Abstract
Although the potential value of RNA as a target for new small molecule therapeutics is becoming increasingly credible, the physicochemical properties required for small molecules to selectively bind to RNA remain relatively unexplored. To investigate the druggability of RNAs with small molecules, we have employed affinity mass spectrometry, using the Automated Ligand Identification System (ALIS), to screen 42 RNAs from a variety of RNA classes, each against an array of chemically diverse drug-like small molecules (~50,000 compounds) and functionally annotated tool compounds (~5100 compounds). The set of RNA–small molecule interactions that was generated was compared with that for protein–small molecule interactions, and naïve Bayesian models were constructed to determine the types of specific chemical properties that bias small molecules toward binding to RNA. This set of RNA-selective chemical features was then used to build an RNA-focused set of ~3800 small molecules that demonstrated increased propensity toward binding the RNA target set. In addition, the data provide an overview of the specific physicochemical properties that help to enable binding to potential RNA targets. This work has increased the understanding of the chemical properties that are involved in small molecule binding to RNA, and the methodology used here is generally applicable to RNA-focused drug discovery efforts.
Introduction
The importance and functional role of RNA is becoming increasingly apparent. Although only 1%–2% of the human genome encodes proteins, 70%–90% is transcribed into RNA. 1 The remaining noncoding RNA (ncRNA) has been implicated in regulating transcription, translation, RNA modification, chromatin structure modification, and alteration of mRNA stability across biological processes. 2 RNAs have been implicated in a variety of human diseases, and many RNAs form complex three-dimensional structures, making them possible targets for small molecules. 3 To date, various classes of ncRNA have been targeted by small molecules. 4 Even so, the limited number of small molecules targeting RNA leads to a dearth in knowledge of the chemical matter needed for RNA-targeted drug discovery.
Drug discovery has traditionally focused on the intersection of target- and phenotype-based approaches. While both approaches are necessary, a target-based approach enables rapid medicinal chemistry optimization of potency and selectivity by first focusing on the union of chemical matter with a validated target. 5 The modulation of disease phenotype is independent of and (in target-based approaches) subsequent to defining a structure–activity relationship (SAR). Here, we take a target-based approach to identify small molecule binders to RNA targets, using the Automated Ligand Identification System (ALIS). ALIS is an affinity-selection mass spectrometry platform for the high-throughput screening (HTS) of small molecules binding to macromolecules 6 that has recently been validated for identifying small molecule–RNA interactions, using a variety of bacterial riboswitches and their known ligands as test cases.7,8 ALIS is an “indirect” affinity-selection mass spectrometry technique that utilizes a size-exclusion chromatography (SEC) separation step to isolate the target–ligand complex away from unbound components and employs liquid chromatography–mass spectrometry (LC-MS) to release and identify the ligand. Indirect approaches such as ALIS avoid the issue of gas-phase detection and validation and are generally applicable to a wide variety of targets. 9 Indirect approaches differ from direct approaches, which require detection of the target–ligand complex in the gas phase by the mass spectrometer. Direct approaches can provide definitive evidence of complex formation, but the interactions in the gas phase may not always be relevant to biological interactions that occur in solution. 9
We report the use of the ALIS platform for the identification of small molecule RNA binders across a broad range of disease-relevant and structured RNA. First, 42 RNA targets were identified from literature that represented various therapeutic areas and RNA classes, such as bacterial and viral ncRNA elements, mammalian lncRNA, structural elements in the 3′ or 5′ untranslated region (UTR) of mammalian mRNA, G-quadruplexes, domains of ncRNA known to bind to RNA-binding proteins for function, RNA repeat elements, small nucleolar RNA (snoRNA), and noncoding splice variants. Each of these RNA targets was screened against a Diversity Library (~50,000 chemically diverse compounds) and our internal tool compounds 10 (herein called “Functionally Annotated Library”; ~5100 compounds intended for phenotypic screening). This approach was distinct from “library versus library” screening, as done using the Inforna method, in that it did not require knowledge of RNA folding and only required that small molecule libraries for testing meet general compatibility standards for LC-MS detection. 11 Machine learning based on the initial set of RNA binders from the primary screen was used to generate a compound collection enriched in RNA-binding properties (~4000 compounds) as attested to by a higher hit rate for binding to RNA in subsequent screens. From this unprecedented set of small molecule–RNA-binding data, we analyzed the physicochemical properties and chemical features necessary for selective RNA-targeting compounds and contrasted these with protein-targeting compounds. We anticipate that lessons learned, such as features enriched in RNA-binding compounds, will facilitate the further identification of small molecules targeting RNA for therapeutic applications and drug discovery.
Materials and Methods
Selection of RNA
Human genome annotation (ENSEMBL) was screened for single-nucleotide polymorphisms (SNPs) mapping within ncRNA transcripts. Transcripts were prioritized for further study based on (1) association with an SNP associated with a human disease or condition through genome-wide association studies (GWAS); (2) literature, histone-mark, or RNA-seq evidence of expression of a discrete RNA or RNA domain of <2 kb from the locus; (3) literature evidence of expression of a functional RNA motif; and (4) literature evidence of an in vitro or in vivo assay for function of an RNA expressed from the locus. In addition, a diversity set of structured RNAs of disease relevance was selected by curation, including viral and bacterial regulatory RNAs, transcripts from candidate RNAopathies with Mendelian inheritance, and known regulatory motifs within human mRNA UTRs.
Preparation of RNA
RNA transcripts were prepared through in vitro transcription by Life Technologies (Carlsbad, CA) or generated by collaborators (Harvard Medical School) using the AmpliScribe T7 High Yield Transcription Kit (Lucigen Corporation, Middleton, WI) as described
12
and further purified by gel filtration in a Superdex 200 Increase 10/300 GL column.
