Abstract
Introduction:
Periodontal disease (PD) is closely linked to systemic health, with established associations with chronic conditions (eg, diabetes, cardiovascular disease). However, most predictive models rely solely on dental data, limiting the consideration of systemic factors such as medical conditions.
Objectives:
This study aimed to enhance PD risk prediction by using linked electronic dental records (EDRs) with electronic health records (EHRs) and machine learning (ML).
Methods:
We used EDR data from 20,946 adult patients at Temple University School of Dentistry’s (2022–2023) axiUm®, linked with medical data (physician documented) from the Pennsylvania Health Share Exchange. The dataset includes demographics, dental diagnoses, medical history, medications, procedures, and social determinants of health. The target variable was PD. Because EHR data are not research ready, extensive preprocessing was required (eg, 1 patient may have 400+ medical codes, which ML/statistical models cannot process directly). To prepare for artificial intelligence/ML, we developed 5 automated feature reduction approaches to retain rich information while reducing variables. After preprocessing, 106 features were retained as independent variables. ML models (Gaussian Naive Bayes, Random Forest, LightGBM, XGBoost) were trained using cross-validation across 5 experimental strategies, including (1) features selected via chi-square test, (2) raw data (without extensive processing), (3) aggregated data, (4) systemic disease complexity system, and (5) EHR-only data. Model performance was assessed using sensitivity, specificity, and area under the curve (AUC).
Results:
The chi-square–selected features yielded the best performance: 85% specificity, 67% sensitivity, and 84% AUC. Although adding medical conditions did not significantly improve overall performance, key conditions (eg, cardiovascular diseases, endocrine/metabolic disorders, renal diseases, respiratory conditions, hematologic disorders, etc) contributed meaningfully to PD risk prediction. EDR factors (oral hygiene, periodontal treatment, brushing, flossing, smoking, and American Society of Anesthesiologists classification) dominated prediction.
Conclusion:
Although dental factors remained dominant predictors, strong systemic–oral health associations were observed. Future studies should validate these findings by integrating medical and dental records.
Knowledge Transfer Statement:
The results of this study can guide clinicians and policymakers in identifying patients at increased risk of periodontitis by integrating medical and dental records. This approach supports earlier interventions and highlights the importance of systemic health in oral disease management. It also demonstrates the potential of artificial intelligence–based prediction models to improve personalized care and promote interdisciplinary collaboration for better overall health outcomes.
Keywords
Background
Oral and systemic health are closely interconnected, with strong evidence linking periodontal disease (PD) to conditions such as cardiovascular disease, diabetes, arthritis, adverse pregnancy outcomes, and respiratory illnesses (Huang et al 2024; Zhao et al 2025). Some studies suggest a bidirectional relationship, in which PD may not only be associated with systemic diseases but could also contribute to their progression (Centers for Disease Control and Prevention 2024). Given these connections, early identification and prevention of PD are essential for improving both oral and overall health (Isola et al 2023). Predictive models offer a promising approach to identifying high-risk individuals before disease onset or progression (Toma and Wei 2023; Khalifa and Albadawy 2024). While electronic dental record (EDR) data provide valuable insights, existing prediction models primarily focus on dental factors, overlooking critical medical information from their physician’s office, such as diagnoses, medications, and procedures in electronic health records (EHRs) (Teza et al 2023; Ferrara et al 2024; Tokede et al 2024; Swinckels et al 2025). Integrating EHR with EDR may offer a more comprehensive patient profile, yet medical and dental records remain siloed due to system incompatibilities, data standardization issues, and lack of interoperability (Beserra et al 2022; Gurupur et al 2024). As a result, to the best of our knowledge, no existing prediction model effectively uses linked EHR and EDR data to improve PD risk assessment (Farina et al 2023; Tokede et al 2024; Swinckels et al 2025).
Various PD prediction models have been developed for the past 2 decades using different methodologies, including rule-based systems and artificial intelligence (AI)–driven machine learning (ML) (Farina et al 2023; Ferrara et al 2024; Swinckels et al 2025). Rule-based systems rely on predefined expert-driven conditions that lack adaptability, whereas ML models learn from large datasets to identify complex patterns and interactions (Tokede et al 2024). Unlike rigid rule-based approaches, ML continuously refines predictions based on new data, leading to more personalized and accurate risk assessments (Farina et al 2023; Swinckels et al 2025). However, ML models are only as effective as the quality and completeness of the training data (Sanyal et al 2021). Ensuring data accuracy, standardization, and integration is crucial for developing robust predictive models (Budach et al 2022). EHR data offer significant advantages over surveys and registry datasets, including real-time availability, longitudinal tracking, and integration of medical, behavioral, and demographic factors (Rudin et al 2020; Ghildayal et al 2024). By incorporating these additional variables, ML-based predictive models have the potential to enhance the detection, prevention, and early intervention strategies for PD (Bashir et al 2022; Zhang et al 2025).
Despite promising advances, existing prediction models lack access to up-to-date medical histories from linked EHRs, relying instead on self-reported histories, which studies have shown to be unreliable (Sulieman et al 2022). In our previous work, we extracted free-text EDR data into structured formats, phenotyped PD diagnoses across multiple record sections, and improved data quality before developing a PD prediction model (Patel et al 2022b; Patel et al 2022c). Using only EDR data, our model demonstrated moderate–high classification performance based on PD severity rather than a binary outcome (Patel et al 2022b). The present study aimed to improve PD prediction by integrating linked EHR and EDR data, incorporating medical histories, medications, and procedures. We hypothesize that incorporating comprehensive medical data will enhance model performance and provide deeper insights into systemic contributors to PD risk.
