Abstract
Since the cell-based cDNA microarray (CBCM) technique has been a useful tool for gain-of-function studies, many investigators have used CBCMs to identify interesting genes. However, this method requires better-established conditions to ensure high reverse transfection efficiency without cross-contamination. Therefore, we optimized CBCM techniques through various means. We determined that Lipofectamine 2000 was the most appropriate transfection reagent by evaluating eight commercialized reagents, and we determined that the most effective concentrations for printing solution constituents were 0.2 M sucrose (to yield a final concentration of 32 mM) and 0.2% gelatin (to yield a final concentration 0.075%). After examining various combinations, we also determined that the best concentrations of cDNA and transfection reagent for optimal reverse transfection efficiency were 1.5 µg/5 µL of cDNA and 5.5 µL of Lipofectamine 2000. Finally, via a time course, we determined that 72 h was the most effective reaction duration for reverse transfection, and we confirmed the stability of cDNA spot activity of CBCMs for various storage periods. In summary, the CBCM conditions that we have identified can provide more effective outcomes for cDNA reverse transfection on microarrays.
Introduction
The reverse transfection cell microarray (RTCM) technique was developed to combine genetic functional studies with high-throughput screening (HTS), following the reporting of the lipid-DNA method (LDM). 1 The advantages of the RTCM method include that it reduces time-consuming processes and allows fast analyses; furthermore, it enables cell phenotype visualization and cell-based image analyses. As commonly used tools, nucleic acid chips and protein chips have been utilized primarily for analyzing gene expression and regulation, genetic networks, and signaling pathways; however, the technique has limitations. As an ex vivo assay, the quantities of mRNA present often do not reflect the expression levels of the proteins they correspond to, and changes in protein expression levels are frequently not caused by mRNA-level events. The RTCM method does, however, offer clear results as a cell-based assay, allowing the observation of cytological changes and facilitating cellular-level analyses.