13
Transcript integrity and purity were verified using LabChip GX Touch/GX Touch HT (CLS137031, PerkinElmer, Inc., Branford, CT). Before ALIS screening, each RNA was prepared at 10 µM in annealing buffer (10 mM Tris, pH 7.4, 137 mM NaCl, 27 mM KCl, 2 mM MgCl2) and annealed by heating to 95 °C for 5 min and then cooling to 25 °C at 3 °C/min in a thermocycler. Preliminary experiments were done to verify annealing conditions as previously reported7,8 (i.e., MgCl2 did not degrade the RNA during annealing; control ligands were optimized in annealing conditions), and an example is shown in
ALIS Experiments
ALIS Configuration
The ALIS 2D LC-MS system configuration used in these studies has been previously described.6–9,14–16 Samples were prepared and equilibrated as described below. Samples were placed into the ALIS system autosampler and chilled to 4 °C. SEC (column dimensions: 2.1 mm ID × 50 mm length, prepared by proprietary media) was performed at 4 °C column temperature using 700 mM ammonium acetate (NH4OAc) running buffer at pH 7.5. An isocratic pump (G1310A, Agilent, Santa Clara, CA) fitted with an online degasser (G1322A, Agilent) was used for eluent delivery at 300 µL/min for a ~20 s chromatography run. Then, a UV detector (G1314A, Agilent, with a G1313 microflow cell) was used to analyze the SEC eluent for RNA–ligand complexes at 230 nm, and a valving system directed the complex to a reverse phase chromatography (RPC) column (Targa-C18, Higgins Analytical, Mountain View, CA; column dimensions: 0.5 mm ID × 50 mm length, 5 μm packing), allowing for direct coupling of the SEC and RPC separations. Ligands were dissociated from the complex (low pH, 40 °C column temperature) and eluted into the mass spectrometer using a gradient of 0%–90% acetonitrile in water (0.2% formic acid) over 2.5 min using a capillary binary pump (G1376A, Agilent) for eluent delivery at 20 µL/min. A mass spectrometer (Exactive Orbitrap, Thermo Scientific, Waltham, MA) was utilized for detection, providing high resolution (100,000 resolving power) and high-accuracy m/z detection (mass error <5 ppm without internal calibration) and allowing for exact mass and formula confirmation for the previously bound compounds.
Sample Preparation for High-Throughput Mixture-Based ALIS Screening against Large Small Molecule Libraries
Compounds for screening were pooled into mixtures of 500 compounds at a concentration of 20 μM/compound in DMSO and then diluted to 1 μM/compound in the above annealing buffer. The compound mixtures (3 µL) were combined with the previously annealed RNA (3 µL of 10 μM) and equilibrated by incubation at room temperature for 30 min. Two rounds of mixture-based samples were run in ALIS with an injection volume of 2.0 µL/injection. Invertase was used to measure nonspecific binding. Compounds yielding reproducible binding in both rounds and not producing signal in the invertase counterscreen were considered ALIS hits. Binding of hits was confirmed in ALIS as single, pure compounds.
Proteins analyzed in the high-throughput Protein-Array ALIS (PA-ALIS) approach were prepared in orthogonal mixtures of five proteins each at 5 μM individual concentration.5,17 Each target was screened in duplicate as two “orthogonal pools” in which each target was mixed with different partner proteins. This approach enabled hits that selectively bind to a single target to be identified, as a selective binder will appear in the two orthogonal pools containing that target.
Construction of Naïve Bayes Models for RNA Binding and the RNA-Focused Library
Naïve Bayes models were built for binders of target RNAs detected in the initial screening collection using calculated physicochemical, topological (ECFP4
18
), and biological (HTSFP
19
) descriptors in Pipeline Pilot v17.0. HTSFPs were assembled using calculated z scores from 344 in-house assays. Model accuracy was assessed by calculating the area under the curve for the receiver operator characteristic (AUC ROC) after leave-one-out cross-validation (
Additional naïve Bayes models classifying selective binding to RNA targets were built from detected binders, where classification of selective binding was defined by binding a single RNA target versus many targets. The RNA-Focused Library was trained on primary hits from a subset of 32 of the original 42 RNA targets in the original screen due to experimental practicality.
Construction of Naïve Bayes Models to Compare RNA and Protein Binders
In order to compare the chemical properties enriched for binders of RNA versus protein targets, we built naïve Bayes models classifying binding to proteins or binding to RNAs using calculated physicochemical and topological descriptors. Protein-binding data for each compound were assembled from historical screens performed using the ALIS platform.5,17 Each model output normalized probability weights for every learning feature present in the observed data, normalized by the total occurrences of that feature across the training dataset. 17
PCA of Protein and RNA Binders
Principal component analysis (PCA) based on physical chemical properties can aid in understanding relationships between sets of compounds. 20 We trained a PCA using 14 physicochemical properties as descriptors (molecular polar surface area, molecular weight, number of atoms, number of positive atoms, number of negative atoms, number of rotatable bonds, number of rings, number of aromatic rings, number of ring assemblies, number of stereocenters, number of hydrogen bond donors, number of hydrogen bond acceptors, fraction sp 3 , ALogP), as previously described, 21 and compared ALIS RNA binders, ALIS protein binders, and literature multivalent or small molecule RNA binders from R-BIND 22 with a subset of approved drugs previously described. 21
PCA of RNA and Protein Targets
Dimensionality reduction starting with features extracted from naïve Bayes models of binders versus nonbinders has been previously been used to visualize and cluster target families. 23 In order to compare RNA versus protein targets based on the chemical matter that binds them, we trained a PCA using ECFP4 descriptors as features. A multicategory model was built in Pipeline Pilot using protein and RNA targets, binders, and nonbinders as categories, and ECFP4 as descriptors. Features that were present ≥20 times and that had a normalized probability ≥1.5 (963 features total) were used to train the PCA.
