Abstract
Organoid methodology provides a platform for the ex vivo investigation of the cellular and molecular mechanisms underlying brain development and disease. The high-grade brain tumor glioblastoma multiforme (GBM) is considered a cancer of unmet clinical need, in part due to GBM cell infiltration into healthy brain parenchyma, making complete surgical resection improbable. Modeling the process of GBM invasion in real time is challenging as it requires both tumor and neural tissue compartments. Here, we demonstrate that human GBM spheroids possess the ability to spontaneously infiltrate early-stage cerebral organoids (eCOs). The resulting formation of hybrid organoids demonstrated an invasive tumor phenotype that was distinct from noncancerous adult neural progenitor (NP) spheroid incorporation into eCOs. These findings provide a basis for the modeling and quantification of the GBM infiltration process using a stem-cell-based organoid approach, and may be used for the identification of anti-GBM invasion strategies.
Keywords
Introduction
Glioblastoma multiforme (GBM) is the most aggressive type of brain cancer and is associated with a median survival of ≤15 months from diagnosis. 1 Therapeutic intervention (including surgical debulking, radiotherapy, and chemotherapy) fails to effectively eliminate the entirety of the tumor, and the rate of tumor recurrence remains high. 2 One aspect of GBM biology that poses a major therapeutic challenge is the diffuse GBM cell invasion into normal surrounding brain.3–5 Emerging experimental and histological evidence indicates that GBM cell migration is accompanied by the expression of stem cell markers and can be predictive of patient outcomes.6–8 Therefore, interventions that specifically target the invasive GBM phenotype are highly desirable. However, the molecular mechanisms underlying the infiltrative nature of GBM cells are not well understood. Furthermore, it is challenging to develop experimental strategies that enable the ex vivo investigation of GBM infiltration into brain-like/neural tissue.
Previous studies have shown the invasiveness of xenografted GBM stem-cell-like cells in the rodent brain.9–11 Moreover, confocal microscopy has been used to visualize glioma cell migration in rodent brain aggregates and organotypic brain slice models.12–15 This body of work, and recent advances in organoid technology, 16 motivated the ex vivo development of a throughput-compatible biological assay. We based our methodology on neural tissue infiltration by free-floating human GBM and neural progenitor (NP) spheroids in coculture with mouse embryonic stem cell (mESC)–derived early-stage cerebral organoids (eCOs). Interestingly, we observed a pronounced eCO infiltration by GBM compared with NP spheroids, thus providing an exclusively cell-based experimental approach for the analysis of invasive GBM cellular tissue compartments in three dimensions and real time.
Materials and Methods
Cell Culture
The generation and characterization of the GBM1 and NP1 cell models have been previously described.
17
GBM1 cells were cultured in Neurobasal medium (Gibco, Waltham, MA; 21103-049), supplemented with 0.5× B-27 (Gibco; 17504-044), 0.5× N-2 (Gibco; 17502-048), recombinant human basic fibroblast growth factor (bFGF; 40 ng/mL; Gibco; PHG0024), and recombinant human epidermal growth factor (rhEGF; 40 ng/mL; R&D, Minneapolis, MN; 236-EG). Cells were maintained in a humidified incubator with 5% CO2 at 37 °C on poly-
Green fluorescent protein (GFP)-expressing lentiviral particles were produced with pFUGW plasmid using a third-generation lentiviral system and HEK 293T cells. Viral particles were added to the cells at a multiplicity of infection (MOI) of 3.07. Spheres were generated through seeding and aggregation of 500 single GBM1 or NP1 cells (each cell line cultured in its respective media as stated above) in ultra-low-attachment (ULA) 96-well plates (Corning, Tewksbury, MA; 7007).
