Abstract
Organic cation transporters (OCTs) are membrane proteins with relevant physiological (because they accept neurotransmitters as substrate) and pharmacological (because of their interaction with drugs) roles. The human OCTs hOCT1 (SLC22A1/hOCT1) and hOCT2 (SLC22A2/hOCT2) are highly expressed in hepatic (hOCT1) and in renal and neuronal tissue (hOCT2), suggesting a possible role in modulating neurotransmitter activity in the liver, kidney, and brain, and their clearance from the blood. Even though there are several data demonstrating that OCTs are regulated under various patho-physiological conditions, it remains largely unknown which proteins directly interact with OCTs and thereby influence their cellular processing, localization, and function. In this work, using a mating-based split-ubiquitin yeast two-hybrid system, we characterized the potential interactome of hOCT1 and 2. It became evident that these OCTs share some potential interaction partners, such as the tetraspanins CD63 and CD9. Moreover, we confirmed interaction of hOCT2 with CD9 by fluorescence-activated cell sorting coupled with Förster resonance energy transfer analysis. Together with other proteins, tetraspanins build “tetraspanins webs” in the plasma membrane, which are able to regulate cellular trafficking and compartmentalization of interacting partners. While CD63 was demonstrated to mediate the localization of the hOCT2 to the endosomal system, we show here that co-expression of hOCT2 and CD9 led to strong cell surface localization of the transporter. These data suggest that tetraspanins regulate the cellular localization and function of OCTs. Co-localization of CD9 and hOCT was confirmed in tissues endogenously expressing proteins, highlighting the potential biological relevance of this interaction.
Keywords
Introduction
Organic cations (OCs) are substances that at physiological pH bear a positive charge. Important neurotransmitters such as dopamine and histamine and many drugs such as cetirizine, cimetidine, and trospium belong to this class of substances. Because of their charge, OCs cannot freely pass the plasma membrane and need specialized transport systems to cross the cell membrane. The organic cation transporters (OCTs) represent such a transport system. OCTs are polyspecific, electrogenic, pH- and Na+-independent, but voltage-dependent bidirectional transporters for OCs with species- and tissue-specific expression. 1 Indeed, OCTs are considered to be an important regulator of monoamine neurotransmitter concentration in the CNS,2–5 and, because of their high expression in secretory organs such as the liver (OCT1) and the kidneys (OCT2), they play an important role in OC excretion. This fact is of particular importance considering their interaction with drugs such as the antidiabetic metformin and the anticholinergic drug trospium, whose excretion is mediated by OCT1 and OCT2.6,7 In the case of intracellular active drugs, OCTs are important also for determining drug efficacy (as suggested for antiretroviral drugs, 8 for example) and unwanted side effects, as demonstrated for example for platinum derivatives, which are untypical OCT substrates.9,10 Polymorphisms of OCT have been demonstrated to have important pathophysiological implications, 11 and to affect drug efficacy 12 and side effects. 13 It is well-known that OCT activity is regulated by various protein kinases 14 and that their expression is markedly changed under specific pathological conditions.15,16 There is scarce information, however, on which proteins directly interact with OCTs, potentially influencing their cellular processing, localization, and function. In this work, we have used a mating-based split-ubiquitin system (mbSUS) screening specific for plasma membrane proteins to identify direct interaction partners of human OCT1 and 2 paralogs (hOCT1 and 2). Among others, the tetraspanin CD9 was identified as a common interaction partner of hOCT1 and 2. The interaction of CD9 with hOCT2 was confirmed using fluorescence-activated cell sorting (FACS) coupled with Förster resonance energy transfer (FRET) analysis, and localization of overexpressed and endogenous proteins was investigated in expression systems and human tissues.
Materials and Methods
DNA Constructs
The full-length hOCT2 (SLC22A2, NM 003058) cloned in the expression vector pRc/CMV (hOCT2) was a generous gift of H. Koepsell (University of Würzburg, Würzburg, Germany). Human CD9 was amplified from pExpress-1-CD9 (clone IMAGp998A1815788Q; imaGenes, Berlin, Germany) by PCR. For FRET–FACS analysis and some transfection experiment, hOCT2 and CD9 were amplified by PCR and inserted in the plasmids pEGFP-N3, pYGFP-C1 or N1, and pECFP-N1 or C1 (Clontech-Takara Bio Europe, Saint-Germain-en-Laye, France), with XhoI and BamHI sites to obtain constructs tagged with green fluorescent protein (GFP), yellow fluorescent protein (YFP), or cyan fluorescent protein (CFP). The primers used in this work were obtained from MWG (Ebersberg, Germany) and are listed in Supplemental Table S1 . CD9 was also inserted into a pcDNA6.F9 vector containing a FLAG and 6xHis-Tag 17 via MluI and NotI sites. All constructs were verified by automated sequencing.
Mating-Based Split-Ubiquitin System
Yeast two-hybrid systems are extensively used to identify protein–protein interactions (PPIs). In this work, we used the mating-based split-ubiquitin system (mbSUS), a particular form of yeast two-hybrid assay, in which PPIs can happen also in the plasma membrane, to identify potential interaction partners of hOCT1 and hOCT2. Using this technique, we already identified the tetraspanin CD63 18 and the lysosomal-associated protein transmembrane 4A (LAPTM4A) 19 as interaction partners of hOCT1–3 and demonstrated that hOCT2 can form oligomers. 20 The mbSUS is described in detail in those publications. Briefly, mbSUS screening of potential PPIs was performed using full-length complementary DNA (cDNA) of hOCT1 or hOCT2 cloned in-frame into the expression vector pMetYCgate containing the C-terminal half of ubiquitin (Cub) 21 as bait and the N-terminal half of ubiquitin (NubG)-fused human cDNA kidney library cloned in the pPR3-N vector (Dualsystems Biotech, Schlieren, Switzerland) as prey. Bait–prey interactions result in ubiquitin reconstitution, cleavage by ubiquitin-specific proteases, and release of the protA-LexA-VP16 peptide transcription factor, which allows yeast growth on minimal media (see Suppl. Fig. S1 ). 18 To identify interactors, plasmid DNAs were isolated with the Zymoprep Yeast Plasmid Miniprep Kit (Zymoresearch, Freiburg, Germany); identified prey clones were reintroduced into yeast, together with the bait (hOCT1 or hOCT2), and growth of the resulting co-transformants was assayed for growth on selective medium. Isolated plasmid DNAs were retransformed in Escherichia coli XL-1-Blue, isolated with the Zyppy Plasmid Miniprep Kit (Zymoresearch), and verified by automated sequencing (screen forward primer: 5′-GTC GAA AAT TCA AGA CAA GG-3′).21,22 The assays included positive and negative controls.18–20 Furthermore, to investigate which part of hOCT2 is important for interactions with the tetraspanin CD9, a truncation strategy similar to that used in studies on hOCT2–CD63 interaction 18 was used. Briefly, CD9 (Cub–CD9) was used as bait, while full-length hOCT2 or several truncated forms of the transporter were fused to a NubG fragment and used as prey. Growth of yeast cells in minimal media was possible only on interaction of the Cub–CD9 and NubG–hOCT2 fused proteins, and consequent activation of reporter genes for histidine and adenine. The primers used in the mbSUS are listed in Supplemental Table 1 .