Methods
Datasets
We obtained data from 20,946 patients using linked EHR and EDR. The EDR dataset was retrieved from Temple University Kornberg School of Dentistry’s axiUm® database. We included adult patients who underwent at least 1 comprehensive oral examination or periodic oral evaluation with complete documentation of periodontal charting findings and periodontal diagnoses between January 2021 and December 2022. Medical information for these dental patients was obtained from their EHR through the Health Share Exchange. The final dataset comprised cross-sectional data, including patient demographics, dental diagnoses, periodontal charting findings, and dental treatments from the EDR, as well as medical history, procedures, lab values, and medication records from the EHR documented by physicians (for additional details, see Appendix: Linkage Process). See Figure 1 for the overall workflow.
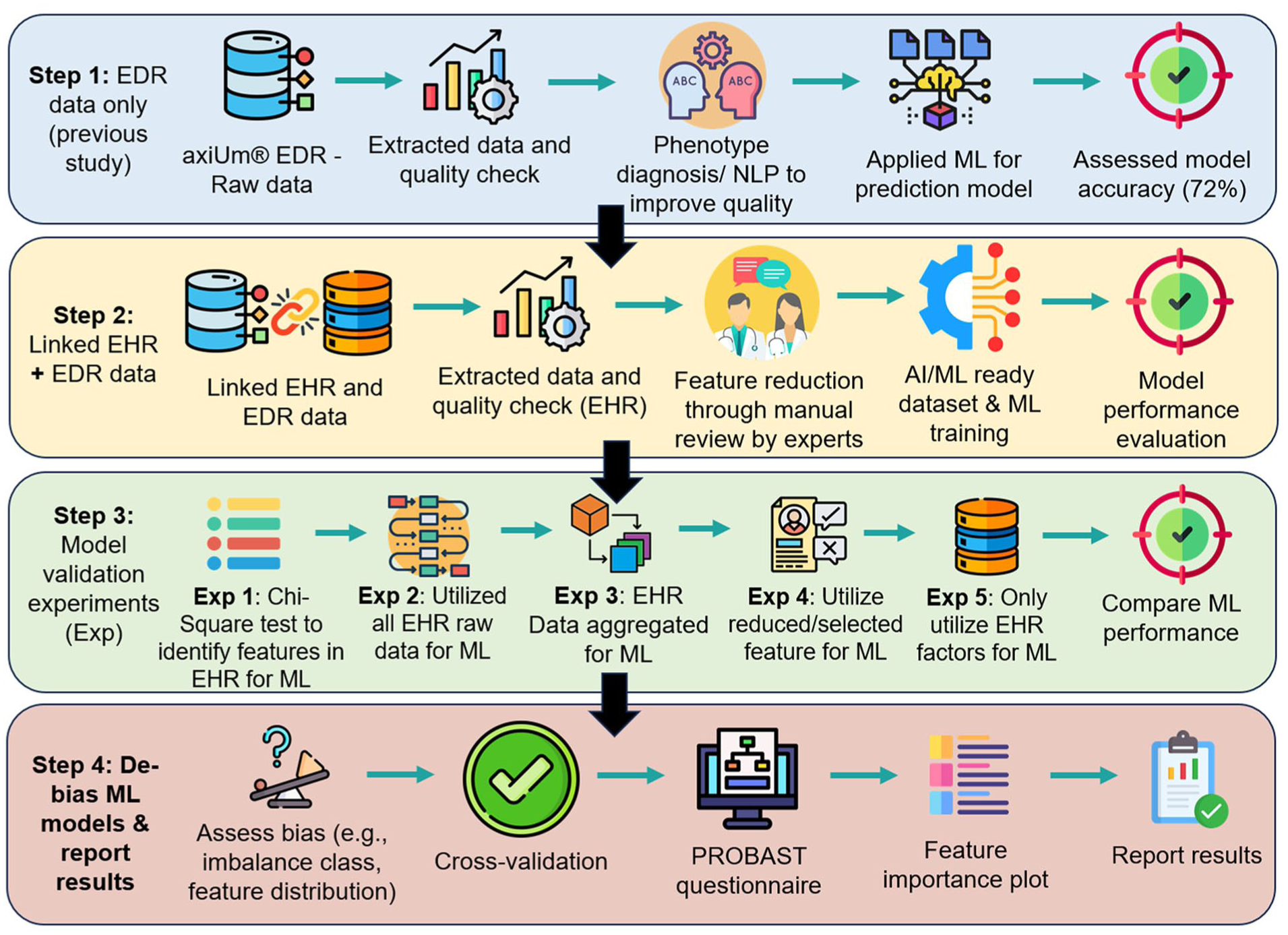
Overall workflow and study design.
Prediction versus Association
We clarify the distinction between prediction and association, as terminology may vary across clinical dentistry, dental research, and data science domains. Our study does not use longitudinal data suited for causal inference or prognostic modeling. Instead, we apply ML in a diagnostic framework to estimate the presence and severity of PD based on cross-sectional EHR/EDR data. This aligns with the diagnostic prediction framework outlined by van Smeden et al (2021), in which the goal is to infer a current unknown state using observable features, not to forecast future outcomes. Since predictions are made on “unseen” test data, the process qualifies as a prediction rather than a simple association, as the ML method uses distinct training and testing datasets. The model classifies PD status and severity in the test set, which it has not seen during training. This approach is common in data science, especially with cross-sectional data.
We recognize concerns about using “prediction” in nonlongitudinal contexts. Breiman (2020) described this as a divide between explanatory modeling, which assumes data-generating mechanisms, and algorithmic modeling, which focuses on predictive performance. ML falls into the latter category, data driven and flexible, often surfacing meaningful associations without mechanistic explanations (Breiman 2020). The approach proposed in this study aligns with this predictive tradition. As Kass (2021) noted, explanation and prediction can coexist, depending on the modeling goal. Statistical modeling, in this context, involves both quantifying uncertainty and assessing utility, even in the absence of causal claims (Kass 2021).