Cerebral Organoid Development and eCO/GBM and eCO/NP Assays
Mouse cerebral organoids were developed using R1 mESCs as described in Lancaster et al., 18 with minor modifications, starting from differentiation day 6. R1 mESCs were cultured on mitotically inactivated mouse embryonic fibroblasts (MEFs) in media consisting of DMEM high glucose (DMEM-HG; Gibco; 11960-044) supplemented with 15% (v/v) knockout serum replacement (KSR; Gibco; 10828-028), 0.1 mM minimum essential medium nonessential amino acid solution (MEM-NEAA; Gibco; 11140-035), 2 mM glutamine (Gibco; 10378-016), 1 mM sodium pyruvate (Gibco; 11360-039), 0.1 mM β-mercaptoethanol (Sigma; M3148), and leukemia inhibitory factor (LIF; 1000 U/mL; Millipore, Darmstadt, Germany; ESG1106) at 37 °C with 5% CO2. Cells were trypsinized once they reached 70%–80% confluency and separated from the MEF cells by differential adherence of these to gelatinized plates. Subsequently, 2000 dissociated R1 mESCs were plated in ULA 96-well plates. The cell suspension was cultured in differentiation media consisting of Glasgow MEM (GMEM; Sigma; G5154) supplemented with 10% KSR, 2 mM glutamine, 1 mM sodium pyruvate, 0.1 mM MEM-NEAA, 0.1 mM β-mercaptoethanol, and 10 μM SB431542 (Sigma; S4317) for 4 days, as previously described. The resulting embryoid bodies were transferred to ULA 24-well plates and cultured in neural induction media consisting of DMEM/F12 medium, N2 supplement (1:100), 1× GlutaMAX, 1× MEM-NEAA, and 1 μg/mL heparin (Sigma; H3149). On day 6, contrary to the Lancaster et al. 18 protocol, organoids were not added to Matrigel droplets but continuously grown in ULA 24-well plates. Neural induction media was removed (on day 6) and replaced with differentiation media consisting of a mixture of Neurobasal medium and DMEM/F12 medium (1:1) supplemented with N2 supplement (1:200), B27 supplement without vitamin A (1:100; Invitrogen; 12587010), 3.5 μL/L of β-mercaptoethanol, insulin (1:4000; Sigma; I9278-5), GlutaMAX (1:100), and MEM-NEEA (1:200). On day 9, the differentiation media was exchanged for differentiation media containing B-27 supplemented with vitamin A (1:100; Gibco; 17504-044). The eCOs were incubated for 2 days before being transferred to ULA 96-well plates containing either GBM1 or NP1 premade spheroids, which were generated 24 h before t0. eCO/GBM1 and eCO/NP1 were cocultured in either GBM or NP media, respectively, for 48 h. Following that time point, differentiation media containing B-27 supplemented with vitamin A was added in a proportion of 1:1.
For real-time imaging, ULA 96-well plates containing eCOs and GBM1 or NP1 spheroids were transferred to an IncuCyte ZOOM live cell imaging system (Essen Bioscience, Ann Arbor, MI) at t0. Phase contrast images were acquired for all conditions every 30 min over a period of 3 days using a 10× objective. Real-time movies were compiled using the IncuCyte ZOOM software package.
Immunohistochemistry
The eCO/GBM1 and eCO/NP1 fusion organoids were fixed with 4% (w/v) paraformaldehyde for 15 min at 4 °C. The tissues were subsequently washed three times with phosphate-buffered saline (PBS), resuspended in 30% (w/v) sucrose solution, and allowed to sink for 24 h at 4 °C. Once the organoids had sunk, they were embedded in OCT (VWR, Radnor, PA; 361603E) in cryomolds and sectioned on a cryostat (20 μm thickness). Images of every section of each individual organoid were taken, and quantification of the GBM1 or NP1 compartments was achieved through color thresholding of the GFP-positive cells using ImageJ (standard settings). Migration distances were measured using ImageJ (freehand line tool).