Cell Culture and Transfection
Human embryonic kidney (HEK293) cells (CRL-1573; American Type Culture Collection, Rockville, MD) were grown in Dulbecco’s modified Eagle medium (DMEM) containing 3.7 g/l NaHCO3, 1.0 g/l D-glucose, and 2.0 mM L-glutamine (Biochrom, Berlin, Germany), gassed with 8% CO2 at 37 °C. Streptomycin (100 mg/l), penicillin (100 U/ml) (Biochrom), 10% fetal bovine serum (FBS), and, only for transfected cells, 0.8 mg/ml G-418 (PAA Laboratories, Coelbe, Germany) was added to the medium. For FACS coupled with FRET (FACS–FRET) analysis, 1–3 µg of cDNA per donor and acceptor construct was transfected in HEK293 cells by the calcium phosphate method, as previously described in Grabner et al.; 19 and 24–36 h post transfection, FACS–FRET measurements were performed as already described in Schulze et al. 18
Madin-Darby canine kidney (MDCK) cells were transfected with hOCT2–EGFP using Nanofectamine according to the manufacturer’s recommendation (PAA Laboratories). Neomycin resistance selection was performed with increasing concentrations of geneticin (0.2–0.8 mg/ml). A single clone was isolated, grown, and used for transfection experiments with either control vector pcDNA3.1 or CD9 in the pcDNA6.F9 vector using the Lipofectamine 2000 method, as recommended by the manufacturer (Invitrogen, Thermo Fisher Scientific, Waltham, MA). Transfected MDCK cells were cultured in minimal essential medium (MEM; Biochrom) containing 10% FBS, penicillin, streptomycin, and 0.8 mg/ml geneticin (PAA Laboratories) at 37 °C and 8% CO2.
FACS–FRET Analysis
For FACS–FRET measurements, a LSRII (BD Bioscience, Heidelberg, Germany) flow cytometer equipped with 405 nm, 488 nm, and 633 nm lasers was used as described in Banning et al. 23 and Schulze et al. 18 Briefly, enhanced cyan fluorescent protein (ECFP) fluorescence and FRET were detected in transfected HEK293 cells after excitation at 405 nm. ECFP emission was measured at 440 nm, while the FRET signal was detected at 525 nm. Enhanced yellow fluorescent protein (EYFP) fluorescence was measured after excitation at 488 nm; emission was detected at 530 nm. At least 1000 CFP/YFP-positive cells were measured in the gate after correction for the background.
Fluorescence Microscopy
For fluorescence microscopy, hOCT2–GFP and CFP–CD9 plasmids were co-transfected in HEK293 cells using Lipofectamine 2000 (Invitrogen). Twenty-four hours later, transfected cells were fixed with 2% paraformaldehyde (PFA). DNA was labeled with Hoechst 33342 (Sigma-Aldrich, St. Louis, MO). Images were acquired using a Zeiss Axiovert 200 M microscope equipped with a Plan-Apochromat 100× (1.4 NA), a multiline argon laser, and a Zeiss AxioCamHR digital camera (Zeiss, Oberkochen, Germany). For analysis of co-localization, the Axiovision co-localization module was used (Zeiss). Expression of hOCT1 and 2 and CD9 in normal human nasal epithelial cells grown on filters, and in human dorsal root ganglia (DRGs), was investigated using specific antibodies raised against hOCTs (MCA34557; AbDSerotec, Kidlington, UK; and Biermann et al. 24 ) or against human CD9 (BD Biosciences, or Epitomics, Abcam, Berlin, Germany). To do this, human nasal epithelial cells or 5 µm thick cryosections prepared from human DRGs were fixed in 4% PFA. After 5 min incubation with 0.2% TritonX100 in Dulbecco’s phosphate-buffered saline (PBS; Biochrom, Berlin, Germany), samples were incubated at room temperature in PBS containing 10% normal goat serum (pH 7.4) for 60 min and then at room temperature for 90 min or overnight at 4 °C with affinity-purified primary antibodies [hOCT MCA34557 1:100 and CD9 Epitomics 1:1000 in 1% bovine serum albumin (BSA) in PBS]. PBS was used to wash the samples; after washing, the samples were incubated at room temperature for 45 min with 1% PBS-diluted secondary antibodies [Alexa Fluor 488 goat anti-rabbit-immunoglobulin (Ig) or Alexa Fluor 594 goat anti-mouse-Ig, 1:1000; Invitrogen] and with 4′,6-diamidino-2-phenylindole (DAPI) at a 1:1000 dilution in 1% BSA in PBS. Samples were rinsed with PBS, coverslipped with Fluoromount (Sigma-Aldrich), and evaluated by epifluorescence microscopy (Observer Z1 with Apotome technology; Zeiss). Negative control slides (without primary antibody or in the presence of the control peptide) were also analyzed.
MDCK cells expressing hOCT2–GFP and CD9 grown on cover slips for 48 to 72 h were fixed in 4% PFA. After fixation, the cells were rinsed three times with PBS and incubated with 0.1% Triton X-100 for 3 min. After extensive rinsing with PBS, blocking of unspecific binding sites by 60 min incubation at room temperature with 10% BSA (Sigma-Aldrich) in PBS was performed. Cells were then incubated overnight at 4 °C with anti-CD9 (BD Biosciences) diluted 1:100 in 1% BSA in PBS. After three washing steps in PBS, the secondary antibodies (goat-anti-mouse Alexa Fluor 594) at a 1:1000 dilution and DAPI at a 1:2000 dilution in 1% BSA in PBS were incubated for 60 min, followed by five more washing steps in PBS. Finally, cells were mounted in Fluoromount (Sigma-Aldrich), and fluorescence photographs were taken with an Axiocam camera mounted on an Axiovert 100 microscope (Zeiss) using Axiovision software.
Human Samples
Lumbar position 4 (L4) human DRGs were obtained from the National Disease Research Interchange (NDRI). The use of human tissue was approved by the Institutional Review Board of Ohio State University, and the NDRI obtained informed consent from donors of human tissue. Human nasal epithelial primary cells were freshly isolated from nasal tissue of patients undergoing surgery at the University Hospital Heidelberg. The use of human material was approved by the ethics committee of the University of Heidelberg (S136/2016).
Chemicals
All substances and standard chemicals were obtained from Sigma-Aldrich or Merck (Darmstadt, Germany), except when specified differently.
Statistical Analysis
Data are presented as mean values ± standard deviation (SD), with n referring to the number of samples used in the experiments. Analysis of variance (ANOVA; with Tukey posttest) was used to prove statistical significance of the effects using GraphPad Prism 5.3 (GraphPad Software Inc., San Diego, CA). A P-value < 0.05 was considered statistically significant.