Therefore, our objective is not causal inference or prognosis but identifying variables associated with PD severity and assessing their predictive value using real-world data. Many studies in medical informatics and data science domains have used cross-sectional data to develop prediction models (Daines et al 2019; Antwi et al 2020). While “association” or “correlates” may better describe the relationships, the use of ML to estimate unobserved outcomes from observed features justifies a careful use of the term “prediction” in a diagnostic sense.
Data Processing for an AI/ML-Ready Dataset
Although EDR and EHR data contain valuable clinical information, they are primarily collected for patient care rather than research or AI/ML applications (Tsai et al 2020). As a result, significant preprocessing is required before they can be used for statistical analysis or ML. PD diagnoses, for example, may appear in free-text clinical notes and structured fields or can be obtained from periodontal charting data when structured diagnosis codes are missing (Patel et al 2022c). This inconsistency arises because dentists are reimbursed based on procedure codes rather than diagnoses, leading to variable completeness in the EDR. In addition, critical patient information, such as social habits, is often documented in free text, further complicating data extraction (Patel et al. 2022b).
To optimize EDR data for research and AI applications, our previous studies developed, tested, and validated automated tools to
convert free-text data into structured categories or codes for statistical and ML analysis (Patel et al 2022a; Patel et al 2022c; Patel et al 2023),
assess and enhance the completeness and depth of recorded information (Patel et al 2022b),
evaluate the concordance between diagnoses documented in different sections of the EDR (eg, structured codes vs clinical notes) (Patel et al 2022a; Patel et al 2022c), and
perform data imputation to manage missing values (Patel et al 2022b).
These preprocessing steps produced high-quality, structured EDR data, ensuring its suitability for PD research and AI-driven predictive modeling.
EHR Data Processing
Like EDR data, we cleaned and preprocessed EHR data to ensure its suitability for ML and statistical modeling. Many patients had more than 400 diagnosis codes ((International Classification of Diseases[ICD]), procedures, and medications, making the dataset highly dimensional and impractical for direct ML applications. Excessive features can lead to overfitting, computational inefficiencies, and reduced interpretability, necessitating feature reduction techniques to retain essential clinical information while improving model performance (Ying 2019).
To address this concern, we developed feature-reduction methods to streamline EHR data while preserving key clinical insights. For instance, medical diagnoses are often recorded under multiple ICD codes for the same condition. Diabetes mellitus, for example, may appear as follows: E11.9: Type 2 diabetes mellitus without complications E11.65: Type 2 diabetes mellitus with hyperglycemia E13.9: Other specified diabetes mellitus without complications
To enhance interpretability and consistency, we collapsed these variations into a single feature labeled “diabetes” to ensure accurate modeling without redundancy. Implementing feature reduction techniques improved computational efficiency, reduced noise, and allowed the model to focus on clinically meaningful variables, ultimately enhancing predictive accuracy (for additional details, see Appendix: Feature Reduction for EHR Data).
Target Variable/Dependent and Independent Variables for ML
Our target variable was periodontitis, classified into healthy, mild (stage I), moderate (stage II), and severe (stages III and IV). We did not consider grading of the disease in the analysis (for data quality and the distribution of PD across various stages, see Appendix Table 1; for aggregated PD classification for ML and methods for improved data quality and calibration practices at Temple University Kornberg School of Dentistry, see Appendix section page 2). We obtained the target variable from the EDR using our phenotyping approach as described in our previous study (Patel et al 2022c), while independent variables were retrieved from linked EHR/EDR datasets. These included demographics (age, gender, race, ethnicity), insurance status, medical history, medications, dental history, treatments, oral hygiene habits (brushing, flossing), and behavioral factors (smoking, alcohol, drug use). We also included biomarkers such as HbA1c levels and systolic and diastolic blood pressure values for each patient. We did not include any periodontal findings or any markers that identify disease in the independent variable list. To minimize confounding, we removed highly correlated features such as diagnosis and medications using the correlation coefficient model (see Appendix Fig 1). The final dataset included 109 independent variables for ML analysis.
Data Imputation
To address missing values in our data, we used multiple imputations by chained equations to estimate missing values while preserving variable relationships, minimizing information loss, and creating a complete dataset for ML (van Buuren and Groothuis-Oudshoorn 2011). Because we evaluated XGBoost with other key ML models, incorporating imputation was necessary, particularly since models such as Gaussian Naive Bayes (GNB) and standard Random Forest (RF) do not handle missing data effectively (Bashir et al 2022; Bates et al 2024). Hence, imputation allowed us to retain a large portion of our dataset (for additional details, see Appendix: Data Imputation & Missingness and Appendix Table 2). This ensured that we did not introduce bias and potentially skew the performance of the model.