For immunohistochemistry, the sections were equilibrated in PBS, permeabilized with PBS containing 0.2% (v/v) Triton X-100 for 10 min, and incubated for 1 h at room temperature with a PBS blocking/staining solution containing 0.03% (v/v) Triton X-100 and 10% (v/v) FBS. Next, the sections were incubated at 4 °C overnight with the following primary antibodies: anti-TuJ1 (1:1000; Cambridge Bioscience, Cambridge, UK; 801202), anti-GFAP (1:200; DAKO, Carpinteria, CA; Z0334), anti-Ki-67 (1:200; Abcam, Cambridge, UK; Ab16667), anti-SOX2 (1:150; Cell Signaling, Danvers, MA; 35795), anti-VIMENTIN (1:200; DAKO; M0725), anti-NESTIN (1:200; Millipore; MAB5326), anti-MMP2 (1:50; Sigma; HPA001939), and anti–matrix metalloproteinase 9 (MMP9) (1:200; Abcam; ab38898). VIMENTIN and NESTIN primary antibodies were human specific, showing no cross-reactivity with mouse cells. The cells were then incubated for 1 h at room temperature in the dark with the following secondary antibodies: Alexa-fluor-488 goat anti-mouse (1:200; Molecular Probes, Waltham, MA; A11029) and Alexa-fluor-Cy3 (1:400; Jackson ImmunoResearch, West Grove, PA; 711-165-152). Nuclei were stained with (DAPI; 1 μg/mL; Sigma; D9542), and imaging was carried out using an EVOS digital inverted fluorescence microscope (Life Technologies, Waltham, MA) and a Nikon A1R confocal microscope.
Statistical Analysis
Data were analyzed by two-sided unpaired Student t-test and expressed as mean ± standard deviation (SD). N represents independent eCO/GBM and eCO/NP hybrids. p values of ≤0.01 were considered highly significant.
Results and Discussion
To establish an experimental approach that was amenable to real-time imaging and medium throughput (in a 96-well format), we separately generated eCOs and GBM spheroids ( Fig. 1A ). The latter were generated from an established patient-derived GBM cell model (GBM1), which is characterized by stem-cell-like features, including NESTIN/SOX2 coexpression, as well as clonal growth, and in vivo tumorigenicity/invasion capacity.17,19 Homogenous GBM spheroids were generated by aggregation of 500 GBM1 cells