Abstract
Purpose
This study aims to identify specific genotypes within the UK Biobank (UKB) cohort contributing to a genetic predisposition for bladder cancer (UBC). It highlighted the impact of environmental exposures and the broader role of certain genes in UBC development, offering a comprehensive understanding of the genetic basis for UBC susceptibility.
Experimental Design
Leveraging the rich data from the UKB- a longitudinal study involving participants across the UK-the primary outcome was the presence of UBC, determined using ICD-10 and ICD-9 codes. The study employed rigorous Genome-Wide Association Study (GWAS) protocols, Phenome-Wide Association (PheWAS) frameworks, and gene-level pleiotropy analyses. Quality control measures were applied, such as single-nucleotide polymorphisms (SNP) missingness and minor allele frequency thresholds. Polygenic Risk Score (PRS) evaluations were also conducted based on the Mavaddat score using UKB's high-density genome-wide SNP dataset.
Results
Our GWAS identified significant associations between UBC risk and genetic variants, notably in the PSCA and TERT genes. The UGT1A polymorphism was found to be protective against UBC, particularly in heavy smokers. The PheWAS framework linked UBC-predisposition polymorphisms to other conditions, such as prostate cancer.
Conclusions
Our GWAS identified significant associations between UBC risk and genetic variants across loci, including PSCA, TERT, TACC3 and TMEM129. The protective effect of the UGT1A variant against UBC, especially concerning tobacco exposure, suggests the potential for genetic-based preventive strategies in UBC management.
Introduction
Urothelial bladder cancer (UBC) is identified as the sixth most frequent cancer in males and ranks tenth in the overall global cancer incidence for both sexes. According to the most recent GLOBOCAN statistics, 2020 registered a cumulative incidence of 573,000 new cases, resulting in 213,000 mortalities. 1 While a majority of UBC cases are thought to be associated with external risk factors such as tobacco smoking, occupational and environmental exposure. 2 Nearly 12% of underlying risk is estimated to be attributed to genetic susceptibility. 3 Evidence supporting the genetic component of UBC is seen in the highly penetrant gene variants BRCA2, ATM, MLH1, and MSH2. 4
Genome-wide association studies (GWAS) have identified several single-nucleotide variations (SNPs) associated with UBC risk.3,5 However, cumulative genetic risk contribution and combined effects of risk exposure may have more important clinical implications for risk stratification and efforts to achieve early cancer detection and screening programs.
While GWAS have been used to explain the substantial risk of UBC development, post-GWAS studies have focused on delineating the functional mechanism of SNPs and their tumor-promoting or tumor-suppressing effects or biological consequences behind risk loci. 6 Herein, we describe the prevalence of germline cancer risk variants among a large cohort of BC patients and employ functional annotation tools to infer the biological roles of these variants in UBC development. In addition, we analyze environmental, occupational, and non-modifiable risk factors and their association with UBC development within the context of germline cancer risk.
Methodology
Population
This investigation utilized both genetic and phenotypic datasets from the United Kingdom Biobank (UKB), a longitudinal, population-based study encompassing participants from England, Scotland, and Wales. The UKB provides a rich data repository, including a diverse array of phenotypic markers, imaging data, healthcare-related information, and genotypic data acquired through whole-genome sequencing. This study represents a case-control retrospective observational study to determine the comparative prevalence of germline variants between patients with and without UBC. All eligible controls from the UKB cohort were included to maximize statistical power and population representation. While a 1:4 case-control matching design could have been used, the unmatched design retained flexibility for exploratory analyses and minimized potential bias, consistent with prior GWAS utilizing UKB data. The study cohort was delineated based on the International Classification of Diseases, Tenth Revision (ICD-10) and Ninth Revision (ICD-9), with the primary outcome of interest being the presence of UBC, further elucidated by tumor histology (refer to Supplementary Table 1). Control subjects were designated as individuals without any cancer history, as indicated by ICD-9/10 codes, self-reporting, or physician diagnosis within the UKB database. The resultant control group, as determined by our inclusion and exclusion criteria, comprised 375,981 participants. To ensure the robustness of the analysis and minimize confounding due to population stratification, the study primarily included participants of European ancestry. Ancestry was confirmed using PCA, and all GWAS models included the first ten principal components as covariates. Additionally, this study utilized genome-wide SNP array data from the UKB to conduct the GWAS analysis. While WES data was available for some participants, the genetic data used in this study were derived from genome-wide genotyping arrays, ensuring comprehensive coverage of SNPs across the genome. The GWAS analysis was conducted using genotyping array data from the UK Biobank. While WES data was available for a subset of participants, it was not used in this study.