ML/AI Training, Testing, Debiasing, and Interpretation
We encountered imbalanced data among different periodontitis categories. Therefore, we addressed data imbalance using undersampling and the synthetic minority oversampling technique (SMOTE) to prevent bias toward the majority class and improve generalizability (Junsomboon and Phienthrakul 2017). We trained and tested GNB, RF, LightGBM, and XGBoost models (see Appendix: Machine Learning and Model Explainability). To improve reliability and reduce bias, we applied 10-fold cross-validation and optimized hyperparameters (see Appendix: Handling Overfitting and Appendix Table 3) (Bashir et al 2022; Bates et al 2024). We evaluated bias using the Prediction model Risk of Bias Assessment Tool (PROBAST) and Transparent Reporting of a multivariable prediction model for Individual Prognosis or Diagnosis (TRIPOD) tool, which assesses participants, predictors, outcomes, and analysis (see Appendix: PROBAST Checklist & TRIPOD Checklist) (Collins et al 2021). The PROBAST checklist is designed to assess the risk of bias and applicability of diagnostic and prognostic prediction model studies. The TRIPOD checklist guides the transparent reporting of studies that develop, validate, or update prediction models. Furthermore, ensemble techniques were not considered, as model interpretability through SHAP cannot be reliably performed from ensemble models, as the SHAP values for each component model would need to be interpreted independently, thereby reducing the interpretability of the already complex black-box models (Lundberg and Lee 2017).
Feature Selection Experiments and Model Validation
We conducted 5 experiments to validate the model through feature selection. Although ML models can handle complex datasets with multiple independent variables, overfitting remains a concern (Bashir et al 2022; Bates et al 2024). We conducted these 5 experiments to ensure consistency among the ML models, determine the most informative features, and compare the influence of different feature selection strategies on model outcomes. Although ML models can process high-dimensional data, they are prone to overfitting if irrelevant or redundant features are not removed. By testing various preprocessing and feature aggregation methods, we reduced noise, enhanced model generalizability, and determined the contribution of EHR-derived variables to PD prediction. These experiments helped us explore which features consistently emerged as strong predictors and evaluate how model performance varied when including or excluding medical versus dental information (Bashir et al 2022; Bates et al 2024).
In experiment 1, we used chi-square tests with a significance threshold of P < 0.05 to identify medical conditions that were significantly associated with periodontitis. We then included only those medical diagnoses and medications that showed significant associations in the predictive model. Chi-square testing is often used as a method of feature reduction in ML, where significant variables are kept and nonsignificant variables are excluded (Dissanayake and Johar 2021).
In experiment 2, we imported semiprocessed EHR data with ICD-10 codes without applying feature reduction methods. This helped us determine which medical conditions were highly correlated with PD outcomes before performing any data simplification.
In experiment 3, we used feature reduction methods to aggregate 10,671 ICD codes, 60,570 medications, and 4,316 procedures to 28 unique diagnosis categories, 58 medication categories, and 150 procedure categories. This was performed to minimize noise and eliminate confounders in the dataset, which can often mislead the model and degrade its performance (Ying 2019).
In experiment 4, we ranked patients based on the number of grouped medical conditions and medications in their medical records to measure systemic health complexity. For example, a patient with ICD codes in 5 diagnosis categories received a complexity score of 5.
In experiment 5, we removed all dental factors from the model and used only EHR data to predict PD. This allowed us to assess how well medical history alone could predict PD outcomes.
For each experiment, we split the data into 70% training and 30% testing sets, using stratified sampling to maintain the target variable’s distribution. To compare model performance across these experiments, we measured sensitivity (correctly identify true positives), specificity (correctly identify true negatives), recall (proportion of actual positives correctly identified), precision (proportion of predicted positives that are true positives), F1 score (harmonic mean of precision and recall), accuracy (overall proportion of correct predictions), and area under the curve (AUC) (how well the model differentiates between positive cases from negative cases) (Rainio et al 2024).
Results
Patient Characteristics
The study included 20,946 patients, with most aged 50 to 70 y (45%), Black/African American (45%), and 57% female. Medicaid covered 42% of patients, while 33% were self-pay, and 23% had private insurance. Smoking was reported by 23% of patients, 20% consumed alcohol, and 13% used recreational drugs. Oral health findings showed a mean DMFT index of 5.07 (standard deviation [SD]: 4.6, confidence interval [CI]: [5.01, 5.14]), a DMFS index of 14.15 (SD: 15.08, CI: [13.94, 14.35]), and an average of 24 teeth (SD: 6.59, CI: [24.06, 24.24]) per patient. The mean plaque index was 77 (SD: 29.05, CI: [73.70, 77.72]), and the bleeding on probing score averaged 20 (SD: 21.49, CI: [19.79, 20.48]). Regarding oral hygiene habits, 54% of patients brushed twice daily, while 24% flossed daily, although 20% never flossed. In addition, 25% had tooth mobility, 21% had defective restorations, 25% had bruxism, and 21% had tooth crowding (Table 1).
Patient Demographics and Behavioral and Oral Health Factors in the Study Population.
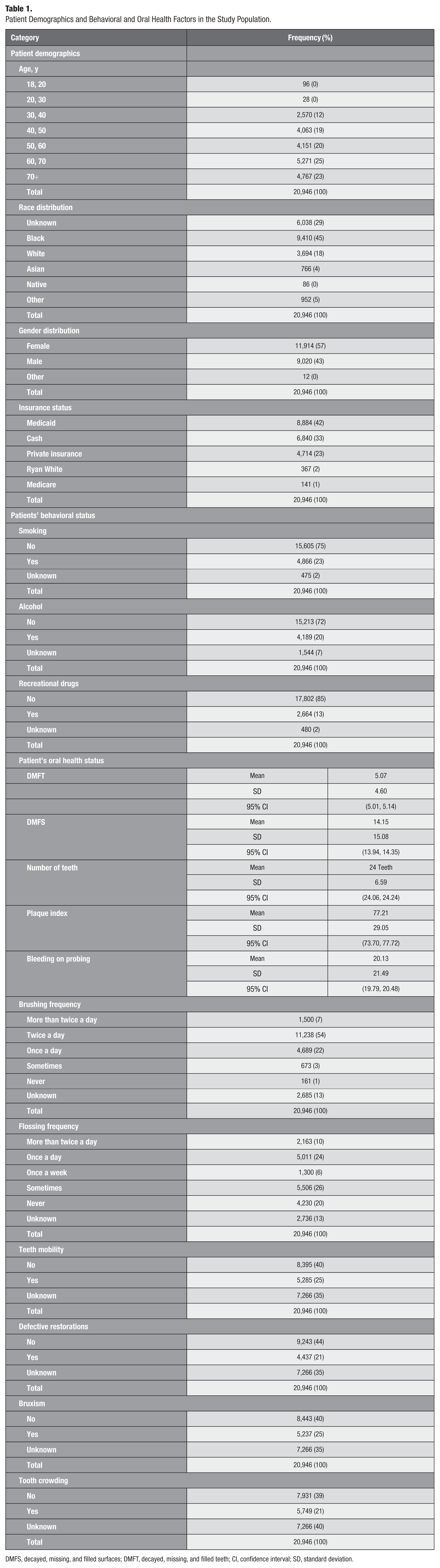
DMFS, decayed, missing, and filled surfaces; DMFT, decayed, missing, and filled teeth; CI, confidence interval; SD, standard deviation.
