Abstract
Hybridoma screening is a key step for the successful generation of high-affinity analyte-specific monoclonal antibodies (MAbs). This work presents an innovative screening method, known as a bacterial macroarray, generated by contact printing of hybridoma cell supernatant samples on a nitrocellulose (NC) membrane initially coated with fluorescein isothiocyanate (FITC)–labeled bacteria. Given that bacterial fixation will be influenced by complex bacterial surface structures, we selected both gram-positive bacteria (Staphylococcus aureus and Listeria monocytogenes) and gram-negative bacteria (Escherichia coli O157:H7 and Cronobacter sakazakii) to optimize the fixation conditions for binding to the NC membrane, such as the aperture of the NC membrane, the concentration of bacteria, the dosage of glycerin in the spotting buffer, and the fixation time and temperature. As a result, we found that a better bacterial macroarray could be developed when the spotting buffer, containing 1011 CFU mL−1 of FITC-labeled bacteria and 15% (V/V) glycerol, was spotted onto a 0.45 µm NC membrane with an incubation of 2 h at 37 °C. Finally, we verified the stability and specificity of the prepared bacterial macroarray by detecting cell cultures with the addition of two MAbs (Escherichia coli O157:H7 MAb E7 and Cronobacter sakazakii MAb 1E9) to simulate the screening experiments. Here, we describe a bacterial macroarray to efficiently screen the targeted antibody-secreted hybridomas.
Introduction
Monoclonal antibodies (MAbs), which have played an indispensable role in clinics and food security 1 in recent years, are now applied as targeted therapies for malignancies, transplant rejection, and autoimmune and infectious diseases, as well as a range of new indications. 2 Therapeutic MAbs and antibody-related products, such as Fc-fusion proteins, antibody fragments, and antibody-drug conjugates (collectively known as monoclonal antibody products), have grown to be the dominant product class within the biopharmaceutical market. 3 . Despite the use of various techniques to produce MAbs, such as chimeric MAbs, 4 transgenic mice,5,6 phage displays, ribosome displays, 7 and covalent displays, 8 the hybridoma technology 9 proposed by G. Kohler of West Germany and C. Milstein of Argentina 10 is the most universal. Hybridoma technology, a method of fusing B cells removed from the spleen of a relevant antigen-challenged animal and SP2/0 myeloma tumor cells, can be used to obtain hybridoma cell lines producing analyte-specific antibodies after screening by the antigen. Hybridoma screening is a crucial step for the successful generation of high-affinity analyte-specific MAbs because the growing of effective monoclonal hybridoma cell strains may be suppressed by other cells in the same well. 11 An indirect enzyme-linked immunosorbent assay (ELISA) method, based on measurement of optical density generated in the chromogenic reaction between an enzyme substrate and an enzyme conjugated with a protein that binds the analyte (antigen or antibody of interest), is commonly used for the screening. However, ELISA is time-consuming and limited to the laboratory for a complete analysis. 12 Thus, rapid and highly sensitive detection methods are needed. With the development of the screening technique of hybridoma cells, many new methods have been developed, including biosensors and droplet-based microfluidics. Lad et al. 13 applied biolayer interferometry to screen hybridoma clones based on disassociation rates using the Octet RED384 platform; this method could identify clones producing high-affinity antibodies missed by ELISA. El Debs et al. 14 used droplet-based microfluidics allowing the functional screening of up to 300,000 individual hybridoma cell clones in less than a day. Nevertheless, these methods have not been widely adopted because of the dependence on expensive equipment and professional operators. A bacterial macroarray, an economical and fast screening technique of hybridoma cells, could displace time-consuming ELISA and other costly techniques.
Being high throughput, rapid, automated, economical, convenient to operate, and highly sensitive, macroarray technology has been rapidly gaining widespread use since the 1980s, including for DNA microarrays,15,16 protein macroarrays17,18 (protein microarrays offer a fascinating possibility to study protein interaction in a massively parallel fashion, including protein-protein,19,20 enzyme-substrate,21,22 protein-DNA, 23 protein-oligosaccharide, or protein-drug interactions 24 ), organize chip, 25 and bacterial macroarrays. A bacterial macroarray, a carrier fixed with whole bacteria for the rapid screening of targeted antibody-secreted hybridomas, will be effective and convenient in the preparation of MAbs. De Marni et al. 26 spotted bacteria exposed to recombinant antibodies on their external surface directly on nanostructured-TiO2 or epoxy slides (purification-independent microarray) for antigen detection; the limit of detection (LOD) of this system for the antigen was 0.4 ng mL−1. The surface of the bacteria is composed of outer membrane proteins, lipopolysaccharides, flagella, capsules, fimbriae, and so on. Compared with antibodies, proteins, amino acids, and other small molecular substances, bacterial fixation on the substrate carrier will be more difficult because of the complex bacterial surface structure and relatively large size. It is generally known that glycerin can benefit the fixation of bacteria on nitrocellulose (NC) membranes. Moreover, the addition of glycerin in the spotting buffer can maintain the complete structure of the protein, retaining the activity of bacterial surface proteins by inhibiting cell dehydration.27,28 Thirumalapura et al. 29 added glycerin to the spotting buffer and developed a bacterial chip for the detection of antibodies produced by bacterial surface antigens with a detection sensitivity of 0.1 µg mL−1. However, the application of bacterial macroarrays in the screening of hybridomas has received little attention thus far.
