Abstract
Introduction:
Oral potentially malignant disorders (OPMDs) can lead to oral cancer, which is one of the most common cancers worldwide. Prevention is crucial in the avoidance of malignant transformations of OPMDs. Artificial intelligence (AI) provides a new and noninvasive tool for analyzing medical data, such as patient data, radiologic images, and clinical photographs. These AI-based tools can help in the decision-making process. However, histological examination is still the gold standard for diagnosing OPMDs.
Objectives:
This systematic review and meta-analysis aimed to investigate the diagnostic accuracy of artificial intelligence on intraoral photographs of patients with OPMDs.
Methods:
A systematic search was conducted on 5 major databases (MEDLINE, Embase, Cochrane Library, Scopus, and Web of Science) on November 10, 2023. Included studies compared AI methods to histology examination as the reference. A quantitative analysis was carried out to assess sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), diagnostic odds ratio (DOR), positive likelihood ratio (LR+), and negative likelihood ratio (LR−) calculated with 95% confidence intervals (CIs).
Results:
Six eligible articles were included, with 898 images out of 4,046 tested using AI-based architectures. Five studies investigated at least 2 AI models. The overall sensitivity, specificity, DOR, LR+, and LR− were 0.94 (95% CI, 0.88 to 0.95), 0.95 (95% CI, 0.85 to 0.98), 212.39 (95% CI, 56.39 to 800.00), 16.89 (95% CI, 5.72 to 48.68), and 0.08 (95% CI, 0.05 to 0.13) for the best-performing AI-based architectures in terms of sensitivity, respectively.
Conclusion:
AI-based diagnostic tools have high negative predictive value that could help identify OPMD lesions using intraoral photographs.
Knowledge Transfer Statement:
This systematic review on AI-based methods to diagnose oral potentially malignant disorders showed that although their high negative predictive value could reduce unnecessary specialist consultations, clinical judgment remains paramount. Further prospective studies are needed to evaluate the integration of AI diagnostics into routine care and screening and policies to enhance efficiency and support early detection and prevention of oral cancer.
Introduction
Oral potentially malignant disorders (OPMDs) are a group of oral mucosal lesions with an elevated risk of progression to malignancy. These disorders comprise diverse diseases with distinct clinical manifestations, histopathological characteristics, and etiological factors (Warnakulasuriya et al 2021). The most important common feature is that all these lesions can lead to oral cancer. Oral cancer is the sixth most common cancer worldwide, with 377,713 new cases and 177,384 deaths every year (Sarode et al 2020; Sung et al 2021). The 5-y survival rate is 34%, as oral cancer is mainly diagnosed at advanced stages (III to IV), probably due to low compliance of patients from lower socioeconomic status (SES) (Abati et al 2020). Lower SES is also correlated with a higher risk of oral cancer (Conway et al 2015). If oral cancer is diagnosed at earlier stages (I to II), the 5-y survival rate increases to 83% (Silverman et al 2010). However, secondary prevention is the most important; if lesions that can develop into oral cancer are diagnosed and treated, many lives can be saved (Feller and Lemmer 2012; Gelband et al 2015).
Visual examination remains the most widely used and cost-effective method for the initial diagnosis of OPMDs. However, its diagnostic accuracy is limited by the subjectivity of clinical assessment and the examiner’s level of experience, which can result in both overdiagnosis and underdiagnosis (Tripathy et al 2023). Warin et al (2022) demonstrated that general practitioners diagnose OPMDs with relatively low sensitivity (0.68; 95% confidence interval [CI], 0.62 to 0.75). Consequently, adjunctive diagnostic techniques such as brush cytology, chemiluminescence, and toluidine blue staining have been introduced to improve diagnostic objectivity. While these methods offer enhanced discrimination between oral cancer and OPMDs, their utility in the accurate identification of OPMDs alone remains limited. At present, biopsy followed by histopathological evaluation continues to be the gold standard for definitive diagnosis and for assessing malignant transformation potential (Jeddy et al 2017). Nonetheless, the invasive nature of biopsy may cause patient discomfort and anxiety, and its application is constrained in low-resource settings due to limited access to specialized care, infrastructure, and trained personnel (Kharche et al 2024). These limitations underscore the urgent need for the development of a novel, noninvasive, cost-effective, and reliable diagnostic modality for OPMDs that facilitates the preliminary screening of patients who are typically underserved or have limited access to health care services.
Recent advances in artificial intelligence (AI) present promising opportunities for the development of novel diagnostic tools to support clinical decision-making. Notably, progress in AI architectures for analyzing unstructured medical data has accelerated in recent years (Szabó et al 2024). The widespread availability and capability of taking high-resolution images of smartphones enable convenient documentation and monitoring of patients, particularly in community and primary care settings. Integrating AI-based image analysis with intraoral photography represents a potentially valuable diagnostic opportunity, offering a low-cost, user-friendly, and accessible solution, particularly beneficial in low-resource settings in which access to specialist care and adequate health care infrastructure is limited.
The aim of this study was to investigate the diagnostic performance of AI-based decision-making tools on regular intraoral photographs that support clinicians in the diagnosis of OPMDs.
Methods
The present meta-analysis was conducted according to the Preferred Reporting Items for Systematic Review and Meta-Analyses (PRISMA) protocols while adhering to the Cochrane Handbook (Cumpston et al 2022). The protocol was registered in the International Prospective Register of Systematic Reviews (PROSPERO) under registration number CRD42023475408.
Eligibility Criteria
The PIRD framework was applied in this meta-analysis: population (P) patients with intraoral images, index test (I) AI, reference test (R) histology, and disease (D) OPMDs with a given confusion matrix for the AI-supported tool. All included articles were available in full text and contained histologically proven OPMD lesions; no specialized photography tool was used (autofluorescence or polarized white light). There were no restrictions on the quality of the analyzed images or language. Reviews, case reports, case series, and descriptive surveys were excluded.