Medical Histories, Medications, and Procedures of Dental Patients
The most prevalent patient diagnoses included hypertension (4,431 cases), hyperlipidemia (2,171 cases), type 2 diabetes (1,603 cases), and gastroesophageal reflux disease (1,805 cases), highlighting the common comorbidities among dental patients. Frequently prescribed medications included fluticasone propionate (2,018 cases), ibuprofen (1,553 cases), amlodipine (1,517 cases), and atorvastatin (1,483 cases), reflecting treatments for inflammatory, cardiovascular, and metabolic conditions, while the most common procedures involved venipuncture and transfusion (1,462 cases), lipid panel tests (1,219 cases), and cardiography (1,095 cases), emphasizing routine monitoring of systemic health in this patient population (see Fig 2 and Appendix Table 4 for medical diagnosis distribution).
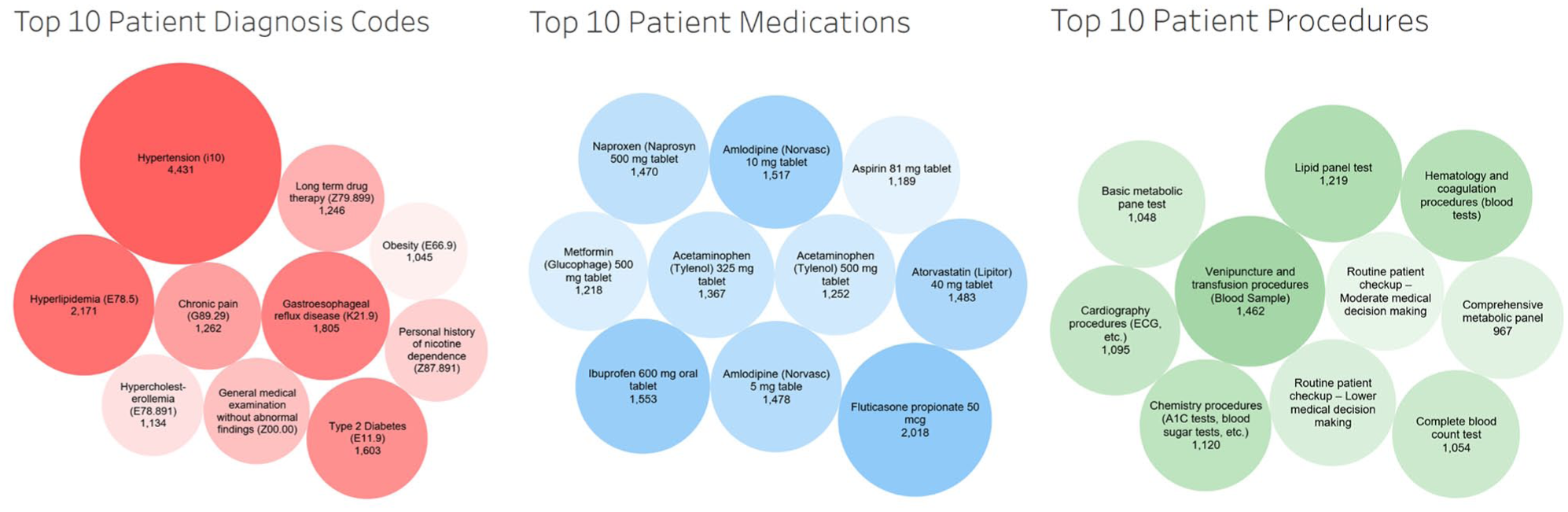
Medical diagnosis, medications, and medical procedures of dental patients.
From the chi-square analysis, we found that PD was associated with cardiovascular diseases (hypertension, ischemic heart disease, myocardial infarction), endocrine and metabolic disorders (type 2 diabetes, hyperlipidemia, hyperglycemia, obesity), renal diseases (chronic kidney disease, kidney failure), respiratory conditions (chronic obstructive pulmonary disease, asthma, respiratory failure), hematologic disorders (hyperkalemia, anemia), gastrointestinal conditions (gastroesophageal reflux disease, hepatitis C, colon polyps), neurologic and psychiatric disorders (anxiety, depression, migraines, epilepsy, posttraumatic stress disorder), and autoimmune conditions (see Appendix Table 5: chi-square results for all factors).
Predictors for Periodontitis Obtained from Linked EHR–EDR Data
As demonstrated in Figure 3, key predictors from the EDR included periodontal findings, American Society of Anesthesiologists severity (ASA) (Describes a patient’s systemic health), smoking, government insurance, self-pay status, fewer prophylaxis procedures, fewer dental visits, older age, high caries risk index, overhanging restorations, and tooth mobility. From the EHR, predictors include features as described above. These findings highlight periodontitis as a systemic inflammatory condition rather than an isolated oral disease. Integrating medical records into periodontal risk assessment enables a more comprehensive, personalized approach to early detection and intervention in both dental and medical settings.