For loss-of-function screens, many investigators have developed siRNA microarray platforms using various transfection reagents and by coating slides with specific substances.2–8 We reported an siRNA microarray system using Lipofectamine 2000, RNAi-Max, poly-
The core concept of the cell-based microarray technique is to maintain high reverse transfection efficiency without allowing cross-contamination by spreading. We have already optimized the conditions for conducting reverse transfections on siRNA microarrays by experimenting with various combinations of substances.9,10,17 However, gain-of-function screens using CBCM techniques are still performed with different conditions. Therefore, we aimed to optimize cDNA reverse transfection conditions by determining the most effective choices for the following experimental parameters: (1) transfection reagent, (2) concentrations of sucrose and gelatin, and (3) combination of cDNA and transfection reagents. We also examined the stability of microarray activity after storage. Optimal final concentrations and other details are summarized in Table 1.
Workflows for Optimizing Printing Solution Conditions for cDNA Reverse Transfection.
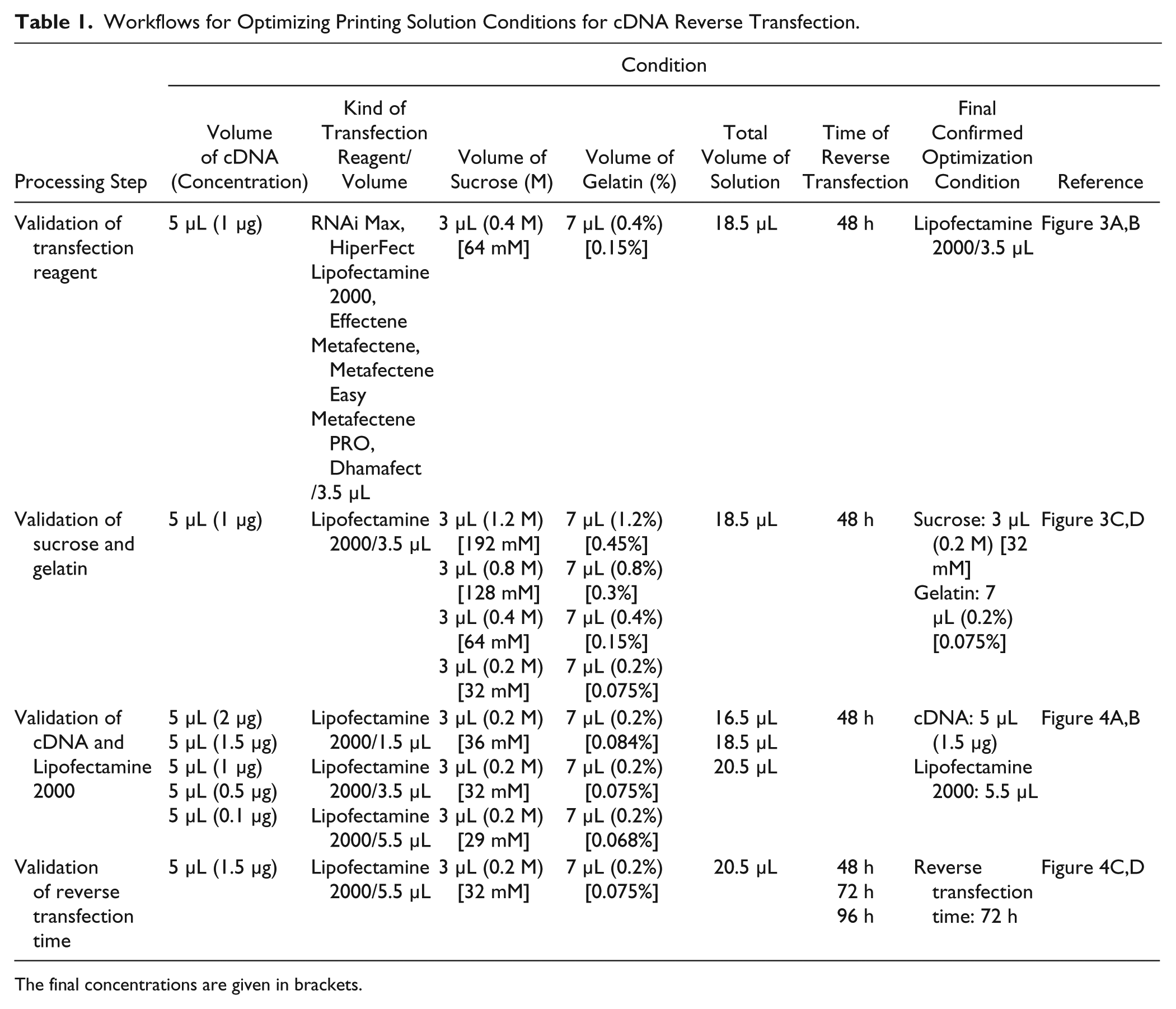
The final concentrations are given in brackets.
Materials and Methods
Processing of CBCM
For the preparation of cDNA solutions, cDNA was transferred into a 384-well plate, and sucrose (Gibco, Invitrogen, Carlsbad, CA) diluted in Opti-MEM (Gibco) was added. Transfection reagent was then added, and the solution was mixed gently. After centrifugation for 1 min at 1000 rpm, the mixture of cDNA and transfection reagent was incubated for 20 min at room temperature. Gelatin (Gibco) diluted in dH2O was then mixed thoroughly into the solution, which was then centrifuged for 1 min at 1000 rpm. The mixture of cDNA was printed onto 0.01% PLL-coated coverslips that were installed in a slide holder. Each cDNA-printed CBCM was moved into a four-well culture dish; prepared cells were then gently dropped onto the CBCM, and were cultured to induce reverse transfection for 2–4 days. After cell fixing and nucleus staining, image acquisition was performed using an automated microscope and analysis software ( Fig. 1 ).