Results
Identification of hOCT1 and 2 Interacting Partners Using mbSUS
To identify proteins potentially interacting with hOCT1 and 2, a mbSUS screening using a human kidney library as prey was performed. The mbSUS is a special yeast two-hybrid system that allows identifying interactions that take place in vivo directly at the plasma membrane, allowing to detect interactions involving integral membrane proteins, membrane-associated proteins, and soluble proteins.18–20 Moreover, in this system, posttranslational modifications such as glycosylation are preserved.21,25 This screening resulted in around 100 and 280 independent clones for hOCT1 and 2, respectively. After sequencing, 24 and 36 potential interaction partners for hOCT1 and 2, respectively, were identified (see Table 1 ). Eleven of these proteins were common to both transporters ( Table 1 ). Some proteins were identified in several independent clones, suggesting a stronger probability that they are real interaction partners. Classification of the identified interaction partners, using the GOTERMMAPPER (https://go.princeton.edu/cgi-bin/GOTermMapper) gene ontology (GO) annotations 26 of the Lewis-Sigler Institute for Integrative Genomics at Princeton University ( Fig. 1 ), shows that they are associated mainly with transport processes, anatomical structures, vesicles-mediated transport, signaling pathways, protein modification, homeostatic processes, and cell adhesion. These results reflect the functional implications of OCTs and reveal which proteins may be involved in posttranslational modification of the transporters and in their cellular trafficking. Of course, further investigations performed with the mbSUS and also in independent systems are necessary to confirm these interactions. The interactions observed here of hOCT with the tetraspanin CD63 and with LAPTM4A have been previously described and characterized in detail.18,19 Since tetraspanins are known to organize the plasma membrane in specific domains by interacting with several proteins, here we will focus on the tetraspanin CD9 and will further investigate its interaction with hOCT using the mbSUS and other independent methods. Interactions between hOCT1–2 and CD9 were confirmed in yeast mating assays ( Fig. 2 ). Figure 2 depicts yeast cells growing on minimal media after mating of OCT coupled with Cub (OCT–Cub) and CD9 coupled with NubG (NubG–CD9), and of OCT–Cub with Nub WT (wild type) as a positive control. No growth of yeast cells could be detected in the negative control experiments (OCT–Cub with the empty NubG vector).
Interactome of hOCT1 and hOCT2 as Identified by the mbSUS.
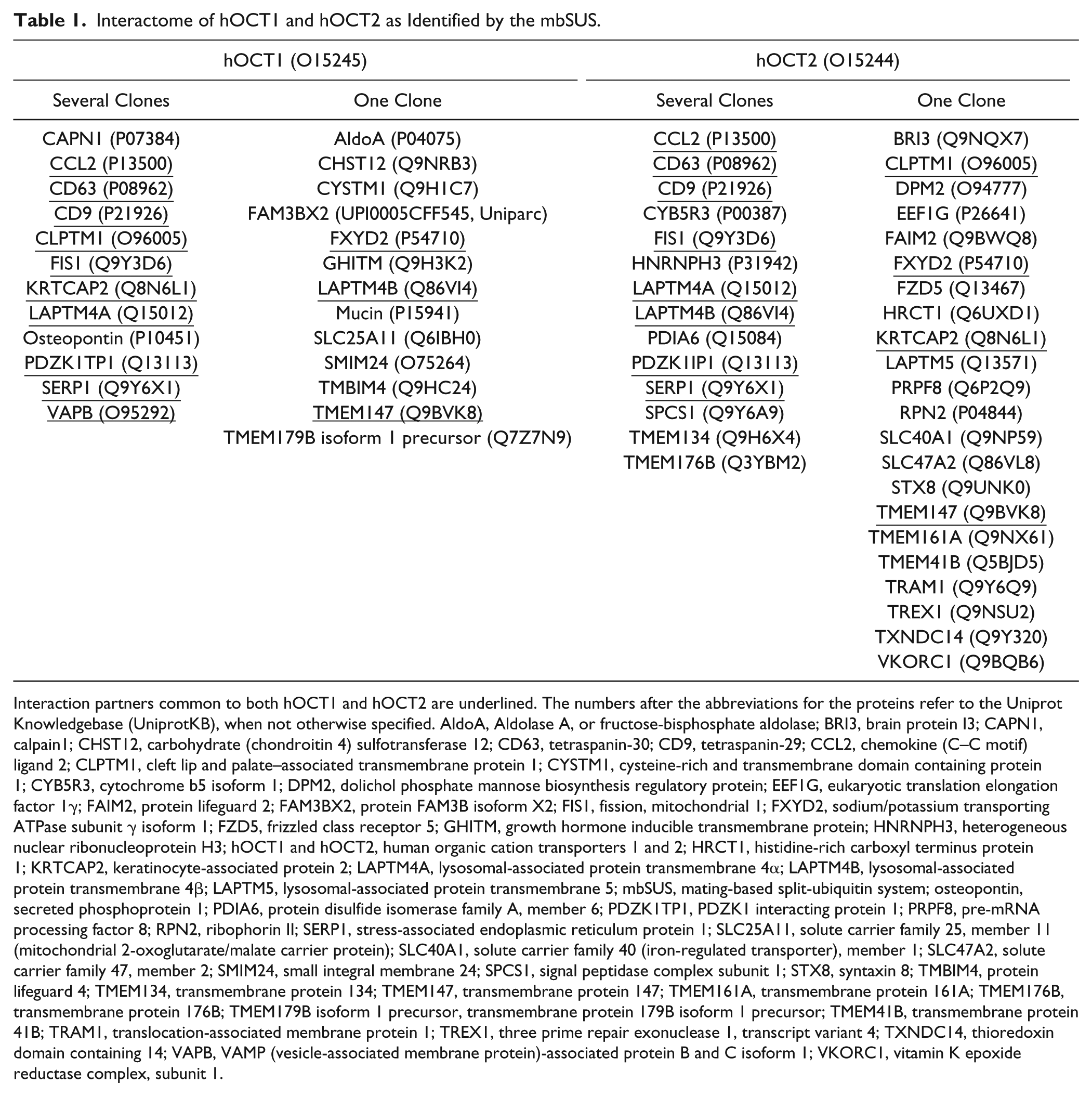
Interaction partners common to both hOCT1 and hOCT2 are underlined. The numbers after the abbreviations for the proteins refer to the Uniprot Knowledgebase (UniprotKB), when not otherwise specified. AldoA, Aldolase A, or fructose-bisphosphate aldolase; BRI3, brain protein I3; CAPN1, calpain1; CHST12, carbohydrate (chondroitin 4) sulfotransferase 12; CD63, tetraspanin-30; CD9, tetraspanin-29; CCL2, chemokine (C–C motif) ligand 2; CLPTM1, cleft lip and palate–associated transmembrane protein 1; CYSTM1, cysteine-rich and transmembrane domain containing protein 1; CYB5R3, cytochrome b5 isoform 1; DPM2, dolichol phosphate mannose biosynthesis regulatory protein; EEF1G, eukaryotic translation elongation factor 1γ; FAIM2, protein lifeguard 2; FAM3BX2, protein FAM3B isoform X2; FIS1, fission, mitochondrial 1; FXYD2, sodium/potassium transporting ATPase subunit γ isoform 1; FZD5, frizzled class receptor 5; GHITM, growth hormone inducible transmembrane protein; HNRNPH3, heterogeneous nuclear ribonucleoprotein H3; hOCT1 and hOCT2, human organic cation transporters 1 and 2; HRCT1, histidine-rich carboxyl terminus protein 1; KRTCAP2, keratinocyte-associated protein 2; LAPTM4A, lysosomal-associated protein transmembrane 4α; LAPTM4B, lysosomal-associated protein transmembrane 4β; LAPTM5, lysosomal-associated protein transmembrane 5; mbSUS, mating-based split-ubiquitin system; osteopontin, secreted phosphoprotein 1; PDIA6, protein disulfide isomerase family A, member 6; PDZK1TP1, PDZK1 interacting protein 1; PRPF8, pre-mRNA processing factor 8; RPN2, ribophorin II; SERP1, stress-associated endoplasmic reticulum protein 1; SLC25A11, solute carrier family 25, member 11 (mitochondrial 2-oxoglutarate/malate carrier protein); SLC40A1, solute carrier family 40 (iron-regulated transporter), member 1; SLC47A2, solute carrier family 47, member 2; SMIM24, small integral membrane 24; SPCS1, signal peptidase complex subunit 1; STX8, syntaxin 8; TMBIM4, protein lifeguard 4; TMEM134, transmembrane protein 134; TMEM147, transmembrane protein 147; TMEM161A, transmembrane protein 161A; TMEM176B, transmembrane protein 176B; TMEM179B isoform 1 precursor, transmembrane protein 179B isoform 1 precursor; TMEM41B, transmembrane protein 41B; TRAM1, translocation-associated membrane protein 1; TREX1, three prime repair exonuclease 1, transcript variant 4; TXNDC14, thioredoxin domain containing 14; VAPB, VAMP (vesicle-associated membrane protein)-associated protein B and C isoform 1; VKORC1, vitamin K epoxide reductase complex, subunit 1.