In this study, we developed a bacterial macroarray for the rapid screening of high-affinity analyte-specific MAbs secreted by a hybridoma cell strain. Bacteria were labeled with fluorescein isothiocyanate (FITC) to produce visible bacterial spots. Thus, we can control and optimize the experiment process rapidly. Moreover, we optimized the fixation conditions of bacteria on an NC membrane, including the aperture of the NC membrane, the concentration of bacteria, the dosage of glycerin in the spotting buffer, and the fixation time and temperature. Finally, we simulated the screening experiments by adding two MAbs (Escherichia coli O157:H7 MAb E7 and Cronobacter sakazakii MAb 1E9) to the cell culture to verify the sensitivity and specificity of this bacterial macroarray. In summary, a method that can be used to rapidly screen hybridoma cell strains secreting high-affinity analyte-specific MAbs was developed.
Material and Methods
Bacterial Strains and Cultivations
The pathogenic bacterial strains used to develop the bacterial macroarray in this study are listed in
FITC Labeling
Bacterial strains listed in
Preparation of Bacterial Macroarray
All foodborne strains (
Optimization of Bacterial Macroarray
Selection of NC Membrane for Bacterial Macroarray
The specific surface area and porosity of the NC membrane will affect the adsorption to bacteria and possibly alter the detection performance, so the assessment of pore sizes of NC membranes is crucial. 31 In this work, FITC-labeled Escherichia coli O157:H7 (E. coli O157:H7), Cronobacter sakazakii (C. sakazakii), Staphylococcus aureus (S. aureus), 32 and Listeria monocytogenes (L. monocytogenes) 33 were fixed on NC membranes of different pore sizes (0.22 µm, 0.45 µm, 0.8 µm, and 5.0 µm). Different surface structures of these NC membranes was observed under the scanning electron microscope (Philips XL30, Netherlands). To find the optimal pore size of NC membranes for different bacterial strains, all strains were spotted on NC membranes with three biological replicates.
Optimization of the Concentration of the FITC and FITC-Labeled Bacteria
To find the optimal concentration of FITC to label bacteria, a series of final concentrations (0.1, 1, 10, 50, 100, and 400 µg mL−1) of FITC were used to label the bacteria; the fluorescence intensity of labeled bacteria was measured by a multifunctional microplate reader (SpectraMax/n2, Molecular Devices, Sunnyvale, CA). Taking the difference of the surface structure between gram-negative bacteria and gram-positive bacteria into account, two gram-negative bacteria strains (E. coli O157:H7 and C. sakazakii) and two gram-positive bacteria strains (S. aureus and L. monocytogenes) were selected to evaluate the optimal concentration of FITC by flow cytometry.
FITC-labeled E. coli O157:H7 and FITC-labeledS. aureus, adjusted to 108 CFU mL−1, 109 CFU mL−1, 1010 CFU mL−1, and 1011 CFU mL−1, were spotted on the NC membranes and incubated at 37 °C for 2 h. The images of the spotted bacterial macroarray were captured under an ultraviolet lamp after washing twice with PBST (PBS with 0.25% Tween-20, pH 7.4).
Selection of Optimal Fixation Time and Temperature and the Dosage of Glycerin
FITC-labeled E. coli O157:H7 and FITC-labeled S. aureus, resuspended with spotting buffer containing a series of concentrations (V/V) of glycerin (0, 5%, 10%, 15%, and 20%), were spotted on NC membranes. The images of the spotted bacterial macroarray were captured under an ultraviolet lamp. The corresponding concentration of the highest fluorescence intensity and uniformity was considered as the optimal glycerin dosage in this bacterial macroarray. The bacterial macroarray, spotted with spotting buffer containing the optimal dosage of glycerin, was colonized under different incubation conditions (4 °C, 12 h; 37 °C, 2 h; and 55 °C, 2 h). After washing twice with PBST, the images of the spotted bacterial macroarray with different treatments were captured under an ultraviolet lamp.