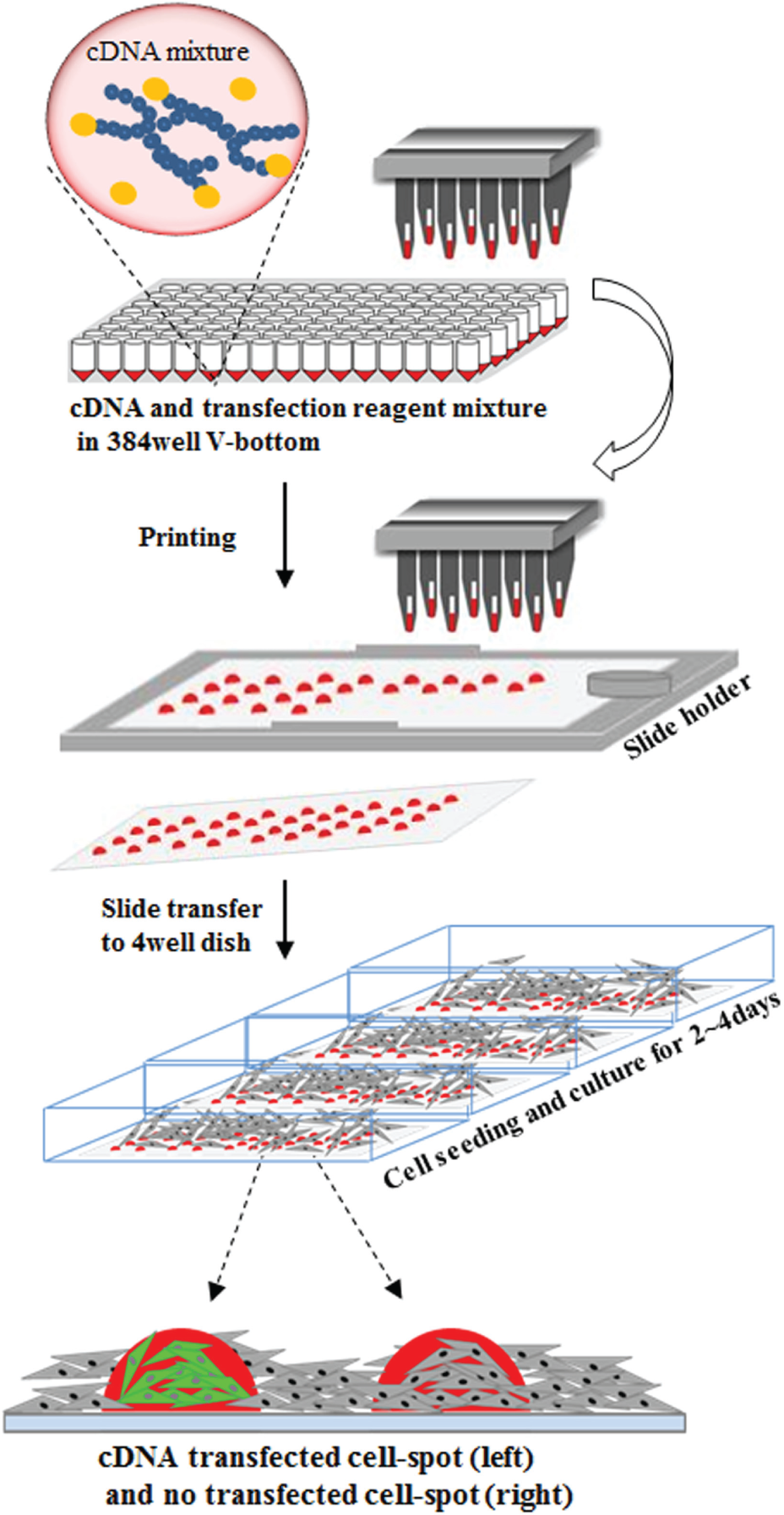
Schematic diagram of CBCM. cDNA mixtures prepared in V-bottomed 384-well plates were printed onto glass coverslips, using a stealth pin-based microarray printer. After the cDNA array was transferred to the four-well dish, cells were seeded and cultured for 2–4 days to induce reverse transfection on the spots.
Cell Culture
HEK293 cell lines were maintained in Dulbecco’s modified Eagle’s medium (DMEM; Gibco) supplemented with 10% (v/v) fetal bovine serum (Gibco) and 1% (v/v) penicillin-streptomycin (Gibco), at 37 °C, in a 5% CO2 incubator.
Preparation of Plasmid
The pEGFP-N3 plasmid, tagged with green fluorescent protein (GFP) driven by the cytomegalovirus (CMV) promoter, was purchased from Clontech (Mountainview, CA) and purified using the QIAprep Spin Midiprep Kit (Qiagen, Hilden, Germany). The concentration was determined using the NanoVue Spectrophotometer (GE Healthcare, Pittsburgh, PA), and plasmid DNA having a 260 nm/280 nm absorbance ratio of >1.8 was used.
Preparation of PLL-Coated Glass Coverslips
Autoclaved glass coverslips (24 × 60 mm; Marienfeld, Lauda-Königshofen, Germany) were dipped for 2 h in 0.01% PLL (Sigma, St. Louis, MO) diluted in dH2O at room temperature. PLL-coated coverslips were washed with dH2O, and dH2O was then removed via slide chamber centrifugation. This was repeated three times. After the last washing step, PLL-coated coverslips were dried on a clean lab bench overnight and were then stored in a desiccating chamber, at room temperature (22–25 °C), until further use.