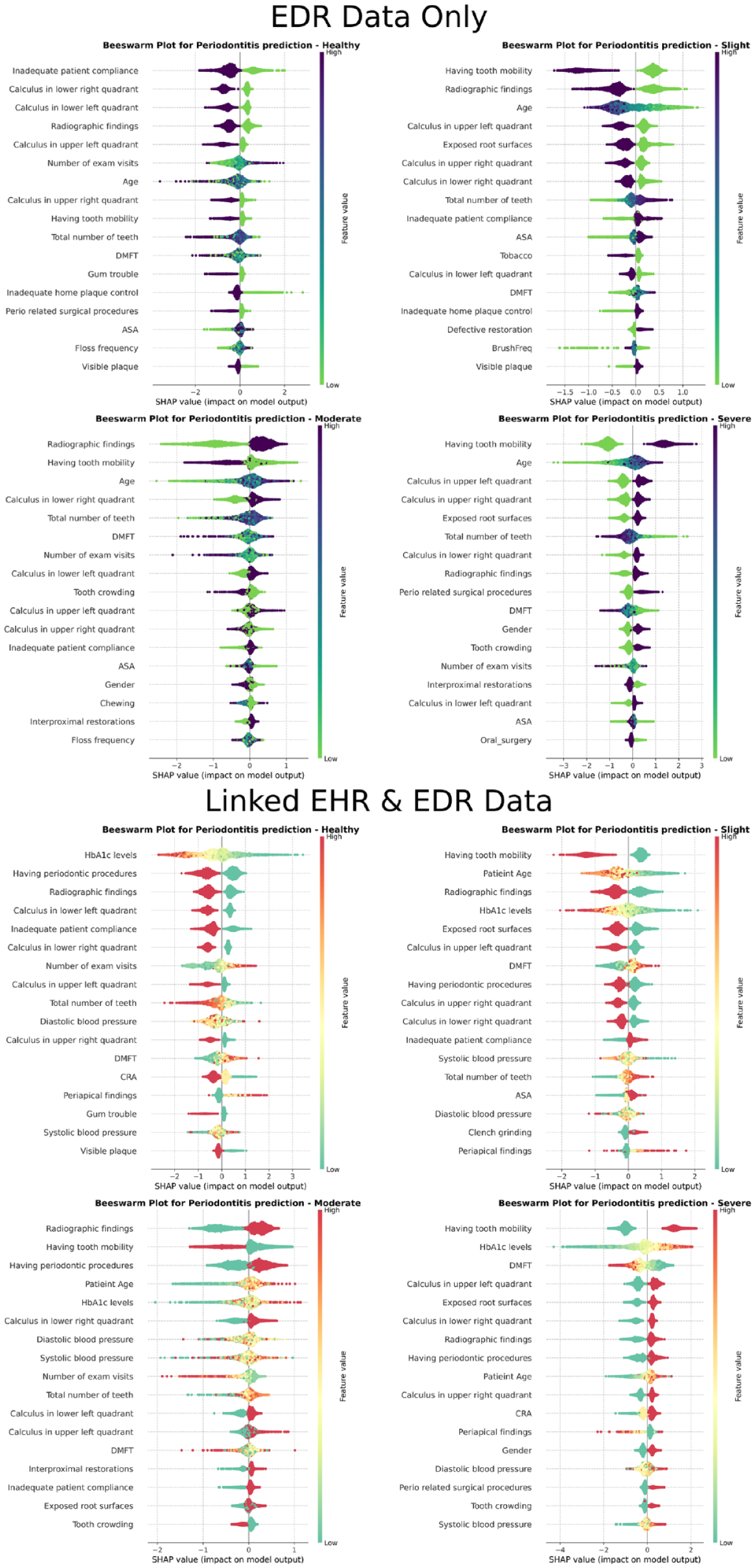
Beeswarm plot for machine learning interpretability and factors predicting periodontitis.
ML Performance and Evaluation
XGBoost outperformed all other ML models (see Appendix Table 6 for the comparison matrix). As shown in Table 2, the model performance metrics were relatively consistent across experiments 1 through 4. Experiment 1 yielded the best performance (chi-square test for EHR feature selection + ML). Using only EDR data, the model achieved an overall AUC of 84% and an F1 score of 82%, with a sensitivity of 66% and a specificity of 86%, indicating strong predictive performance but moderate sensitivity in detecting periodontitis severity levels (Table 2). When incorporating both EDR and EHR data, the overall accuracy and sensitivity remained at 81% and 66%, respectively; however, the AUC improved to 86%. Despite the small improvement, it added medical features to the prediction, especially in severe periodontitis cases (such as HbA1c levels and blood pressure). Therefore, adding EHR data provided valuable insights into the contribution of medical factors to the prediction.
Scores Summarizing the Model Performance Metrics across the Various Experiments Using XGBoost.