Information Sources
Five major databases were searched for the systematic review on November 10, 2023: MEDLINE (via PubMed), Embase, CENTRAL (The Cochrane Central Register of Controlled Trials), Scopus, and Web of Science. No filters or restrictions were used. Two domains were presented in the search key: the first one included the index test (AI) and the second one included the disease (OPMD). Different search queries were used for each database (Appendices 1 to 5). Eligible studies were identified by reviewing titles and abstracts; backward and forward citation chasing of eligible articles was also performed to identify all studies.
Selection Process
The EndNote 20.6 software package was used for record management. Two independent reviewers (D.H., Á.F.) performed the selection after duplication removal. The first selection was based on the titles and abstracts of the published original papers. After conflict resolution by a third independent reviewer (P.M.), the eligibility of the remaining full texts was investigated by the same 2 independent reviewers. At this stage, an expert in the field (A.B.) helped resolve conflicts.
Data Collection
Among the eligible articles, 2 independent authors (D.H., Á.F.) collected data on decision-making in the OPMD classification of the AI-supported tool and filled out a customized data extraction sheet designed for this review. The data analysis units covered sensitivity, specificity, and the confusion matrix. In addition, the following information was collected: first author, publication year, study period, teaching, validation, testing sample size, teaching settings (epochs, batch size, and learning rate), training methods, and augmentation types. Information (number of layers and parameters) on the AI-based models used was collected from the internet if not provided in the article. Some data about the tested AI-based architectures were collected from the internet if not included in the article. The sources for this information were https://github.com and https://pytorch.org.
Risk-of-Bias Assessment
The quality of the eligible studies was assessed by 2 independent authors (D.H., Á.F.). The risk-of-bias assessment was conducted with the Quality Assessment of Diagnostic Accuracy Study-2 (QUADAS-2), designed to evaluate the quality of diagnostic accuracy studies. This tool contains 4 domains: patient selection, index test, reference standard, and flow and timing. Each domain is assessed in terms of the risk of bias, and the first 3 are also evaluated for applicability.
Data Synthesis
