Abstract
Transmembrane proteins function within a continuous layer of biologically relevant lipid molecules that stabilizes their structures and modulates their activities. Structures and interactions of biological membrane–protein complexes or “memteins” can now be elucidated using native nanodiscs made by poly(styrene co-maleic anhydride) derivatives. These linear polymers contain a series of hydrophobic and polar subunits that gently fragment membranes into water-soluble discs with diameters of 5–50 nm known as styrene maleic acid lipid particles (SMALPs). High-resolution structures of memteins that include endogenous lipid ligands and posttranslational modifications can be resolved without resorting to synthetic detergents or artificial lipids. The resulting ex situ structures better recapitulate the in vivo situation and can be visualized by methods including cryo-electron microscopy (cryoEM), electron paramagnetic resonance (EPR), mass spectrometry (MS), nuclear magnetic resonance (NMR) spectroscopy, small angle x-ray scattering (SAXS), and x-ray diffraction (XRD). Recent progress including 3D structures of biological bilayers illustrates how polymers and native nanodiscs expose previously inaccessible membrane assemblies at atomic resolution and suggest ways in which the SMALP system could be exploited for drug discovery.
Keywords
Introduction to Native Nanodiscs
Drug discovery increasingly relies on obtaining accurate information from atomic-resolution structures, screening campaigns, and ligand binding assays of a relevant target state. For membrane proteins this has not been achievable in general because the lipids are stripped off with detergents for screening assays and structure determination, but this compromises stability and relevancy to the target. Alternatively, targets are presented in cellular or vesicular forms that do not allow high-resolution structures to be determined. This presents an important problem that needs to be overcome as membrane proteins play critical roles in most disease pathways and drug resistance mechanisms and would benefit from easier structure-aided design.
A major development has paved the way to overcome this obstacle. It is now known that membrane proteins do not need to be divorced from their membrane microenvironment to be characterized by structural and functional assays. Instead, a native nanodisc comprising a section of biological membrane can be excised from cellular membrane fractions in an intact state. Second, there is a specialized group of amphipathic copolymers that are uniquely suited to this task ( Fig. 1 ). All have polar and apolar sidechains arranged along a linear sequence and can bind and insert nonspecifically into membranes before gently liberating their contents into discs with diameters between 5 and 50 nm. The universe of potential polymers is much larger than the protein universe and is less well understood or developed. Hence, the search is on for the best polymers and experimental approaches to understand and exploit the wide diversity of memteins, particularly those of therapeutic interest.