Screening of Simulated Hybridomas
The principle of the bacterial macroarray for the screening of hybridomas is shown in Figure 1 . The prepared bacterial macroarray was blocked with blocking solution (PBS containing 3% nonfat dried milk) for 30 min at 37 °C and washed three times with PBST. Simulated hybridoma supernatant, diluted with MAbs E7 (4 mg mL−1) of E. coli O157:H7 and 1E9 (2.15 mg mL−1) of C. sakazakii with DMEM high-glucose medium containing 20% serum for a final concentration of 1:1000, was incubated with the bacterial macroarray at 37 °C for 30 min and rinsed three times with PBST; the targeted antibody contained in the supernatant will combine with the corresponding bacteria. The bacterial macroarray was incubated with goat anti-mouse-HRP-conjugated secondary antibody containing 0.2% nonfat dried milk and 2% polyvinyl pyrrolidone (PVP) at 37 °C for 30 min, then rinsed three times with PBST, followed by coloration with HRP–3,3-diaminobenzidine for 5 min in a shaker. Finally, the reaction was stopped by adding distilled water, and the result was visualized by the naked eye. Herein, the prepared bacterial macroarray can be used to detect the sample containing the MAbs of interest in 2 h.
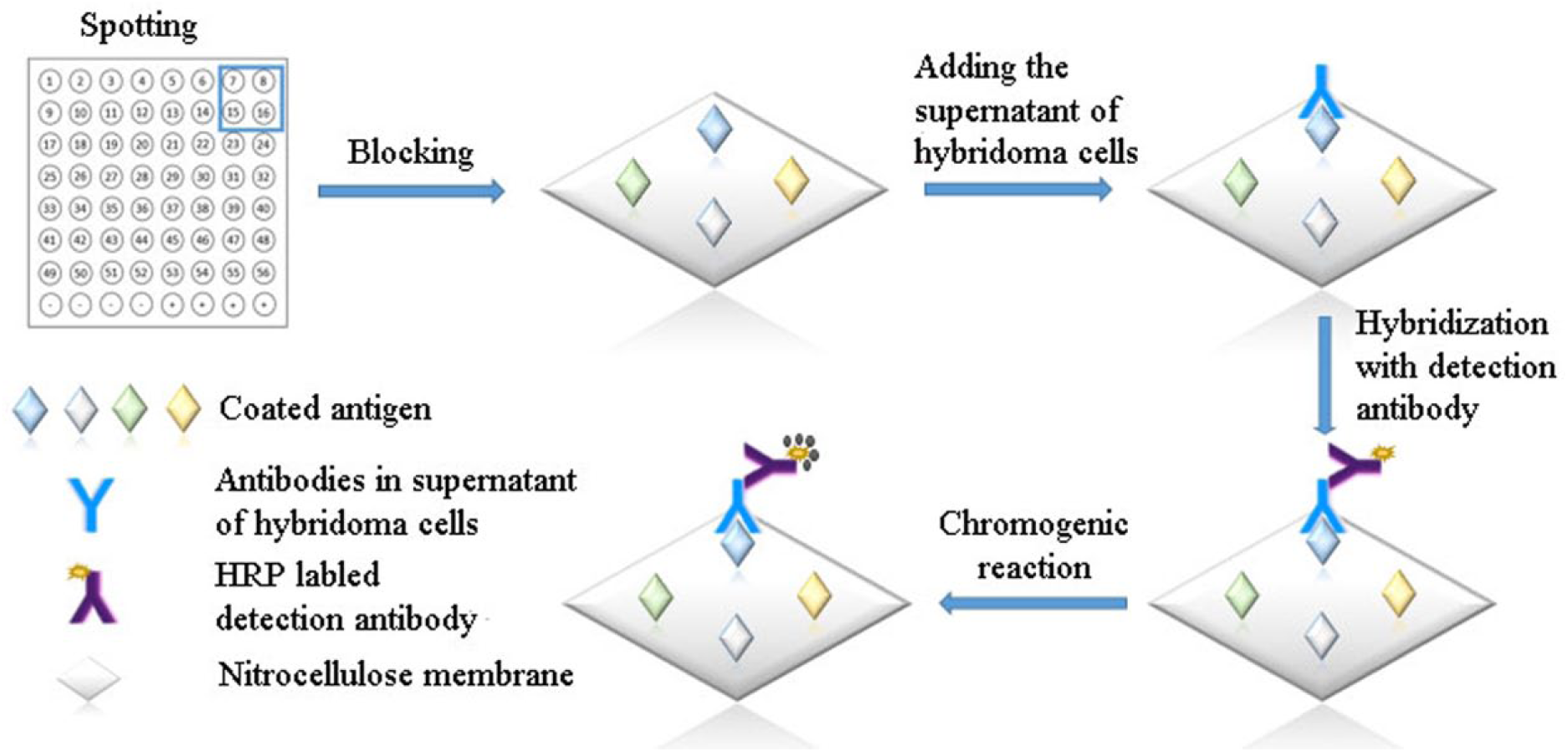
Schematic diagram of the principle of bacterial macroarrays.