GWAS analysis
We adhered to the standard data manipulation and analytical protocols as endorsed by the UKB research analysis framework and as previously described in the literature as the gold standard for GWAS (https://ukbiobank.dnanexus.com). Given that the presence of UBC is conceptualized as a dichotomous trait, a linear mixed model was employed in GWAS to optimize statistical power while simultaneously controlling Type I errors due to case-control imbalance. Before initiating GWAS, preliminary quality control (QC) measures involved evaluating the missingness of SNPs and individual data, conducting a QC cross-check between recorded gender and X-chromosomal data, and setting a minor allele frequency (MAF) threshold of 0.05.
In the initial phase of the analysis, age, sex, and body mass index (BMI) were incorporated as covariates. Subsequent to the GWAS, clumping analysis was conducted utilizing PLINK2 software, focusing on variants with p-values less than 5 × 10^-8 and a linkage disequilibrium (LD) R^2 threshold of 0.1 for identifying independent GWAS loci. LD score regression analyses were performed to estimate SNP-based heritability, ascertain potential confounding bias, and evaluate genetic correlation. As initially outlined, Q metrics were rigorously applied before the GWAS execution. Beta effect sizes and ORs were calculated relative to the reference alleles. For example, the T alleles of rs2294008 in PSCA and the T allele of rs17863783 in UGT1A were used as effect alleles, with the C and G alleles as references.
Phenome-wide association framework (PheWAS)
We set out to determine whether genetic predisposition to UBC is also associated with the risk of other phenotypes. The GWAS models were adjusted for age, sex, BMI, and the first ten genetic principal components to account for population stratification. Smoking status was not included in the primary GWAS model to maintain focus on genetic susceptibility, but it was analyzed in subsequent interaction and subgroup analyses. 7 This model was used to evaluate the association of all SNPs with each trait under the assumption of additive allelic effects by PLINK2. P values were corrected with multiple tests using the false discovery rate (FDR) with an adjusted threshold of 0.05. We estimated the odds ratio (OR) between UBC predisposition polymorphisms and multiple diseases utilizing the PheWAS R package for all significant associations. (R version 3.6.1) The PheWAS analysis included 11 SNPs identified from the GWAS, which met the genome-wide significance threshold (p < 5.31 × 10−8). These SNPs were located within four loci: PSCA (6 SNPs), TERT (2 SNPs), TACC3 (1 SNP), and TMEM129 (1 SNP). The analysis also included the UGT1A variant (rs17863783: G > T) based on its association with bladder cancer risk in our study. The PheWAS was conducted using the R PheWAS package, with significant associations determined after multiple testing corrections (FDR q < 0.05). All genetic variant positions and annotations are based on the GRCh38 (hg38) genome reference build.