Printing for CBCM
Prepared cDNA solutions were printed onto 0.01% PLL-coated glass coverslips using SMP9 stealth pins (Telechem, Atlanta, GA) and a high-throughput microarray printer (Genomic Solutions, Ann Arbor, MI), at 22–25 °C, 55%–65% relative humidity. Printed spots were 200–300 µm in diameter, with a 1000 µm spot-to-spot interval. cDNA-printed microarrays were stored in a desiccating chamber at room temperature (22–25 °C) until further use.
Reverse Transfection by CBCM
HEK293 cells were cultured until reaching 70%–80% confluence in 25T or 75T flasks. Cells were then detached from the flasks via incubation with 0.05% trypsin-EDTA (Gibco), and were seeded (at 1 × 106/4 mL per slide) with assay media (Opti-MEM-glutamax, 5% [v/v] fetal bovine serum, 1% [v/v] penicillin-streptomycin; Gibco) onto the CBCM, which was then transferred to a four-well rectangular dish (ThermoFisher Scientific, Waltham, MA). The cells were then cultured at 37 °C in a 5% CO2 incubator for 2–4 days to induce reverse transfection.
Imaging Acquisition and Analysis
With the development of cell image–based screening methods, various automated microscope scanners equipped with a high-resolution camera such as a Zeiss Observer 1 (Carl Zeiss Inc., Thornwood, NY) or a scanR high-content imager (Olympus, Tokyo, Japan) have been developed for microarray application,18,19 and the confocal microscope manufactured by Molecular Devices (Sunnyvale, CA) can be used in this study. For imaging acquisition, cells were washed with 1× phosphate-buffered saline (PBS) once, and were fixed with 4% (w/v) paraformaldehyde in 1× PBS for 10 min. Cells were then washed again with 1× PBS, and were incubated with 5 mM DRAQ5 (1:2000 in PBS; Biostatus, Leicestershire, UK) for nucleus staining. Confocal images were acquired using an ImageXpress Ultra point scanning confocal microscope (Molecular Devices) equipped with the following: (1) four solid-state lasers for simultaneous excitation at 405, 488, 561, and 635 nm; (2) a galvanometer for x axis scanning; and (3) a stage for y axis scanning. Each cell spot was scanned at 10× or 20× magnification, using a Nikon air immersion objective