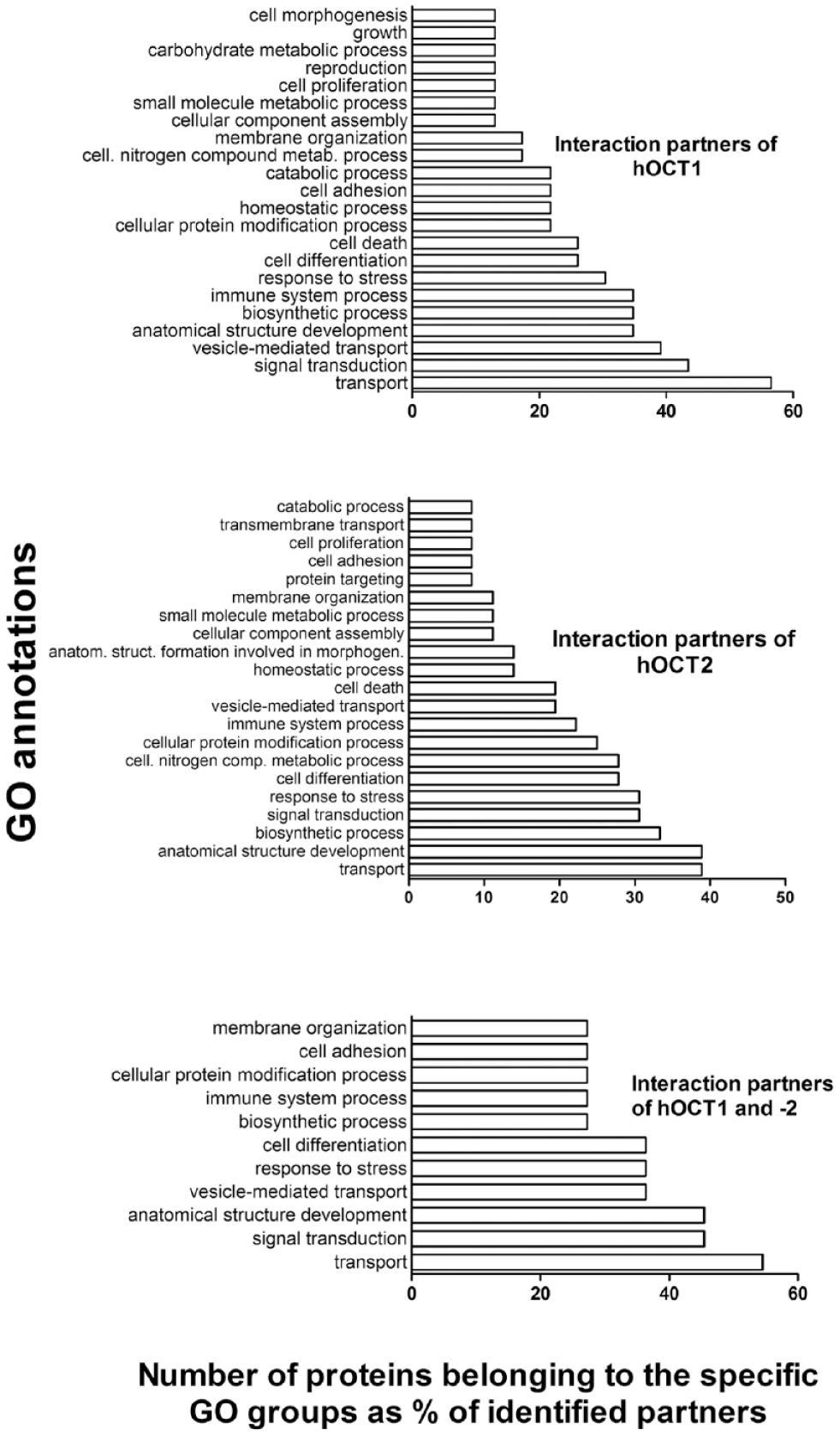
Classification of the interaction partners of human organic cation transporter 1 (hOCT1; upper panel) and hOCT2 (middle panel), and of hOCT1 and 2 common partners (lower panel), according to their gene ontology (GO) annotations. Groups containing more than two interaction partners are shown. GO annotations were according to the generic GOTERMMAPPER (https://go.princeton.edu/cgi-bin/GOTermMapper) classification 26 of Princeton University, NJ.

Example of interaction of CD9 with human organic cation transporter 2 (hOCT2) in the mating-based split-ubiquitin system (mbSUS). Reporter gene activation was shown as growth of yeast cells on minimal medium without adenine and histidine. Yeast mating assay demonstrates interaction between the N-terminal half of ubiquitin (NubG)–CD9 and hOCT2–Cub (the C-terminal half of ubiquitin) (colony formation
Identification of the Interaction Domain with the mbSUS
For the identification of the domains of hOCT2 involved in the interaction with CD9, the mbSUS was again applied. As bait, CD9–Cub was used, while as prey, several different hOCT2 truncations fused to NubG were used. These four truncated hOCT2 contained the first (a) 168, (b) 212, (c) 266, or (d) 379 amino acids (aa) of the transporter sequence ( Figure 3 ). Since every truncation shows an interaction with CD9, it can be supposed that CD9–hOCT2 interaction involves a region of hOCT2 located in the first 168 aa sequence, which contains the N-terminus, the first and second transmembrane domain, and the large extracellular loop between them.