Gene-level genetic pleiotropy analysis
To establish the influence of SNPs on multiple phenotypic traits and assess the broader impact of specific genes on the development of UBC, we measured the degree of pleiotropy utilizing the SNP2gene function of FUMA to map SNPs in GWAS to a gene with 100 genome phase 2 EAS as a reference panel. 8 FUMA is a bioinformatics tool that integrates many data sources, including LD structure, functional scores, and chromatin interactions, to correlate associated variants with pertinent genes. Initially, FUMA identifies significant independent variants and the surrounding genomic regions based on LD structure. These variants are then annotated with tools and databases such as ANNOVAR, CADD, RegulomeDB, and Hi-C data.9–11 Following this, the annotated variants are allocated to genes based on their position, eQTL association, and chromatin interaction using the default parameters set by FUMA. For evaluating variant-level pleiotropy, we determined the count of associated traits that met a genome-wide significance threshold (p < 5.31 × 10−8). Functional mapping was conducted using the 1000 Genomes Phase 2 as a reference panel, aligned with the population-specific parameters used in this study. We identified deleterious coding SNPs, either exonic or splicing with CADD score >20.00 and expression quantitative trait loci (eQTLs) of defined tissue types (FDR ≤ 0.05 for analysis) within the 3'UTR. We included deleterious coding SNPs, either exonic or splicing with CADD score >20.00 and expression quantitative trait loci (eQTLs) of defined tissue types (FDR ≤ 0.05 for the analysis). To obtain insight into the putative biological mechanism of prioritized genes, we utilized the GENE2FUNC process to annotate these genes in a biological context. Genomic positions and allele changes provide precise identification of the variants, and these details were incorporated into the logistic regression models to account for the effect of allele frequencies. Herein, we describe the prevalence of germline cancer risk variants among a large cohort of UBC patients and employ functional annotation tools to infer the biological roles of these variants in UBC development.
Polygenic risk score (PRS) evaluation
Phenome-wide association analysis (PheWAS) was conducted using the PheWAS R package (version 3.6.1), and the PheWAS catalog was accessed at https://phewascatalog.org for reference and mapping of PheCodes. PRS was derived based on the Mavaddat score using the available UKB high-density genome-wide SNP data set. PRS for UBC was constructed using all SNPs previously identified by GWAS to contribute to UBC risk. Quality control on target samples using PLINK using MAF (0.05), missing samples and genotype of call rates 0.1, exclusion of variants failing Hardy-Weinberg equilibrium test at the 10 × 10−6. The PRS for UBC was constructed using the SNP identified utilizing the hard threshold approach (Supplementary Figure 1). 12 SNPs were selected based on their independent association with UBCa, determined via LD clumping (r² < 0.1). In traditional GWAS, SNPs are manually selected, and effect sizes (weights) are predefined based on the GWAS results. However, in our PRSice-2 analysis, the process is automated, and the effect sizes (weights) are automatically derived from the GWAS summary statistics file used in PRSice-2. The SNPs for the PRS model were selected from the “significant_variants_prs.txt” file generated by the UK Biobank, which includes SNPs with varying P-values and effect sizes. PRSice-2 relies on the GWAS summary statistics file and applies statistical thresholds (p < 0.05) and LD clumping to identify independent SNPs for inclusion in the final PRS model. The PRS model incorporated age, sex, BMI, and smoking status as covariates and was calculated using the additive genetic model, where the score represents the sum of weighted risk alleles carried by an individual. The PRS model incorporated age, sex, BMI, and smoking status as covariates and was calculated as the sum of weighted risk alleles. Detailed methodology can be found in Supplementary Material 1.
