Abstract
Using genetically modified animals to model neurodevelopmental conditions helps better our understanding of biology underlying these conditions. Animal research has unique characteristics not shared with clinical research, meaning systematic review methods must be adapted to this context. We aim to evaluate the quantity, characteristics, and reporting quality of systematic reviews which synthesise research using genetically modified animals to model neurodevelopmental conditions. On 23 January 2023, we searched PubMed, Embase, and the Web of Science Core Collection to identify systematic reviews of genetic neurodevelopmental condition animal research where the modified gene was one in a list of 102 genes associated with neurodevelopmental conditions identified through large-scale exome sequencing or Fmr1, Mecp2, or Ube3a. Two independent reviewers screened studies based on full text and assessed the reporting quality of relevant reviews using an adapted version of the PRISMA checklist (PRISMA-Pre). Twelve review publications met our criteria. We found mixed levels of reporting: items such as identifying the publication as a systematic review in the title, search strategies, and funding sources being well reported, and others such as protocol registration and data sharing less well reported. We also identified 19 review registrations via PROSPERO, most of which remain unpublished after their anticipated end dates. Systematic reviews are limited by lack of reporting. Increased awareness of reporting guidelines may help authors increase the transparency and reproducibility, and therefore the reliability, of their systematic reviews.
Keywords
Introduction
Up to 40% of neurodevelopmental conditions (NDCs) presenting with intellectual disability (ID), epilepsy, and autism are caused by single genetic alterations, often occurring de novo in that individual (Brunet et al., 2021). These single gene alterations provide an opportunity to develop animal models harbouring those alterations, which may give insights into NDC pathophysiology and treatment more generally, including NDCs with polygenic or environmental causes. Such models can be used to investigate associated cellular, circuit, and behavioural traits. Researchers have developed numerous genetically modified animal models to study NDCs, and research using these models plays a vital role in developing our understanding of the biology underlying NDCs (basic research) and in the testing of potential drug treatments (preclinical research). However, despite extensive research, successful translation of laboratory findings to the clinic is rare (Pankevich et al., 2013; Silverman et al., 2022).
Systematic review provides a well-developed method for identifying areas of poor methodological quality or high risk of bias within research literature. Systematic review is a research method used to summarise and appraise all available evidence related to a pre-specified topic (Egger et al., 2001) and can identify areas for improvements, which might increase internal validity and research rigour and reproducibility.
Clinicians have used systematic reviews to inform evidence-based healthcare since the 1980s. Findings from such reviews have led to considerable improvements in the way clinical trials are conducted and reported (Plint et al., 2006). Over the last two decades, researchers have adopted systematic review methods to summarise and appraise evidence from laboratory animal studies (de Vries et al., 2014) and have achieved similar success in research improvement (McCann et al., 2016; Ramirez et al., 2017). However, the fundamental differences between clinical and animal studies mean that systematic review methodologies must be appropriately adapted (Hunniford et al., 2021). The development of several tools has been instrumental in supporting researchers conducting systematic reviews and meta-analyses of animal studies, including checklists for assessing the risk of bias (Hooijmans et al., 2014) and reporting quality (Macleod et al., 2004) of laboratory animal studies, and a protocol template for systematic reviews of animal intervention studies (de Vries et al., 2015). Despite this, reporting quality of animal systematic reviews is low (Hunniford et al., 2021; Mueller et al., 2014). Currently in development is an extension to the Preferred Reporting Items for Systematic reviews and Meta Analyses (PRISMA) guidelines specifically designed for preclinical systematic reviews (PRISMA-Pre). PRISMA-Pre is a set of reporting guidelines that aim to improve the reporting quality of preclinical systematic reviews (Hunniford et al., 2021).
Here, we aim to evaluate the quantity, characteristics, and reporting quality of systematic reviews which synthesise research using genetically modified animals to model NDCs. We assessed the reporting quality of included systematic reviews using the PRISMA-Pre checklist. We were interested in (1) models with alterations in any of the 102 high-confidence genes identified via large-scale exome sequencing by Satterstrom et al. (2020) and (2) models of Fragile X syndrome (FXS), Rett syndrome (RTT), and Angelman syndrome which have alterations in the Fmr1, Mecp2, and Ube3a genes, respectively. These additional 3 genes are not included in the list of 102 genes; however, they are extensively researched monogenic NDCs.
This review does not synthesise evidence from existing reviews. The rationale behind this review is that by identifying the quantity of and assessing the reporting quality of existing systematic reviews in this area, we can inform guidance on how future systematic reviews within this research field should be conducted. Findings from this review have been used to inform the development of a living evidence summary of research using genetically modified animals to model NDCs, a preliminary protocol for which is available on the Open Science Framework (OSF; Digital Object Identifier [DOI]:10.17605/OSF.IO/GFTZP).