Results
Initial ALIS Screening and Building of a Small Molecule RNA Dataset
A set of 42 RNA targets (
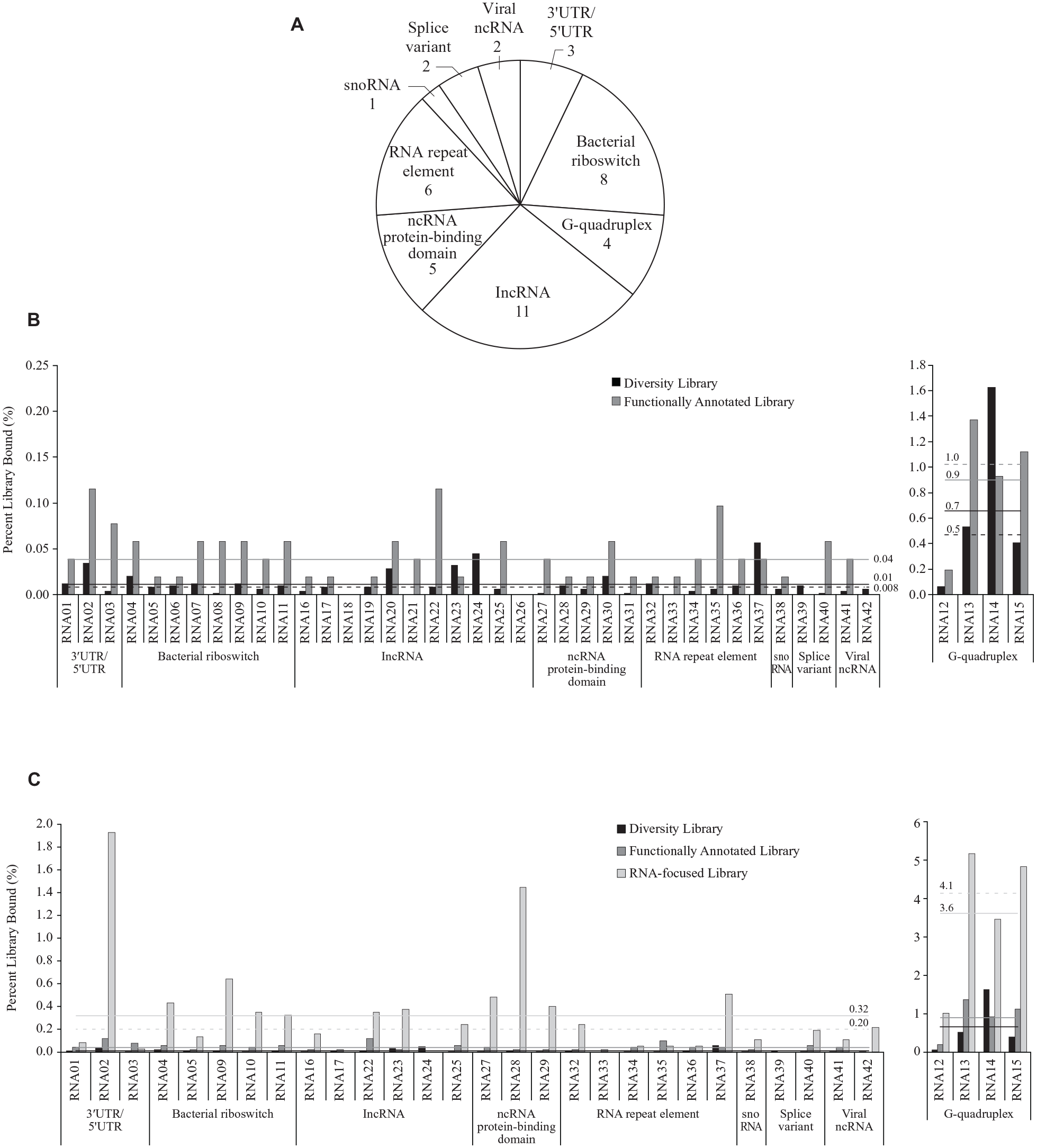
(
Of the 42 RNA targets screened, only two targets (RNA18 and RNA26) failed to bind any compounds from the screening collections (
Though the ALIS platform was able to identify small molecule binders for diverse RNA targets, we wondered whether the observed low hit rate across targets was due to inherent bias in the interrogated screening libraries that are traditionally used for protein-targeted drug discovery. The Functionally Annotated Library, assembled using pharmacogenomic data, and the Diversity Library, assembled from synthetic and commercial acquisition efforts, have historically both been applied toward the discovery of protein-binding ligands. We hypothesized that applying cheminformatic approaches to our combined primary binding data (
We generated naïve Bayesian models trained on RNA binders versus nonbinders using calculated physicochemical properties, chemical fingerprints, and biological fingerprints as descriptors.18,19 Model accuracy was assessed by calculating the AUC ROC score. High model AUC ROC scores supported model accuracy for several descriptors, and the model based on ECFP4 fingerprints performed best (
We evaluated the performance of the RNA-Focused Library in a second screen against a subset of 32 of the initial RNA targets (
The performance of the RNA-Focused Library, however, was variable across RNA targets. Of the 32 RNA targets rescreened, 24 targets resulted in an enriched number of binders, while 4 targets (RNA17, RNA24, RNA33, RNA39) yielded a similar or worse hit rate with the RNA-Focused Library compared with the previous libraries screened. One RNA target (RNA26, not shown), which did not have any binders from the initial screen against the Diversity and Functionally Annotated Libraries, also did not have any binders from the new RNA-Focused Library. Because the RNA-Focused Library hit rate was often greater than but uncorrelated with the initial screening library hit rate across targets, we concluded that our machine learning models selected compound features that generally promoted binding to RNA and were not strongly biased toward any one target. Furthermore, this study was aimed to identify compound features that promote general RNA binding instead of trying to optimize binding to each specific RNA target.