Results
Selection of NC Membrane for Bacterial Macroarray
As shown in Figure 2A , the images of the bacterial macroarray, photographed under an ultraviolet lamp by a camera (microlens AF-S VR Micro-Nikon 105 mm f/2.8G IF-ED, Nikon D90, Tokyo, Japan), indicated that the fluorescence intensity on the 0.45 µm, 0.8 µm, and 5.0 µm membranes was higher than that on the 0.22 µm membrane. As shown in Figure 2B , it was significant that the total fluorescence intensity on the 0.45 µm membrane was higher than that on the 0.8 µm and 5.0 µm membranes; thus, the 0.45 µm membrane was used in the subsequent experiments. Scanning electron micrographs of the NC membranes indicated that the surface structure of the NC membrane became looser with increased pore size ( Fig. 2C ). Generally, the size of bacteria is approximately 0.5 to 5 µm; thus, it is difficult to fix the bacteria on the NC membrane with a smaller pore size (such as 0.22 µm). In conclusion, both the membrane morphologies and total fluorescence intensity suggested that the sensitivity (bacterial fixation) of the 0.45 µm NC membrane was higher than that of the 0.8 µm and 5.0 µm NC membranes. Thus, the 0.45 µm NC membrane was used in this bacterial macroarray in the following experiments.
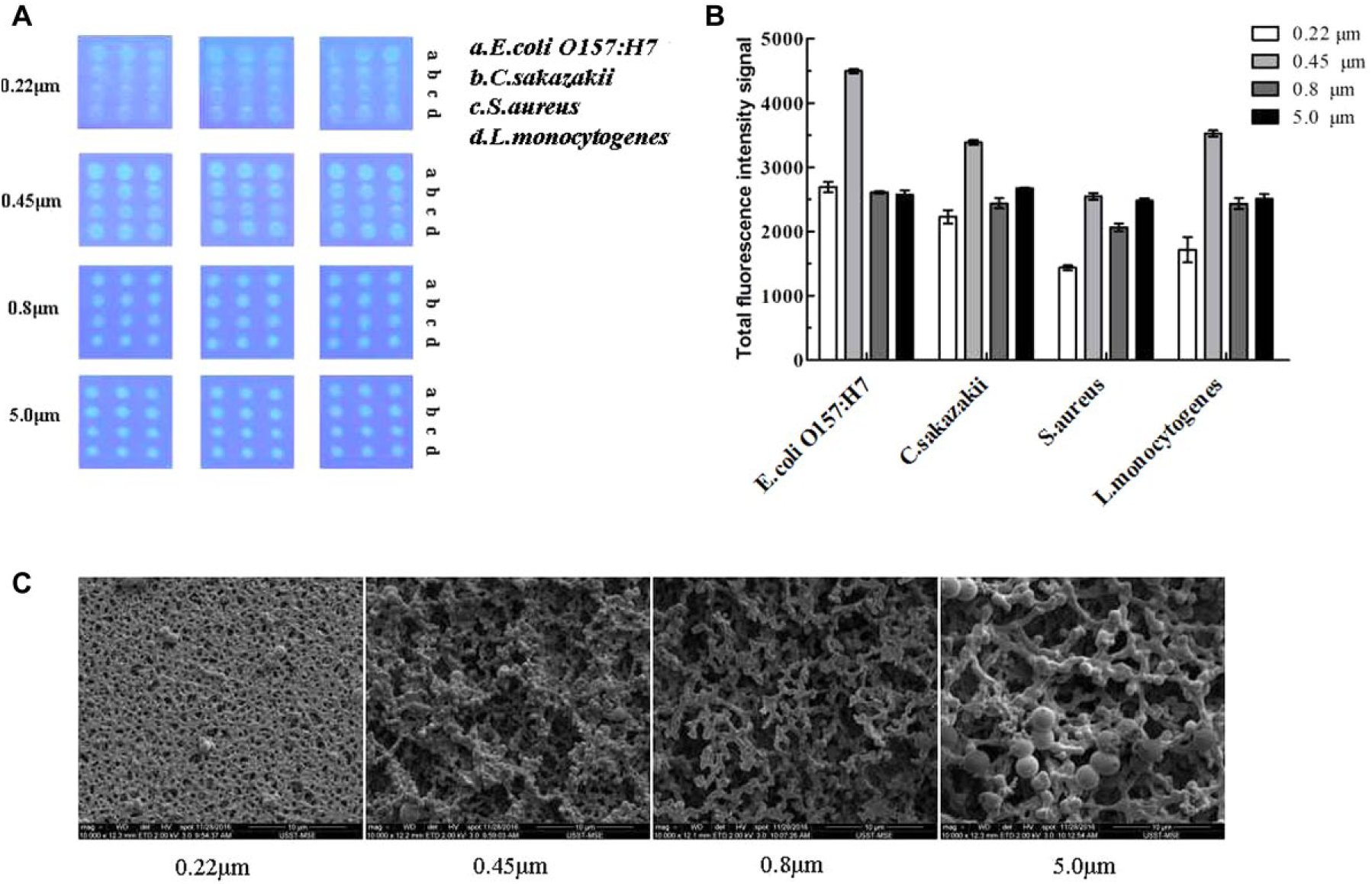
Selection of nitrocellulose (NC) membrane. (
Optimization of the FITC Concentration and FITC-Labeled Bacteria
Figure 3A provides images of FITC-labeled E. coli O157:H7 (a) and FITC-labeled S. aureus (b) with fluorescence microscopy; bacteria labeled with green fluorescence can be observed distinctly. Figure 3B shows the fluorescence intensity of the four FITC-labeled bacteria strains (E. coli O157:H7, C. sakazakii, S. aureus, and L. monocytogenes). Clearly, with increased FITC concentration, the fluorescence intensity of the sample spots increased correspondingly. However, the increased rate of the fluorescence intensity decreased when the FITC concentration was within the range of 200 to 400 µg mL−1. Hence, 400 µg mL−1 was the optimal concentration used in the subsequent experiments. The results analyzed by flow cytometer ( Fig. 3C ) indicated that the average labeling rates of E. coli O157:H7, C. sakazakii, S. aureus, and L. monocytogenes were 88.97%, 88.39%, 89.34%, and 84.66%, respectively ( Fig. 3D ). A high and stable labeling rate of FITC can guarantee the accuracy and stability of this bacterial macroarray.