Methods
Protocol registration
Stage 1 of this Registered Report received peer review via Peer Community In Registered Reports (PCI-RR) and is preregistered on OSF (DOI: https://doi.org/10.17605/OSF.IO/952QK). At the time of preregistration, we had completed preliminary searches and optimised our search strategy to inform the development of our study protocol.
Deviations from the preregistration
We have made the following changes from stage 1 of our Registered Report:
Introduction ‘characterised by’ has been changed to ‘presenting with’. ‘autism spectrum condition’ has been changed to autism to align with best practices when referring to autism outlined in ‘Avoiding Ableist Language: Suggestions for Autism Researchers’ (Bottema-Beutel et al., 2021). We had originally described this work as an umbrella review. However, as we do not synthesise results from the systematic reviews we have included, we thought it more appropriate to label our work as simply a review of systematic reviews. To account for this change and to improve grammar, ‘we aim to conduct an umbrella review to identify’ has been changed to ‘we aim to evaluate’. Numbers (1) and (2) have been added to improve the readability of the sentence describing our genes of interest. Following acceptance of our stage 1 Registered Report, we made the decision to include reviews investigating animal models with Ube3a gene alterations. Our rationale for this decision is that, upon searching the literature, we realised that (similar to Fmr1 and Mecp2) Ube3a models are extensively used in NDC research and appear to be more extensively reviewed. Alterations in Ube3a are highly associated with the human condition Angelman syndrome. To account for this change, we added mention of Angelman syndrome to the introduction. ‘identifying the quality and reporting quality’ has been changed to ‘identifying the quality of and assessing the reporting quality’ to improve clarity of our methods. ‘preliminary protocol for which has been preregistered’ in relation to future work informed by this project has been changed to ‘preliminary protocol for which is available’ as our living evidence summary protocol is not an official preregistration but rather a living document in an Open Science Framework project.
Methods Systematic search dates have been added. ‘databases’ has been changed to ‘data sources’ as this is a more accurate description of these resources. ‘autism spectrum condition’ has been changed to ‘autism’ (see introduction deviations). We have added that the full search terms and PRISMA-Pre checklist are also available in the stage 1 Registered Report. Author initials are added to the methods where appropriate to attribute author contributions. We planned to use in house code to retrieve full texts but in practice for this dataset found EndNote’s full text retrieval function more convenient. The number of interlibrary loans we required to access all full texts has been added. We added details on how SyRF displays records in a random order and reviewers are unaware of other reviewers’ decisions. As we are concerned with reporting quality in our work, we thought it appropriate to mention our own use of randomisation and blinding. Angelman syndrome models have been added to the inclusion criteria (see introduction deviations). Our stage 1 Registered Report did not report the complete methods we would use to search for and present PROSPERO data, only mentioning that we would contact authors of relevant PROSPERO registrations. We added these details in stage 2. Additionally, due to time constraints, we were unable to contact authors individually. However, we did search for published versions of PROSPERO registrations marked ongoing in case they had been published.
Bibliographic search
On 23 January 2023, EW conducted a systematic literature search on three electronic data sources: PubMed including Medline (accessed via National Center for Biotechnology Information (NCBI)), Embase (accessed via Ovid), and Web of Science Core Collection.
Our search strategy includes three components: (1) broad terms related to NDCs, ID, epilepsy, and autism, and associated genes; (2) terms related to animal models, and (3) terms related to systematic reviews or meta-analyses. Terms used to identify animal models were taken from van der Mierden et al. (2022) and terms used to identify systematic reviews and meta-analyses were taken from Langendam et al. (2021). Full search terms are given in Appendix 1 and the stage 1 Registered Report.
Where citations appeared in multiple databases, EW removed duplicate versions of the citation using the Automated Systematic Search Deduplicator (ASySD) tool (Hair et al., 2023).
Screening
We uploaded our search results, with duplicate citations removed, to the Systematic Review Facility (SyRF) platform (RRID:SCR_018907; Bahor et al., 2021) for screening, data extraction and management of records. SyRF displays records to reviewers in a random order, and reviewers were unaware of the decisions or annotations made by other reviewers, or in the case of reconciliation, unaware of which reviewer made which decision or annotation.
Two independent reviewers (EW and GC) screened each publication for inclusion, and any disagreements were reconciled by a third independent reviewer (MM). We planned one round of screening, where we screened the full texts of all studies retrieved from our searches against our inclusion and exclusion criteria, to avoid potentially excluding systematic reviews where the decision for inclusion rests on information contained in the full text but not in the abstract (Wilson et al., 2023a).