lens with specific filter sets for Alexa488 and Alexa635. Image quantification was performed using the “cell scoring” and “count nuclei” application modules of the MetaXpress software package (Molecular Devices) to evaluate the proportion of GFP fluorescent-positive cells by intensity, nuclei numbers (average 300–500 cells), and size ( Fig. 2 ). GraphPad Prism 5 software was used to determine statistical significance for comparisons of two groups, by one-way or two-way analysis of variance (ANOVA) and the Bonferroni method. Differences with p < 0.05 were considered statistically significant.
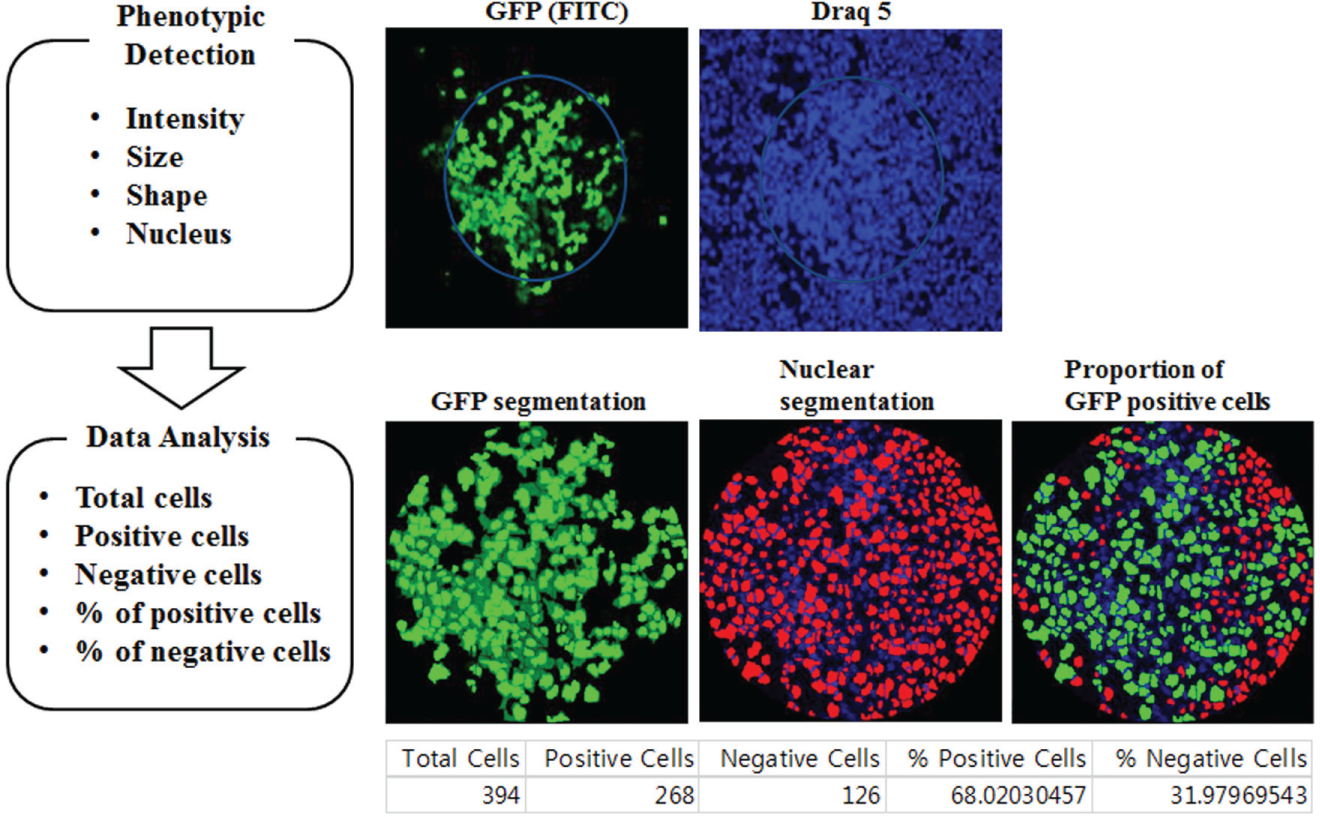
Image acquisition and analysis. Cell spot images were acquired with two fluorescent channels, which were filtered with Alexa488 for green fluorescence and 635 nm for DRAQ5. We used the cell scoring method to analyze images. First, nuclei were counted, and then how many cells were expressed by a GFP signal based on certain criteria, such as intensity threshold and size of pixel.
Results and Discussion
Determination of the Most Effective Gene Delivery Reagent for CBCM
We determined the optimal transfection reagent for achieving effective GFP expression via microarray printed from cDNA solutions by evaluating different transfection reagents. For the selection of the best transfection reagent, 5 µL of cDNA (1 µg), 3 µL of 0.4 M sucrose (for a final concentration of 64 mM) diluted in OptiMEM media, 3.5 µL of a transfection reagent (diluted in dH2O), and 7 µL of 0.4% gelatin (for a final concentration of 0.15%) were combined to create a printing solution. The transfection reagents tested were RNAi-Max and Lipofectamine 2000 (Gibco); HiPerFect and Effectene (Qiagen); Metafectene, Metafectene Easy, and Metafectene Pro (Biontex, Munich, Germany); and Dharmafect (Dharmacon, Lafayette, CO). Three reagents—Lipofectamine 2000, Effectene, and Metafectene Pro—showed high efficiency of GFP expression. However, the cross-contamination phenomenon of “spreading” was observed for Effectene and Metafectene Pro. Therefore, Lipofectamine 2000 was identified as the best transfection reagent for CBCM procedures ( Fig. 3A , B ).