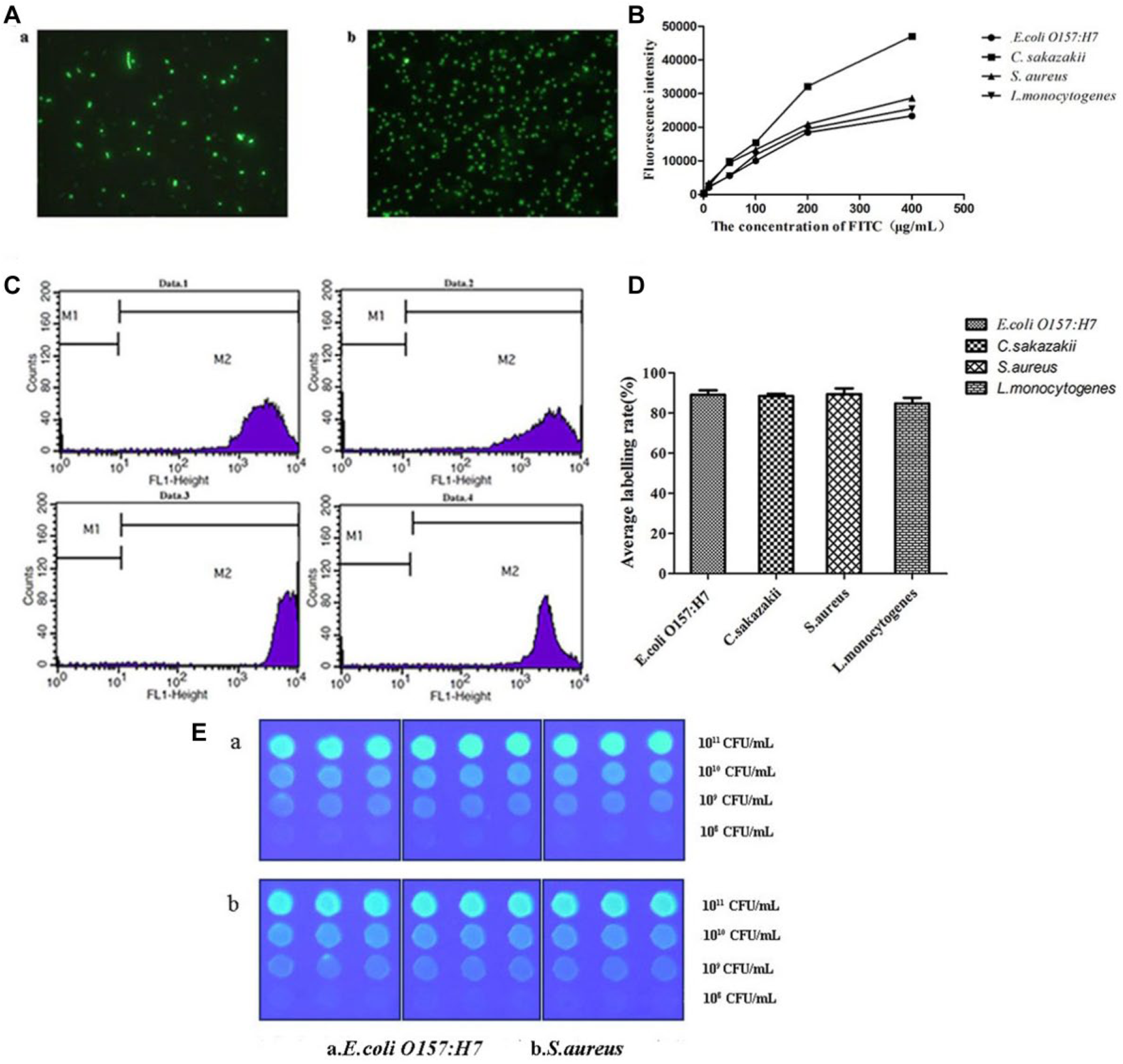
Optimization of the fluorescein isothiocyanate (FITC) concentration and FITC-labeled bacteria. (
As shown in Figure 3E , the fluorescence intensity was higher when the spotting concentration of both FITC-labeled E. coli O157:H7 and FITC-labeled S. aureus was 1011 CFU mL−1. With increased FITC-labeled bacterial concentration, the fluorescence intensity of the spots increased correspondingly, which reflected a better fixation of FITC-labeled bacteria on the NC membrane. The results of the bacterial macroarray for both gram-positive bacteria (S. aureus) and gram-negative bacteria (E. coli O157:H7) showed that the fluorescence intensity was the strongest at the dose of 1011 CFU mL−1 ( Fig. 3E ). Hence, we added 1011 CFU mL−1 FITC-labeled bacteria to the spotting buffer in the following experiment.