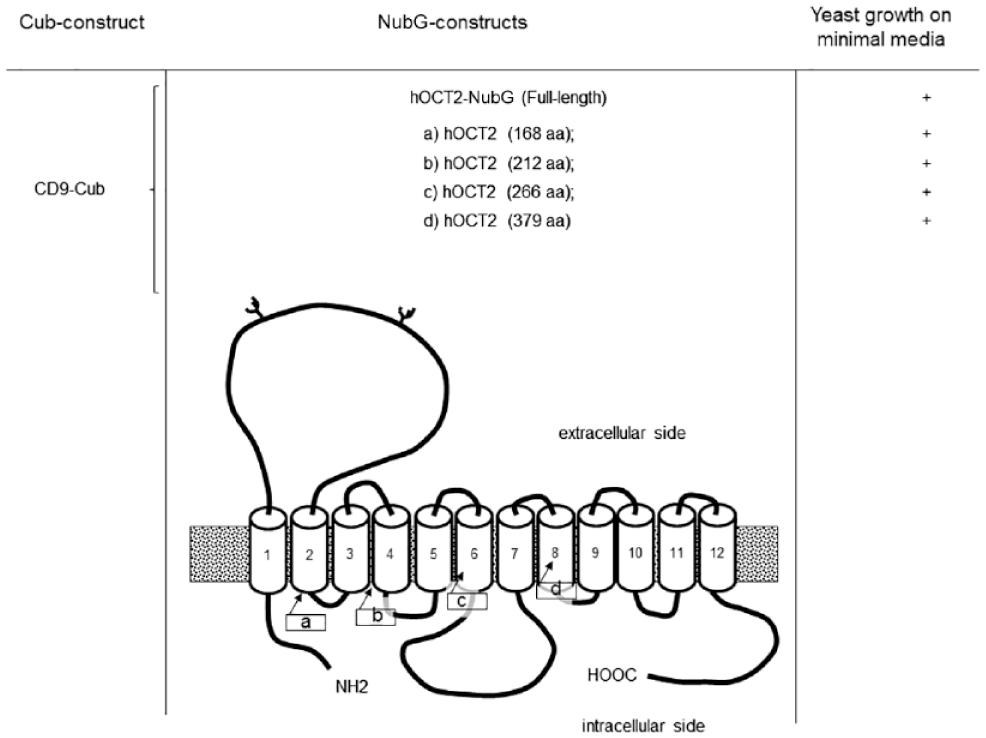
Identification of hOCT2 region interacting with CD9 by the mbSUS. CD9–hOCT2 (human organic cation transporter 2) interaction domains identified by the mating-based split-ubiquitin system (mbSUS). Truncation strategy to identify the hOCT2 part involved in the interaction with CD9 by the mbSUS. As bait, CD9 fused to the C-terminal part of the ubiquitin was used, while as prey, the full-length hOCT2 or four different truncations (
CD9 Interacts with hOCT2 In Vitro in HEK293 and MDCK Cells
The spatial proximity of CD9 and hOCT2 is an indication of probable interaction between the two proteins. Co-localization experiments after transfection of hOCT2–GFP and CD9–CFP plasmids in HEK293 cells show an extensive expression of both proteins in the plasma membrane. To a minor degree, a co-localization also can be detected in some intracellular vesicles ( Fig. 4A ). Since hOCT2 is mainly expressed in polarized cells such as renal proximal tubule cells, the co-localization of hOCT2 and CD9 was investigated also in the MDCK cells, a known model of polarized cells. Again, transfection of CD9 in MDCK cells stably expressing hOCT2–GFP resulted in co-localization of the two proteins mainly in the plasma membrane ( Fig. 4B ). CD9 and hOCT2 seem to co-localize in membrane protrusions (microspikes) resembling filopodia or microvilli (enlargements of Fig. 4B ).
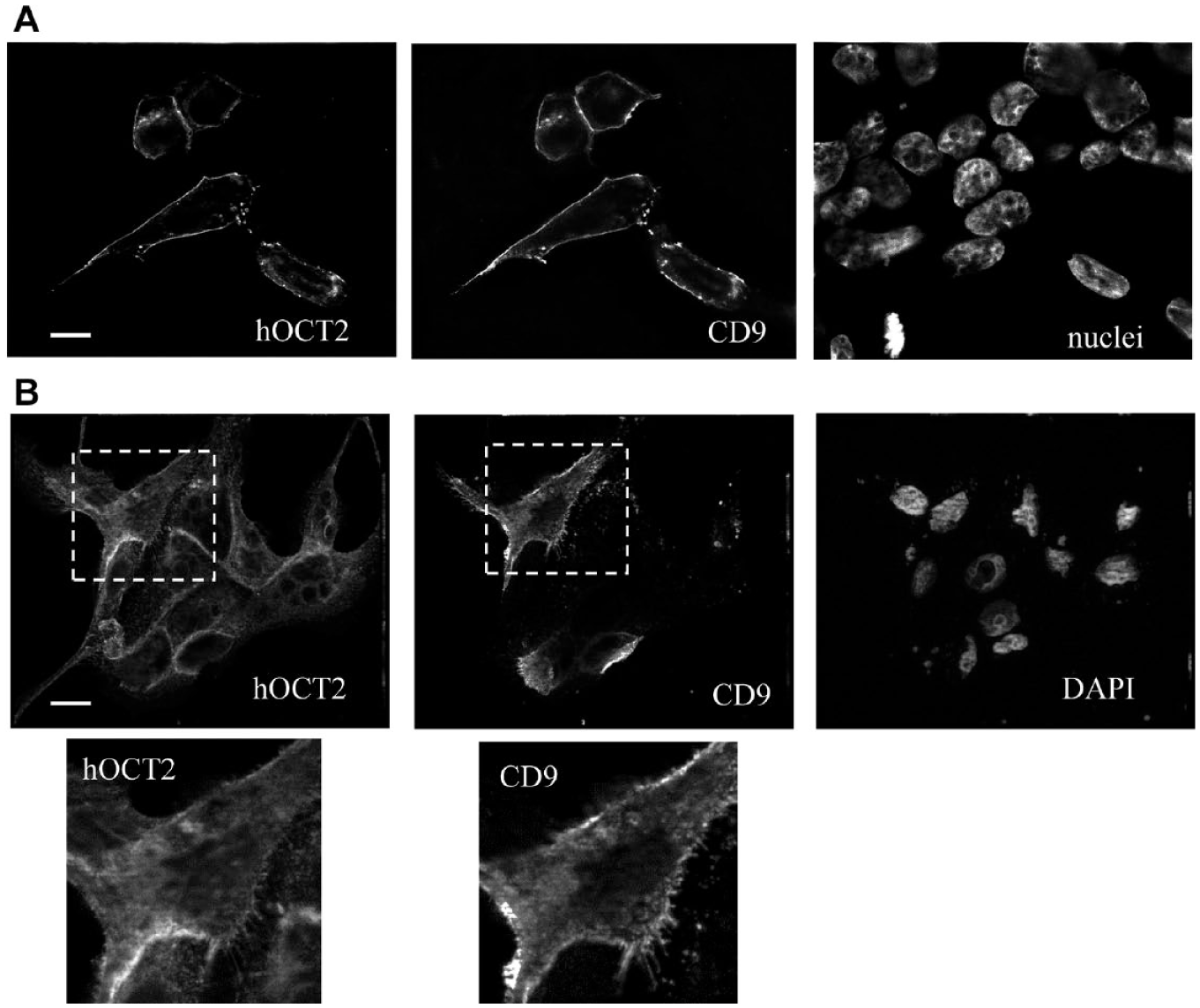
(
The interaction between CD9 and hOCT2 was also investigated by FACS–FRET analysis using N- or C-terminus YFP-tagged CD9 (YFP–CD9 or CD9–YFP, respectively) and N- or C-terminus CFP-tagged hOCT2 (CFP–hOCT2 or hOCT2–CFP), which were transfected in HEK293 cells, respectively. CFP–hOCT2 and YFP–CD9 or CD9–YFP co-expression resulted in a significant FRET signal ( Fig. 5 ). These data suggest a physical interaction between CD9 and hOCT2, mainly localized near the N-terminus of hOCT2.