EW retrieved full text PDF files using the find full text feature in EndNote 20 (with institutional subscription) or via hand-searching. Where we could not access the full text using our institutional subscriptions, we requested the full text via interlibrary loan. In total, we used 24 interlibrary loans. Where the full texts of relevant articles were not written in English, we planned to use Google Translate. However, our search did not return any relevant non-English-language publications.
Inclusion and exclusion criteria
Studies were screened according to the criteria outlined below:
Study design – We only included systematic reviews or meta-analyses that include animal studies, either as a review limited to animal studies or those which included them alongside other study types (e.g. clinical studies). We excluded all other study designs.
Animal models – We only included systematic reviews synthesising research using genetically modified animals to model NDCs where the modified gene appeared on the list of 102 genes identified via large-scale exome sequencing by Satterstrom et al. (2020); or genetically modified animal models of FXS, RTT, or Angelman syndrome; or other genetic models of NDCs characterised by ID, epilepsy, and autism. Animal models may be of any species. A diverse range of other models of NDCs are available but are not the focus of this review and will not be included.
Publication type
We included systematic reviews published in (1) peer-reviewed journals, (2) as conference abstracts, or (3) as preprints (where they are identified in searches). We did not search dedicated preprint servers. We placed no restriction on publication date or language.
Data extraction
Two independent reviewers (EW and MM) conducted conduct data extraction. Discrepancies between reviewer decisions were reconciled by a third independent reviewer (ESS). We carried out data extraction using the SyRF platform and collected the following information:
Bibliographic data
We extracted the names of first authors, year of publication, title, and DOI of each included review.
Characteristics
To understand the purpose and scope of included systematic reviews, we extracted the following characteristics from each:
The aim of each systematic review and the primary research question each review seeks to ask.
Whether the review only included animal studies, or also included clinical or in vitro studies.
Which animal models the review included.
The total number of studies included in the systematic review.
The total number of studies investigating relevant animal models.
Reporting quality
There are two broad approaches to evaluating the quality of systematic reviews: addressing the completeness of reporting or assessing the risks of bias arising from the approaches which were taken. Precise evaluation of the second requires completeness of the first, so these are overlapping but distinct.
Here, we assessed the completeness of reporting (reporting quality) of each included systematic review using the 46-point checklist developed by Hunniford et al. (2021). The checklist is adapted from the PRISMA guidelines for systematic reviews and is more specific to systematic reviews of animal studies. However, the checklist is not currently an official extension to PRISMA. Although it is not recommended to use the general PRISMA guidelines to assess in vivo systematic review reporting quality, the adapted checklist for preclinical systematic reviews has been designed for this purpose. There are no validated tools for assessing risks of bias in systematic reviews of in vivo research, and we will not conduct a formal assessment of risk of bias.
The checklist items are written in full in Appendix 2 and in the stage 1 Registered Report.
PROSPERO search
In addition, we searched the PROSPERO database to identify the status of ongoing but unpublished preregistered systematic reviews. On the 25 September 2023, we searched PROSPERO using the following terms: neurodevelopment OR neurodevelopmental OR autism OR autistic OR ASD OR intellectual disability OR epilepsy. We limited this search to reviews of animal studies for human health protocols.
Two reviewers (EW and GC) screened each of the search results based on our inclusion criteria, and disagreements were reconciled by a third reviewer (ESS). For each relevant review, EW noted the data of registration, expected start date, expected end date, and current stage of the review. We had planned to contact authors of PROSPERO registrations to establish if their review has been published; however, we did not complete this due to time constraints. We did, however, search for published versions of preregistrations labelled ‘ongoing’ in the PROSPERO system.
Data synthesis
We did not conduct a meta-analysis or perform statistical analyses on our data. We have presented a descriptive summary of the bibliographic, characteristics, and reporting quality data extracted from each included systematic review. We scored each included systematic review using the PRISMA-Pre checklist and have provided summary graphs detailing which items of the checklist each review met. In addition, we assessed which tools are currently being used to conduct systematic reviews in this field, including the tools used to screen studies (Checklist Item 6a), extract numerical data (Checklist Item 17a), and measure study quality or risk of bias (Checklist Item 19).