Two-by-two contingency tables were extracted from the studies, containing true-positive, false-positive, false-negative, and true-negative values. The classical approach in diagnostic meta-analysis is to calculate pooled sensitivity and specificity. Some studies also included images of oral cancer to investigate the classification of oral cancer alongside OPMD and disease-free cases. In these cases, sensitivity and specificity were calculated based on the ability of the model to differentiate between OPMD and non-OPMD. The bivariate model of Reitsma et al and Chu et al was fitted (Reitsma et al 2005; Chu and Cole 2006) This approach takes into account the dependency between sensitivity and specificity. Diagnostic odds ratios (DORs) were also calculated for each study.
Sensitivity and specificity of the included studies were plotted as summary estimates of sensitivity and specificity and the corresponding 95% confidence and prediction regions on a hierarchical summary receiver-operating characteristic (HSROC) plot. The confidence region contains the pooled sensitivity and specificity (more exactly, 1-specificity) in 95% of the cases. The prediction region contains the true sensitivity and specificity (more precisely, 1-specificity) of a new study in 95% of the cases; hence, it provides excellent insight into heterogeneity.
All statistical analyses were performed in R (Team RC. R: A Language and Environment for Statistical Computing; Vienna, Austria R Foundation for Statistical Computing, 2021), and the code was heavily based on the online tool described by Freeman et al (2019). The CIs of proportions were calculated with the Wilson scoring method, whereas the CIs of the likelihood ratios were calculated with the Koopman asymptotic score method (Koopman 1984), both using the R package DescTools (Signorell A. 2017). Figure 1 summarizes the outcomes and calculation methods.
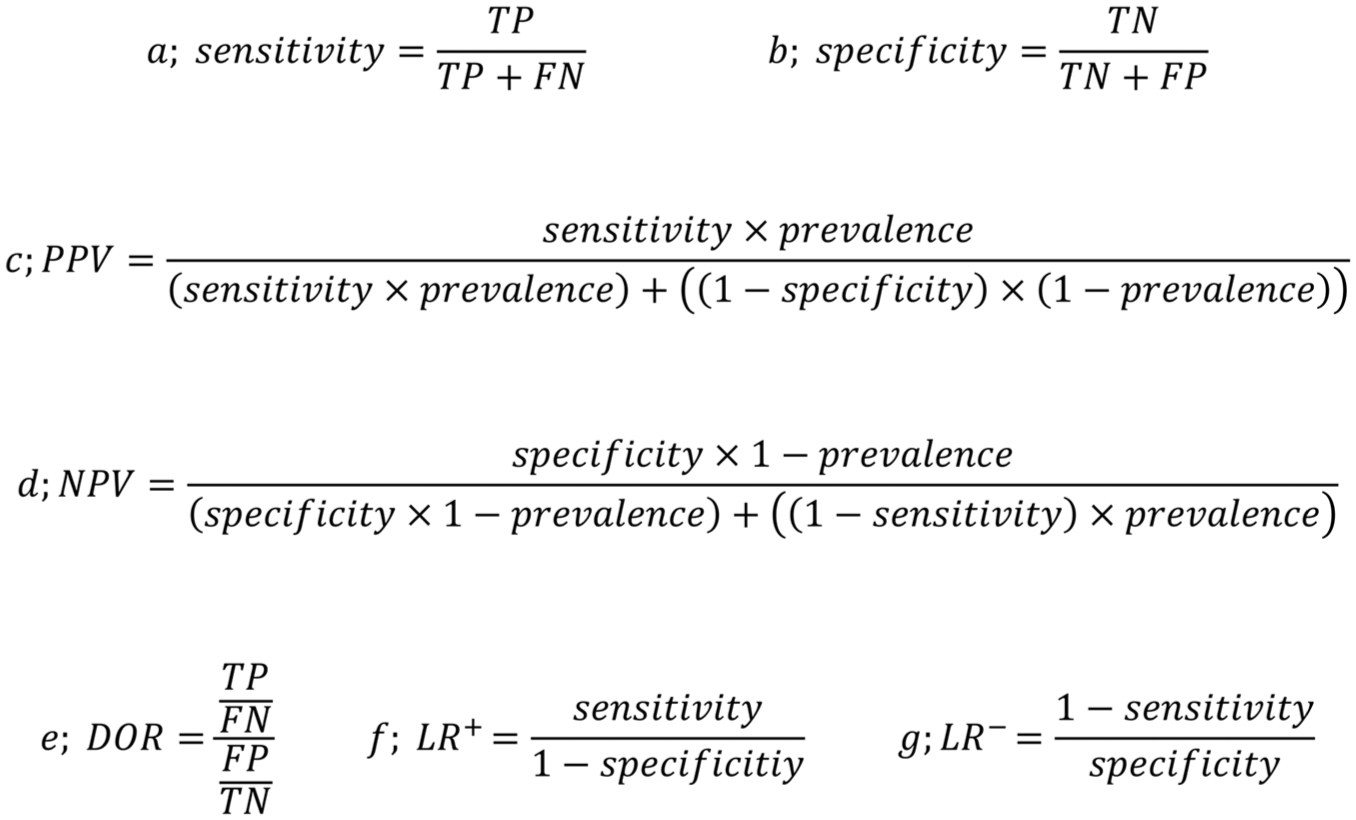

Calculation method of (
Results
Search and Selection
The systematic search of 5 databases identified 11,819 articles. The selection process is illustrated in the PRISMA flowchart (Fig 2). After removal of duplicates, 8,264 articles remained for title and abstract selection. Of these, 125 were eligible for full-text selection. For 22 articles, only conference abstracts were available and were excluded from the analysis. The full text of 3 articles could not be retrieved. One hundred sixteen articles did not meet the eligibility criteria, with the reasons for exclusion shown in Appendix 6. Finally, 6 articles were eligible for quantitative and qualitative analysis (Jurczyszyn and Kozakiewicz 2019; Lin et al 2021; Sharma et al 2022; Warin et al 2022; Keser et al 2023; Achararit et al 2023). No other eligible articles were found through citation chasing or manual search (Appendix 7). The inter-reviewer agreement assessed by Cohen’s kappa was 0.91 and 1.00 in the title-abstract and full-text selection phases, respectively.
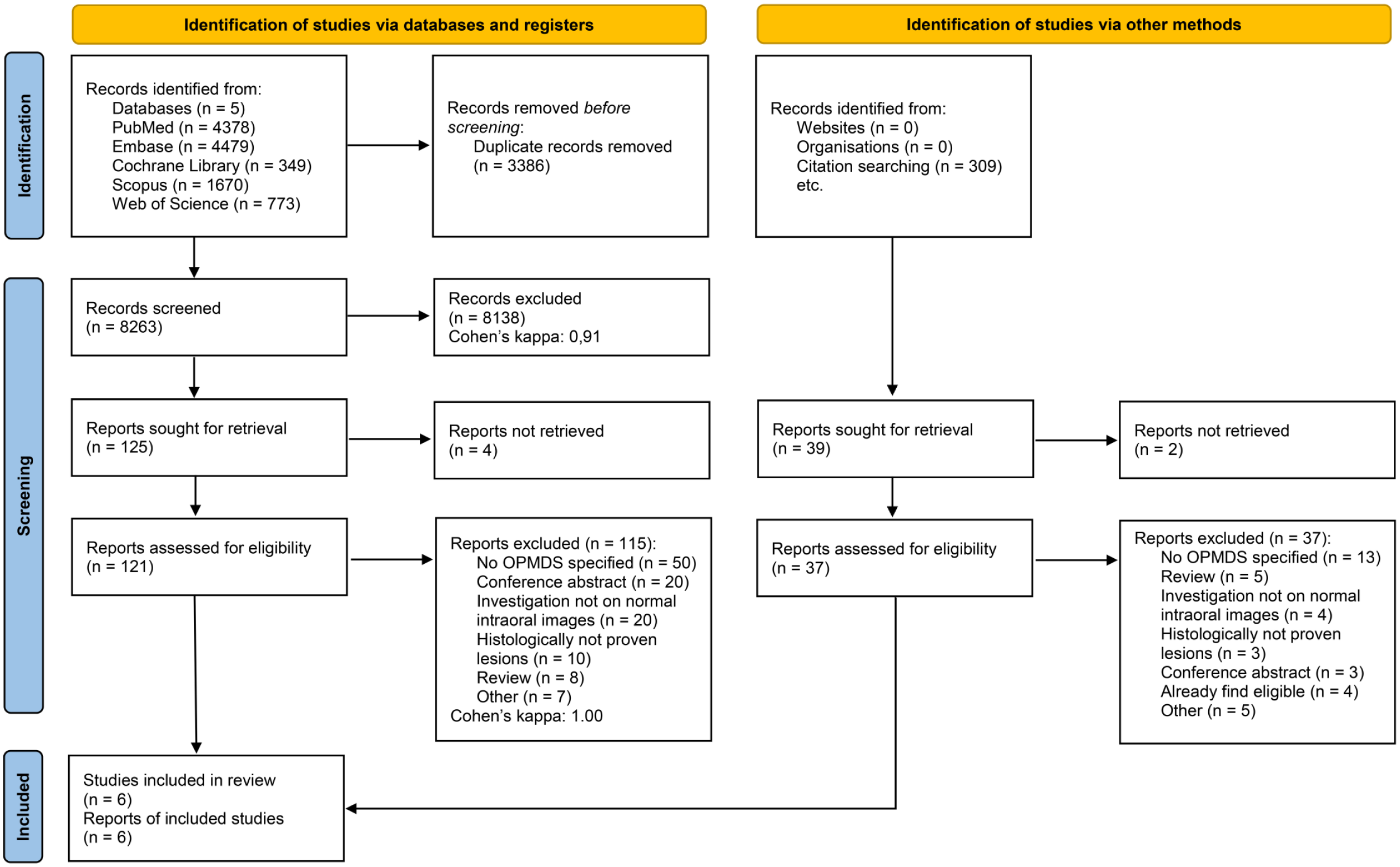
PRISMA selection flow chart.
