation-edited NMR experiments to confirm binding of 20 hits. 152 Warner et al. determined x-ray structures of four of the hits in complex with the riboswitch, the highest-affinity binder among which is shown. 162 Right: Al-Hashimi’s group used virtual screening to identify ligands that bind to HIV-1 TAR RNA. 167,178 The highest-affinity binder identified from EBVS is shown with its competitive dose to displace 50% of the Tat peptide value or CD50. 178
Garavís et al. identified fragments from a fragment library of 355 fluorinated compounds that bound to telomeric repeat-containing RNA (TERRA) using 19F NMR. 160 These lncRNAs, transcribed from telomeres, contain an average of 34 r(UUAGGG) repeats that fold in vivo into G-quadruplexes. 160 TERRA promotes heterochromatin formation at telomeres, protecting telomere ends from the DNA damage response, and its expression is elevated in various human cancer cells. 160,161 The group acquired 1D 19F NMR spectra with and without a CPMG T2 filter on cocktail samples containing eight fragments each. Upon addition of the target molecule to compounds, signals of binders broadened while signals of nonbinders did not change. The group ultimately identified 20 compounds from the NMR screen and validated the hits using 1H NMR STD experiments. 160 All seven hits thermally stabilized TERRA2, as determined by circular dichroism (CD) melting experiments. 160 Taken together, this showed that fragment-based screening can be applied to identify ligands targeting RNA G-quadruplexes. 160
On certain occasions, a combination of fragment-based methods may elucidate different aspects of ligand–RNA binding. For example, one screen examined fragments binding to the thiamine pyrophosphate (TPP) riboswitch and interfering with the riboswitch mechanism, modulating downstream protein synthesis.
152,162
The Escherichia coli TPP riboswitch contains a 78-nucleotide aptamer domain that binds to TPP and causes structural changes in a 90-nucleotide expression domain, sequestering the ribosome binding site and leading to translational attenuation.
152
Cressina et al. screened a library of 1300 fragments by competition equilibrium dialysis and obtained 20 hits.
152
The group confirmed binding of these 20 hits by WaterLOGSY and T2 relaxation-edited NMR experiments. Warner et al. solved crystal structures of four of the fragments in complex with the TPP riboswitch using small-angle x-ray scattering (SAXS) experiments (
Predictive Approaches by Virtual Screening
The high cost of HTS in time and resources has led to an increased use of virtual screening to identify new scaffolds against therapeutic targets. 163,164 Virtual screening samples the chemical space and removes inactive compounds before experimental testing using in vitro assays. 163 One strategy employed by virtual screening is the docking of large virtual libraries of compounds to the 3D structure of a target. 163,164 Molecular docking estimates the binding mode of a ligand in a receptor as well as its binding affinity. 164 Several algorithms and scoring functions for protein–ligand docking have been adapted for RNA docking. However, challenges with modeling the charged structure and conformational flexibility of RNA targets limit their success, especially considering that RNA can undergo large conformational changes upon ligand binding. 165 -167 In addition, relatively few experimental RNA–ligand structures are available to develop scoring functions calculating the free energy of ligand binding. 165,166 Docking algorithms developed to address these issues are discussed in detail below.