stably expressing GFP. To maximize experimental throughput, we used a short mESC differentiation period (12 days) that suffices to induce neural tissue identity in eCOs 18 prior to the coculture assay ( Fig. 1A ). Qualitative immunohistological analysis showed the presence of the neuronal marker TuJ1 and distinct GFAP-positive cells within the eCOs, thus confirming neural lineage commitment at day 12 ( Suppl. Image 1 ). Notably, we observed that the subsequent coculture of GBM spheroids and eCOs for 48 h resulted in the spontaneous incorporation of GBM1 spheres into eCOs in 100% of cases, based on a total of 114 specimens across seven different experimental sets ( Fig. 1B ). Hybrid organoid formation was preceded by GBM spheroid/eCO fusion (as visualized by confocal microscopy after a 16 h coculture period; Fig. 1C ). These results indicate a high reproducibility of GBM/eCO hybrid organoid formation.
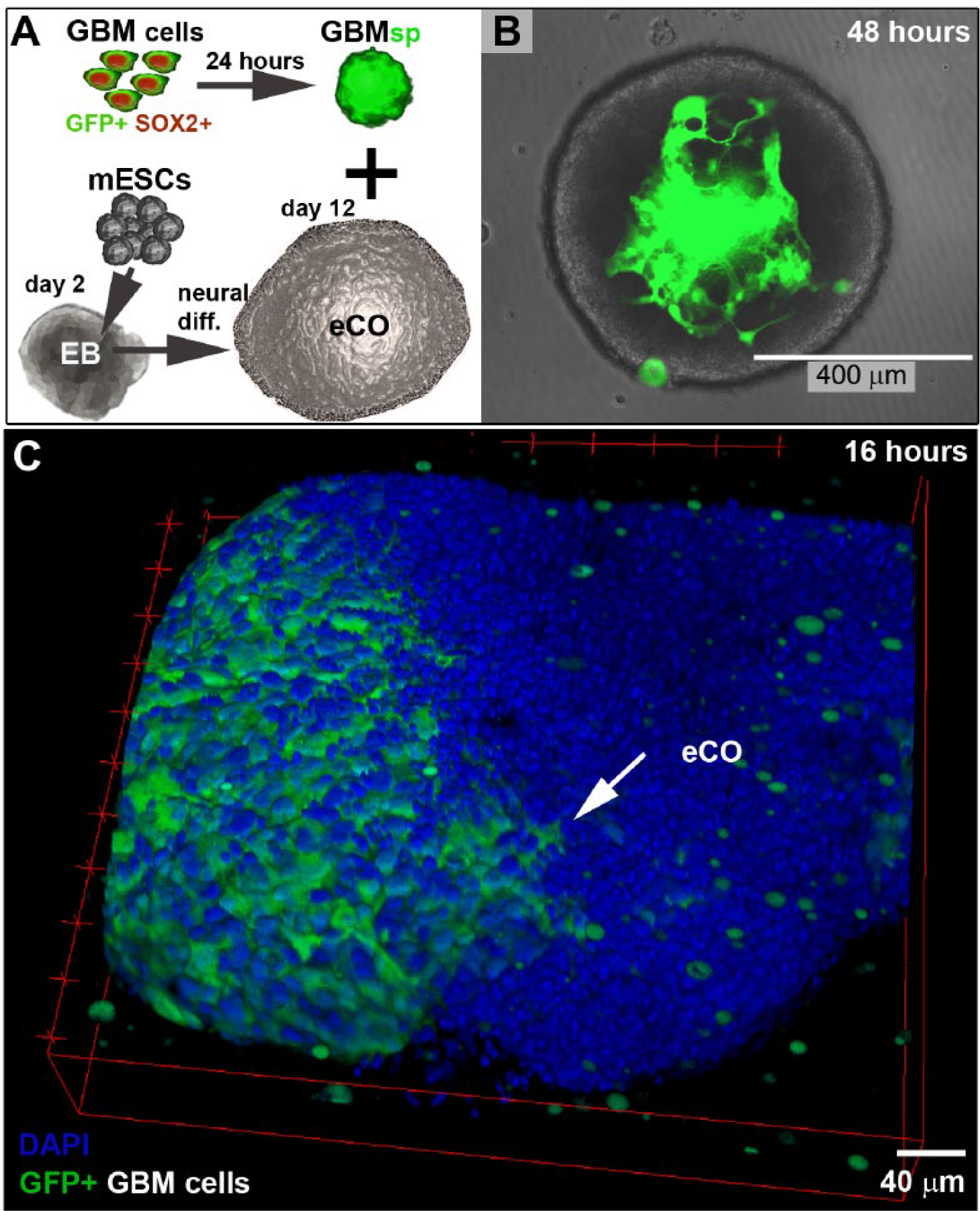
Investigation of neural tissue infiltration by free-floating GBM compared with NP spheroids through the coculture with mESC-derived eCOs. (
Based on the spontaneous infiltration of the eCOs by GBM cellular compartments, we next sought to address the dynamics of this process and whether the hybrid organoid phenotype was dependent on the cellular context (i.e., GBM vs NP). To this end, we utilized an adult brain–derived NESTIN and GFP-expressing NP line (NP1; originating from epilepsy surgery excess tissue 17 ) ( Suppl. Image 2 ) and generated 500-cell spheroid aggregates from GBM and NP cells, respectively. In agreement with reports describing fused cerebral organoids,20,21 time-lapse microscopy experiments revealed that both GBM and NP spheroids first attached to and then fused with and ultimately became incorporated into eCOs ( Fig. 2A , Suppl. Movies 1–4 ; note that we refer to incorporated GBM and NP spheroids as GBM and NP compartments). However, the GBM spheroids showed a rapid incorporation with eCOs (resulting in hybrid eCO/GBM compartments <18 h), whereas NP incorporation required a prolonged period (≥24 h; 96% ± 3.7; n = 70).