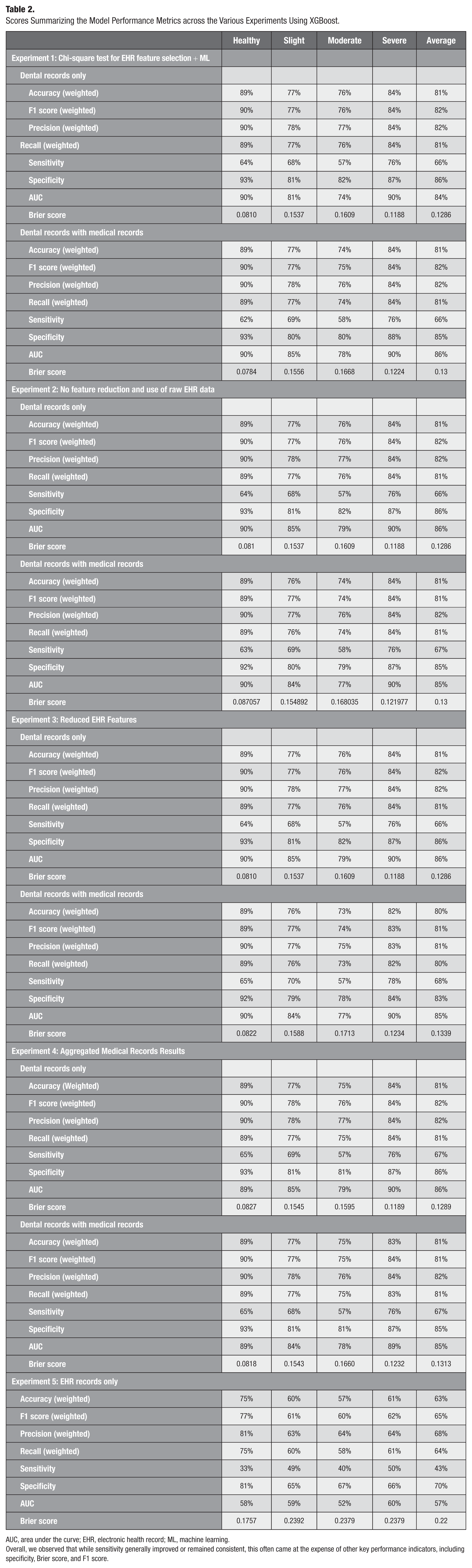
AUC, area under the curve; EHR, electronic health record; ML, machine learning.
Overall, we observed that while sensitivity generally improved or remained consistent, this often came at the expense of other key performance indicators, including specificity, Brier score, and F1 score.
Experiment 5 (only EHR features) revealed that many medical conditions were significantly associated with PD; some emerged as key predictors, and some appeared in the top 20 features. This model achieved an F1 score of 65% and a specificity of 70%, demonstrating that even with only EHR data, it can deliver satisfactory performance in predicting PD. SHAP analysis further revealed that the model successfully identified meaningful associations between diabetes, heart disease, and periodontitis, reflecting its ability to detect key known correlations within the EHR data (Huang et al 2024). However, when EDR data are introduced, the model tends to prioritize dental features only, which are stronger indicators of the disease outside of biomarkers such as HbA1c levels of blood pressure. This suggests that although EHR data hold valuable information, they may require more nuanced integration with EDR data (for more details, see Appendix Figs 2 and 3 and the Beeswarm plots) (Lundberg and Lee 2017).
Across experiments 1 to 4, we observed a mean feature similarity of 95% with experiment 1, showing the highest overlap with the other experiments, indicating that its feature set captured most of the key predictors of PD. Please see Appendix Experiments Feature Similarities for more details.
Discussion
This study highlights the importance of PD extending beyond the oral cavity, underscoring its connection to overall systemic health. While dental factors remained dominant in prediction, integrating medical history improved ML performance and provided additional insights. Our models show that men are more likely to develop severe PD than women are. They also suggest that the White/Caucasian population is less susceptible to periodontitis than the Black/African American population. These results align with our previous studies (Patel et al 2022b). Given the well-established connections between periodontitis and systemic diseases such as cardiovascular disease, diabetes, and respiratory illnesses, a siloed health care approach is no longer adequate. Patients with underlying conditions may benefit from proactive periodontal screenings to prevent disease progression. Medical professionals should also incorporate periodontal health into the management of chronic diseases, particularly for high-risk populations (Huang et al 2024). However, implementing integrated care faces significant barriers, particularly the lack of interoperability between EHR and EDR (Beserra et al 2022). Most health care systems operate in separate domains, limiting real-time, data-driven risk assessment that accounts for both oral and systemic health (Rudman et al 2010). While some institutions have adopted integrated medical–dental EHR platforms, these systems are expensive and inaccessible to many providers. In addition, historical patient data stored in standalone dental records remain difficult to incorporate into newer platforms (Tsai et al 2020).
Finally, despite the structured nature of EHR data, we encountered challenges in adapting these datasets for AI/ML. Patients often had more than 400 separate codes for various medical conditions, medications, medical procedures, and laboratory tests, resulting in high-dimensional data with redundancy and noise. We addressed this issue using feature reduction methods to retain key medical predictors while enhancing model efficiency and interpretability. Our feature reduction method addresses a critical gap in transforming high-dimensional data into clinically interpretable, low-dimensional representations suitable for ML and statistical modeling. Unlike existing methods that prioritize harmonization or black-box dimensionality reduction (Bashir et al 2022), this tool integrates ontology-driven mappings and expert-informed heuristics to preserve clinical meaning across diagnoses, medications, and procedures. By automating a task that is usually manual and prone to errors, this method makes it easier to use real-world clinical data for building prediction models, supports teamwork across different fields, and helps bring AI-ready data for research.
To advance integrated care, efforts should focus on strengthening data linkage frameworks between EHR and EDR (Rudman et al 2010; Huang et al 2024). Health information exchanges centralize medical data across health care providers; however, similar infrastructures are also needed for dental records (Rudman et al 2010; Huang et al 2024). Standardizing data formats and adopting common terminologies for diagnoses could facilitate seamless integration (Gurupur et al 2024). In addition, AI-driven tools could help extract and harmonize unstructured data from both domains, improving predictive modeling and patient risk stratification.