Results
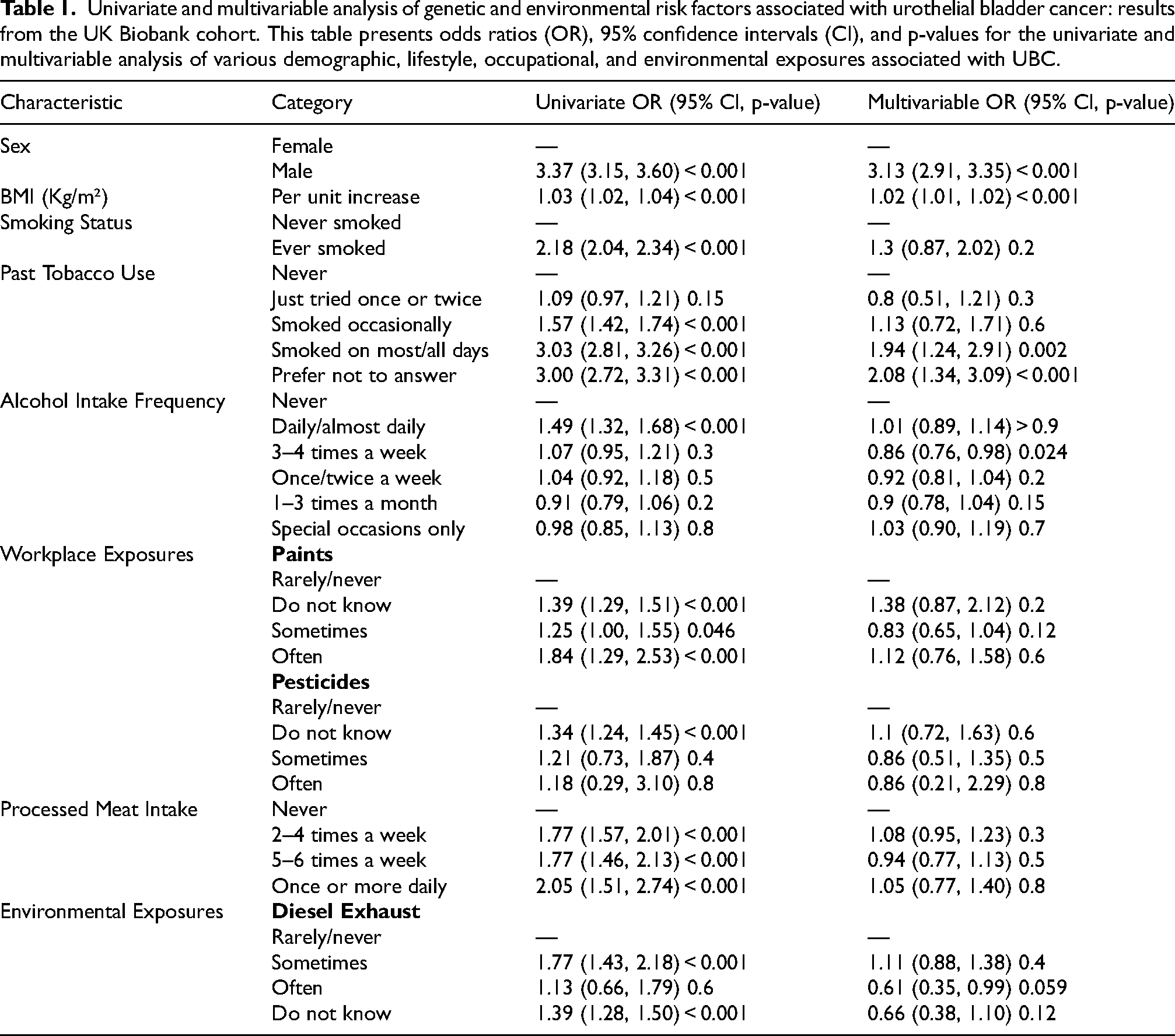
The total number of UBC cases was 5071 after quality control, and the total number of controls was 375, 419. The mean (SD) age at recruitment of the UBC patients and controls was 62.03(± 5.93) and 55.52 (± 8.12) (p < 0.001); BMI 28.17 (±4.68) and 27.38(±4.79) (p < 0.001), respectively. In terms of exposure, UBCs were more likely to be current smokers 16.19% vs 10.38% (p < 0.01) or have a history of previous smoking 49.33% vs 33.05% (p < 0.01). In addition, UBC patients were living in areas of high nitrogen oxide 44(± 15.74) vs. 26.84(7.63) (p > 0.001) but not particulate matter air pollution 16.17(1.86) vs. 16.25(±1.90) (p = 0.2). Both UBC and non-cancer controls reported similar rates of proximity to major roads (7.07% vs. 7.62%), night work (1.57% vs 1.12%), pesticide exposure (0..07% vs 0.06%), total sugar intake (g) 127 ((±60.44) vs 129 (±58.27), current alcohol use (92.24% vs 91.61%), however, UBC were less likely to report history of no meat intake (6.21% vs 9.55%, p < 0.001). Supplementary Table 2 represents characteristic differences among the two cohorts. After adjusting for age, sex, BMI, and smoking status high, no significant associations between frequent exposure to paints (OR 1.38, p = 0.6), pesticides (OR 0.86, p = 0.8), workplace chemical exposure (OR 1.23, p = 0.2) and workplace cigarette smoke (OR 1.02, p = 0.8), diesel exhaust (OR =1.11, p = 0.4), meat (OR 0.94, p = 0.8) and sugar intake (OR 1.0, p = 0.12), night shift work (OR = 0.96, p = 0.8) and particulate matter (OR 0.95, p = 0.083)and nitrogen oxide air pollution(OR 1.0, p = 0.4) and UBC were found. There remained a strong association between the male sex (OR 3.13, p < 0.001), smoking status (OR 1.94, p = 0.002), and a small increase in UBC risk with increasing BMI (OR 1.02, p < 0.001). Surprisingly, frequent alcohol intake of 3–4 times per week was found to be protective against the development of UBC (OR 0.86, p = 0.024) (Table 1).
Univariate and multivariable analysis of genetic and environmental risk factors associated with urothelial bladder cancer: results from the UK Biobank cohort. This table presents odds ratios (OR), 95% confidence intervals (CI), and p-values for the univariate and multivariable analysis of various demographic, lifestyle, occupational, and environmental exposures associated with UBC.
Genome-Wide association analysis of UBC
We found 11 SNPs that meet the threshold for GWAS (p < 5.31 × 10−8) significant for the association between variants and UBC. (Figure 1). These included six individual SNP within the PSCA gene, 2 SNP within TERT, 1 SNP in TACC3, TMEM129. All the above-mentioned SNP have been previously reported to be associated with UBC progression development, such as an overexpression