Optimization of the Fixation Time and Temperature and the Dosage of Glycerin of the Spotting Buffer
Figure 4A shows the effect of the three different fixation conditions (4 °C 12 h, 37 °C 2 h, and 55 °C 2 h) of FITC-labeled E. coli O157:H7 (a), C. sakazakii (b), S. aureus (c), and L. monocytogenes (d). Not surprisingly, the fixation ability of bacteria on the NC membrane was the lowest when the bacterial macroarray was colonized at 55 °C for 2 h, which may result from the denaturation of the NC membrane at the higher temperature. The result of average fluorescence intensity is shown in Figure 4B ; there is no significant difference for E. coli O157:H7, C. sakazakii, or S. aureus in the immobilization conditions of 12 h at 4 °C and 37 °C for 2 h. However, the best fixation effect was found when the bacterial macroarray of L. monocytogenes was incubated at 37 °C for 2 h. Hence, we chose to fix FITC-labeled bacteria on the NC membrane at 37 °C for 2 h in the following experiments.
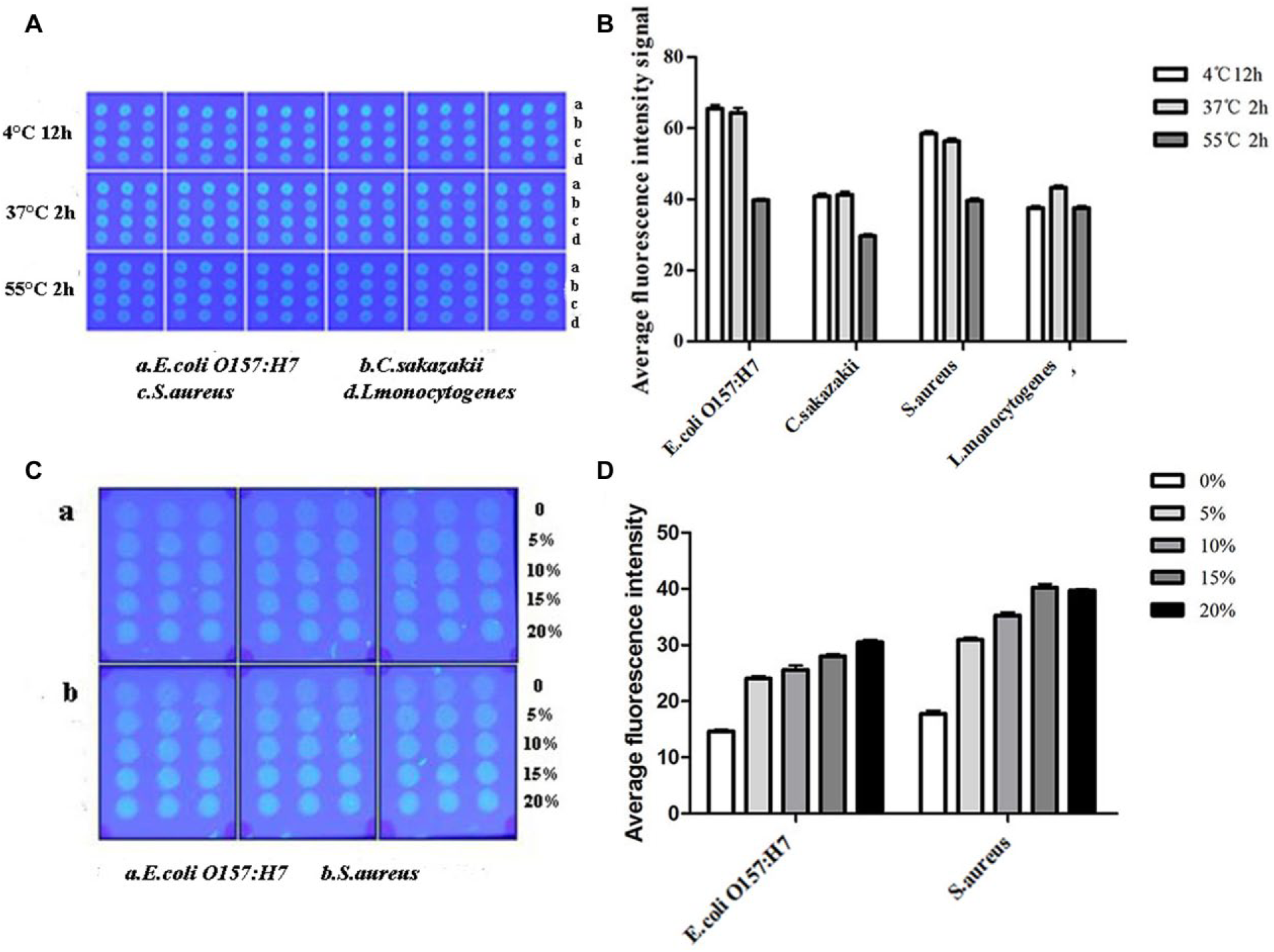
Optimization of the fixation time and temperature and the dosage of glycerin of the spotting buffer. (
As shown in Figure 4C , the “coffee ring,”34–36 a common phenomenon in the preparation of macroarrays, could be observed in this bacterial macroarray. However, the coffee ring disappeared when the supplementation of glycerin was 10% to 20%. Within a specific range, with increased glycerin concentration, the uniformity of the sample spots increased correspondingly. Figure 4D shows that the average fluorescence intensity of both E. coli O157:H7 (a) and S. aureus (b) increased with the increased dosage of glycerol (0%–15%). Generally, although the addition of glycerol in moderation can protect the bacterial surface structure attached on the NC membrane, higher concentrations of glycerin may cause blockage of the arrayer. Therefore, the addition of 15% glycerol in the spotting buffer was optimal in the preparation of this bacterial macroarray.