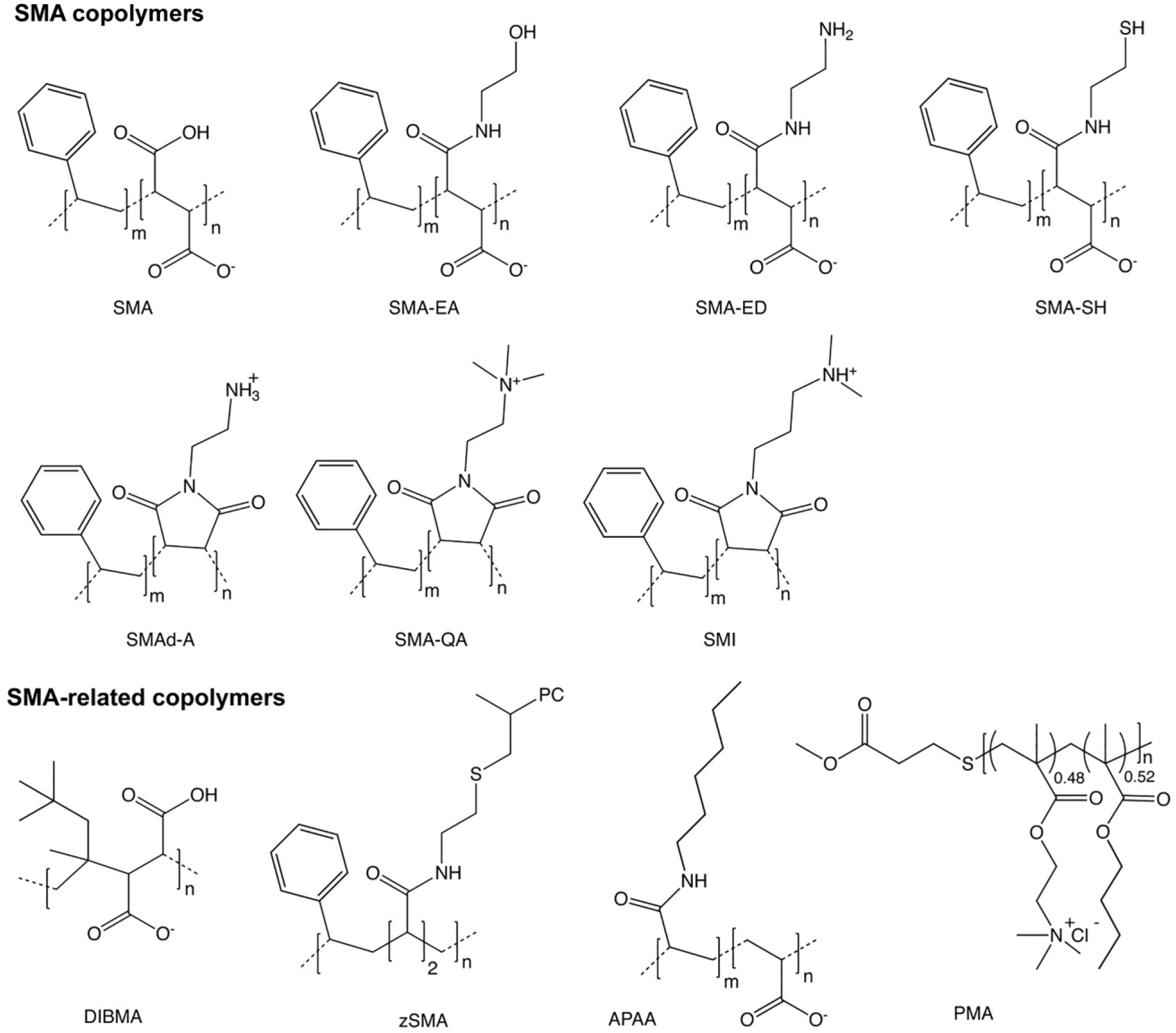
In order to help bring conformity to this rapidly expanding field, new scientific terms have been developed by the research community to describe core concepts. Native nanodisc refers to nanometer-scale discs that originate from biological as opposed to synthetic membranes. The newly introduced term memtein means the minimal unit of membrane structure and function that includes membrane protein bound to a continuous layer of biological lipid molecules, whether peripherally or integrally bound. 1 SMALP is an abbreviation for styrene maleic acid lipid particle, which is a bilayer-containing nanodisc that may or may not contain protein or native lipids. 2
Copolymers That Solubilize Memteins
Most of the copolymers used to solubilize membranes and form native nanodiscs contain nonalternating sequences of styrene (S) and maleic anhydride sidechains. The latter groups can be conveniently hydrolyzed to yield the activated form with charged maleic acid (MA) groups ( Fig. 1 ). 3 The ratio of S/MA groups is generally between 2:1 and 3:1 ( Table 1 ), as this provides an optimal degree of hydrophobicity for bilayer insertion and polarity for membrane solubilization. The polymer can be added to membrane fractions at a critical concentration of around 1%–2% w/v to spontaneously form nanodiscs. The discs are water soluble, nonviscous, and stable after prolonged storage or reconstitution from freeze-drying without compromising the activity of, for example, a G-protein-coupled receptor (GPCR). 4 Commercially available SMA reagents from Polyscope (Geleen, Netherlands) include XIRAN 25010 and 30010, which have S/MA ratios of 3:1 and 2.3:1, respectively. Total Cray Valley (USA) offered SMA2000 and SMA3000 copolymers that contain S/MA ratios of 2:1 and 3:1 (and are similar to Lipodisq reagents) until 2018 when their SMA portfolio was bought by Polyscope, which offers a full range of SMA products. Thus far, the SMA(2:1) type copolymers appear to be the most popular and have successfully solubilized bacterial, fungal, mammalian, and plant membranes, and yield 10 nm nanodiscs that include monomeric and multimeric proteins having between 1 and 48 transmembrane α-helices.5–7
SMA and SMA-Related Polymers Used to Prepare Nanodiscs.
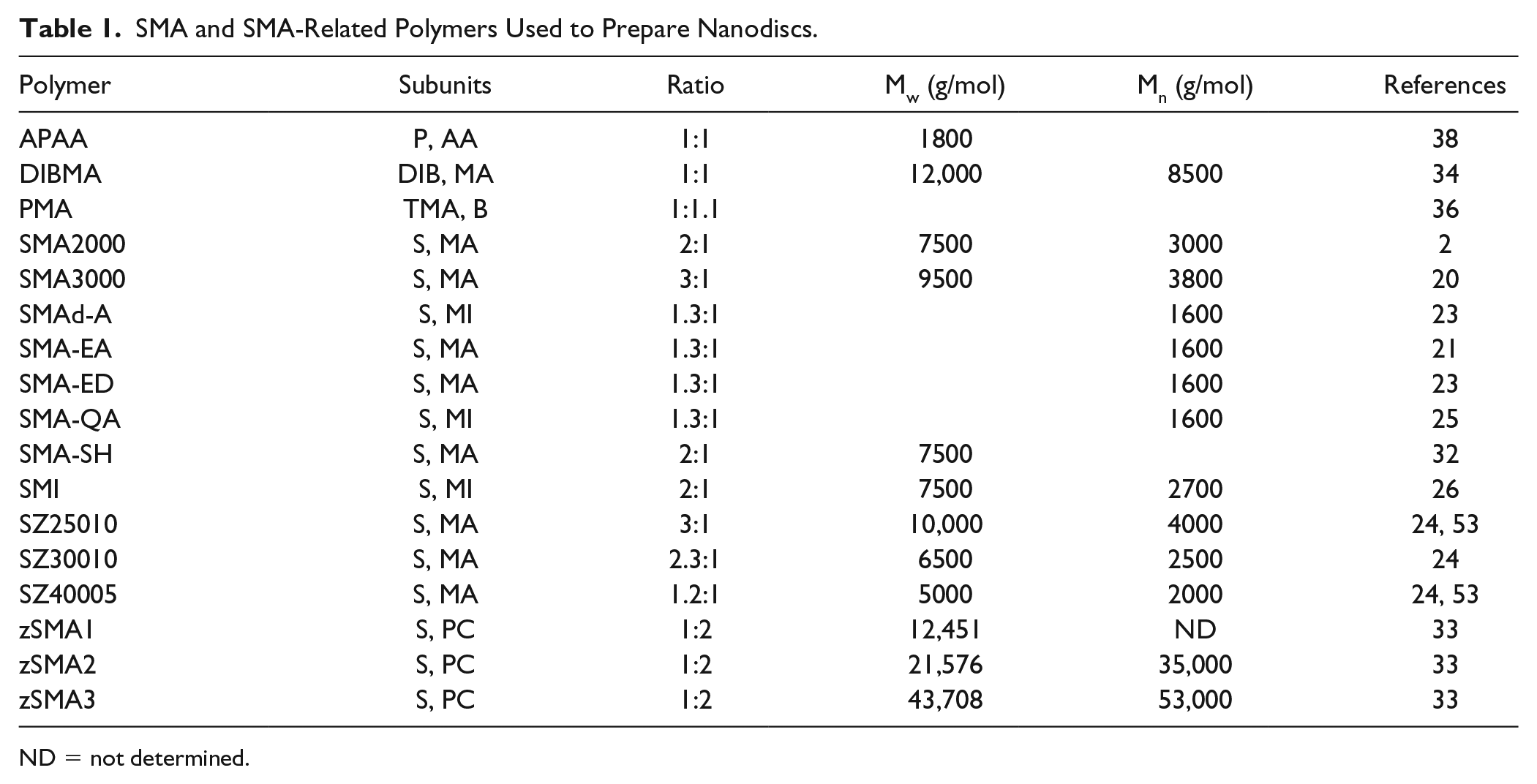
ND = not determined.