Results
Systematic search results
Our systematic searches of PubMed, Embase, and Web of Science Core Collection returned a total of 1,753 records (441 from PubMed, 640 from Embase, and 672 from Web of Science Core Collection; Figure 1). 428 records were removed using the ASySD tool, leaving 1,325 unique records remaining. Of these, only twelve records met our inclusion criteria via full text screening. Ten of the included records were peer-reviewed journal articles and two were conference abstracts. We did not identify any preprints through our searches. The same systematic review was identified as a conference abstract (Zhang and Spruyt, 2022) and a peer-reviewed journal article (Zhang et al., 2021). As our evaluation is primarily concerned with reporting quality of these publications, we assessed the conference abstract and journal article as separate publications.

Flow chart showing the number of studies identified through searches of (a) PubMed, Embase, and Web of Science Core Collection and (b) PROSPERO, and the number of studies included after screening.
Characteristics of included reviews
All the reviews included in this evaluation were published between 2016 and 2022 (Table 1). Most of the reviews, including both conference abstracts, were published in 2022.
Characteristics of the 12 systematic reviews included in this evaluation.

SR = Systematic review. MA = Meta-analysis. N included studies shows the total number of studies included in each review and the total number of animal studies (in brackets).
indicates a journal article and conference abstract of the same review.
Only two reviews (a journal article and a conference abstract) conducted a meta-analysis alongside their systematic review. The remaining 10 included only qualitative synthesis of evidence.
Three of the reviews (all journal articles) included studies with animal or human participants, while the remaining reviews included only animal studies. None included in vitro or in silico work alongside animal studies.
Each of the reviews covered a distinct research question. The review aims and research questions were coded into the following topics: sleep disturbances; microbiome differences; stress (specifically the hypothalamic-pituitary-adrenal axis); neurobiology; sensory differences (specifically auditory and olfaction); model suitability; and biomarker identification (Table 1).
A variety of genetic alterations were included in the reviews evaluated, but this only represented 11 of the total 105 genetic modifications of our interest. Many reviews looked at multiple genes. The genes assessed were Fmr1 (6 reviews); Mecp2 (5 reviews); Ube3a, Pten, and Shank3 (4 reviews each); Adnp (2 reviews); and Cacna2d3, Chd8, Kcnma1, Nrxn1, and Tbr1 (1 review each). Fmr1, Mecp2, and Ub3ea alterations are strongly associated with FXS, RTT, and Angelman syndrome, respectively. In addition, Pten, Shank3, Adnp, and Nrxn1 are associated with Cowden syndrome, Phelan-McDermid syndrome, Helsmoortel-Van der Aa syndrome, and Pitt-Hopkins-like syndrome 2, respectively (SFARI Gene, 2024). Cacna2d3, Chd8, and Tbr1 are associated with autism and ID generally but not with any named syndromes.
The number of studies included in each review varied greatly, ranging from 13 studies to 531 (mean = 98, median = 26).
Evaluation of reporting quality
The PRISMA-Pre checklist items are in six categories divided by the section of a manuscript which they refer to: title, introduction, methods, results, discussion, and other. A PRISMA-Pre checklist for abstracts does not yet exist, so we were mindful when evaluating the two conference abstracts and have marked reporting as not applicable for items where reporting would be unfeasible, for instance, the inclusion of a PRISMA flow diagram or summary table of included studies.
Items related to reporting in the title
Eleven out of the 12 reviews identified the report as a systematic review, and 11 identified that the report contained animal data (Figure 2).

Level of reporting of items within the title section.
Items related to reporting in the introduction
In the introduction section, all the reviews described the human condition being modelled, but only six reviews provided an explicit statement of the questions being addressed (Figure 3). None of the included reviews focused on the effects of interventions, so the reporting item related to describing the biological rationale for testing the intervention was not applicable for any of the reviews.

Level of reporting of items within the introduction section.
Only two reviews indicated whether a protocol was registered (both were registered a priori on the PROSPERO platform). However, only one indicated whether any deviations were made to the protocol (Figure 4).