Molecular Recognition with a Driven dynamics OptimizeR (MORDOR) is a program developed for induced-fit docking of ligands to RNA. 165 In this methodology, the ligand and receptor undergo conformational changes to yield an optimal fit. 165,168 MORDOR uses molecular simulations to model ligand and RNA flexibility during docking and calculates a score using the total energy of the complex, which includes a solvation term. 165 This method was used to screen for compounds against the human telomerase RNA, and the binding of a subset was subsequently confirmed by NMR STD experiments. 169
Daldrop et al. applied DOCK 3.5.54 to screen for compounds binding to a guanine riboswitch (GRA). 170 The group adapted a scoring function to RNA–ligand binding by using RNA-specific parameters to calculate van der Waals and electrostatic energies. 170 Crystal structures were determined for three of the ligands in complex with GRA that had been identified by docking. 162 Lang et al. reported an updated version of DOCK, DOCK 6, that added AMBER generalized Born and Poisson–Boltzmann implicit solvent models to physics-based scoring functions. 166 The docking of 70 RNA–ligand complexes using these solvent models, in combination with explicit water molecules and counterions, improved the success rate of reproducing experimentally determined structures. 166 Furthermore, DrugScoreRNA and LigandRNA are two knowledge-based, stand-alone functions for scoring ligand–RNA complexes that derive potentials from experimentally determined 3D structures of ligand–RNA complexes. 171,172 While DrugScoreRNA uses distance-dependent potentials to score RNA–ligand complexes, LigandRNA uses distance- and angle-dependent potentials. 171,172 When tested on a set of 42 complexes, a combination of LigandRNA and DOCK 6 correctly identified ligand poses in 47.6% of cases compared with 35.7% for LigandRNA or DOCK 6 alone and 31.0% for DrugScoreRNA. 172
In addition to the above, Morley and Afshar developed RiboDOCK for docking ligands to RNA. 173 The scoring function of the program includes both attractive and repulsive potentials. The attractive potentials account for parallel π-π aromatic stacking (Sarom) and interactions between a positively charged ligand carbon and RNA acceptor (SposC−acc), such as a guanidinium carbon and RNA carbonyl oxygen. The repulsive interactions account for donor–donor (Sdon–don) and acceptor–acceptor (Sacc–acc) repulsion. This program was then used to identify compounds that target the bacterial ribosomal A site. 173,174 An updated program based on RiboDOCK, rDock, was later released. 175,176 Terms that were added to the rDock scoring function include a van der Waals potential and attractive and repulsive potentials, in addition to external restraint terms such as cavity and pharmacophore restraints. 175,176 When tested with a set of 56 RNA–ligand complexes, rDock predicted at least one correct pose in 98% of cases. 175
An alternative approach to virtual screening for ligands against RNAs combines NMR spectroscopy with computational methods to generate ensembles of conformations of an RNA. Molecular dynamics (MD) force fields used to model the conformational space of RNAs have had limited accuracy. The Al-Hashimi group carried out an 80 ns MD simulation on NMR structures of HIV-1 TAR RNA and then selected 20 conformers (dubbed TARNMR-MD) that agreed with experimentally determined NMR residual dipolar couplings (RDCs) for elongated TAR RNA. 167,177 RDCs report on the angle between internuclear bond vectors in a biomolecule and an external magnetic field. 178,179 The group applied virtual screening of 51,000 small molecules to each of the 20 conformers using the ICM docking program (Molsoft) and experimentally validated six of them for binding to TAR and inhibiting its interaction with Tat. 167
Al-Hashimi’s group later generated an ensemble of 20 TAR structures using four sets of RDCs and 8.2 μs MD simulation, and then carried ensemble-based virtual screening (EBVS) using ICM to identify TAR binders.
178,180
They assembled three small-molecule libraries of hits and nonhits from experimental HTS for docking, and then assessed the performance of EBVS by analyzing the enrichment factor (EF) and area under the curve (AUC) of a receiver operating characteristic (ROC) curve.
178
EF measures the number of true positives in a subset of compounds relative to random screening of the database.
181
An ROC curve is a plot of the true-positive rate against the false-positive rate and measures the ability of docking to separate active compounds from inactive compounds, while AUC values range from 0.5 for random selection to 1.0 for perfect enrichment.
182
EBVS of 2% of nonhits in a library consisting of 78 hits and 103,349 nonhits identified 42% of hits for an EF of 21 and AUC of 0.88. Similar results were obtained for the other two libraries. Hit compounds in cell-based assays were significantly enriched, with AUC values ranging from 0.91 to 0.94.