Selectivity of Small Molecule–RNA Interactions across RNA and Protein Targets
Using the entire set of binding data from the initial and expansion RNA screens, we interrogated how binding of small molecules compared across the panel of 42 RNA targets screened. We defined an RNA-selective compound as one that bound to only 1 RNA target across our panel of 42 RNA targets. While many compounds bound multiple RNA targets (
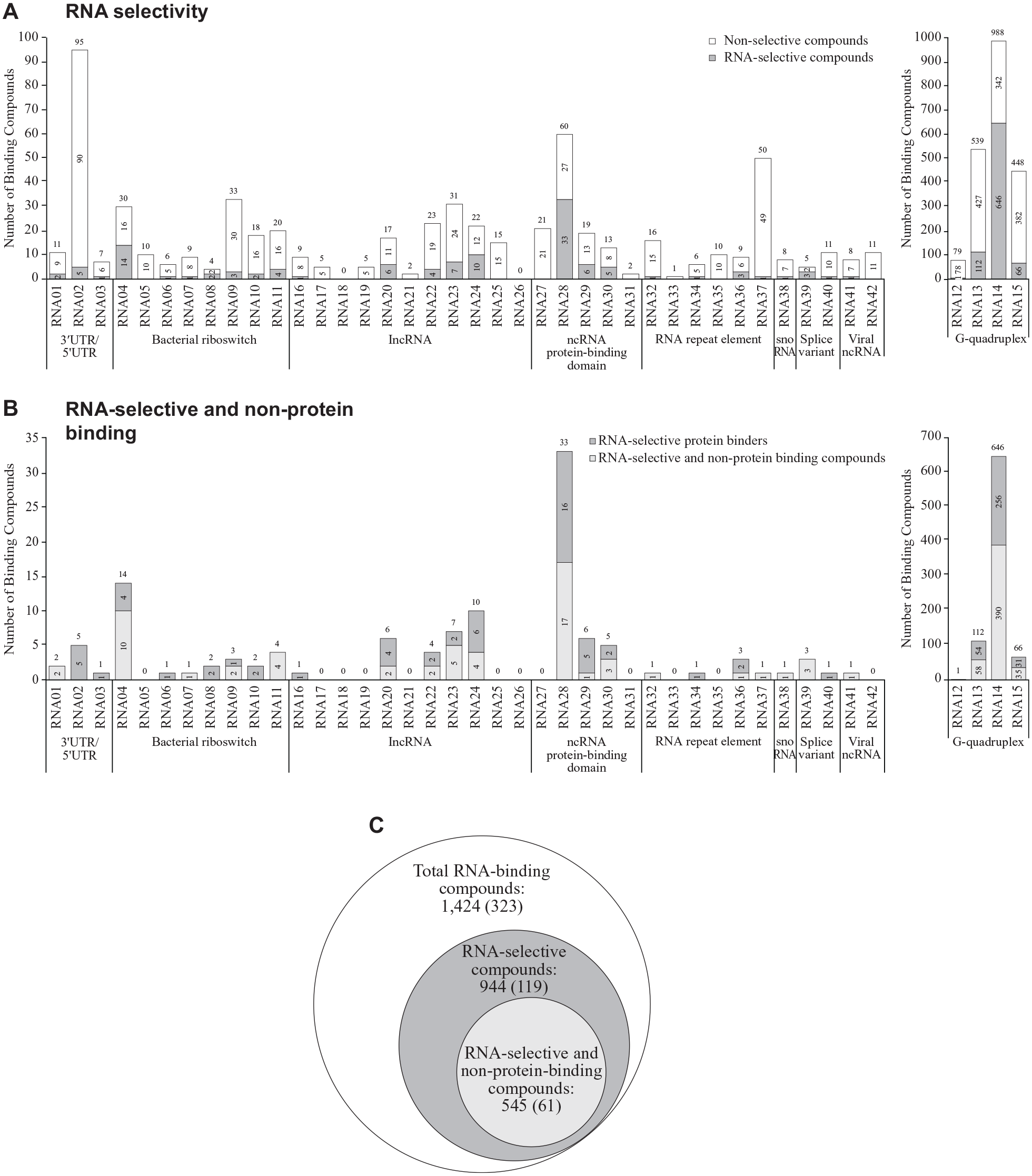
(
Having demonstrated the ability to identify binders with RNA selectivity within our panel, we next questioned whether or not these RNA-selective compounds bound to any known proteins. We used historical internal ALIS binding data collected at our company to identify any protein targets for our RNA-selective compounds. We found that 545 compounds of the 944 RNA-selective compounds also did not bind any known protein targets (57.7%;
In order to determine if binding promiscuity across RNA targets also indicated promiscuity across protein targets, we assessed the binding of our three libraries of compounds against protein targets screened in ALIS (internal historical data). A comparison of the number of RNA targets to the number of protein targets bound by each compound revealed that the degree of promiscuity across RNA targets was not correlated with the degree of promiscuity across protein targets (
It is important to note that we defined RNA selectivity as binding to a single RNA target in our panel and identified compounds that did not bind any known protein targets. However, binding interactions revealed by ALIS can have a range of affinities (Kd < 10 μM). Therefore, compounds may be binding with varying affinities to the identified RNA and protein targets. Determination of compound–target affinities may uncover “nonselective” compounds that in fact have a much higher affinity to a single target compared with all other targets. Furthermore, historically more protein targets have been screened in ALIS compared with RNA targets, making the selectivity criteria for proteins currently more rigorous than those for RNAs through this technique.