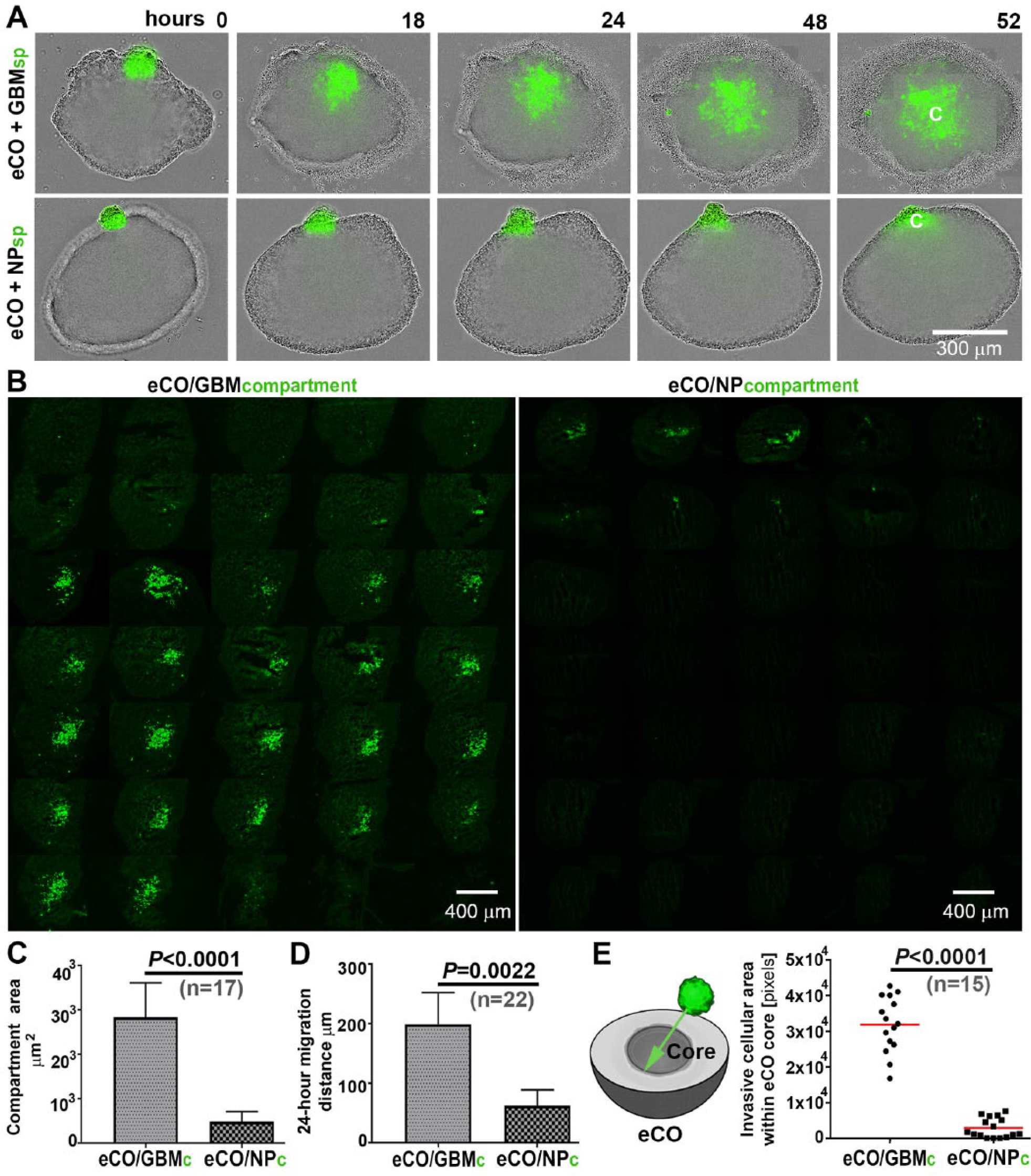
Comparison of GBM and NP spheroid infiltration of eCOs. (
Furthermore, histological analysis and image quantification showed a striking phenotypic difference between eCO/GBM and eCO/NP hybrid organoids ( Fig. 2B ). When compared with their NP counterparts, the GBM compartments within eCO/GBM hybrid organoids were significantly larger (≥5-fold) after a 52 h coculture period ( Fig. 2C ). In addition, within the organoids, GBM cells displayed a significantly increased migration capacity at the 24 h time point ( Fig. 2D ). GBM cells consistently infiltrated the inner layers (core) of eCOs more effectively than their NP counterparts at the experimental end point ( Fig. 2E ). Further qualitative immunohistological analysis of inner-layer GBM compartments (using human-specific antibodies for GBM compartment visualization) showed that the infiltrative GBM cells are characterized by coexpression of VIMENTIN and MMP2 proteins, which have been previously implicated in glioma invasion ( Fig. 3A , B ; MMP9 immunopositivity was rarely detected; Suppl. Image 3 ).22,23 In addition, the GBM cells expressed the known “stemness” markers NESTIN and SOX2 ( Fig. 3C ). Expression of the cell cycle marker Ki-67 was low or absent in the invasive GBM cell population, indicating a migratory rather than proliferative GBM cell phenotype ( Suppl. Image 4 ).