The types and concentrations of polymers as well as the buffer and temperatures used influence resultant disc sizes, as can be seen by small angle x-ray scattering (SAXS). 8 Comparative studies of different SMA types reveal that molecular weights in the range of 10 kDa are most effective at solubilizing the monomeric, dimeric, trimeric, and tetrameric states of the reaction center of Rhodobacter sphaeroides. 9 Large protein oligomers can be solubilized by the addition of longer SMA polymers or additional lipids to the membrane fraction to produce 50‒100 nm diameter discs. The polymer will incorporate virtually any molecule inside a biological membrane into a SMALP, and can adapt to accommodate large oblong assemblies. 7 Smaller discs with 6–10 nm diameters can be made using the SMA(3:1) polymer to solubilize liposomes. 10 The species of associated lipids are detectable by nuclear magnetic resonance (NMR) 2 or mass spectrometry (MS) methods.11,12 The lipid molecules can exchange between nanodiscs through collisions, 13 and new lipid or protein molecules can subsequently be incorporated into the disc or removed. 14 The heterogeneous polymer belt is rarely seen, but its envelope can be observed by small-angle neutron scattering (SANS) using hydrogenated and deuterated lipids. 15 The styrene rings pack against the lipid tails as seen by their intermolecular interactions in NMR spectra. 15 Most polymers used to make nanodiscs are around 5–10 kDa in size and exhibit sequence and length polydispersities that limit resolution of the interactions. This has stimulated the development of polymer synthesis approaches including reversible addition-fragmentation chain transfer to provide greater homogeneity.16,17
SMA polymers are distinguished from classical detergents by their mild solubilizing activity, which is essential for minimizing any perturbation of the memtein18,19 and allowing the bound lipid poses to be maintained.15,20 While some classical detergents may solubilize more total protein, the specific yield of biological lipid-bound multimers in SMALPs is higher. The optimal pH range at which the original SMA copolymers are most effective at solubilizing membranes is between 7 and 9, with an optimum around 8. The maleic anhydride group can be derivatized with an ethanolamine group to produce a SMA-EA copolymer that solubilizes liposomes over a broad pH range, and by varying temperature, salt, and cation concentrations it is possible to generate nanodiscs with diameters between 10 and 50 nm.21,22 The 15N-edited NMR spectra of cytochrome b5 protein in SMA-EA nanodiscs are well resolved and indicate a folded state. The addition of lanthanides including Yb3+ induces the alignment of large discs with external magnetic fields to allow tilt angles of transmembrane helices to be determined. The attachment of an ethylenediamine group to SMA yields a zwitterionic SMA-ED form that forms nanodiscs at pH values below 5 and over 7. 23 Once dehydrated the resultant SMAd-A copolymer can solubilize dimyristoyl phosphatidylcholine (DMPC) vesicles at pH 6 or below. Both ethylenediamine-derivatived SMAs tolerate high salt and divalent cation concentrations. Low salt levels can increase polymer solubility further, 24 although higher salt may be necessary for membrane solubilization in the presence of millimolar divalent cations. These cations otherwise bind the negatively charged MA groups and can lead to precipitation of polymer-cation complexes.
Positively charged derivatives of SMA are also able to solubilize membranes into discs. Modifying SMA by the addition of quaternary amide groups produces a cationic SMA-QA copolymer that effectively solubilizes vesicles into discs. Their diameters range from 10 to 30 nm and depend on the amount of added DMPC, pH values between 2 and 10, and high ionic levels. 25 The incorporation of a positively charged group in SMA copolymers allows nanodiscs to be made from membranes at pH values under 7.8 and in high divalent cation concentrations. 26 The resultant styrene maleimide (SMI) discs have ~6 nm diameters and maintain stability at 80 °C.
A ratio of 2:1 for S/MA ratios appears most effective at solubilizing as this perturbs the lipid bilayer least, 15 although other polymer types will behave differently. The ZipA cell division protein can be prepared using SMI, although at slightly reduced yields. 26 Solubilization of the adenosine A2A receptor using SMI does not compromise ligand binding activity, although some proteins can bind nonspecifically to polymers through electrostatic attractions and hydrophobic interactions. As a range of anionic SMA and cationic SMI copolymers are available, their potential influences on the activity of a specific target can be compared to minimize any nonspecific interactions.