Properties of RNA Targets Favorable for Small Molecule Binding
For RNA to serve as a small molecule druggable target, RNA function must be mediated by secondary or tertiary structure,2,29 as opposed to sequence. We found that there is no correlation between RNA target size and compound binders (
To further interrogate this point, we looked to the G-quadruplex class of RNA. These G-rich RNAs can form well-defined, stable structures with four guanines interacting through Hoogsteen bonding in a planar manner around a central monovalent cation, particularly K+, which coordinates between the G-quartets.31,32 These structured RNAs are associated with several biological processes, including transcript processing and translational control. In our study, the relevance of the G-quadruplex structure for compound binding was investigated by screening these RNA targets under conditions that promote the G-quadruplex structure (i.e., high K+) as well as conditions that disfavor G-quadruplex formation (i.e., high Na+ in place of K+), as confirmed by circular dichroism under the same conditions.
32
Interestingly, we found that the many compounds that bound to the folded G-quadruplex structures did not bind to the same sequence under conditions that disfavor quadruplex formation (
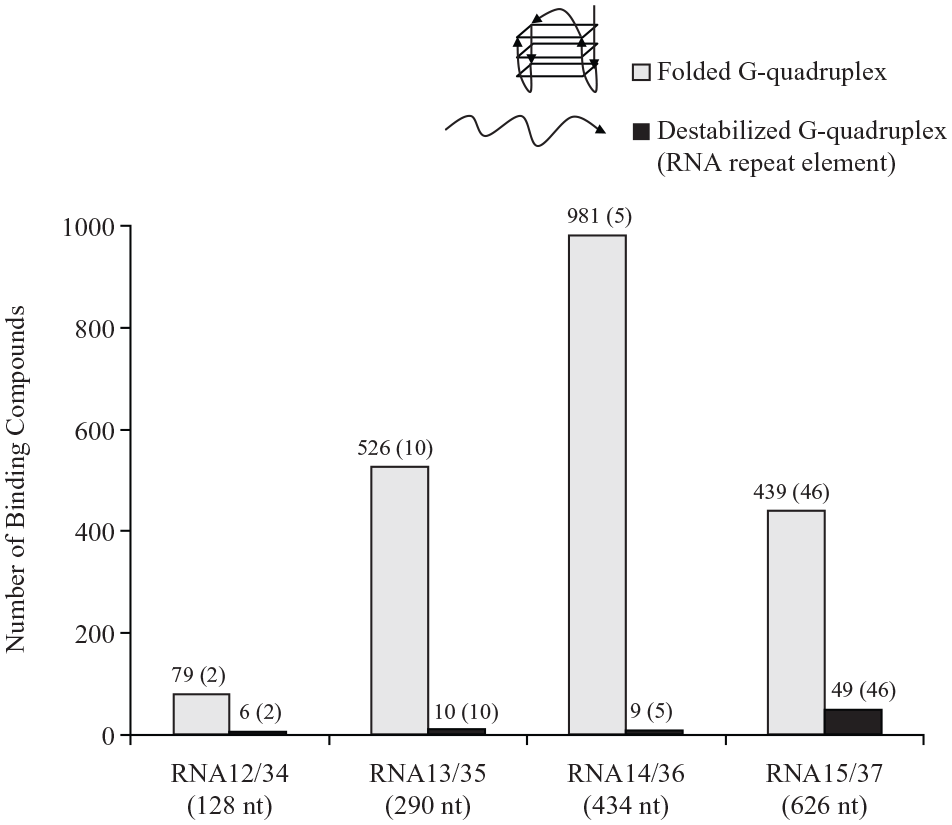
The number of binding compounds for folded versus unfolded G-quadruplex RNA. The same RNA sequence was stabilized for G-quadruplex formation (or not) using different salt conditions (i.e., high K+ concentration for G-quadruplex formation; high Na+ concentration for destabilized conditions). These RNA targets were screened in ALIS against the Diversity, Functionally Annotated, and RNA-Focused Libraries. The number of overlapping compounds between the folded and unfolded states is indicated in parentheses.
Through this work, we found no correlation between linear RNA size and compound binding and identified a strong example of structure dependence for compound binding. While it is known that small molecules can bind the major and minor grooves of RNA via intercalation or base-pair complementarity, for target specificity and strong pharmacology we aimed to target the tertiary structure of folded RNA by binding in the diverse pockets created by higher-order folding. The trends and requirements seen here suggest that we have, in fact, targeted structured RNA. However, further structural analysis is necessary to confirm these hypotheses.