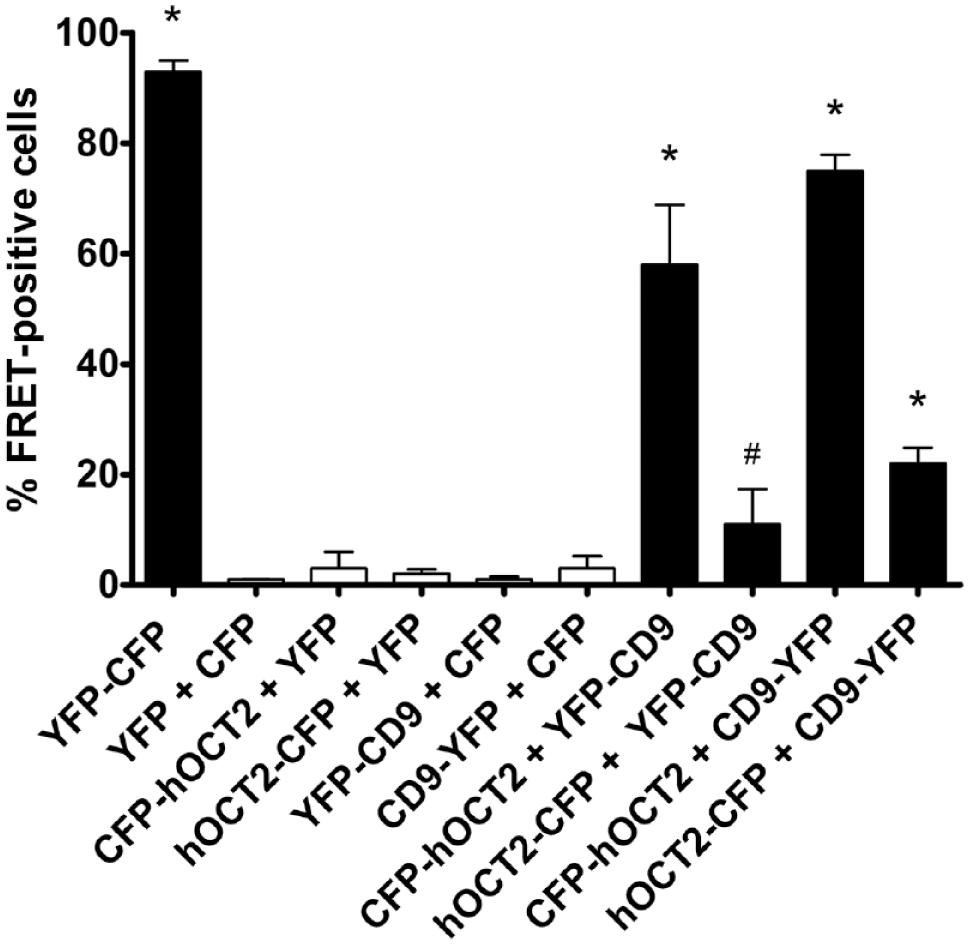
Identification of human organic cation transporter 2 (hOCT2) interaction domain with CD9 by fluorescence-activated cell sorting and Förster resonance energy transfer (FACS–FRET) in human embryonic kidney (HEK293) cells. HEK293 cells transfected with the cyan fluorescent protein–yellow fluorescent protein (CFP–YFP) fusion protein (YFP–CFP, positive control, black column); with CFP and YFP, CFP–hOCT2 or hOCT2–CFP and –YFP, and YFP–CD9 or CD9–YFP and –CFP (negative controls, white columns); and with CFP–hOCT2 or hOCT2–CFP and YFP–CD9 or CD9–YFP (black columns) were analyzed on a LSRII (BD Bioscience) flow cytometer. Shown are mean values ± standard deviation of at least four independent transfections. *Statistically significant differences for all other groups. #Statistically significant differences for all other groups except CFP–hOCT2 or hOCT2–CFP and –YFP, and YFP–CD9 or CD9–YFP and –CFP [analysis of variance (ANOVA) with Tukey posttest, P < 0.05].
Co-expression of CD9 and hOCT in Human Tissues
After having demonstrated that CD9 and hOCT can interact, we have investigated whether these proteins are expressed together in human tissues known to express hOCT1 and/or 2, such as epithelial nasal cells 27 and dorsal root ganglia, 28 respectively. In human epithelial nasal cells, CD9 and hOCT strongly co-localize in the plasma membrane ( Fig. 6A and B ). Since the OCT antibody we used cannot discriminate between OCT1 and 2 (our own data), and since these cells express both transporters, 27 we are not able to determine whether CD9 co-localizes with only one type of transporter here. In DRGs, for technical reasons, it is not possible to label for hOCT2 and CD9 at the same time, since both primary antibodies used for these experiments are from mouse. Therefore, we show two distinct pictures for hOCT2 and CD9 expression in this tissue. As evident from inspection of Figure 7 , OCT2 is expressed in satellite cells of DRGs, in which expression of CD9 also can be observed. Therefore, it can be concluded that CD9 and hOCT in some tissues are co-expressed, and probably in such tissues, there can be a direct interaction between these two proteins.