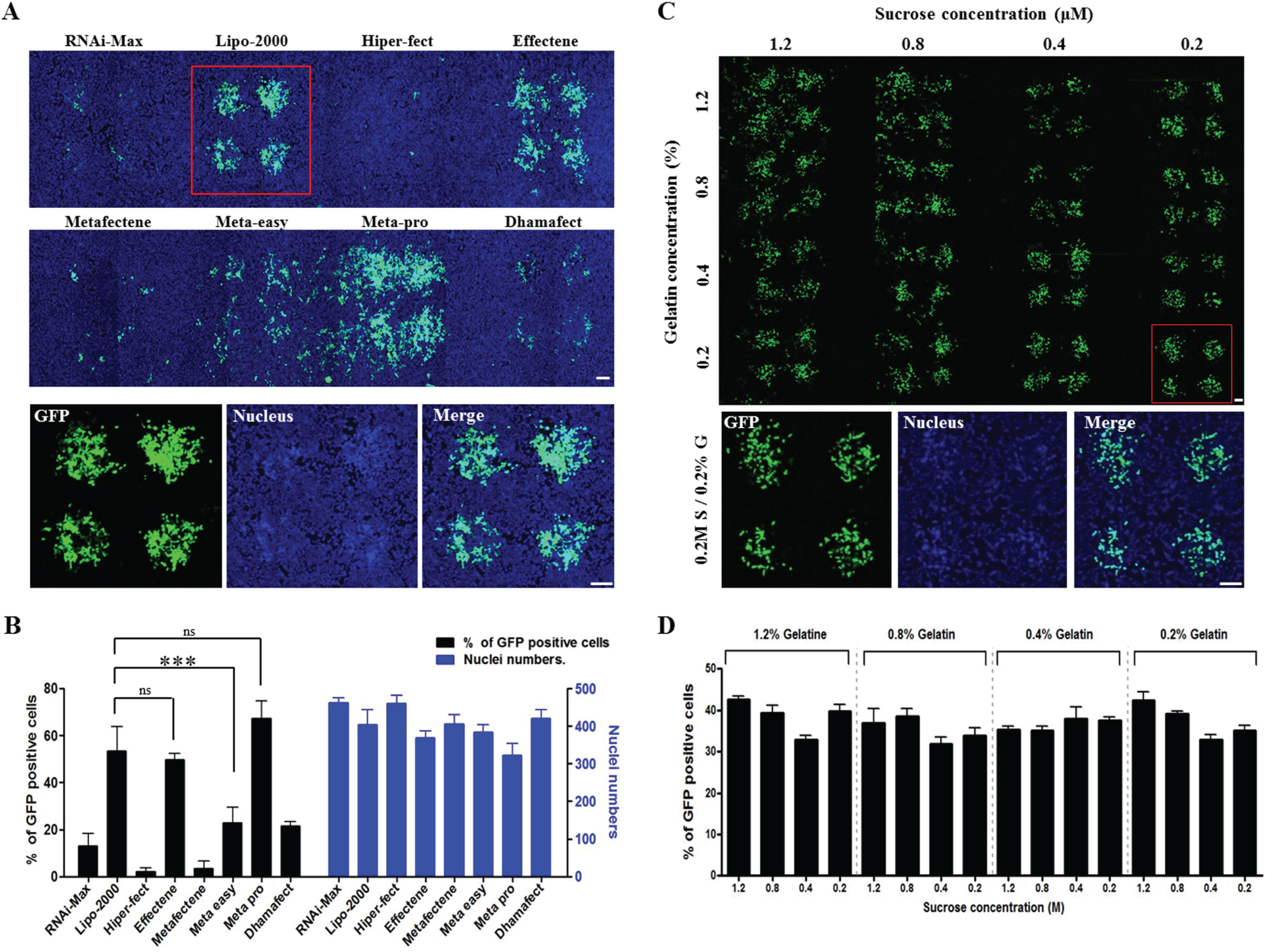
The determination of optimal transfection reagent and sucrose/gelatin concentrations for cDNA reverse transfection efficiency in HEK293 cells. (
Determination of Optimal Sucrose and Gelatin Concentrations
Appropriate concentrations of sucrose and gelatin in the cDNA transfection solution help to maintain spot circular formation after printing, and help prevent the cDNA spot from being swept away by media for cell seeding. As cell-friendly materials, sucrose and gelatin induce cell attachment and growth on the cDNA spot. Therefore, we performed titrations of sucrose and gelatin to determine their optimal concentrations for cDNA reverse transfection efficiency. For this investigation, 5 µL of cDNA (1 µg), 3 µL of 0.2–1.2 M sucrose (for final concentrations of 32–192 mM) diluted in OptiMEM media, 3.5 µL of Lipofectamine 2000, and 7 µL of 0.2%–1.2% gelatin (for final concentrations of 0.075%–0.45%) diluted in dH2O were combined to make printing solutions. Our results showed no significant differences in GFP expression levels based on sucrose and gelatin concentrations. However, the only conditions where stable GFP expression on the cDNA spot occurred without cross-contamination (via spreading) were those with low concentrations of sucrose (below 0.4 M, with a final concentration below 64 mM) and gelatin (below 0.4%, with a final concentration below 0.15%). We determined that the optimal concentrations for cDNA transfection were 0.2 M sucrose (for a final reaction solution concentration of 32 mM) and 0.2% gelatin (for a final reaction solution concentration of 0.075%) ( Fig. 3C , D ).