Characteristics of RNA-Binding Compared with Protein-Binding Compounds
We next used our comprehensive small molecule–RNA dataset (
Consistent with the findings of Morgan et al.,
22
we observe that the majority of RNA binders and protein binders overlap within drug-like chemical space, although there are a few RNA-binding and protein-binding compounds that occupy space that is distinct from the drug-like compounds (
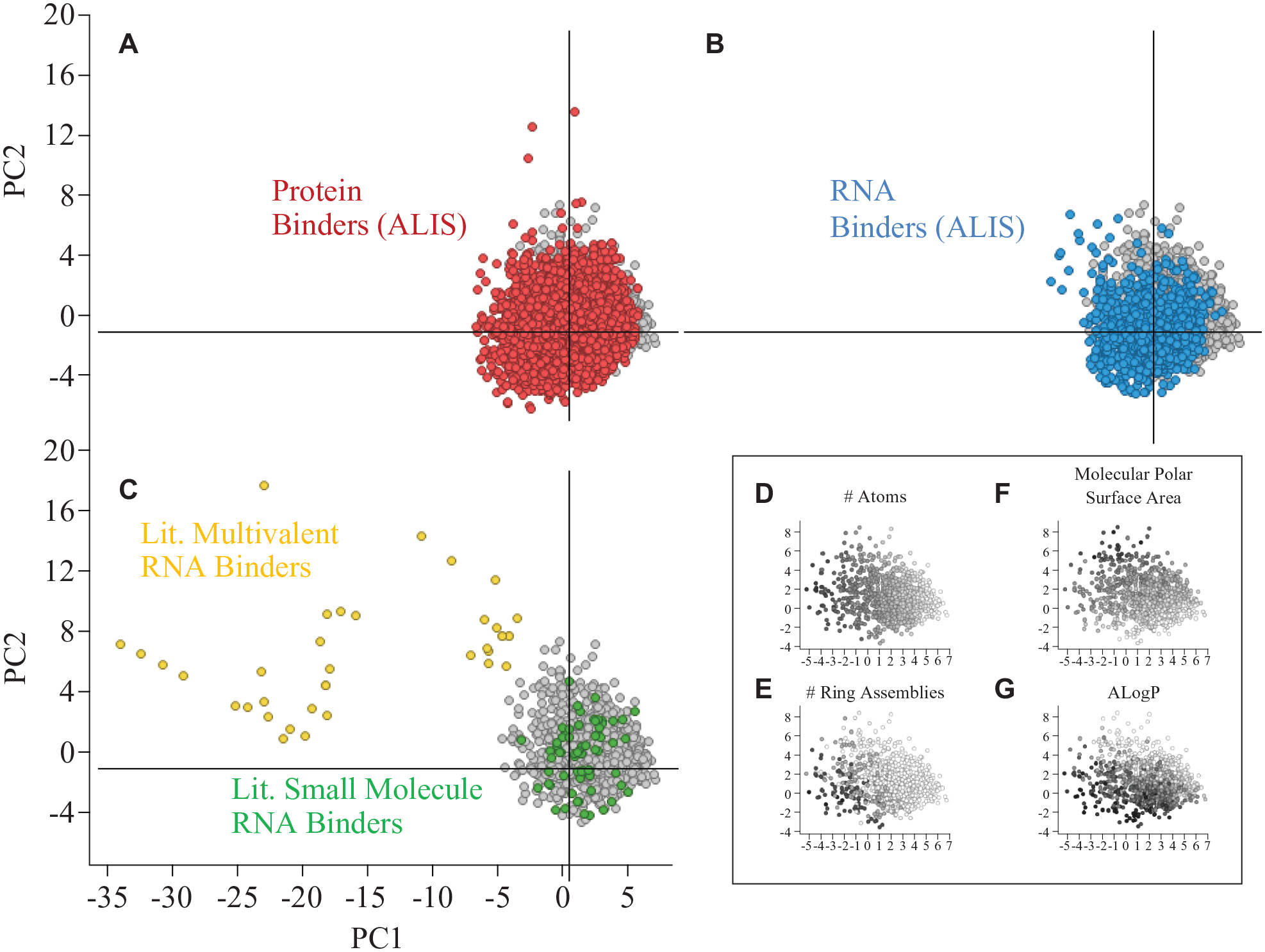
Comparison of physical chemical space occupied by protein binders, RNA binders, and drugs. In all plots, the same subset of approved drugs (gray) is depicted for comparison. (
We then further explored two properties of RNA and protein binders that significantly contributed to the PCA: molecular weight and AlogD. Using CHEMGENIE,
10
our company’s biochemical and pharmacogenomic database, we assembled historical protein ALIS binding data for RNA screening library compounds. We then calculated physicochemical properties for each compound in the screening collection and trained naïve Bayesian classification models to identify key differences in feature weights for RNA or protein binding (
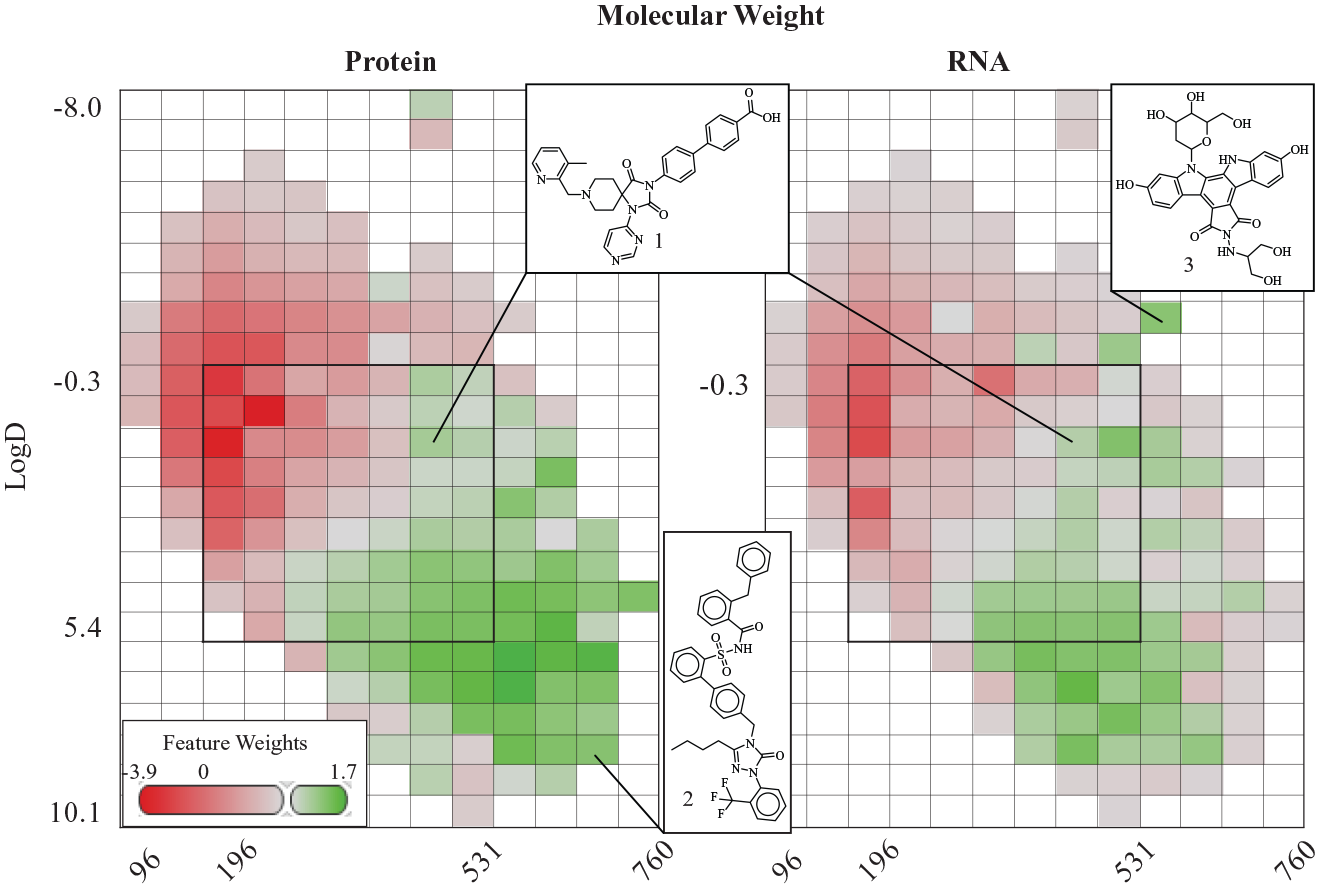
Feature score maps for physicochemical modeling of protein and RNA binders. Screening library compounds were binned based on calculated molecular weight (x axis) and LogD (y axis), and the color of each heat map cell represents the model normalized. Example compounds are displayed (1: a spirohydantoin inhibitor of prolyl hydroxylases; 34 2: triazolinone sulfonamide dual angiotensin 1/2 receptor antagonist 37 that does not bind RNA; 3: an indolocarbazole analog of NB-506 38 that had no detected protein binding in ALIS) and regions satisfying Lipinski’s rule of 535,36 are bound with bolded lines.