Our results demonstrate that interdisciplinary collaboration is another crucial step in providing patient care. Shared risk assessments, referral systems, and co-management strategies could improve patient outcomes by addressing periodontal and systemic health in tandem (Mohd Norwir et al 2025). Policy-level changes, such as insurance reimbursement models that incentivize preventive periodontal care for high-risk medical patients, could further drive integration efforts (Huang et al 2024).
Our findings align with prior research and demonstrate the predictive power of ML models for assessing periodontitis risk. However, most existing models rely solely on dental data, limiting their ability to incorporate systemic health influences. While linked EHR data provided valuable insights, dental factors remained the primary predictors. Compared with other studies that incorporated limited risk factor information, our study provided new knowledge about associated medical conditions and medications with periodontitis (Farina et al 2023; Ferrara et al 2024; Swinckels et al 2025).
This study has limitations. First, this study used only cross-sectional data. However, longitudinal data analysis will provide superior results to determine the correlations between oral and systemic health. Next, the accuracy of ICD-coded diagnoses in EHRs and procedure and diagnosis codes within EDRs can vary, and misclassification or missing data could influence feature importance rankings. Finally, the study results may not be generalizable as the dataset was used from one institute’s EHR. Our future studies will include (1) natural language processing–extracted clinical diagnoses with ICD codes to improve reliability, (2) conduct multisite studies to improve generalizability, and (3) conduct longitudinal studies to determine correlations between systemic and oral health.
Conclusion
This study highlights the potential of linking EHR and EDR for PD prediction. While dental factors remain the most influential predictors, systemic conditions contribute to disease risk, reinforcing the need for interdisciplinary collaboration in managing periodontitis. Efforts should focus on improving interoperability between EHR and EDR, optimizing AI/ML models for integrated datasets, and expanding large-scale data linkages to study various research questions aiming to connect systemic health with oral health.
Supplemental Material
sj-docx-3-jct-10.1177_23800844251408849 – Supplemental material for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records
Supplemental material, sj-docx-3-jct-10.1177_23800844251408849 for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records by J.S. Patel, M. Tellez, R. Katiyar, N.N. Al-Hebshi, R. Santana, R.M. Yucel and A. Ismail in JDR Clinical & Translational Research
Supplemental Material
sj-docx-4-jct-10.1177_23800844251408849 – Supplemental material for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records
Supplemental material, sj-docx-4-jct-10.1177_23800844251408849 for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records by J.S. Patel, M. Tellez, R. Katiyar, N.N. Al-Hebshi, R. Santana, R.M. Yucel and A. Ismail in JDR Clinical & Translational Research
Supplemental Material
sj-pdf-1-jct-10.1177_23800844251408849 – Supplemental material for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records
Supplemental material, sj-pdf-1-jct-10.1177_23800844251408849 for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records by J.S. Patel, M. Tellez, R. Katiyar, N.N. Al-Hebshi, R. Santana, R.M. Yucel and A. Ismail in JDR Clinical & Translational Research
Supplemental Material
sj-pdf-2-jct-10.1177_23800844251408849 – Supplemental material for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records
Supplemental material, sj-pdf-2-jct-10.1177_23800844251408849 for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records by J.S. Patel, M. Tellez, R. Katiyar, N.N. Al-Hebshi, R. Santana, R.M. Yucel and A. Ismail in JDR Clinical & Translational Research
Supplemental Material
sj-png-5-jct-10.1177_23800844251408849 – Supplemental material for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records
Supplemental material, sj-png-5-jct-10.1177_23800844251408849 for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records by J.S. Patel, M. Tellez, R. Katiyar, N.N. Al-Hebshi, R. Santana, R.M. Yucel and A. Ismail in JDR Clinical & Translational Research
Supplemental Material
sj-png-6-jct-10.1177_23800844251408849 – Supplemental material for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records
Supplemental material, sj-png-6-jct-10.1177_23800844251408849 for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records by J.S. Patel, M. Tellez, R. Katiyar, N.N. Al-Hebshi, R. Santana, R.M. Yucel and A. Ismail in JDR Clinical & Translational Research
Supplemental Material
sj-png-7-jct-10.1177_23800844251408849 – Supplemental material for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records
Supplemental material, sj-png-7-jct-10.1177_23800844251408849 for Periodontitis Prediction Model Using Linked Electronic Health and Dental Records by J.S. Patel, M. Tellez, R. Katiyar, N.N. Al-Hebshi, R. Santana, R.M. Yucel and A. Ismail in JDR Clinical & Translational Research
Footnotes
Acknowledgements
We would like to express our sincere gratitude to Dr Margaret Grisius and Dr Lorena Baccaglini, program directors at the National Institute of Dental and Craniofacial Research (NIDCR), for their thoughtful review and valuable feedback throughout this project. We gratefully acknowledge Mr Manan Patel for his programming support and Mr Ryan Brandon for his invaluable assistance in extracting data from the axiUm system. We also thank the Temple University Institutional Review Board (IRB) office for reviewing and approving this study under the exempt category (IRB No. 28321-0006). Finally, we thank Health Share Exchange for providing us with the dental patients’ linked medical records.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This publication is supported by the NIH-NIDCR’s U01DE033259 award.
Data Availability Statement
This study uses real patient records from EHR and EDR, which contain patient identifiers. Due to patient privacy and institutional security requirements, this data cannot be shared publicly. Researchers interested in accessing the dataset should contact the first and senior author, Dr Jay Patel, to explore potential collaboration. Access will be granted only after completion of the required documentation, IRB approval, a data use agreement, and a business associate agreement.
A supplemental appendix to this article is available online.