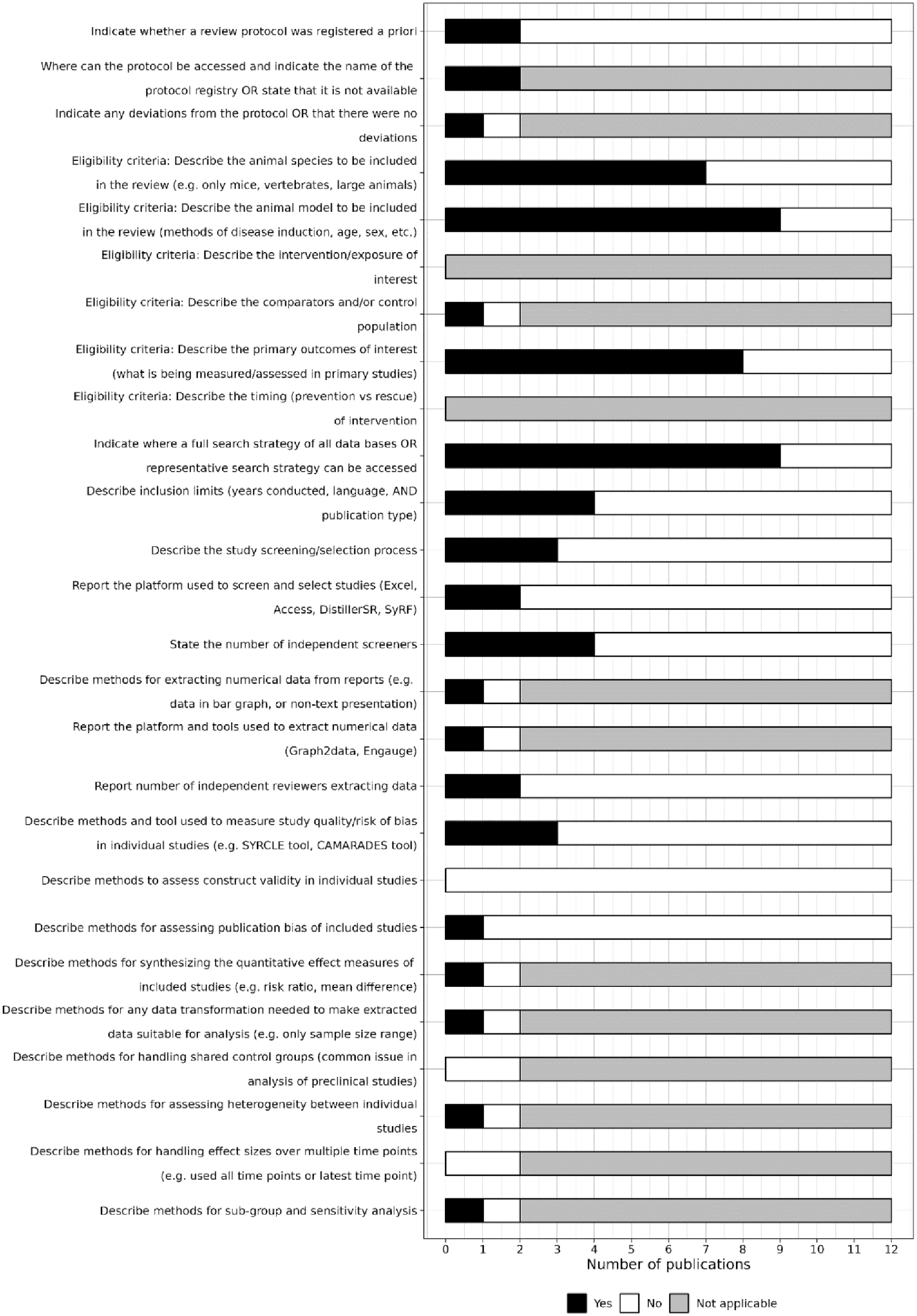
Level of reporting of items within the methods section.
Items related to reporting in the methods
The eligibility criteria related to animal species, models, and outcomes of interest were reported in seven, nine, and eight of the reviews, respectively (Figure 4). Items relating to reporting interventions of interest and the timing of intervention delivery were not applicable to any of the reviews.
Nine of the 12 reviews gave the full search strategy of all databases searched, and only four described inclusion limits added to the search (Figure 4). Three reviews described the study selection process, two reported the platform used for screening (Rayyan and Microsoft Excel), four reported the number of independent screeners, and two reported the number of reviewers extracting data.
Three reviews reported methods for assessing the risk of bias of the studies they included (one used both Collaborative Approach to Meta-Analysis and Review of Animal Data from Experimental Studies (CAMARADES) and SYstematic Review Centre for Laboratory animal Experimentation (SYRCLE) tools, one used only the SYRCLE tool, and one used an unknown tool), none reported methods for construct validity assessment, and only one reported the methods for assessing publication bias (Figure 4).
Several of the reporting criteria for the methods section are related to meta-analyses, so were only applicable for the two reviews that reported a meta-analysis. One review reported eligibility criterion related to any controls or comparators, described the methods of data extraction, reported the platforms or tools used to extract numerical data (Rayyan), and described methods for synthesising the quantitative effect measures of included studies, any required data transformations, heterogeneity assessment, and sub-group or sensitivity analysis (Figure 4). The other review, which showed limited reporting, was a conference abstract and may have been affected by abstract word limits. Neither of these meta-analysis reported methods for handling shared control groups or effect sizes over multiple time points.