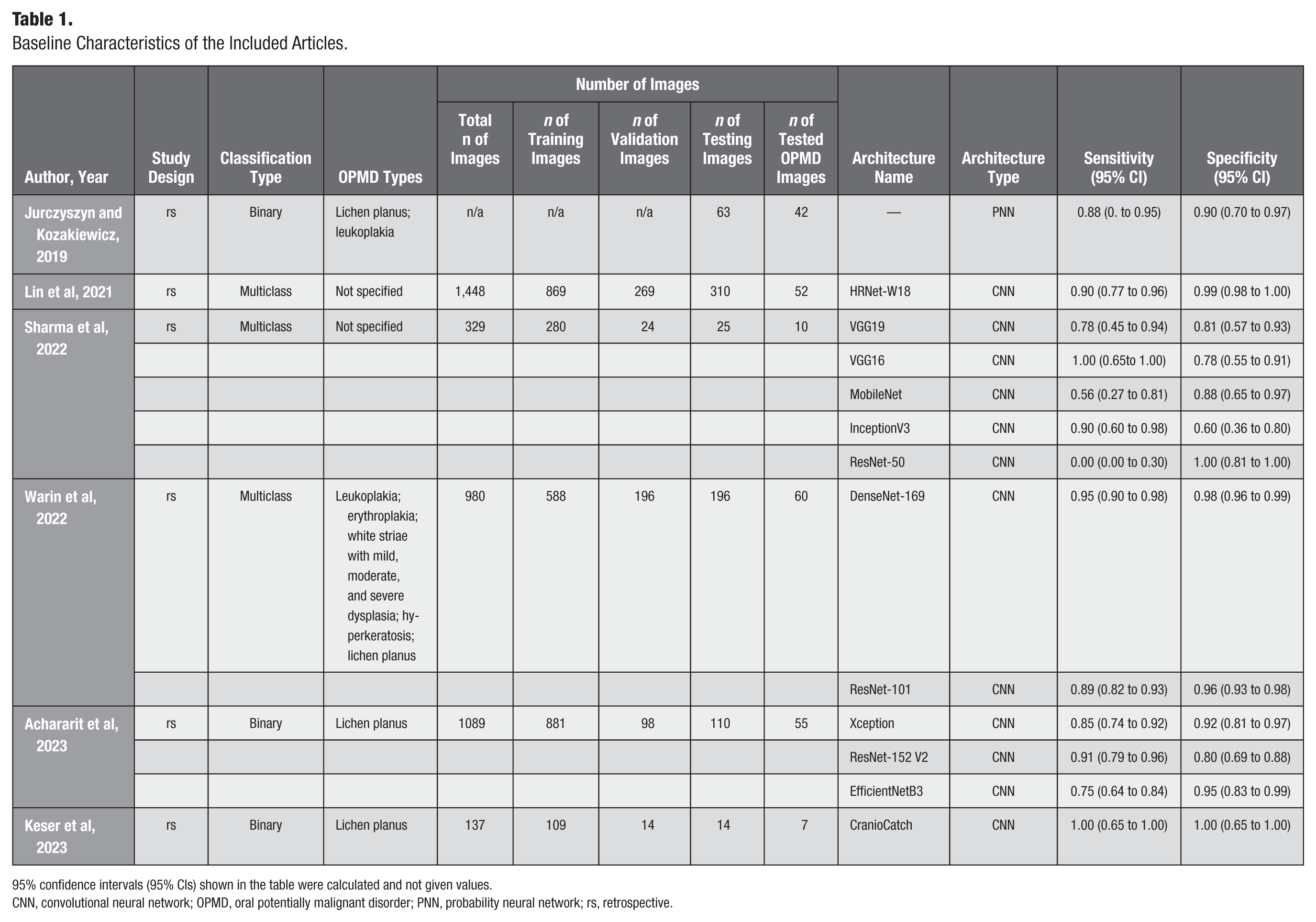
Six retrospective articles were included in this meta-analysis. Four of the studies were conducted in Asian countries (China, Thailand, and India), and the remaining 2 studies were conducted in Europe (Poland and Turkey). Binary and multiclass classification models were evenly used in the studies. Binary classification models distinguish between 2 classes, while multiclass classification models categorize data into 3 or more classes. The teaching setting was well described, and all 6 eligible studies provided essential information on how many times it went through the data (epochs), how much data were processed at once (batch size), and how quickly the model adjusted its parameters (learning rate). This information is important for understanding how the model was trained, allowing researchers to reproduce the result and compare performance or improve the model in future work. A total of 697 images were tested with AI-based architectures from a total amount of 3,983 images. Different types of OPMD were tested (Jurczyszyn and Kozakiewicz 2019; Warin et al 2022) in the included articles. Only 1 type was investigated (Achararit et al 2023; Keser et al 2023). The baseline characteristics of the participants recruited in the articles are summarized in the Table 1.
Baseline Characteristics of the Included Articles.
95% confidence intervals (95% CIs) shown in the table were calculated and not given values.
CNN, convolutional neural network; OPMD, oral potentially malignant disorder; PNN, probability neural network; rs, retrospective.
Sensitivity and Specificity
The 6 eligible articles included 1,512 OPMD images, of which 216 were tested. The sensitivity and specificity of the AI-based architectures tested ranged from 0.00 to 1.00 and from 0.60 to 1.00, respectively. These data indicate pronounced variability in results. Due to the overlapping population within a study, the best-performing AI-based diagnostic tools in terms of sensitivity were chosen, and a quantitative analysis was performed from the data of these AI-based models, which showed an overall 0.93 (95% CI, 0.88 to 0.95) sensitivity and 0.95 (95% CI, 0.85 to 0.98) specificity (Table 1). However, the sensitivity and specificity of all of the tested AI-based tools were also calculated and visualized (Appendix 8).
Positive and Negative Predictive Value
Positive predictive value (PPV) and negative predictive value (NPV) were calculated for the best-sensitivity AI-based models in the 6 eligible architectures. The overall PPV and NPV were 0.42 and 0.99, respectively. The global prevalence of OPMDs was used to calculate the overall PPV and NPV (Abati et al 2020).