Highlighted in
The unique RNA–small molecule binding dataset that we have generated offered us the opportunity to compare RNA and protein targets with respect to the chemical matter that binds (and does not bind) them. We built naïve Bayesian models for each of our RNA and protein targets, based on the binders and nonbinders in our three small molecule libraries. We then extracted the chemical features from these models that were most enriched in binders for at least one of the targets and used these features across all targets to train a PCA (
To further investigate the specific features that are important for RNA binding, we built naïve Bayesian models trained on RNA binders versus nonbinders, with chemical features (ECFP4) as descriptors (cross-validated AUC ROC = 0.805). In this case, each chemical feature was assigned a normalized probability related to its enrichment within a model category of interest; a positive probability for a given feature indicated that molecules possessing that feature tend to bind RNA, while a negative probability indicated a feature to be among nonbinders. We analyzed chemical features with the highest and lowest model weights to find several features that were enriched for overall RNA binding (features i–xii;
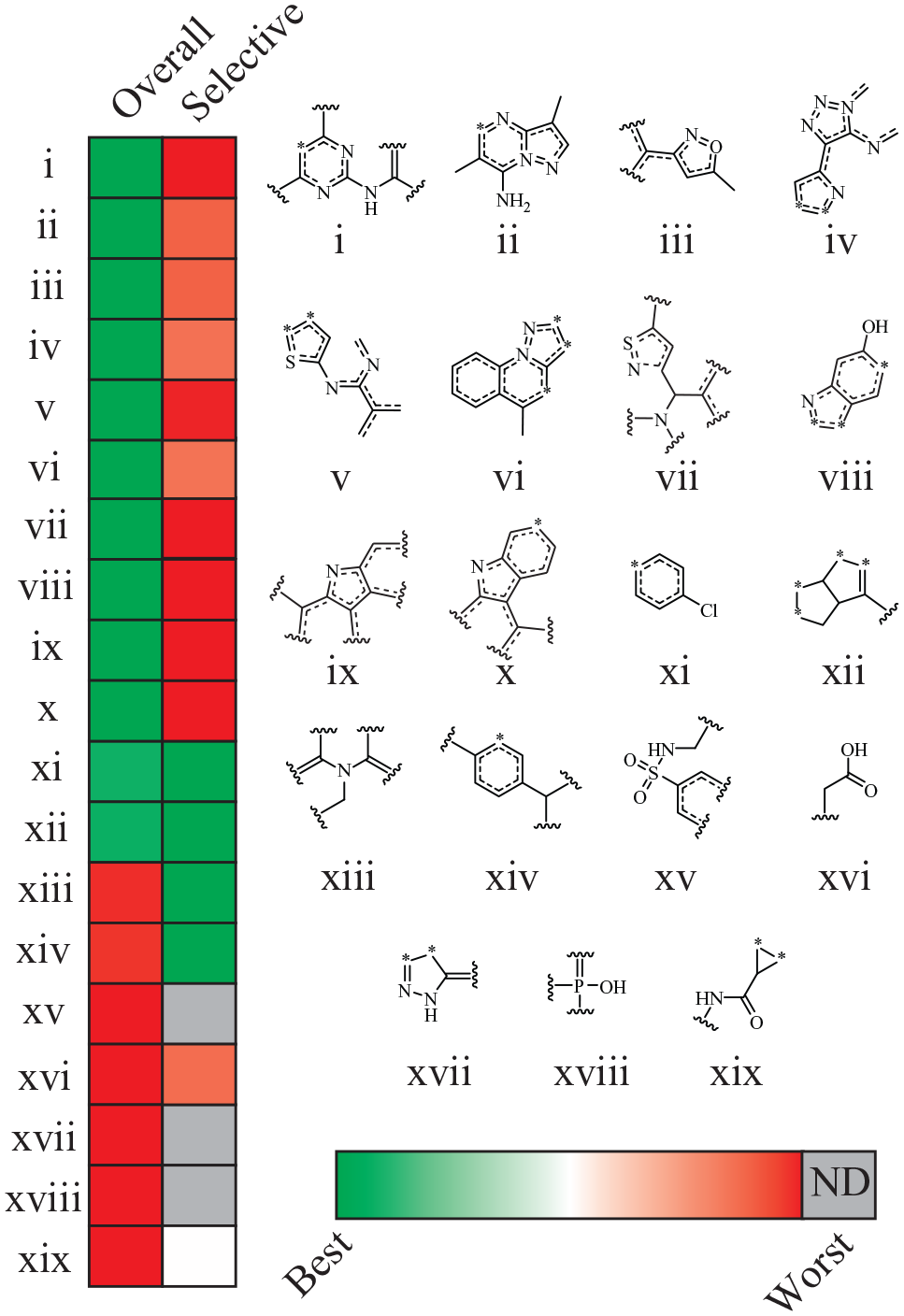
Chemical substructures enriched in specific binders, promiscuous binders, and nonbinders.