The mild solubilization of membranes by SMA in a lipid aspecific manner is due to the nonselective insertion of styrene groups into the bilayer. 2 However, there is a preference for bilayers that possess some disorder and short or unsaturated acyl chains, 27 with ordered membranes being more difficult to solubilize. 26 Both plasma and intracellular membranes in live cells are solubilized in sequential order by SMA molecules that progressively penetrate each bilayer. 28 The outer perimeter is breached first and then the inner membranes based on the stepwise release of fluorescent proteins.
Lipid rafts can be turned into nanodiscs using SMA for protein analysis. For example, T-cell membranes are fragmented by adding SMA(3:1) and can then be immunoprecipitated to isolate complexes of glycosylphosphatidylinositol-anchored proteins and Src family kinases, as well as ordered lipids including cholesterol, phosphatidylserines, sphingomyelins, ceramides, and monohexosylceramides. 29 Whether the positions of these ordered lipids or other lipids such as phosphatidylinositol, phosphatidylglycerol (PG), phosphatidylcholine (PC), and phosphatidylethanolamine (PE) can be determined remains to be seen, but is likely to regulate associated receptors.
The length and concentration of copolymer as well as temperature changes or lipid addition provide ways to toggle which membranes are incorporated into SMALPs. 2 This control could allow organelle-specific memteins to be collected from cells or membrane mixtures. 30 Comparative analysis of 12 SMA copolymers and conditions using the photosynthetic assembly from spinach thylakoid membranes reveals those that generate discs with native activity and subunit composition. In this case, the most efficient polymers include SMA1440, XIRAN 25010, XIRAN 30010, SMA PRO10235, and SMA1735. Ways to simplify the separation and detection of the assemblies include variations in buffer conditions and the removal of more loosely bound peripheral membrane proteins. 31
Functional derivatives of SMA can be synthesized by attaching a thiol-containing sidechain to the MA groups, as demonstrated with the SMA-SH polymer. This contains an amino-mercapto-ethane group to which fluorescent dyes or molecular tags can then be conjugated 32 to provide tags for affinity purification or detection by fluorescence methods. Such derivatives offer promising strategies for purifying native complexes from endogenous sources of membranes. Incorporation of a PC headgroup within a zwitterionic “zSMA” polymer 33 allows for the solubilization of membranes into 10–30 nm diameter discs. The lack of carboxylic acids in zSMA lends itself to a wider range of solution conditions, including low pH and high concentrations of cations that would otherwise bind the acidic groups.
Like SMA, a related copolymer that contains diisobutylene and MA residues also solubilizes membranes directly and is referred to as DIBMA. The replacement of the styrene group with an aliphatic moiety removes the ultraviolet (UV) light-absorbing signal and is compatible with UV detection of bound protein by circular dichroism methods. DIBMA allows particularly gentle disruption of membranes to yield large and labile memteins within 12–29 nm discs. 34 The association of DIBMA with millimolar concentrations of divalent cations enhances solubilization of anionic membranes and reduces nanodisc size. 35 A set of polymethacrylate (PMA) copolymers have been designed to include a distinct backbone with various ratios of hydrophobic and cationic sidechains and also lack a UV signal. 36 Treatment with PMA produces DMPC discs with 17 nm diameters and fragments bacterial membranes into particles with diameters of about 100 nm. 37 Further variations of the hydrophobic group yielded a series of alkyl polyacrylic acid (APAA) polymers. 38 Derivatization of this polymer with butyl, pentyl, or hexyl groups and adjusting its concentration induces disc diameters of between 7 and 17 nm. The form with the longest acyl chain yields the most total membrane protein in this case. Such studies can be used to derive structure–activity relationships for solubilization of memteins and could inform the rational design of tailored copolymers for solubilizing specific target states or extremely large or labile complexes.
Structures of Memteins
Dozens of different membrane proteins have been solubilized from bacterial, yeast, insect, and mammalian cells using SMA for structural and biophysical analysis over the past decade. The recent high-resolution structures of several memteins reveal the positions not only of contacts between protein subunits but also of bilayers of lipid and interdigitating posttranslational modifications, representing a major step forward.
Cryo-Electron Microscopy Developments
The multidrug exporter AcrB was solubilized by a homogenous SMA(2:1) preparation with minimal monomers present for resolution of its membrane-bound conformation at 3.0 Å resolution. 39 The structure displays the 12 nm diameter structure of the asymmetric trimer as it sits in the Escherichia coli membrane. The SMALPs containing His-tagged wild-type AcrB protein could be purified in a single step over a Ni-affinity column before being applied to grids for cryo-electron microscopy (cryoEM) data collection. Included in the structure are 31 lipid molecules that decorate the trimeric protein and form a bilayer organized in hexagonal patterns of lipids in the inner leaflet and ordered disarray in the outer leaflet ( Fig. 2 ). The polymer groups and lipid headgroup identities cannot be discerned, although more homogeneous polymers may address this limitation.