DOR
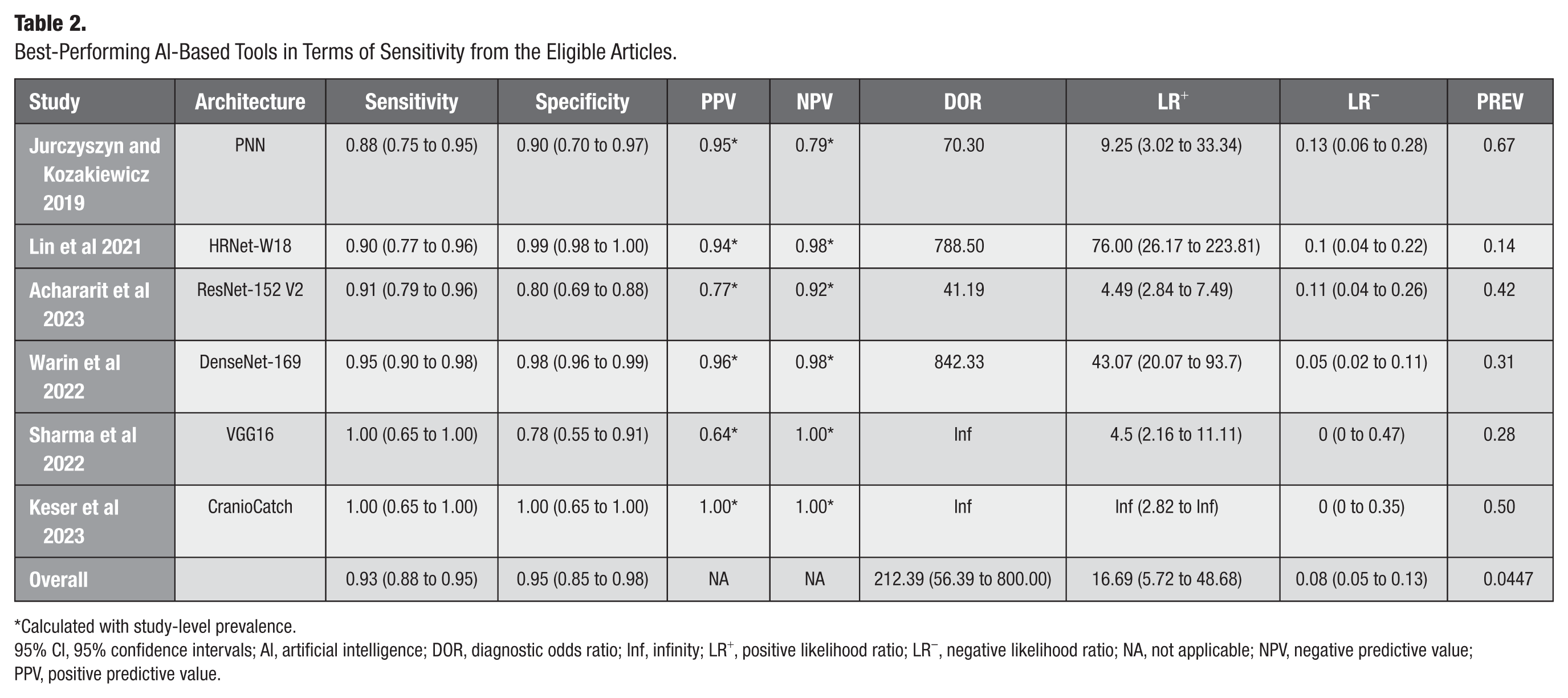
The DOR was calculated based on the 2 × 2 confusion matrix. The DOR also showed high diversity, ranging from 41.19 to 842.33. The same AI-based architectures with the best sensitivity were selected from the studies; the DOR was 218.32 (95% CI, 59.98 to 794.60) (Table 2).
Best-Performing AI-Based Tools in Terms of Sensitivity from the Eligible Articles.
Calculated with study-level prevalence.
95% CI, 95% confidence intervals; AI, artificial intelligence; DOR, diagnostic odds ratio; Inf, infinity; LR+, positive likelihood ratio; LR−, negative likelihood ratio; NA, not applicable; NPV, negative predictive value; PPV, positive predictive value.
Likelihood Ratio
Positive and negative likelihood ratios were calculated for the best-sensitivity AI-based models, which were 16.69 (95% CI, 5.72 to 48.68) and 0.08 (95% CI, 0.05 to 0.13), respectively (Table 2).
HSROC
HSROC visualized data heterogeneity for the best-sensitivity AI-based models. The heterogeneity of the data was considered moderate (Fig 3).
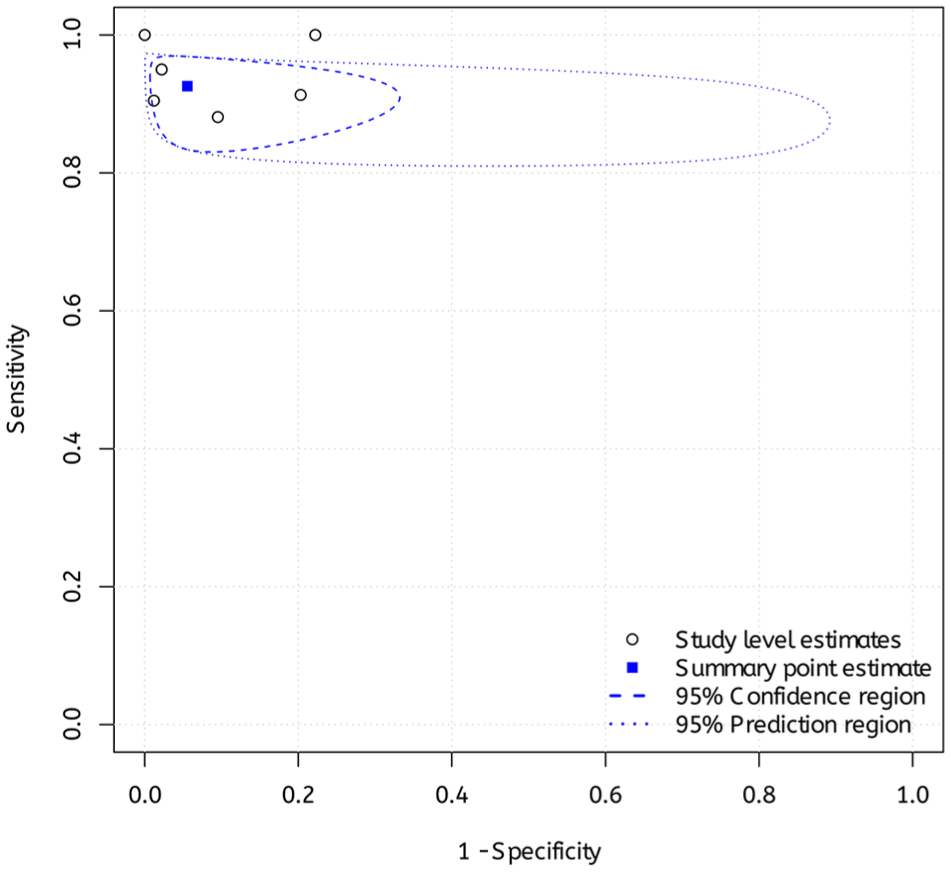
Hierarchical summary receiver operating of the best performing artificial intelligence–based architectures in terms of sensitivity.