178
Altogether, these studies demonstrate that EBVS may be used to identify bioactive compounds that bind to HIV-1 TAR RNA (
Databases of Known RNA-Binding Ligands
An enormous amount of data describing ligand–RNA interactions is available, but unfortunately it is challenging to mine. To aid future RNA drug discovery attempts, it would be helpful to compile all protocols and experimental data in searchable databases. This knowledge can be used to identify scaffolds of interest for RNA-binding optimization or tools for screening approaches. Several freely available databases (Inforna, 19 SMMRNA, 183 R-BIND, 184 G4LDB, 185 and NALDB 186 ) documenting a catalog of small molecule–RNA interactions exist, with insight ranging from basic selectivity and affinity measurements to detailed compound parameters and experimental insight. Inforna (https://disney.florida.scripps.edu/software/) is a database of experimentally determined, privileged RNA motif–small molecule interactions alongside interactions reported in the literature. 19 The motifs, or structural elements, within an RNA target of interest are compared with the database of interactions, informing lead compounds for further investigation and a chemical similarity searching feature to expand those leads. 19 SMMRNA and R-BIND (https://rbind.-chem.duke.edu/) were/are interactive databases that displayed multiple binding parameters, including molecular weight, hydrogen donor and acceptor count, XlogP, number of rotatable bonds, and number of aromatic rings, 183,184 and SMMRNA also housed 2D and 3D structures for ~770 unique RNA-binding compounds. 183 The G-quadruplex ligand database, G4LDB, was a second database that focused on reported G-quadruplex ligands. 184 It contained >800 G-quadruplex ligands with ~4,000 activity records. The data set consisted of physical properties, 3D ligand structures and design, in vitro and/or in cell binding, and activity data. 185 NALDB (http://bsbe.iiti.ac.in/bsbe/naldb/HOME.php) provides detailed information about small molecules targeting all nucleic acids. 186 It contains more than 3500 ligand entries and, similar to the databases mentioned above, provides data sets consisting of ligand physical properties, 3D ligand structures, and activity data. 186
RNA-Binding Scaffolds
While the interest in screening RNA structures to identify small molecules able to bind them/modulate their biology has initially remained essentially academic, this picture has drastically changed. There is an increased interest of the pharmaceutical industry in assessing the possibility of drugging RNA targets using corporate compound collections. This has not only triggered a change in the type of RNA target structures under investigation but also increased the chemical diversity reported to interact with RNA.
The 2008 comprehensive review by Thomas and Hergenrother 187 constitutes an outstanding reference point to assess the evolution of chemical equity studied for its capacity to interact with RNA. By the time this review was published, there was still a clear focus to use reasonably characterized available RNA binders like aminoglycosides, oxazolidinone antibiotics, aromatic bases, and intercalating agents as starting point points to develop more specific binders. The late 1990s and early 2000s have witnessed the initiation of more thorough screening efforts to identify novel chemotypes with improved selectivity and toxicity profiles as well as better potential for further optimization toward therapeutic applications. Early attempts in screening large compound libraries demonstrated a low hit rate and yielded hits with challenging properties. 188 Some interesting developments were achieved in identifying more suitable chemistry already by the mid-2000s, as exemplified by MS-based efforts by Ionis scientists, 189 resulting in the optimization of a benzimidazole series of HCV-IRES IIa binders 190 possessing low micromolar activity in an HCV replicon assay together with minimal cellular toxicity. Other hit-finding strategies like fragment-based approaches have also been applied early on to RNA targets, resulting in the confirmation of some privileged scaffolds as illustrated in an NMR screen to identify E. coli A site rRNA binders reported by AbbVie researchers. 191 This study confirmed benzimidazoles and 2-amino-benzimidazoles as privileged scaffolds for RNA interactions but also highlighted 2-amino-quinoline and 2-amino-pyridine scaffolds as alternative leads with good potential for optimization.
The emergence of miRs as potential therapeutic targets and miR biogenesis inhibition as a potentially interesting mechanism of action for small molecules has triggered several developments in the screening approaches to identify binders for specific Dicer and Drosha sites on miR targets. The Disney group has successfully applied its combined 2DCS and cheminformatics platform to identify numerous inhibitors against primary (pri) and precursor (pre) miRs as well as binders of intronic and expanded repeats implicated in different neurological disorders. Although 2DCS was initially validated using aminoglycosides, 192 this approach has enabled the identification of miR binders across a range of chemotypes. While a dimerization strategy to optimize the hits into cell-active and -selective miR biogenesis inhibitors has often been applied by the Disney group, 193 several hits were demonstrated to be useful as monomeric binder, including hits based on 1,8-diamino-2,7-naphtyridine 194 and again 2-substituted benzimidazole. 117 A recent development of this technique has allowed the Disney group to study the potential of clinical compounds as well as compounds targeting specific protein classes like kinase inhibitors to also bind and potentially modulate RNA. 112,195 In particular, three topoisomerase inhibitors (doxorubicin, epirubicin, and mitoxantrone) bound RNA, with an optimal motif identified in oncogenic miR-21, the levels of which are downregulated upon compound treatment. 181 While a broader outcome still needs to be published, these studies shed an interesting light on a potential secondary pharmacology, centered on RNA modulation, that compounds developed against protein targets might possess and that, to this day, has been difficult to study and exploit and thus has been largely overlooked.