Simulation of the Application of the Bacterial Macroarray
To verify the performance of this bacterial macroarray in practical applications, we simulated the screening of the prepared bacterial macroarray by detecting cell cultures with the addition of two MAbs (E. coli O157:H7 MAb E7 and C. sakazakii MAb 1E9). MAbs of E7 (4 mg mL−1) and 1E9 (2.15 mg mL−1) were gradient diluted with PBS for a concentration of 1:1000, 1:2000, 1:4000, 1:8000, 1:16,000, 1:32,000, 1:64,000, and 1:128,000. As shown in
Figure 5
, the bacterial macroarray was able to detect the culture medium containing MAb (E7) of E. coli O157:H7 (15 and 16 in
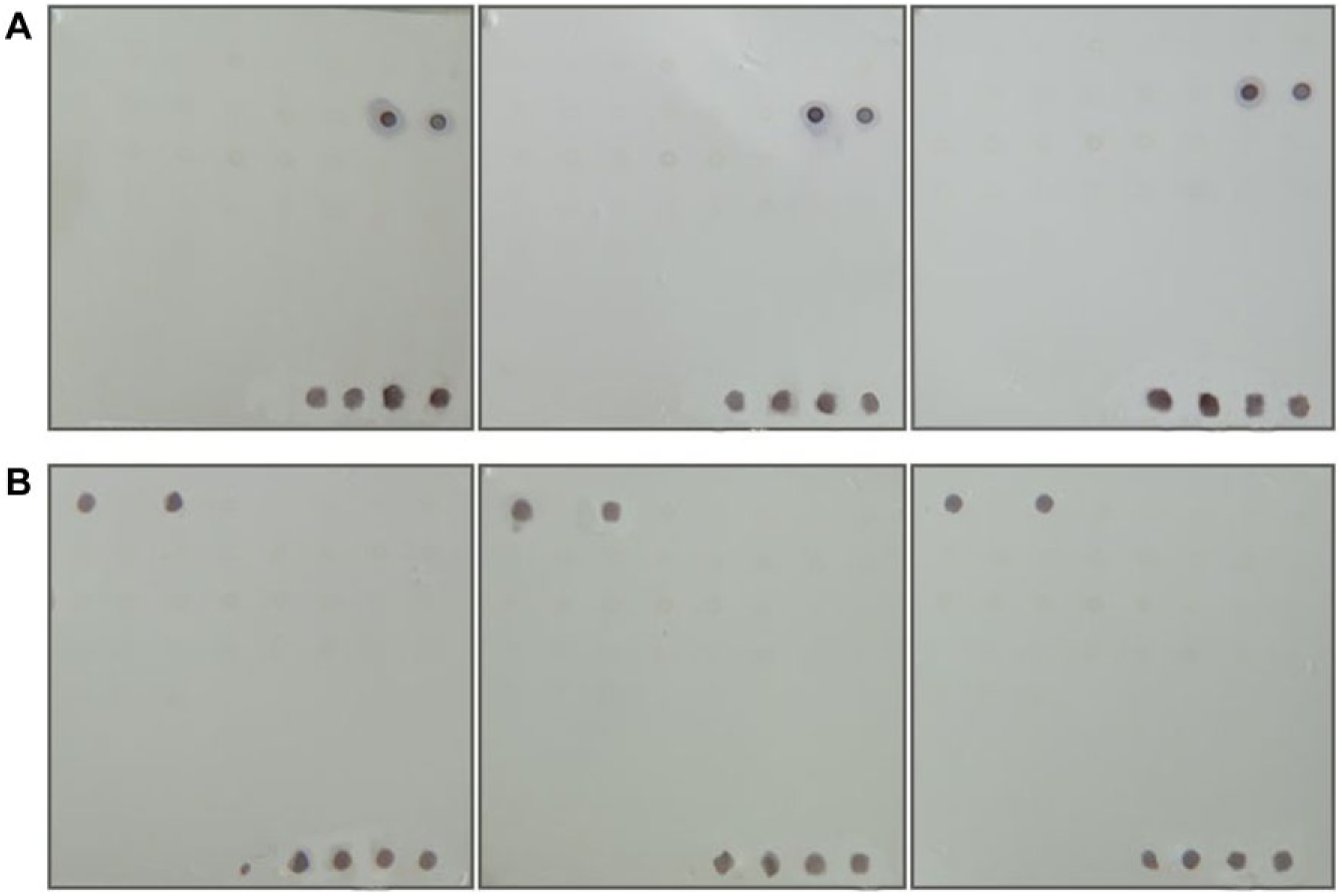
Screening results of the bacterial macroarray. (
Discussion
In this study, an effective and visual bacterial macroarray, spotting 56 common FITC-labeled foodborne pathogenic bacterial strains, was developed for the screening of MAbs-secreted hybridomas in the preparation of MAbs. We found that 400 µg mL−1 was the optimal concentration of FITC to label both the gram-negative bacteria (E. coli O157:H7 and C. sakazakii) and gram-positive bacteria (S. aureus andL. monocytogenes). Then, we optimized the spotting conditions of the labeled bacteria on the NC membrane. As a whole, spotting buffer, containing 1011 CFU mL−1 of FITC-labeled bacteria and 15% (V/V) glycerol, was spotted on a 0.45 µm NC membrane and incubated for 2 h at 37 °C. Afterward, the prepared bacterial macroarray was used to precisely detect cell cultures containing the MAbs of E7 (E. coli O157:H7) and 1E9 (C. sakazakii) within 2 h with LODs of 31.25 ng and 125 ng, respectively. These results show that this bacterial macroarray is feasible and time saving for the screening of hybridoma cells secreting specific MAbs. Although this bacterial