Determination of Optimal Ratio of cDNA and Transfection Reagent
To obtain high-efficiency outcomes with lipid-based gene transfection techniques, the ratio of cDNA quantities and transfection reagent amounts in the printing solution is an important factor. Therefore, we performed reverse transfection using cDNA printing solutions consisting of different amounts of Lipofectamine 2000 and cDNA, as follows: 5 µL of cDNA (0.1–2 µg), 3 µL of 0.2 M sucrose (with final solution concentrations of 29–36 mM) diluted in OptiMEM media, 1.5–3.5 µL of Lipofectamine 2000, and 7 µL of 0.2% gelatin (with final solution concentrations of 0.068%–0.084%) diluted in dH2O. In this investigation, the solutions containing 2 and 1.5 µg of cDNA, combined with 5.5 µL of Lipofectamine 2000, showed high transfection efficiency. Because we saw no significant differences in outcomes when using either 1.5 or 2 µg of cDNA ( Fig. 4A , B ), we decided that 1.5 µg of cDNA was the optimal amount (because it uses less cDNA and achieves the same outcome).
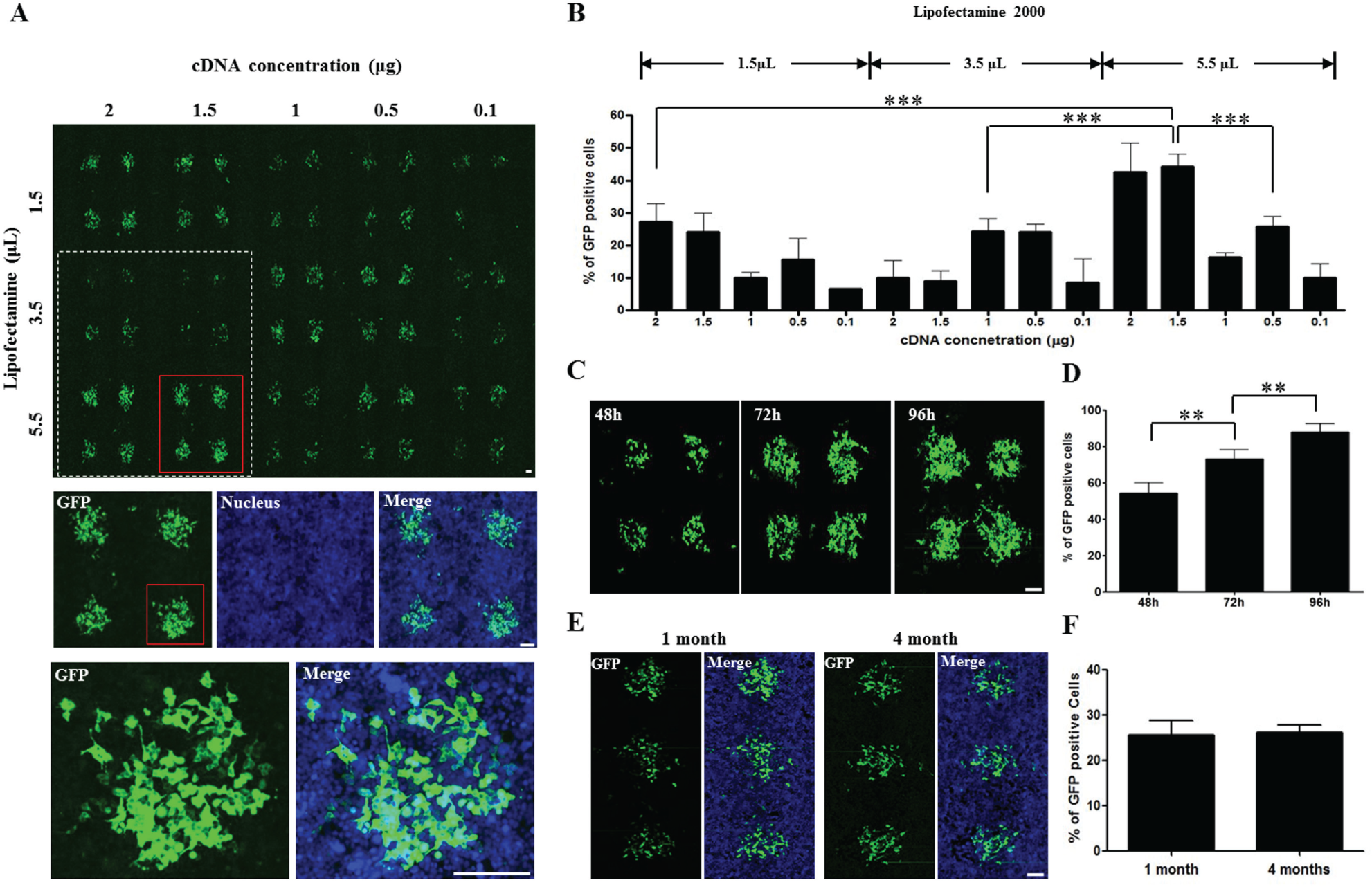
Reverse transfection optimization by quantitative combination of cDNA and Lipofectamine 2000, the determination of optimal reverse transfection duration time, and cDNA spot stability in HEK293 cells. (
Determination of Optimal Reverse Transfection Time
To determine the optimal duration for cDNA reverse transfection time, 5 µL of cDNA (1.5 µg), 3 µL of 0.2 M sucrose (for a final solution concentration of 32 mM) diluted in OptiMEM media, 5.5 µL of Lipofectamine 2000, and 7 µL of 0.2% gelatin (for a final solution concentration of 0.075%) diluted in dH2O were combined to make a printing solution. This was then used to induce reverse transfection over a time course of 48, 72, or 96 h after cell seeding. GFP expression levels were efficient for all time conditions. However, we decided that 72 h was the best duration time for general reverse transfection; this was because the 48 h condition showed lower efficiency than the 72 h condition, and the 96 h condition displayed the overgrowth/cross-contamination problem ( Fig. 4C , D ).
Stability of Reverse Transfection Activity on CBCM