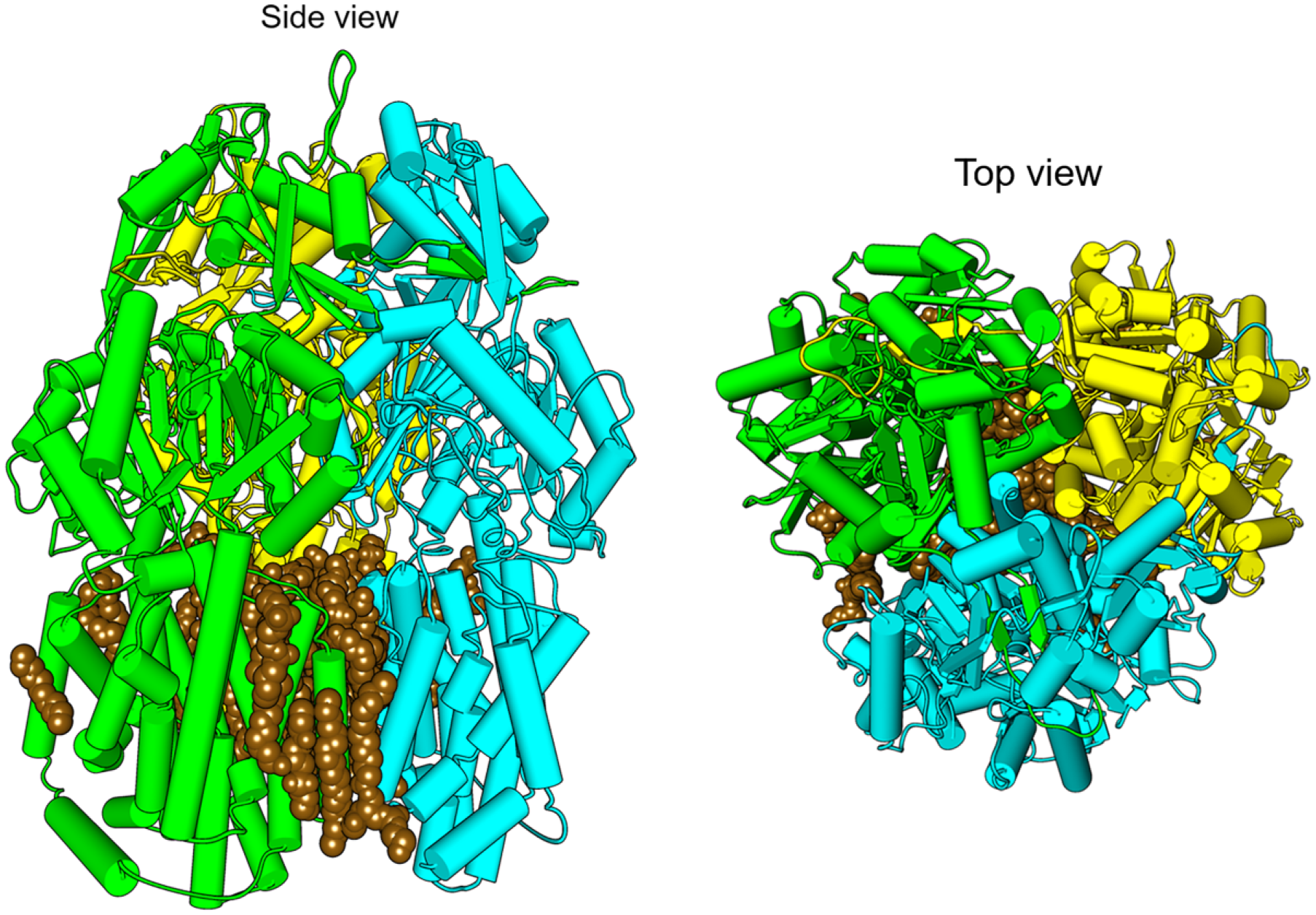
The cryoEM structure of trimeric wild-type AcrB. The protein solubilized intact with SMA(2:1) retains its membrane microenvironment, as seen from the top and side perspectives, with subunits colored yellow, green, and blue, with 24 hexagonally arrayed phospholipids in the central bilayer colored brown in native (EMD-7074, PDB ID: 6BAJ). 39
The structure of the Alternative Complex III (ACIII) memtein in a native nanodisc was determined at 3.4 Å resolution. 7 This could be accomplished using either 1% SMA3000 or XIRAN 25010 copolymer to solubilize the 464 kDa complex from Flavobacterium johnsoniae. After a 1 h incubation the insoluble material could be removed by high-speed centrifugation and the complex could be purified over Ni-NTA metal affinity resin, to which it binds inherently. After this point no detergent or SMA was needed, and the protein was ready for downstream analysis. The SMALP is able to accommodate the 9 × 13 nm supercomplex of six protein subunits including 48 transmembrane helices and accessory cytochrome c oxidase. The structure displays the poses of acylated residues responsible for photosynthetic electron transport, heme groups, iron–sulfur clusters, and biological phospholipids ( Fig. 3 ). These studies demonstrate the potential of the SMA technology for structural biological studies of intact membranes and illustrate its applicability to resolving diverse protein and lipid assemblies.