Overall sensitivity, specificity, PPV, NPV, DOR, and LR+, LR−, and HSROC were calculated and visualized for the best-performing AI-based architectures in terms of specificity (Appendices 11 to 13).
Risk-of-Bias Assessment and Publication Bias
In terms of risk of bias, 5 of the 6 eligible studies were rated low. Jurczyszyn and Kozakiewicz (2019) did not provide full information about the AI-based diagnostic tool used; thus, the risk of bias and the applicability of the index test yielded high risk. Detailed results on applicability and risk of bias can be found in Appendix 14. Because of the limited number of studies, publication bias could not be assessed.
Discussion
To the best of the authors’ knowledge, this is the first meta-analysis to systematically evaluate the diagnostic accuracy of AI-based tools specifically for OPMDs using intraoral images. The studies analyzed demonstrated that AI-based diagnostic tools could help diagnose OPMDs in everyday practice. This meta-analysis showed high sensitivity, specificity, NPV, LR+, and LR− for the proposed AI-based models in the diagnosis of OMPDs. However, these findings need to be interpreted with caution due to the low number of eligible articles. This finding demonstrates that AI-based diagnostic models could have the potential of distinguish images of OPMDs from normal mucosal lesions, likely benign or malignant. Nevertheless, further studies are needed on this topic to strengthen this finding.
Identifying these lesions may be more complex in clinical practice due to factors such as lesion localization and other types of diseases, including paraneoplastic mucosal diseases, pemphigus vulgaris, or mucous membrane pemphigoid and certain types of Candida infection (do Carmo et al 2014; Binnie et al 2024). One of the critical responsibilities of a dentist is oral cancer screening, a procedure that can prove challenging in the absence of sufficient expertise. As OPMDs can lead to oral squamous cell carcinoma, early diagnosis and adequate treatment of OPMDs can decrease the risk of malignant transformation, thus reducing the number of new cases of oral cancer (Warnakulasuriya 2020).
The systematic search also identified several articles that investigated OPMDs or other oral lesions (González and Quintero Rojas 2021; Tanriver et al 2021) However, these could not be used in the meta-analysis because they boosted their teaching datasets with images gathered from search engines such as www.google.com/imghp. In the authors’ opinion, they cannot be certain that photographs collected from the internet are OPMDs, which can significantly affect the results.
The data analysis of 6 studies emphasized the ability of AI-based diagnostic tools to diagnose OPMDs. However, the diagnostic performance varied widely, with sensitivity ranging from 0.00 (95% CI, 0.00 to 0.30) to 1.00 (95% CI, 0.65 to 1.00) and specificity from 0.60 to 1.00. This variability likely reflects a combination of factors, including differences in AI model architectures, training procedures, dataset quality, and image characteristics. For example, in the article by Sharma et al 2022, ResNet50, a deep and complex convolutional neural network with the highest number of layers and parameters among the models analyzed, showed a sensitivity of 0% (95% CI, 0.00 to 0.30) (Sharma et al 2022). The researchers selected training parameters (50 epochs, batch size of 16, and learning rate of 0.001) for all AI-based models. It is likely that, for ResNet50, the data quality, quantity, or the uniform training settings were suboptimal. Beyond this specific case, broader variability across studies can be attributed to heterogeneity in data sources (eg, single center vs multicenter), inconsistency in image acquisition protocols, and lack of standardized preprocessing and augmentation methods. Furthermore, the authors did not report essential information regarding lesion size, severity, or subclassification, all of which can significantly affect diagnostic performance. Larger or more clinically evident lesions are generally easier to recognize, whereas subtle lesions, such as simple leukoplakia, may be misclassified as benign hyperkeratosis, leading to underdiagnosis. The absence of a unified evaluation framework, including differences in validation strategies and performance metrics, further contributes to this inconsistency and underscores the need for comprehensive benchmarking and reporting standards.
In this meta-analysis, the proposed AI-based architectures demonstrated an exceptionally high NPV of 1.00, suggesting strong performance in correctly identifying disease-free cases. However, PPV was notably low at 0.42, indicating that a substantial proportion of positive predictions were false positives. This limitation has important implications for clinical usability. A low PPV reduces clinicians’ trust in AI outputs and may result in unnecessary patient anxiety, redundant follow-up visits, and inefficient use of health care resources. While false positives might not always lead directly to invasive interventions such as biopsies, they do increase the burden on clinical workflows and may contribute to overdiagnosis. In settings with limited access to specialists, a high false-positive rate could divert attention from higher-risk patients or contribute to alarm fatigue. Nevertheless, this may have a beneficial effect in uncertain cases, as such encounters can motivate patients to adhere to regular follow-up visits and enhance early detection efforts. Therefore, despite the promising safety profile indicated by the high NPV, the clinical utility of these models remains limited without further refinement. Improving PPV through better dataset stratification, inclusion of lesion-level annotations, and advanced model calibration techniques is critical to enhancing diagnostic precision and real-world applicability. Despite these challenges, AI technologies hold significant promise in improving access to specialist care for at-risk populations.