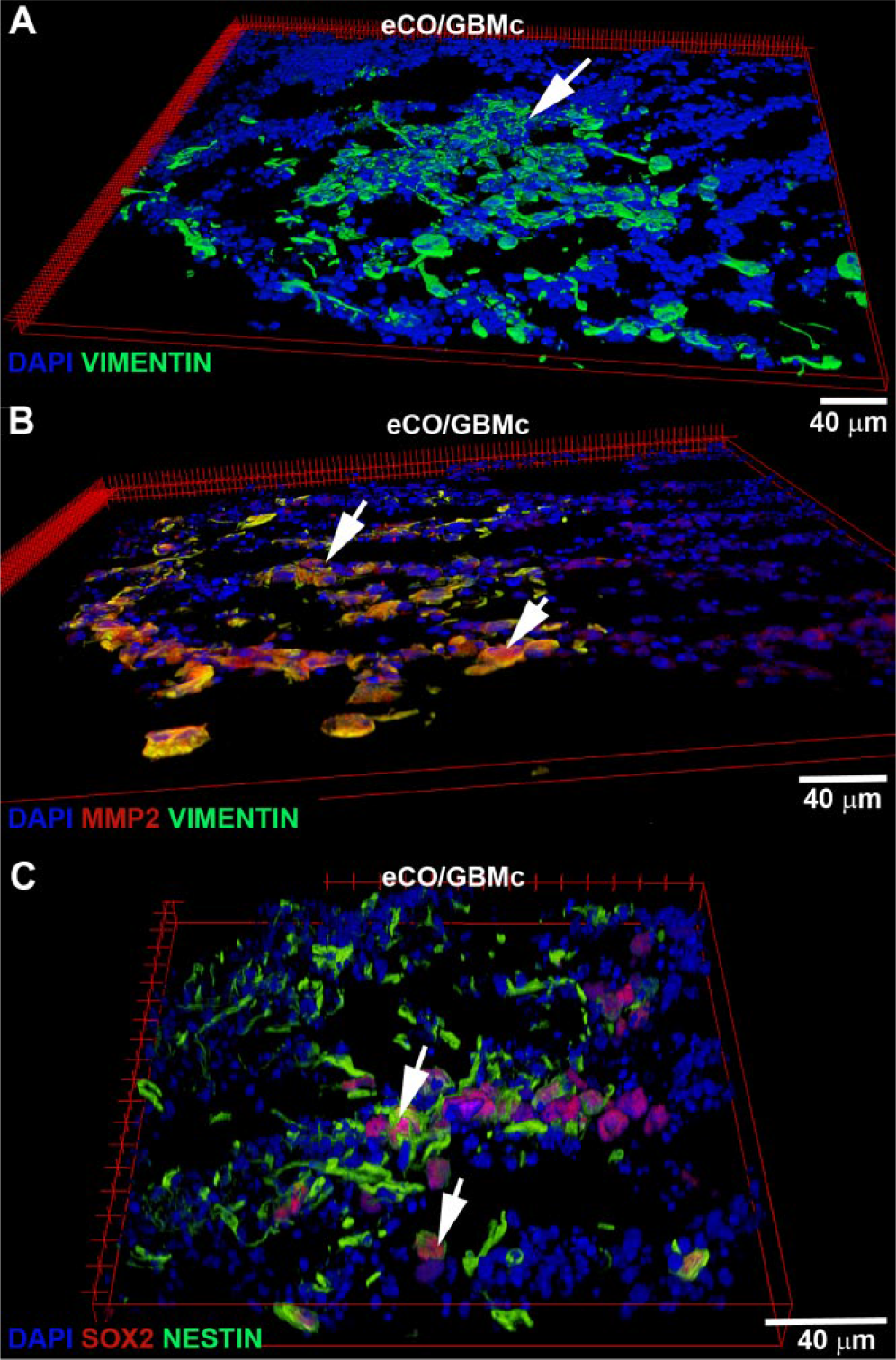
Representative confocal microscopy immunofluorescence images (volume view) of the invasive GBM cellular compartment in histological sections of self-aggregated hybrid organoids (eCO/GBM). (
In summary, our technical study demonstrates that coculture of human GBM/NP spheroids and mESC-derived eCOs robustly resulted in the formation of hybrid cellular compartment organoids. We aimed to limit the experimental variability that is often associated with organoid methodology (e.g., reviewed in Jorfi et al. 24 ). For example, long-term-cultured cerebral organoids have been reported to contain heterogeneous architectures and cellular diversity.18,25 Thus, we utilized a short 12-day mESC differentiation period generating eCOs and a well-characterized patient-derived GBM cell model.17,19 Accordingly, we developed a reproducible and self-aggregating ex vivo experimental model system that aims, in real time, to recapitulate the infiltrative characteristic of GBM that makes current therapy so ineffective. The use of standard (inverse) microscopy, live cell imaging, and a medium-throughput (96-well) assay format was made possible by eCO diameters of 300–800 µm. The resulting hybrid organoid sizes would allow for the use of 384-well (optical bottom) microtiter plates in medium- or high-throughput screening applications, and automation of hybrid organoid dispensing into microtiter plates may be an option (note that a gel environment is currently not required for the assay). The protocol described herein may require significant modifications with regard to using the 1536-well screening format.
Importantly, the coculture experiments indicated a significant difference in the infiltration capacity of GBM versus NP cells. Time-lapse microscopy identified the distinct phases of GBM/NP spheroid attachment and fusion, with tissue infiltration invasion exhibited exclusively by GBM cells. This hybrid organoid GBM invasion readout may complement limitations of monocompartment assays, especially regarding identifying and distinguishing between molecular mechanisms that promote GBM cell migration and/or viability (e.g., utilizing chemical and/or genetic screens). The