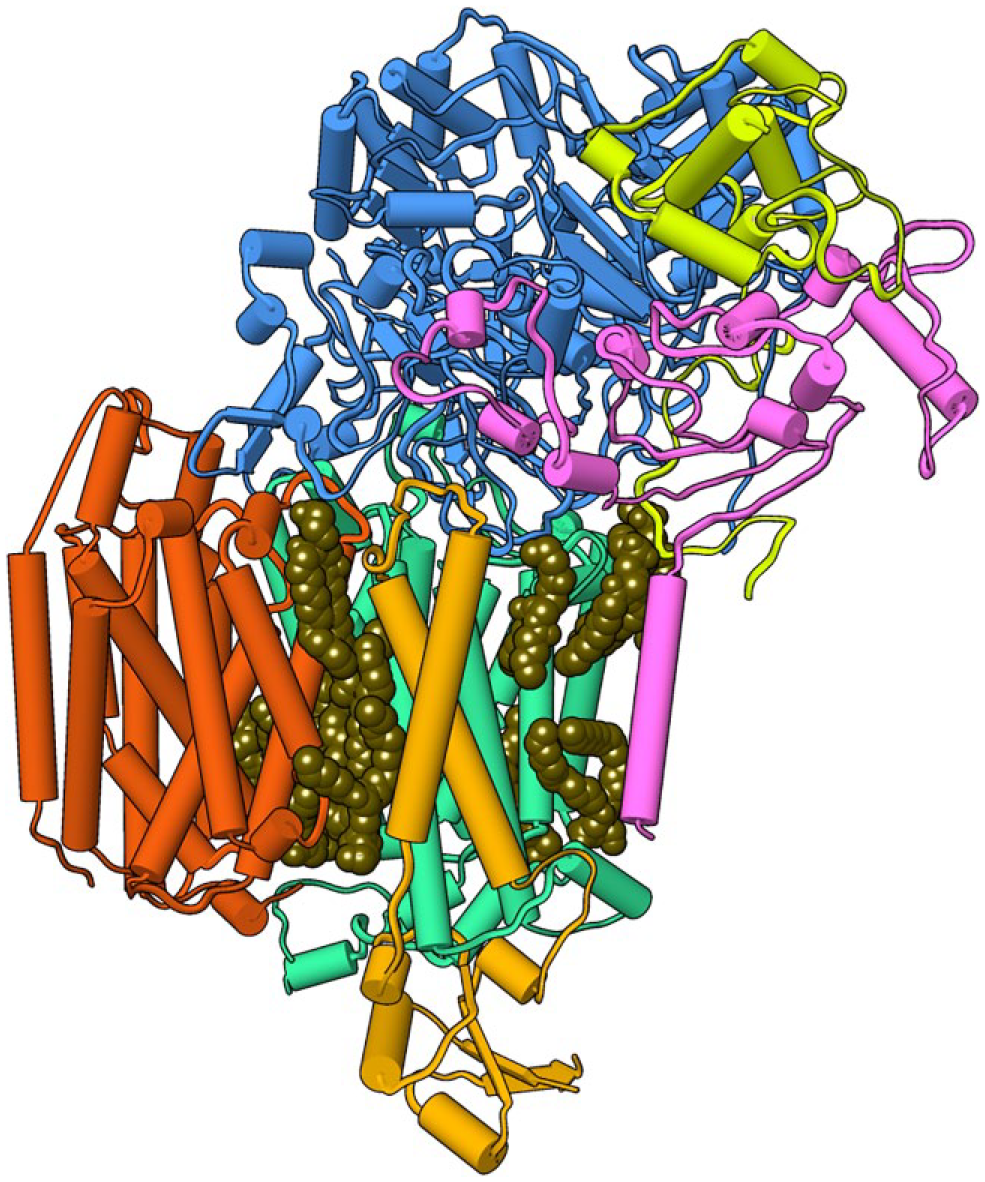
The cryoEM structure of the ACIII photosystem complex after solubilization with SMA(2:1) polymer. The color-coded ActA (purple), ActB (blue), ActC (magenta), ActD (orange), ActE (yellow), and ActF (red) subunits and phospholipids (space filling in brown) are shown (EMD-7286, EMD-7448). 7
X-Ray Diffraction and GPCR Multimers Bound to Lipid
The GPCR superfamily contains many potential targets that have proven difficult to produce in sufficient amounts of stable functionally intact states. Progress here has been demonstrated by the determination of a 2.0 Å resolution structure of a microbial rhodopsin after solubilization with SMA(3:1) copolymer ( Fig. 4 ). 14 The seven transmembrane helices form the expected trimer that binds transretinal as well as a set of the monoolein molecules used for in meso crystallization, although no biological lipids are bound tightly enough to be retained. The positions of such lipids remain of interest as they are known to help self-assemble rhodopsin into trimers and contribute to the protein’s structure, flexibility, and photocycle. Subsequent studies reveal that a heterotetrameric GPCR complex that recognizes the growth hormone secretagogue and dopamine binds to G proteins after solubilization with SMA (2:1) treatment. 40 Similarly, the adenosine receptor expressed in Pichia pastoris or HEK 293T cells maintains stability and ligand binding after solubilization with SMA(2:1) even after storage and freeze-thawing. 4 The melatonin and ghrelin receptors can be solubilized with SMA(2:1) or SMA(3:1) to form 13 nm diameter nanodiscs that retain function. 41 The cannabinoid receptor 1 solubilized with SMA(2:1) is recognized by conformationally specific antibodies in surface plasmon resonance (SPR) experiments and retains stability and activity in fluorescence-activated cell sorting (FACS) even after repeated freeze-thaws. 42 The preparations can be used for library screening to find variants with improved binding affinity or stability, indicating utility in high-throughput screening campaigns.