We next questioned what features were enriched in selective RNA binders. Surprisingly we learned that many of the features scoring highest in the overall binding model were among the lowest scoring features in the selective binding model, reflecting their effectiveness in enriching for general binding. Nevertheless, we did identify several substructures enriched among specific RNA binders (features xi–xiv;
In summary, our dataset and analyses have revealed that while many of the physicochemical properties of the RNA-binding compounds are similar to those of the protein-binding compounds, the features that are enriched in RNA binders are distinct from the features enriched in protein binders. Furthermore, combinations of specific features may render compounds to bind one RNA target preferentially over dozens of other targets.
Discussion
By using the ALIS affinity-selection mass spectrometry platform for HTS of 42 varied RNA targets against a total of ~60,000 compounds each, we have generated millions of target–compound interaction data points from which we have identified new drug-like small molecules that bind to RNA. Based on an analysis of small molecule binders of RNA, we have successfully built an RNA-Focused Library of ~3700 small molecules that is enriched for RNA binders compared with previous compound libraries in our collection. Our analysis has revealed compounds that are selective for RNA targets in our panel as well as general RNA binders. Our selective compound set includes compounds that have selectivity for specific RNAs against other RNA targets as well as protein targets. By looking at correlations to RNA target size, we reason that the majority of our compounds may be binding in a structure-dependent manner. Importantly, comparative screening under structured and disfavorable RNA-folding conditions identifies compounds that specifically bind the physiologically relevant folded state.
Our cheminformatics approach has given us an initial understanding of the physicochemical properties and chemical substructures that lead to both general and specific RNA binding. Importantly, many of our identified RNA-targeting small molecules are classically drug-like in their physicochemical properties, implicating their potential function as RNA-targeting therapeutics. Although there are a limited number of examples of small molecules intentionally designed to target RNA, this work provides initial guidance in this regard, and has uncovered selective RNA–small molecule interactions for focused small molecule drug discovery efforts.
The ALIS platform is a flexible approach that has historically been useful in providing an entry point for drug leads. In fact, ALIS has been used routinely to screen large numbers of drug-like small molecules to find binders to several types of protein targets in the past, and binders have often led to functional lead compounds with additional follow-up studies. 5 It is evident that additional structural, cell-based, and functional assays need to be done to clarify the biology of the small molecule–RNA interactions identified in this study. Furthermore, RNA ligandability using small molecules is becoming increasingly important with the development of new technologies such as ribonuclease-targeting chimeras (RIBOTAC), which uses a small molecule binder to RNA to recruit a nuclease for RNA degradation. 40 The ALIS technique used here complements other techniques, such as fluorescent indicator displacement (FID) 41 and selective 2′-hydroxyl acylation and primer extension (SHAPE), 29 that have been used to probe small molecule interactions with RNA. The intent of this study is to establish the ligandability of RNA by small molecules and assess the physiochemical properties and enriched features that govern RNA binding and selectivity. This work provides unique insights into RNA-targeted small molecule libraries and the identification of RNA ligands and will help add to previous efforts to classify the factors involved in RNA–small molecule ligand binding,42,43 thus aiding in RNA-targeted drug discovery efforts toward modulating the function of previously undruggable pathways.
Supplemental Material
Supplementary_Data_final – Supplemental material for Targeting RNA with Small Molecules: Identification of Selective, RNA-Binding Small Molecules Occupying Drug-Like Chemical Space
Supplemental material, Supplementary_Data_final for Targeting RNA with Small Molecules: Identification of Selective, RNA-Binding Small Molecules Occupying Drug-Like Chemical Space by Noreen F. Rizvi, John P. Santa Maria, Ali Nahvi, Joel Klappenbach, Daniel J. Klein, Patrick J. Curran, Matthew P. Richards, Chad Chamberlin, Peter Saradjian, Julja Burchard, Rodrigo Aguilar, Jeannie T. Lee, Peter J. Dandliker, Graham F. Smith, Peter Kutchukian and Elliott B. Nickbarg in SLAS Discovery
Footnotes
Acknowledgements
The authors would like to thank Anne Mai Wasserman and Kerrie Spencer for their advice and support of this research.
Supplemental material is available online with this article.
Authors’ Note
Julja Burchard is currently affiliated with Sera Prognostics, Inc., Salt Lake City, UT, USA. Graham F. Smith is currently affiliated with AstraZeneca, Drug Safety and Metabolism, IMED Biotech Unit, Cambridge, UK.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors except R.A. and J.T.L. are current or former employees of Merck & Co., Inc., and may hold stock or other financial interests in Merck & Co.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: J.T.L. received support from a Merck MINt award, and R.A. was funded by a Pew Latin American Fellowship. All other authors were supported by Merck & Co., Inc.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