Items related to reporting in the results
All 12 reviews reported the number of individual reports included in the review (Figure 5). Nine provided a summary table of individual studies with data and references, and nine also included a PRISMA flow diagram. These items were not applicable for the two conference abstracts. For reporting of study characteristics, 8 reviews reported the animal species, 10 reported model details, 5 reported a measure of the sample size, and 4 reported individual study designs or intentions. Reporting of intervention details was not applicable for any review. Of the three reviews which reported methods to conduct a risk of bias assessment, and one review which reported methods to analyse publication bias, all reported the results of these assessments (Figure 5).

Level of reporting of items within the results section.
The meta-analysis published in a peer-reviewed journal article reported the number of studies included in quantitative analysis, the outcome effects of included studies and associated confidence intervals, a measure of heterogeneity between included studies, and results from sub-group or sensitivity analysis (Figure 5). These items were not reported in the conference abstract.
Items related to reporting in the discussion
Two of the three reviews that conducted a risk of bias assessment discussed the impact of this on the included studies (Figure 6). Six reviews discussed the limitations of the individual studies they included, but only three discussed the limitations of the review itself.

Level of reporting of items within the discussion section.
Items related to reporting in other sections
Finally, 10 reviews (all the journal articles) reported the source of funding for the review, and 4 reviews included a data availability statement (Figure 7).
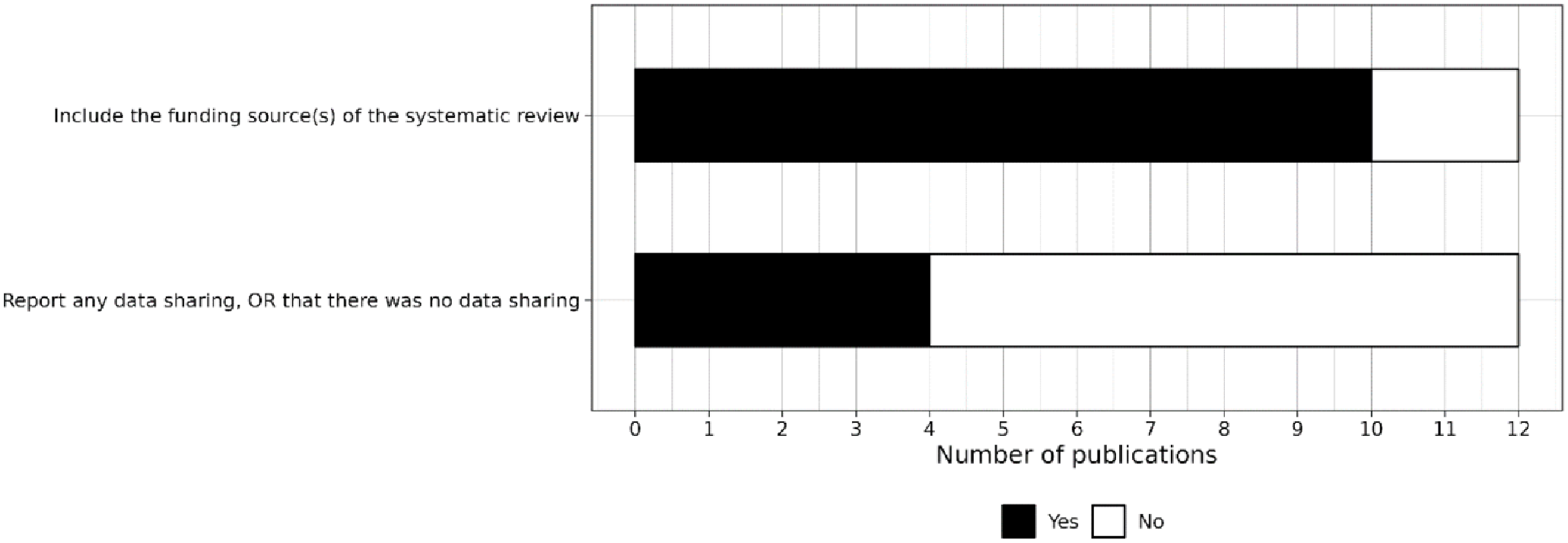
Level of reporting of items within other sections.
Identification of ongoing reviews via PROSPERO
Our search of PROSPERO identified 82 ongoing or completed reviews. After screening, we identified 19 review registrations fitting our inclusion criteria (Table 2). It was often difficult to tell if reviews would include genetic models, so we opted to be overinclusive, only excluding from our analysis if it was clear that genetic models would be excluded. Many of the protocols did not specify specific models of interest, instead often stating that they will include ‘all’ animal models of NDCs or ‘all’ animal models of autism.