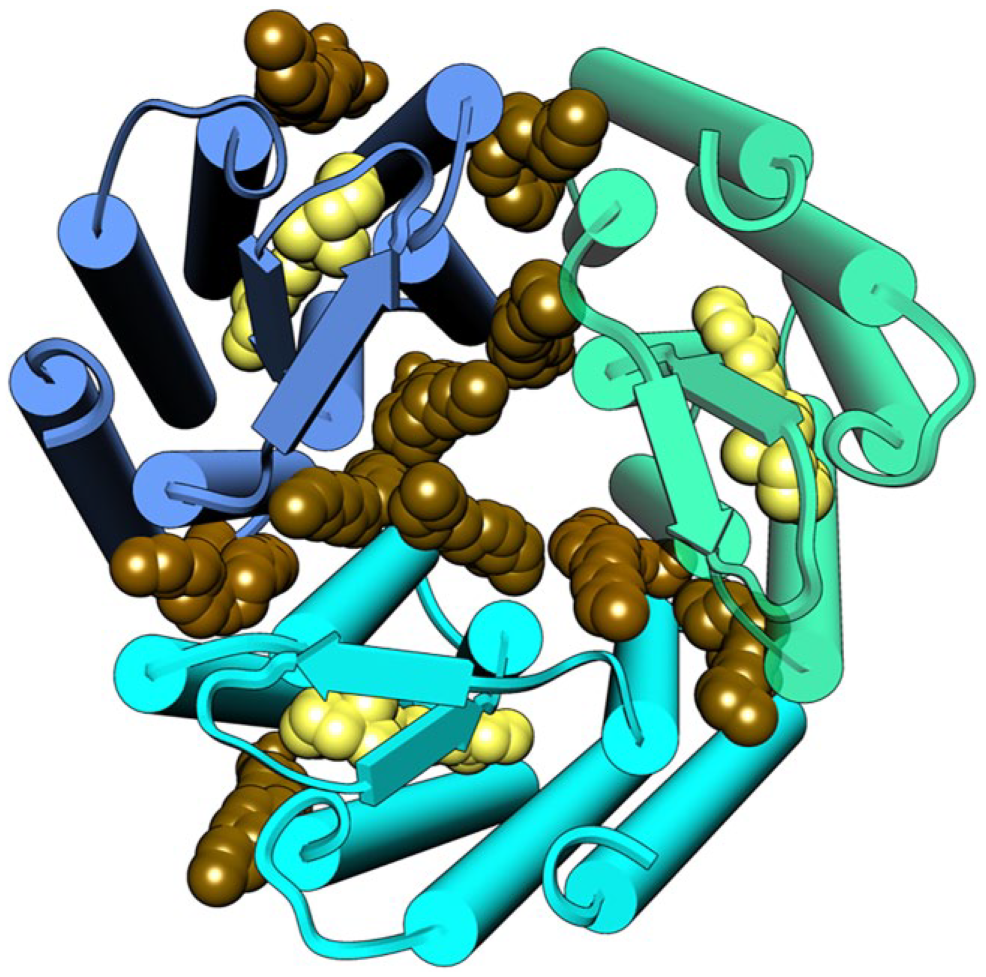
The in meso crystal structure of a rhodopsin trimer bound to monoolein after solubilization with SMA. The three subunits of the trimer (green, aqua, and blue) are shown with nine monoolein lipids (brown) within the interfaces and three retinal molecules (yellow) (PDB ID: 5ITC). 14
EPR and NMR Analysis of SMALPs
The structures, dynamics, and interactions of SMALPs have been resolved by electron paramagnetic resonance (EPR) and NMR spectroscopies.1,20 The coat protein of the Pf1 bacteriophage can be solubilized by SMA(3:1) to reveal stable helices in magnetically oriented 30 nm discs. 43 Their NMR signals are sharper than those found in bicelles or peptide-based nanodiscs. The cytochrome b5 protein solubilized into 50 nm diameter SMA-EA-based discs magnetically aligns and displays a stable transmembrane helical structure. 21 The zinc diffusion facilitator CzcD in 10‒15 nm discs formed by XIRAN 25010 or 30010 polymers retains an annulus of associated lipid 44 and exhibits resolvable amide and methyl NMR peak groups that can then be assigned. The elucidation of novel structures of proteins in nanodiscs by NMR and XRD remains challenging due to the polydispersity of the polymers, prompting the search for more regular polymer configurations.
Mass Spectrometry of Memteins
The detection of lipid species in SMALPs is best performed by MS, as shown by the study of the human equilibrative nucleoside transporter-1 (hENT1) extracted XIRAN 30010 and cholesteryl hemisuccinate from insect cell membranes. Electrospray ionization (ESI) MS data show that the functionally intact state retains 16 PC and 2 PE molecules but not polyunsaturated lipids. 11 Different types of phospholipids associate with each of three different bacterial membrane proteins, ZipA, PgpB, and FtsA, based on liquid chromatography (LC)-MS/MS analysis. 45 The rhomboid protease GlpG solubilizes similarly with DIBMA, SMA(3:1), and SMA(2.3:1) and retains about 50 phospholipid molecules of various types, thus preserving native-like activities and stabilities as well as showing suitability for ligand screening.12,46 The polydispersity and nonselectivity of SMA copolymers may allow for the retention of a broad range of lipid and protein types but also presents technical issues, including peak broadening. High voltages may be needed to release strongly held components within nanodiscs. The development of laser-induced liquid bead ion desorption (LILBID) MS approaches including soft and hard laser pulses allows for the detection of memteins that can be progressively dissociated 47 and could be used resolve novel multimeric states and their compositions.