SMMs have also been successfully used by the Schneekloth group to identify compounds that modulate Dicer processing of pre-miR-21 in vitro, 119 but they more recently broadened their approach to lncRNA Malat-1, identifying binders that specifically recognize the ENE triplex substructure that could yield interesting tools to study Malat-1 function. 196 RNA binders identified by Schneekloth and coworkers have again underlined benzimidazoles as a privileged scaffold, but also evidenced other useful chemotypes like N-substituted carbazoles.
Beyond microarrays, efforts toward identifying oncogenic miR-21 biogenesis inhibitors have been reported by the Garner and O’Keefe groups using their catalytic enzyme-linked click chemistry assay 197 and a DSF-based screen, 84 respectively. Interestingly, both of these groups used natural products/natural product extracts in their compound sets and identified scaffolds that were further characterized, including tetracyclines derivatives and butylcycloheptyl prodiginine. The Al-Hashimi and Hargrove groups have also investigated natural product scaffolds using virtual screening of a dynamic ensemble of HIV-TAR, identifying amiloride derivatives as an interesting set of binders. 17
While some attempts to identify RNA-specific pharmacophores and to define an RNA binder’s chemical space have been disclosed, 20,76,198 these have been based on a relatively limited data set, and there is still a scarcity of data when it comes to systematic screening of a large and diverse compound set with properties in line for subsequent medicinal chemistry optimization. One such attempt has been recently reported by Merck scientists using ALIS, 127 described above. More recently, Merck’s diversity set of ∼50,000 members and a set of functionally annotated compounds (∼5100 compounds) were studied for binding an array of 42 RNA targets, thus probing interactions between drug-like molecules and biologically important RNA structures. 133,199 The binders identified were largely biased toward G quadruplexes (1097 of ∼1420 compounds, or 77%), and only 119 were selective for one of the 42 RNAs screened. Although the data generated are somewhat limited in scope and predictive power for RNA in general because of the bias for G-quadruplexes, these studies did identify features enriched in selective and general RNA binders as well as differences between RNA and protein binders. Further, the physicochemical spaces occupied by the RNA and protein binders overlap and are within a drug-like space. Interestingly, based on the hits identified and using a combination of machine learning and nearest-neighbor selection based on chemical and biological similarity, an RNA-focused library (∼3700 compounds) was assembled. This library was subsequently screened against a subset of 32 of the initial RNA targets, yielding a hit rate of 0.32% (G quadruplexes excluded), a 32-fold improvement compared with the initial diversity set. Collectively, these studies suggest that using quality RNA binders as seeds, chemoinformatic approaches could facilitate the discovery of novel relevant RNA ligands. It will be interesting to see how these compounds advance to in cellulis activity.
In complementary studies, an FID-based screen of the LOPAC library was conducted by Tran and Disney to study its potential for RNA binders, identifying eight hits displaying alkyl pyridinium, indole, 2-phenyl benzimidazole, and 2-phenyl indole scaffolds. Identification of their privileged RNA-binding space identified AU-rich hairpin loops as their preferred binding partners with Kd values ranging from 4 to 160 µM. 91 Similarly, through use of 30,000 small molecules from The Scripps Research Institute’s (TSRI) and National Cancer Institute’s (NCI) collections, they were computationally analyzed for potential to bind RNA, yielding a 1987-compound set that was both chemically diverse and drug-like. From these screens, 239 novel RNA binders were identified. Analysis of the hits identified 13 privileged scaffolds that include phenyl imidazolines, 2-aminopyrimides, 2-indoles, benzimidazoles, and anilines as RNA binders, with all hits falling within the drug-like space upon comparison of their physiochemical properties to FDA-approved drugs. Further investigation identified a privileged RNA-binding partner in the SL1, SL2, and SL3 stems of the 3′ UTR of HCV that, upon binding, inhibited viral replication in an HCV replicon assay by a mechanism not previously shown before. 76
While affinity selection will yield binders that will require thorough follow-up characterization in order to validate the potential functional activity of the identified hits, the wealth of information generated in that way will certainly be very informative to refine our views of both the chemical and target space of RNA binders and help expand the chemical diversity useful for RNA modulation. At a stage where our understanding of how to rationally design and optimize small molecules with the potential to selectively modulate RNA function is still in its infancy, additional similar efforts will be useful to help medicinal chemists fully realize the potential of RNA as both a primary and a secondary pharmacologic target.