To check the stability of CBCM through different durations of storage, cDNA-transfectant spots were printed using optimized mixture conditions, as follows: 5 µL of cDNA (1.5 µg), 3 µL of 0.2 M sucrose (for a final solution concentration of 32 mM) diluted in OptiMEM media, 5.5 µL of Lipofectamine 2000, and 7 µL of 0.2% gelatin (for a final solution concentration of 0.075%) diluted in dH2O. CBCMs that had been stored in a desiccating chamber for 1 and 4 months were used to induce reverse transfection for 48 h. We confirmed that no significant decreases in GFP expression efficiency resulted from long-term storage at these durations ( Fig. 4E , F ). Furthermore, we observed similar results with CBCMs that had been stored for longer than 6 months (data not shown).
The CBCM technique is a useful tool for genetic functional studies because of its advantages, such as miniaturization of HTS, cell-based visualization, and cellular-level analysis. However, more precise CBCM conditions are required to satisfy the needs of various assays and cell lines. Therefore, we sought to optimize CBCM conditions by examining various parameters, such as transfection reagent selection, sucrose and gelatin concentrations, cDNA–transfectant reagent ratios, and duration of reverse transfection reaction in HEK293 cells. We also attempted to apply these results to other cell lines, such as HeLa and U2OS, and observed reverse transfection efficiencies (data not shown); however, lipid-based transfection methods have been defined for various cell lines. We recently reported results for cell-defined lentivirus microarrays (CDLMs), 20 and we expect lentivirus-based cDNA microarrays using hydrogel-coated slides 11 to be an alternative for cell types where reverse transfection via CBCM is difficult.
In conclusion, the CBCM technique is a powerful tool for gain-of-function studies, and we have developed an optimized protocol for this technique. These results will provide fundamental resources for the development of more effective cell-based microarray systems.
Footnotes
Declaration of Conflicts of Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: this work was supported by s National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIP) (NRF-2014K1A4A7A01074642), Gyeonggi-do, and by a grant from the Bio-industry Technology Development Program (no. 312062-05).