Modeling Native Nanodiscs
Models are emerging that illustrate the complexities of membrane interactions of SMA polymers and peripheral membrane proteins. The polymer–lipid events can be seen in trajectories during molecular dynamics simulations, as illustrated by the process of SMA(2:1) and SMA(3:1) molecules binding, destabilizing, and perforating a DMPC bilayer model as they extract lipids into nascent discs. 48 The interactions of KRAS, a small GTPase, with a bilayer of PC and phosphatidylserine molecules reveals the presence of three major orientations on the bilayer differentiated by conformational changes in flexible elements. 49 Single-molecule fluorescence resonance energy transfer (FRET) measurements in SMALPs taken from mammalian cell membranes indicate three matching conformational states. The convergence of the simulated and experimental data suggests that multiple signaling states form membranes and are separated by small energy barriers. The oncogenic form with a G12V mutation appears to favor a state with an accessible effector binding loop, suggesting an oncogenic mechanism that is altered by mutation or stabilized by specific interactions with lipids.
Future Directions for Membrane Proteins
Many targets remain to be resolved. Full-length ion channels solubilize into nanodiscs upon SMA treatment to display symmetric tetramers arranged around central pores. 50 The hKCNH5 and hKCNQ1 α-subunits of the neuronal and cardiac voltage-gated potassium (Kv) channels as well as the regulatory KCNE1 β-subunit can be transferred from mammalian cells into SMALPs and exhibit improved behavior over detergent solubilized forms, including greater stability and ease of concentration. The channels appear to arrange themselves randomly on electron microscopy grids to expose diameters of 12‒15 nm. The exposed sidechains of spin-labeled KCNE1 protein in SMA(3:1) nanodiscs appear to be more dynamic than those that are held inside the membrane based on EPR studies. 51 Moreover, SMA-based solubilization increases the signal to noise and phase memory time in double electron–electron resonance spectra, thus improving distance information.
Tetraspanins constitute a challenging set of targets due to their multimeric networks, glycosylation and palmitoylation patterns, and small soluble domains that are stabilized by disulfides. A survey of the feasibility of purifying 15 tetraspanins expressed in Saccharomyces cerevisiae into SMALPs found that 5 could be obtained with high yields by solubilization in three SMA types. although others were less fruitful, 52 highlighting the need to develop tools for efficient solubilization and purification of oligomeric networks of transmembrane proteins that express at low levels.
The plant metabolon responsible for the biosynthesis of the glucoside dhurrin can be placed in SMALPs. Its subunits form a supercomplex in microsome membranes that can be purified in an intact state with multiple copies of three transmembrane proteins and a soluble glycotransferase protein. 53 This use of SMA(2:1) results in discs with diameters of 10‒25 nm, while conventional detergents destroy the integrity of the supercomplex that binds to and is modulated by interactions with its soluble partner and negatively charged phospholipids. These and many other molecular machines that operate within membranes have yet to be resolved at atomic resolution, justifying efforts to continue to improve amphipathic polymer formulations and ensure compatibility with structural and biophysical tools to characterize memteins and their mechanisms.
In conclusion, a wealth of information about the production of native nanodiscs using SMA-related polymers has been generated by more than 150 published studies by the SMALP network, paving the way forward for drug discovery applications. An expanding range of polymer types and methods have been optimized for solubilizing transmembrane targets from virtually any cell type to stabilize their structures and retain cellular lipids that could modulate function.54 Importantly, the SMALP system allows the preparation of endogenous heteromultimeric protein–lipid complexes that are too labile to survive detergent extraction or that cannot be assembled from recombinant sources. Challenges remain, including the isolation of endogenous complexes that are of very low abundance and those that are transiently bound to soluble regulatory and effector proteins. There are technical issues that require optimization, and not all memteins behave well in SMALPs, although improved polymers are in development. These will address the realities that some polymers can exhibit nonspecific associations with proteins, cations, and biosensor surfaces, and these interactions can affect performance in some assays. Although polydispersity is useful for solubilizing heterogeneous or undefined membrane compositions, it also poses challenges for obtaining high-resolution data of monodispersed states. Disc sizes can also be variable, and the preparation of uniform samples for structure–function studies is desirable and increasingly feasible. Polymers with novel tags that allow endogenous memteins to be conveniently purified and visualized are being designed. Measuring the concentration of polymers in solution can be accomplished with fluorescent tags that allow interactions to be detected. 41 Such tags will likely prove useful in ligand screening assays and a wider array of biophysical and structural methods and tools for proteomics and metabolomics. 55 Those methods already demonstrated to be compatible with the SMALP system include circular dichroism, cryoEM, FACS, EPR, MS, NMR, SANS, size exclusion chromatography, SPR, and XRD techniques, as well as commonly used purification systems. A growing list of proteins, which increasingly includes drug targets, have been incorporated into nanodiscs. The development of improved polymers for native nanodisc formation will facilitate increased resolution and higher-throughput screening for ligands by methods including NMR. 56 This progress is intended to enable the discovery of novel ligands and effective lead molecules for native-state membrane protein targets. Partnerships between academia and industry will bring this next generation of polymers and allied assay systems for native nanodiscs into the mainstream of drug discovery.
Footnotes
Acknowledgements
The authors thank Stefan Scheidelaar, PhD and Polyscope Polymers (Geleen, The Netherlands) for sharing polymers and methods.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This was funded by the Campus Alberta Innovation Program (RCP-12-002C), Alberta Prion Research Institute/Alberta Innovates Bio Solutions (201600018), and NSERC (RGPIN-2018-04994) grants awarded to M.O. and the Canada Foundation for Innovation and Genome Canada, which support The Metabolomics Innovation Centre (TMIC).