Anticipated start and end times, and time between each, of the 19 review registrations identified via PROSPERO.

The registered review appears in the main review as a journal article.
The registered review appears in the main review as a conference abstract.
The anticipated end date has not past at the time of analysis. Time of analysis (which is also used to calculate days elapsed) was 5 October 2023.
Only two of these reviews were published at the time of our analysis; these were the two reviews from our main search which reported a protocol. The remaining 17 reviews were marked as ongoing. We did, however, identify that the unpublished conference abstract we found in our main review was registered in PROSPERO, although this registration was not reported in the conference abstract.
All the registrations were made between 2018 and 2023. Six registrations, all of which are ongoing, were made on the same date by the same first author. The mean number of days authors anticipated to complete reviews was 291 (minimum 52 days and maximum 715 days). Some of the ongoing reviews anticipated a very quick completion time (less than 100 days).
The two published reviews were registered in 2020 and 2021, and authors anticipated 508 and 395 days, respectively, to complete their reviews. In practice, they took 528 days and 347 days, respectively, from start date to journal submission (Table 2).
Discussion
Our searches identified 10 published systematic reviews, 2 conference abstracts, and 17 ongoing reviews related to genetic animal models of NDCs. Most of the reviews are recent, with the earliest published in 2016. The first PROSPERO registration was in 2018; before 2018, PROSPERO did not allow registration of animal systematic reviews (Pieper and Rombey, 2022). We made the decision to include both Zhang et al. publications (one peer-reviewed article and one conference abstract) in our analysis, as despite the publications referring to the same systematic review research, they are separate publications and their reporting quality can be assessed separately.
From the published literature, we can see that reporting against the PRISMA-Pre checklist was mixed. Reporting in the title to identify publications as (1) systematic reviews and (2) related to animal research was overall good. One of the reasons we chose to screen based on the full text rather than one round of title and abstract screening was that we were unsure if this would cause us to erroneously exclude publications with unclear titles or abstracts.
Within the introduction section, authors consistently reported descriptions of the conditions being modelled, but only half explicitly reported the review aims. The lack of clear reporting in this area made it difficult to code the review aims in our summary table and may impact the use of individual reviews if their purpose is not clear.
Methods and results reporting were mixed. Few reviews reported having a preregistered protocol. Those that did preregister did so through PROSPERO, a dedicated repository for protocols of systematic reviews related to human health. Protocol registration is often considered a fundamental step in the systematic review process, to ensure that the review methodology is transparent and reduces the risk of bias being introduced in the systematic review (Soliman et al., 2020).
Eligibility criteria and search strategies were reported well overall, with room for improvement, while reporting of methods to screen studies was limited. The reporting of the number of studies included, and inclusion of summary tables and PRISMA flow diagrams were better. The small number of reviews that included a meta-analysis limits our ability to interpret reporting of items related to quantitative analysis with any confidence.
Few of the reviews conducted a risk of bias assessment, meaning the potential impact of risk of bias within primary studies was often not discussed. Many tools have been developed to assess the quality of evidence included in systematic reviews of animal data and associated risks of bias, including the CAMARADES checklist (Macleod et al., 2004) and the SYRCLE risk of bias tool (Hooijmans et al., 2014). Only one of the reviews that did not conduct a risk of bias assessment stated their reasoning, that the reporting of the studies they included was too poor to conduct a risk of bias assessment. However, the purpose of a risk of bias assessment is to determine precisely the reporting quality and present this evidence (Soliman et al., 2020). Systematic reviews of animal studies often observe very limited reporting quality and present that many primary studies they include are at high risk of bias (Soliman et al., 2020).
Only half of the reviews discussed the limitations of the studies they had included, and a quarter discussed the limitations of the review itself. Systematic reviews, just like any other form of research, are susceptible to research bias. Systematic reviews often play a key role in research and healthcare decision-making, so it is important that the findings of reviews are as free from bias as possible. As discussed above, protocol registration is one method for mitigating at least some of this bias and improving the transparency of the work; however, this does not negate the need to discuss limitations.
Finally, funding sources of the reviews were reported in all peer-reviewed journal articles, likely because of publishing policies. Data availability statements, which should report whether data are available or not available, were included in less than half of the reviews, despite this also being a requirement for most journals.
Systematic review is only one method used to bring understanding to translational challenges within biomedical science. In NDC research, there are many potential explanations for this translational barrier, including limitations in how research is designed, conducted, and reported (Wilson et al., 2023b), and research that is rooted in the medical model and fails to take into consideration advances in our understanding of NDCs achieved through neuroaffirmative research approaches (Heraty et al., 2023).