A major limitation in the current literature is the substantial methodological heterogeneity across studies. Key differences exist in dataset composition, image acquisition protocols, preprocessing techniques, and data augmentation methods. Moreover, studies use varied validation strategies, some relying exclusively on internal cross-validation, while others use external test datasets, often without standardized dataset-splitting procedures or consistent performance metrics. The lack of benchmarking frameworks and inconsistent reporting of model parameters, such as hyperparameters, feature selection methods, and overfitting-prevention strategies, further impedes reproducibility and cross-study comparisons. Notably, variability in AI architecture selection, training configurations (eg, batch size, learning rate, number of epochs), and evaluation criteria can significantly influence reported performance. A more systematic discussion and control of these methodological differences are essential to ensure interpretability, comparability, and reproducibility across studies and to establish best practices in the development of AI-based diagnostic tools. These issues collectively pose significant challenges to the generalizability and clinical translation of AI-based diagnostic models for OPMDs (Srivastava et al 2014; Wong et al 2021).
The diagnostic performance and clinical utility of AI-based tools for detecting OPMDs are influenced by several critical limitations, including restricted dataset diversity, risk of overfitting, and inadequate consideration of demographic factors. Many models are trained on data from single institutions or homogeneous populations, limiting their generalizability to diverse real-world settings. Insufficient representation of rare OPMD subtypes, along with small sample sizes and poor dataset stratification, increases the risk of overfitting, leading to diminished performance on external data. Furthermore, important demographic variables such as age, sex, and lifestyle-related risk factors are often overlooked, despite their relevance to lesion characteristics and malignancy risk. These methodological shortcomings can introduce diagnostic uncertainty, algorithmic bias, and decreased clinician confidence. These challenges underscore the urgent need for diverse, well-annotated datasets, rigorous external validation, and clinician-centered AI model development to ensure equitable, accurate, and clinically applicable diagnostic support for OPMDs (Virgens et al 2025).
In addition to these methodological concerns, inconsistency in intraoral image quality presents another critical limitation. Differences in image resolution, illumination, focus, angulation, and anatomical framing introduce significant variability, which may adversely affect model performance and generalizability. Inconsistent image quality, labeling practices, and diagnostic criteria across studies further hinder AI reliability and interpretability, often necessitating reliance on conventional diagnostic methods. However, most studies fail to report standardized criteria for image acquisition or quality control. The absence of such standards compromises diagnostic reliability and hinders reproducibility. Establishing and adhering to image quality benchmarks, such as minimum resolution thresholds, uniform anatomical orientation, and exclusion of artifacts, would enhance dataset consistency and improve model training outcomes. Future work should incorporate objective image quality assessment protocols or validated scoring systems to ensure that only diagnostically acceptable images are used for AI development and evaluation (Virgens et al 2025).
The integration of AI-based diagnostic tools into clinical workflows also raises important ethical considerations. One significant concern is the risk of misdiagnosis, which can occur due to algorithmic bias, poor generalizability, or limitations in training data. Such errors may lead to inappropriate management decisions, potentially compromising patient safety. Moreover, there is a risk that clinicians may become overly reliant on AI outputs, particularly when the decision-making process of the system lacks transparency or interpretability. This overreliance could diminish the role of clinical judgment, especially in ambiguous cases. To ensure responsible implementation, AI systems must be rigorously validated, used as adjunctive tools rather than standalone solutions, and integrated with appropriate clinician training and oversight mechanisms. Addressing these ethical and practical concerns is essential to ensure the safe, equitable, and accountable use of AI in the diagnosis of OPMDs (Kharche et al 2024).
The analyzed articles used binary and multiclass classification AI-based models. Binary classification architectures, which can distinguish between only 2 categories, can be used in screening by general practitioners or even in self-screening applications. In this case, the exact differential diagnosis is less crucial than determining whether the lesion warrants referral to a specialist, as discussed in the article by Figueroa et al (2022). In this context, 2 categories can be predefined, namely, “nonsuspicious” and “suspicious” or “nonreferral” and “referral.”
AI-based models with multiclass classification could be used in special care to aid in solving differential diagnostic problems because these architectures can deal with multiple predefined lesions and conditions. According to the current state of science, biopsy is still the gold standard, and further development of multiclass classification models and clinical trials is needed to improve diagnostic performance.
AI-based tools demonstrate potential not only in the classification of OPMDs but also in predicting their risk of malignant transformation, facilitating oral cancer screening and detection, and supporting clinical decision-making in cancer management (Hegde et al 2022; Umer 2024) In a systematic review and meta-analysis, Sahoo et al (2024) evaluated the diagnostic performance of AI-driven tools in detecting OPMDs and oral cancer. A total of 55 studies were included in the qualitative synthesis, encompassing the use of histopathological slides, intraoral photographs, and optical coherence tomography (OCT) images. Of these, 3 studies focused specifically on the identification of OPMDs—2 using intraoral images and 1 using histological images—reporting pooled sensitivity and specificity values of 0.96 (95% CI, 0.92 to 1.00) and 0.93 (95% CI, 0.90 to 0.96), respectively.
Li et al (2024) conducted a systematic review and meta-analysis to comprehensively evaluate the diagnostic performance of AI-based tools in the detection of OPMDs and oral cancer. The study assessed various imaging modalities, including OCT, autofluorescence, and conventional clinical photographs. The analysis of 17 studies revealed that OCT demonstrated the highest diagnostic accuracy (OR = 154.064; 95% CI, 38.83 to 611.34), followed by clinical images (OR = 61.83; 95% CI, 33.42 to 114.39) and autofluorescence (OR = 27.09; 95% CI, 17.85 to 41.11). However, meta-regression analysis showed no statistically significant differences between these subgroups. Following outlier removal using a graphic display of heterogeneity plot, diagnostic performance rankings shifted, with clinical images exhibiting the highest accuracy (OR = 77.77; 95% CI, 25.83 to 234.15), followed by OCT (OR = 56.99; 95% CI, 24.54 to 132.38) and autofluorescence images (OR = 33.91; 95% CI, 16.32 to 70.44), with no statistical significance (P = 0.637; Li et al 2024).