reported experimental approach is limited in being unable to observe migration of GBM cells along blood vessels and into the corpus callosum (evident in patient tumors 26 ). Moreover, the results described are linked to the specific biological materials and protocols employed, and a hybrid organoid environment may substantially differ from that of the adult brain. Additional studies are required in terms of comparing stem-cell-based organoids with other ex vivo glioma migration methodologies, including organotypic slice culture models.13–15 However, eCO/GBM hybrid organoids (and their noncancerous controls) provide the foundation for the future development of high-throughput assays to investigate the underlying molecular mechanisms of invasion and screen for drugs that abrogate this phenotype. Future modifications of the assay may be based on the replacement of R1 mESCs with other (e.g., human) ESCs and/or the replacement of the GBM1 stem-cell-like model with other brain tumor cell models and/or freshly isolated primary cells. Likewise, future studies may investigate hybrid organoid maturation and the development of GBM versus NP compartment phenotypes (e.g., with respect to quiescence and self-renewal capacity) over a prolonged (>2 months) culture period.
Supplemental Material
DS_DISC764623 – Supplemental material for Spontaneous Glioblastoma Spheroid Infiltration of Early-Stage Cerebral Organoids Models Brain Tumor Invasion
Supplemental material, DS_DISC764623 for Spontaneous Glioblastoma Spheroid Infiltration of Early-Stage Cerebral Organoids Models Brain Tumor Invasion by Bárbara da Silva, Ryan K. Mathew, Euan S. Polson, Jennifer Williams, and Heiko Wurdak in SLAS Discovery
Footnotes
Acknowledgements
We thank P. Ceppi, B. K. Irving, M. Lorger, C. Gomes, and A. Tsakiridis for carefully reading the manuscript.
Supplementary material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: H.W. acknowledges support from the MRC New Investigator Award (MR/J001171/1). B.dS., E.S.P., and J.W. acknowledge support from Brain Tumor Research and Support across Yorkshire. R.K.M. acknowledges support from the Leeds Cancer Research UK Centre.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