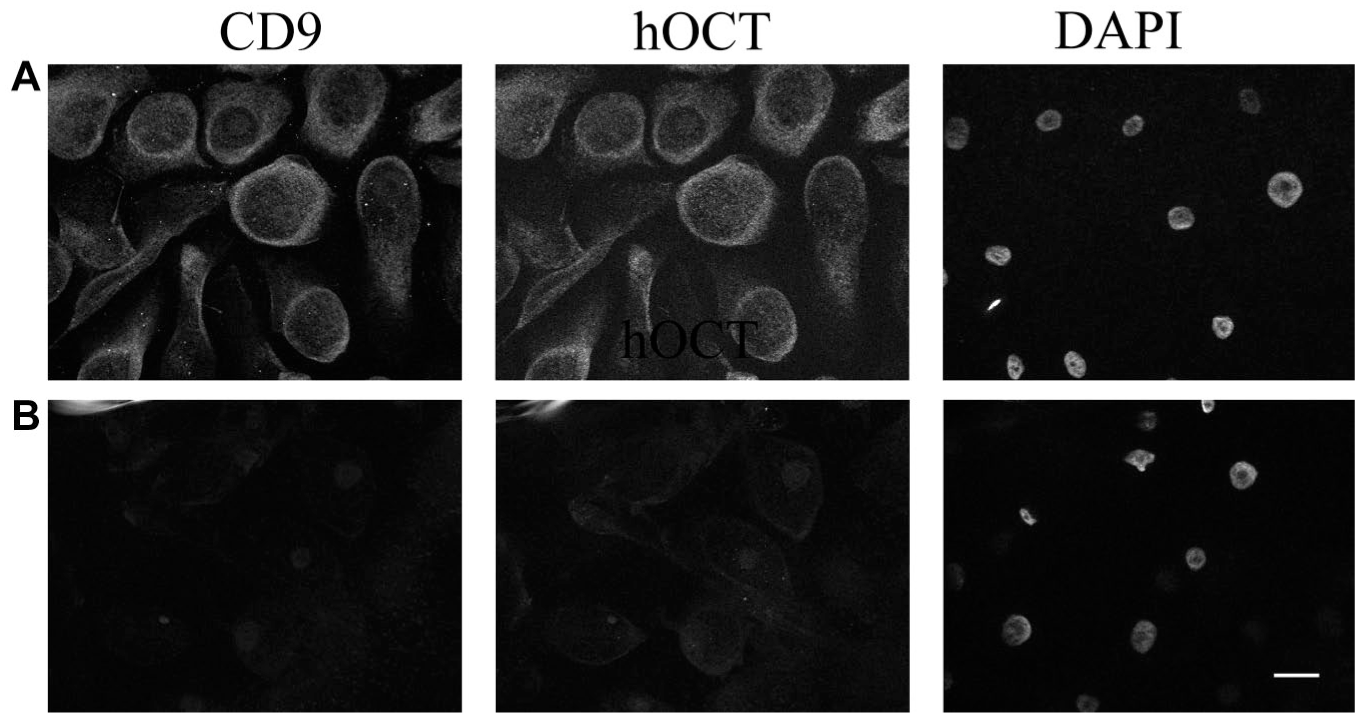
(
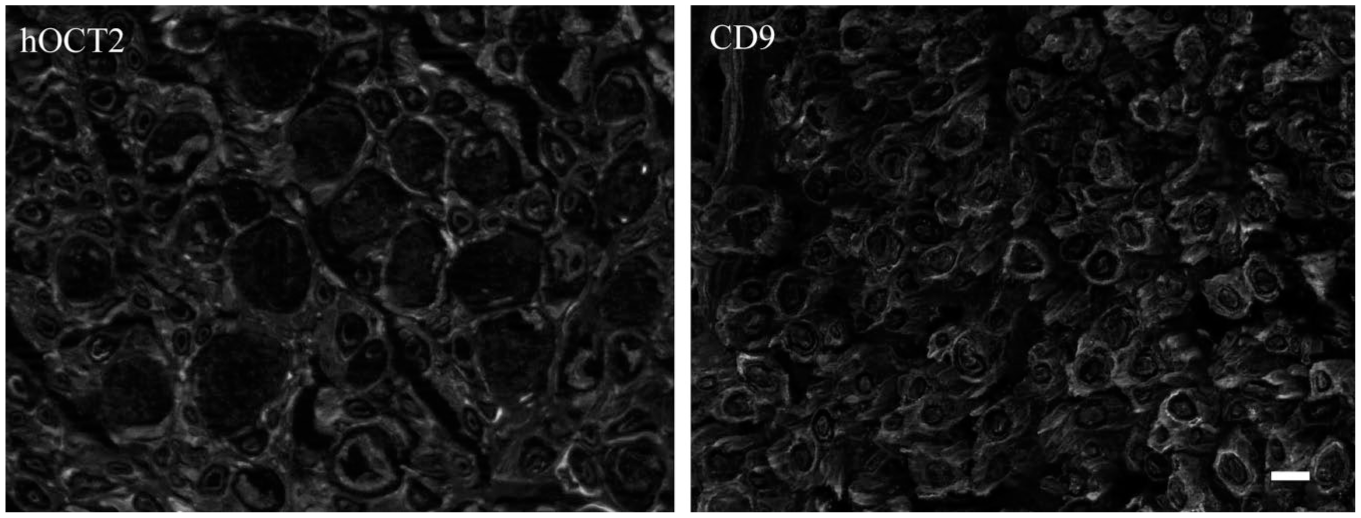
Representative labeling of human organic cation transporter 2 (hOCT2) and CD9 in human dorsal root ganglia. CD9 is expressed in satellite cells. hOCT2 exhibits a similar distribution, although at a lower intensity. Scale bar = 50 µm.
Discussion
Transport across the plasma membrane is essential for vital cellular processes, such as nutrient uptake, handling of waste products and toxic substances, and distribution of cellular products. 29 Transporters, the proteins that mediated these processes, are engaged in various interactions that contribute to establishing their localization, life span, and sensitivity to signals from second messenger systems. 30
Of special interest are PPIs, which have the potential to modulate protein function in a specific fashion. In this work, we have identified potential interaction partners of hOCT1 and 2, membrane transporters with important physiological and pharmacological roles, using the mbSUS, a yeast two-hybrid system that specifically detects interaction partners of membrane proteins in vivo in a eukaryotic organism. Using this system, we identified several potential interaction partners of hOCT1 and 2.
According to GOTERMMAPPER classification, 26 the common interaction partners of hOCT1 and 2 are involved in processes such as transport, signal transduction, development of anatomical structures, vesicle-mediated transport, response to stress, cell differentiation, biosynthetic processes, processes of the immune system, cellular protein modification proteins, cell adhesion, and membrane organization ( Fig. 1 ). Specific interaction partners for hOCT1 are associated with extracellular matrix organization (calpain 1 and osteopontin), generation of precursor metabolites and energy- or nucleobase-containing compound catabolic processes (fructose-biphosphate aldolase), and a mitotic cell cycle (mucin). Specific interaction partners for hOCT2 are associated with DNA metabolic processing (TREX1 and TMEM161A), messenger RNA (mRNA) processing (HNRNPH3 and PRPF8), protein folding (PDIA6), embryo development (FDZ5), vacuolar transport (STX8), cell division (TRAM1), and protein maturation (SPCS1).