Systematic review of animal studies of NDCs is still an emerging activity. There are fundamental differences between clinical and animal research, such as the experimental methods used, number of participants/animals per paper, and number of relevant papers retrieved in systematic literature searches, meaning systematic review methods have to be adapted from the clinical space to fit animal research. Systematic reviews of animal research related to other biomedical conditions are ahead in this space, and many resources have been developed to help researchers conduct their reviews with rigour (de Vries et al., 2014, 2015; Hooijmans et al., 2014; Macleod et al., 2004; Sena et al., 2014). Unfortunately, our findings demonstrate that study authors of the reviews we have included have not engaged with much of this guidance and the available resources.
Systematic reviews are an empirical form of research, and if we are to rely on their findings, they must be conducted rigorously (Sena et al., 2014). When assessing the quality of research, including research like systematic reviews, we cannot directly assess the methodological quality; therefore, we assess the reporting quality as a proxy. Where the reporting of systematic review is limited, our understanding of the review’s methodological quality is limited, and in turn we have less certainty around the reliability of the review’s findings. We identified 17 systematic reviews in progress, demonstrating that this is an area of research growth. Moving forward, we encourage authors to make use of existing resources to maximise the value and impact of their work. Table 3 summarises some key resources for conducting systematic reviews of animal studies.
Resources for conducting systematic reviews of animal studies.

Most of the systematic reviews we identified were published in 2022, and likely conducted during 2020–2021. We cannot know with any certainty if laboratory closures during the COVID-19 pandemic influenced the uptick in reviews conducted and whether this activity will continue to rise. However, it is important that the reporting quality, as well as the design and conduct, of systematic reviews in this area follow the advances in systematic review methodology we see in other areas, including focal cerebral ischaemia, so that they can have similar effects on primary research direction, conduct, and policy (McCann and et al, 2016).
Strengths and limitations of this study
A major strength of our study is that, by preregistering our study using the Registered Report format, our rationale and proposed methods have been peer-reviewed, meaning we have been able to make improvements to our proposed study design prior to beginning our research.
To maximise the sensitivity (recall) of our search, we used broad search terms related to NDCs, ID, epilepsy, and autism, in parallel with terms related to individual associated genes, and screened studies based on full text to avoid erroneously excluding systematic reviews which do not report their inclusion criteria in their abstract.
A limitation, however, is that by only searching primarily English-language databases we may have missed non-English reviews (possibly evidenced by the lack of such reviews in our results).
Although we identified protocols for ongoing reviews via PROSPERO, we did not analyse the reporting of these protocols. In addition, from our results, we know that the majority of published reviews were not preregistered, suggesting that we may be unaware of many more ongoing and unpublished reviews. We used a draft extension of the PRISMA guidelines specifically designed for systematic reviews of animal studies (PRISMA-Pre; Hunniford et al., 2021; also included in Appendix 2) to assess the reporting quality of included systematic reviews. This extension remains in draft, is not yet an official extension to PRISMA and may be subject to change. The checklist was designed to improve the reporting quality of journal articles. In hindsight, it may not have been appropriate for us to apply the checklist to conference abstracts. Conference abstracts have heavily limited word counts and their purpose is to act as short communication pieces where further details are given in presentations (oral or poster).
In addition, many aspects of the PRISMA-Pre checklist were not applicable for this data set. All the systematic reviews identified were interested in mechanistic studies, so intervention-related questions within the checklist were not relevant. In addition, only two of the reviews included a meta-analysis. An improvement to the PRISMA-Pre checklist may involve adding additional subheadings to separate the meta-analysis-related questions from ‘core’ systematic review reporting items.
Conclusion
To conclude, recent years have seen the publication of a diverse range of systematic reviews investigating genetic animal models of NDCs. Systematic evaluation of published research can help summarise research findings and strengthen evidence most effectively when effort is taken to minimise bias within the review itself. Within our sample, the reporting of published systematic reviews is mixed. We rely on clear and transparent reporting as a proxy to determine sound methodological quality. Therefore, researchers conducting systematic reviews in this space should engage with systematic review guidance, where available, to strengthen and ensure the robustness and reliability of their reviews. Increased awareness of reporting guidelines may help authors plan and report their systematic reviews with more details to increase their transparency and reproducibility.
Footnotes
Appendix 1
Appendix 2
Author contributions
Data availability
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: EW is funded by a Simons Initiative for the Developing Brain PhD studentship (SFARI #529085).
Ethics approval
This study only examined data from published systematic reviews and systematic review preregistrations, so ethical approval is not required.