Conclusion
Pharmaceutical drug discovery over the decades has been predominantly directed toward several targets—enzymes, G-protein-coupled receptors, and proteases. 200,201 Expanding this targetable space is a noted priority among academics and companies and is led by omics/profiling initiatives. 202 -204 The continued development of new molecular entities such as ASOs, mRNAs, and small activating (sa) RNAs will breathe new life into numerous previously written off “undruggable” protein targets. 205 -209 RNA (translatable and nontranslatable) displays an enormity of function with roles correlating or even driving disease. 210,211 Prior to 2000, attempts made to develop small molecules that target RNA were wrought with selectivity and affinity challenges. 187 Regardless, RNA targeting small molecules demonstrated some success with various classes of marketed antimicrobials, alongside that of riboswitches that utilize endogenous ligands for modulation. 187 Interest in RNA–small molecule targeting over the last 5–10 years has erupted, with numerous biotech and pharmaceutical companies investing resources in the development of RNA targeting platforms (https://cen.acs.org/articles/95/i47/RNA-drug-hunters.html), a consequence of the development of novel methods and modalities that have broken down historical barriers that rendered this field dormant and controversial (https://www.the-scientist.com/lab-tools/drug-discovery-techniques-open-the-door-to-rna-targeted-drugs-65903). These companies are utilizing several enabling technologies to drive RNA targeting drug discovery, with a good proportion involving target engagement-driven projects that could piggyback on nonclassical protein drug discovery. The selection of correct hit identification approaches is key to the target-directed identification of compounds that have favorable properties for selective RNA targeting. The use of phenotypic assays as a primary screen with follow-up protein target deconvolution is common practice (especially within industry). 212,213 Beyond bacterial riboswitch and spliceswitch screens, deconvolution of RNA targets is less common. 193 A recent report that found RNA rather than protein to be the primary target for an established cancer drug indeed demonstrates the importance of RNA target deconvolution for understanding drug mode of action. 195
Here, we have discussed the different approaches for the target-directed screening of small molecules against RNA targets, many of which can also be applied to other targets such as proteins. Due to the dynamics of RNA, difficulty in associating function–structure in vitro, and noted selectivity challenges with compounds displaying certain properties (e.g., intercalators), there is a preference to use unlabeled screening approaches that can assess target selectivity, affinity, and/or function in a single well. For binding approaches using labeled RNA, selectivity can easily be incorporated in the primary screen by using unlabeled competitor RNA. For unlabeled approaches, this is more difficult and instead cost-effective follow-up assays are required to reduce primary hit collections. Secondary assays preferably assess functional responses in vitro or in cell-based assays such as monitoring RNA–protein binding (e.g., FP, 27 electromobility shift assays [EMSAs], 214 alpha screen, 215 and cat-ELCCA 216 ), RNA levels and structure (e.g., sequencing, RT-qPCR, SHAPE, 217 and rG4-seq 218 ), direct function (e.g., reporter assays 219 and processing assays [splicing, Dicer, or Drosha]), and phenotypic outputs (e.g., differentiation and proliferation). Most of these assays are costly and low throughput and require RNA labeling or further on-target cellular follow-up. Improving small-molecule libraries by biasing against promiscuous RNA binders, furthering our understanding of RNA-binding scaffolds and RNA–small molecule interactions, and developing HTS approaches that can cost-effectively monitor target engagement and selectivity in parallel will help rapidly produce high-quality RNA-binding hits for lead optimization, moving the “modern” RNA targeting field beyond tool development and validation.
Footnotes
Acknowledgments
We also would like to acknowledge the many works that could not be summarized in this review and their contribution to this evolving field.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding for these efforts has been provided by the taxpayers of the United States of America in the form of grants from the National Institutes of Health (R01 GM97455, DP1 NS096898, P01 NS09914, and R33 NS096032 to M.D.D.) and by AstraZeneca.