Warin et al (2022) showed that the sensitivity of the diagnosis of OPMDs differs among general practitioners (0.68; 95% CI, 0.62 to 0.75) and oral and maxillofacial surgeons (0.74; 95% CI, 0.61 to 0.87). There are other possible methods in the evaluation of OPMDs: autofluorescence imaging, reflectance spectroscopy, and vital staining technique. Sharma et al (2021) investigated 63 lesions in 45 patients with the Identafi® 3000 device and toluidine blue staining and compared the results with histological examination. The overall sensitivity and specificity of the violet light were 0.73 (95% CI, 0.59 to 0.86) and 0.46 (95% CI, 0.27 to 0.67), of green–amber light were 0.78 (95% CI, 0.62 to 0.90) and 0.15 (95% CI, 0.04 to 0.35), and of toluidine blue were 0.51 (95% CI, 0.34 to 0.68) and 0.85 (95% CI, 0.65 to 0.96), respectively. A statistically significant association was observed only between the toluidine blue staining and histological results (P = 0.04) (Sharma et al 2021).
Adeoye et al (2021) used a time-to-event AI-based model to examine data from 716 patients with diagnosed oral leukoplakia and oral lichenoid lesions. They used anamnestic, clinical, and pathological data to estimate malignant transformation potential. They highlighted the possibility of applying this AI-based predictive tool in clinical settings, particularly in areas with limited access to specialists.
Research frequently leverages online datasets; however, many are limited in size and derived from commonly used sources such as Kaggle and GitHub. Sahoo et al (2024) revealed that of 55 studies, 26 relied on such databases, underscoring the need for globally integrated data-sharing frameworks to advance AI applications, particularly in complex diseases such as OPMDs and oral cancer, which demand diverse and representative imaging data. High-quality AI development depends on the effective curation of large, well-annotated datasets, with standardized patient selection and image segmentation to reduce variability. Global collaborative efforts, such as The Cancer Imaging Archive, play a crucial role in addressing these challenges by offering access to robust, extensively labeled datasets (Sahoo et al 2024).
The findings from this systematic review and meta-analysis provide qualitative and quantitative evaluations of the current use of AI-based tools in diagnosing OPMDs while highlighting potential directions for future research. Comprehending the capabilities of these AI tools may open new opportunities for early detection and prompt treatment of OPMDs, potentially improving patient outcomes and preventing the development of oral cancer.
Strengths and Limitations
Meta-analysis is placed at the top of the evidence pyramid, but the generalizability of this study is limited. As strict eligibility criteria were applied, only a few articles were found. None of the eligible studies provided information on lesion severity, a potentially critical variable influencing diagnostic accuracy.
In terms of the strengths of this study, a preregistered protocol was followed to ensure transparency and a rigorous methodology was applied. All images used in the eligible studies were biopsy proven, and most articles had a low risk of bias, confirming results of this study.
Implications for Practice and Research
The rapid application of scientific results is of utmost important (Hegyi et al 2020; Hegyi et al 2021). Findings of this study indicate that AI-based diagnostic tools hold potential for assisting general practitioners in evaluating the need for specialist referral. However, these tools are not a substitute for histopathological examination, which remains the gold standard. Given the high negative predictive value observed, a negative result may reduce the immediate need for specialist consultation. Nonetheless, clinical judgment should always guide decision-making. Future prospective studies using standardized imaging protocols, high-quality image acquisition, and large, diverse datasets are essential to rigorously assess the diagnostic performance of AI-based tools and to enable meaningful comparisons across studies. Policy discussions are warranted to explore the integration of AI-driven diagnostics into routine health care and screening programs, with the aim of enhancing efficiency and contributing to the early detection and prevention of oral cancer.
Conclusion
Based on the analyzed studies, AI-based diagnostic tools offer a noninvasive approach that may be capable of screening for OPMDs with high negative predictive values. At its current stage of development, AI may help increase doctor–patient interactions within at-risk populations, potentially leading to more timely diagnosis of lesions. Nonetheless, the ultimate clinical decision must always rest with the physician.
Author Contributions
D. Horvath, contributed to conception and design data acquisition, analysis, and interpretation, drafted the manuscript; A. Fekete, contributed to data acquisition and analysis, critically revised the manuscript; P. Martinekova, contributed to conception and design, data analysis, critically revised the manuscript; S. Kiss-Dala, contributed to conception, data analysis and interpretation, drafted and critically revised the manuscript; E. Abram, contributed to data interpretation, critically revised the manuscript; K. Marton, contributed to data acquisition and interpretation, critically revised the manuscript; A. Zsembery, contributed to conception, critically revised the manuscript; P. Hegyi, contributed to conception and design, critically revised the manuscript; A. Brody, contributed to conception and design, data acquisition and analysis, critically revised the manuscript. All authors gave their final approval and agreed to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jct-10.1177_23800844251403962 – Supplemental material for Artificial Intelligence Helps Diagnose Oral Potentially Malignant Disorders: A Systematic Review and Meta-Analysis
Supplemental material, sj-docx-1-jct-10.1177_23800844251403962 for Artificial Intelligence Helps Diagnose Oral Potentially Malignant Disorders: A Systematic Review and Meta-Analysis by D. Horvath, A. Fekete, P. Martinekova, S. Kiss-Dala, E. Abram, K. Marton, A. Zsembery, P. Hegyi and A. Brody in JDR Clinical & Translational Research
Footnotes
Acknowledgements
There is nothing to declare.
Declaration of Generative AI and AI-Assisted Technologies in the Writing Process
During the preparation of this work, no generative AI or AI-assisted technologies were used.
Ethical Approval
No ethical approval was required for this systematic review with meta-analysis, as all data were already published in peer-reviewed journals. No patients were involved in the design, conduct, or interpretation of this study. The datasets used in this study can be found in the full-text articles included in the systematic review and meta-analysis.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: open-access funding was provided by Semmelweis University. This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Data Availability Statement
All data used in this meta-analysis were extracted from previously published studies. No new primary data were generated. The extracted dataset supporting the findings has been uploaded and will be available in Figshare.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