Members of the hOCT interactome identified by the mbSUS screening are distributed along all cell structures: in the plasma membrane (e.g., CD9, CYSTM1, and PDZK1P1 for hOCT1; CD9, BRI3, FZD5, SLC40A1, and SLC47A2 for hOCT2), in the endoplasmic reticulum (e.g., KRTCAP2, SERP1, VAPB, and TMEM147 for hOCT1; CYB5R3, DPM2, PDIA6, SERP1, SPCS1, RPN2, TMEM41B, and VKORC1 for hOCT2), in the Golgi system (e.g., CHST12 and TMBIM4 for hOCT1; FZD5 and STX8 for hOCT2), in endosomes (e.g., CD63 and LAPTM4A for hOCT1 and hOCT2), in lysosomes (e.g., CD63 and LAPTM4A for hOCT1 and hOCT2), in mitochondria (e.g., FIS1, GHITM, and SLC25A11 for hOCT1 and CYB5R3 for hOCT2), and in peroxisomes (e.g., FIS1 for hOCT1 and hOCT2), suggesting that they are important for transporter cellular processing. Even though the mbSUS is an attractive screening method to identify interaction partners, the results obtained have to be carefully interpreted. Some interaction identified here is probably of minor physiological interest. For example, hOCTs are mainly expressed in polarized cells in a specific membrane domain (e.g., hOCT2 in the basolateral membrane of renal proximal tubules). In these cells, there is also a specific expression of SLC47A2 (MATE2), but in the apical plasma membrane domain. For this reason, an interaction between hOCT2 and MATE2 can happen in the unpolarized yeast system, but it is improbable in renal proximal tubules. The interaction between hOCTs and the potential partners identified by mbSUS should be confirmed by an independent method. Such investigations have been already performed for CD63 and LAPTM4A, two interaction partners for hOCT1 and 2, which have been identified here and in other works.18,19 LAPTM4A seems to be important in the regulation of OCTs by modulating their trafficking to and from the cell membrane and by recruiting it in an endocytotic compartment, where the OCT2–LAPTM4A complex is degraded. 19 The interaction of OCTs with the tetraspanin CD63 seems to be important for transporter membrane distribution. CD63 may play a role in the recycling of OCTs from endosomes to the basolateral membrane. 18 In this work, we further focused on CD9 as an interaction partner of hOCT1 and 2. By mating experiments and by FACS–FRET experiments in HEK cells, we were able to confirm the CD9–hOCT2 interaction. Together with CD63, CD9 belongs to the tetraspanin superfamily. Tetraspanins organize other proteins into a network of multimolecular membrane microdomains, the tetraspanin-enriched microdomains (TEMs) or “tetraspanin web.”18,31 In these TEMs, tetraspanins interact with many other proteins, such as integrins, regulatory proteins, and other transmembrane proteins, influencing signaling pathways, endocytosis, and trafficking processes.18,32–35 The ability of tetraspanins to manage interactions between other proteins and to generate functional complexes suggested the term “molecular facilitators.”18,32,36 The tetraspanins share a common structure with intracellular N- and C-termini, four transmembrane domains and two extracellular (EC) loops (a small EC1 with 13–31 aa and EC2 with 69–132 aa 37 ). Most tetraspanin–protein interactions involve EC2. 37 Together with other tetraspanins, CD9 is a component of extracellular vesicles such as exosomes, structures able to transfer a biological cargo to other sites. 38 CD9 is broadly expressed in several different cells and tissues. For this reason, CD9 can assume many different functional roles, depending on the cells in which it is expressed. 39 CD9 is important in a variety of cellular functions, such as proliferation, adhesion, migration, and signaling, and is involved in sperm–egg fusion, metastasis, and inflammatory reactions. 39 In animal models, CD9 deficiency decreases the metastatic potential of breast cancer cells and the severity of arthritis, but it produces lung inflammation. 39 In this work, we demonstrated that CD9 interacts with hOCT1 and 2, probably in close proximity to the N-terminus of the transporter. Since some tissue expresses CD9 and hOCT1 or 2, this interaction may have a regulatory function on transporter activity. Since hOCTs interact also with other tetraspanins, such as CD63, 18 there is a possibility that a CD9–hOCT–CD63 interaction complex is formed. The interactions of CD9 and CD63 with plasma membrane proteins seem to influence the movement of these proteins to and from specific domains of the plasma membrane: CD9 is known to be important for regulation of trafficking of plasma membrane proteins such as the major histocompatibility complex class II (MHC-II) in dendritic cells, 40 while CD63 is known to regulate the trafficking of hOCT2 to the basolateral membrane domain in renal proximal tubules. 18
The hOCTs are highly expressed in epithelial cells: The co-localization of CD9 and hOCT2 in membrane protrusion of MDCK cells suggests that they are co-expressed in digital junctions, which may represent a transitional contact mechanism between epithelial cells before adherens junctions appear and reinforce the cell–cell contacts. 41 The CD9–hOCT2 interaction also can probably take place in vivo in renal tissue, since we observed an expression of mRNA for CD9 in a preparation of human renal proximal tubules ( Suppl. Fig. S2 ), which are well-known to express hOCT2.18,19
Since transporter function can be regulated by different signaling pathways, 16 by development of cancer, 42 and by inflammatory processes, 43 all conditions in which CD9 can play some role, it can be speculated that CD9–hOCT interactions can have important implications for the physiological and pharmacological functions of these transporters. For example, it can change neurotransmitter concentration in synaptic clefts and change the pharmacokinetics of hepatic- and renal-secreted drugs. In future studies, it should be evaluated whether and how CD9 influences hOCT function.
In conclusion, in this article, we have characterized the protein interactome of hOCT1 and hOCT2, and, using the example of CD9, we have confirmed this interaction, which seems to happen also in human tissues expressing these proteins.
Supplemental Material
DS_DISC859837 – Supplemental material for Identification of the Tetraspanin CD9 as an Interaction Partner of Organic Cation Transporters 1 and 2
Supplemental material, DS_DISC859837 for Identification of the Tetraspanin CD9 as an Interaction Partner of Organic Cation Transporters 1 and 2 by Beatrice Snieder, Sabine Brast, Alexander Grabner, Sven Buchholz, Rita Schröter, Gilles A. Spoden, Luise Florin, Johanna Salomon, Tobias Albrecht, Vivien Barz, Alex Sparreboom and Giuliano Ciarimboli in SLAS Discovery
Footnotes
Acknowledgements
The authors are grateful to Astrid Dirks and Ute Neugebauer for excellent technical assistance.
Supplemental material is available online with this article.
Author contributions
Designed research: GC and SaB. Performed research: SaB, AG, SvB, LF, GAS, GC, BS, RS, and VB. Contributed new reagents or analytic tools: AS, GAS, LF, and TA. Analyzed data: SaB, GC, GAS, and LF. Wrote the article: GC.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Deutsche Forschungsgemeinschaft (CI 107/11-1 to GC, and FL 696/3-1 to LF).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