macroarray could be used in the detection of antibodies in the supernatant of hybridoma cells, we could not estimate the titer of antibodies accurately. Hence, indirect ELISA methods might also be required for auxiliary screening in practical applications. In the future, the relationship between the fluorescence intensity of the macroarray and the titer of the antibodies could be studied to further improve the screening efficiency.
Supplemental Material
SUPPLEMENTARY_INFORMATION-Lijie-Development_of_bacterial_macroarrays_for_the_rapid_screening_of_targeted_antibody-secreted_hybridomas – Supplemental material for Development of a Bacterial Macroarray for the Rapid Screening of Targeted Antibody-Secreted Hybridomas
Supplemental material, SUPPLEMENTARY_INFORMATION-Lijie-Development_of_bacterial_macroarrays_for_the_rapid_screening_of_targeted_antibody-secreted_hybridomas for Development of a Bacterial Macroarray for the Rapid Screening of Targeted Antibody-Secreted Hybridomas by Jie Li, XuZhao Zhai, Chengchao Ding, Yali Liu, Qingli Dong, Dongpo Xu, Xiang Wang, Jingxuan Qiu, Qi Zhang, Jing Pan and Qing Liu in SLAS Discovery
Footnotes
Acknowledgements
The authors would like to acknowledge funding support from the Agricultural Science Promotion Plan of Shanghai (2017, No. 4-4), the Science and Technology Innovation Plan of Shanghai (No. 18495800400), the Natural Science Foundation of China (No. 81572809), and the Natural Science Foundation for Young Scientists of China (No. 81502504).
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: the Agricultural Science Promotion Plan of Shanghai (2017, No. 4-4), the Science and Technology Innovation Plan of Shanghai (No. 18495800400), the Natural Science Foundation of China (No. 81572809), and the Natural Science Foundation for Young Scientists of China (No. 81502504).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