of PSCA found to be associated with bladder, prostate, and pancreatic cancers.13–15 Furthermore, somatic TERT promoter variants in UBC have been associated with worse prognosis and found in 17.1% of patients. Recent studies suggest TERT variants are among UBC's earliest genetic events.16,17 In terms of Beta Effect Size and overall risk of UBC development with associated germline variants, SNP within PSCA (Beta 0.8), TERT (0.26), TMEM129 (0.15), and TACC3 (0.14) were found to be positively associated with the development of UBC. In contrast, variants within UGT1A, THEM6, SLC14A1, LY6K/D were protective. (Figure 1) The T allele of rs2294008 in PSCA was associated with increased UBC risk (OR 1.18, 95% CI: 1.12–1.23), while the T allele of rs17863783 in UGT1A conferred a protective effect (OR 0.66, 95% CI: 0.56–0.77). To put this within odds of development of UBC as compared to individuals with no associated variants, PSCA OR 1.18(95%CI 1.12–1.23) (variant 8:142680513:C: T), TERT OR 1.18 (95% CI 1.12–1.26) (variant 5:1280362: G:A), TACC3 OR 1.15(95%CI 1.09–1.22)(variant 4:1730846:GGGGT: G) and TMEM129 OR 1.15(95% CI 1.09–1.22) (variant 4:1717567:T:C). A sequence variant within UGT1A (UDP glucuronosyltransferase family 1 member A6) (variant 2:233693631: G: T) was among
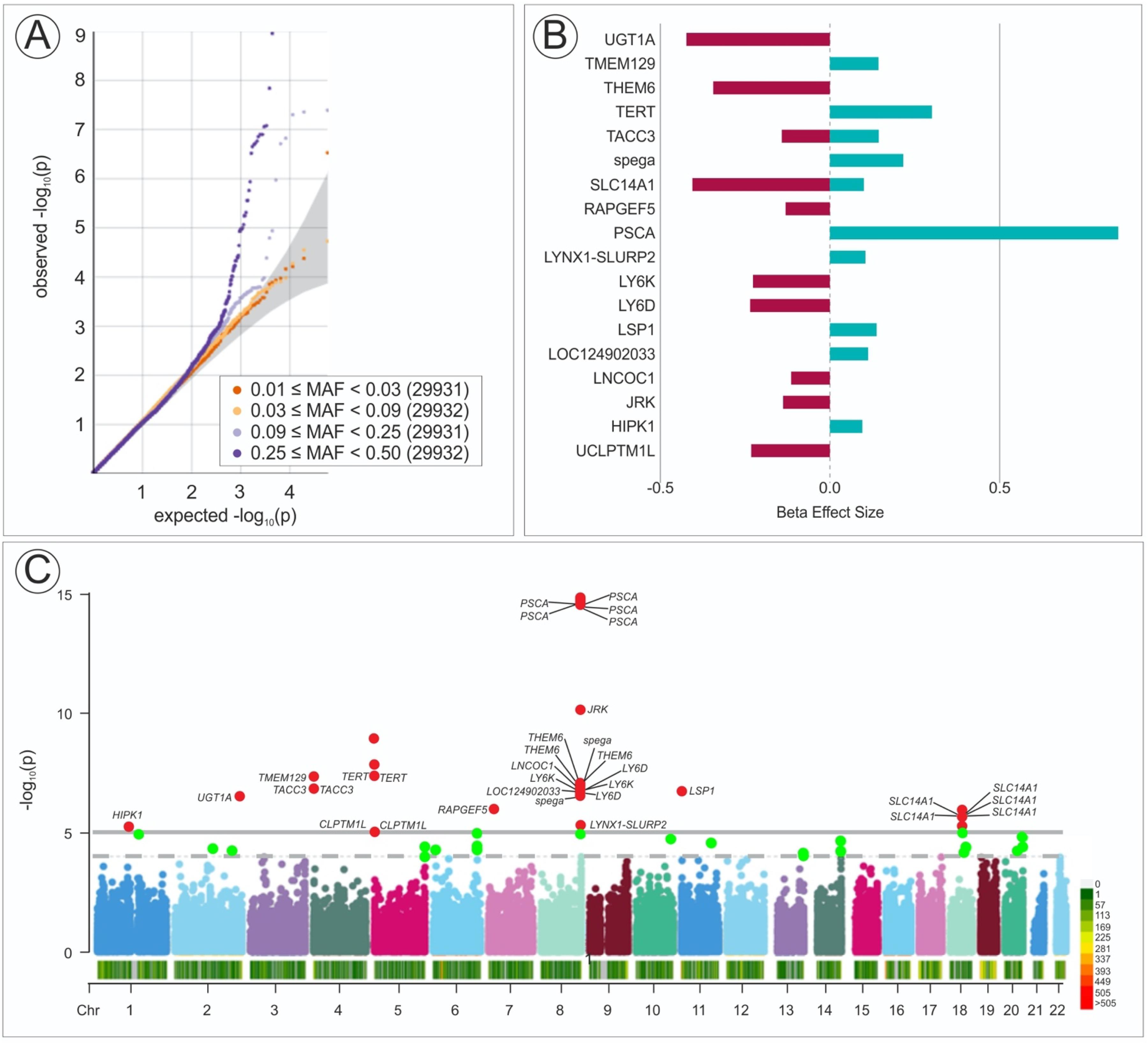
GWAS results from UK Biobank showing top variants associated with UBC. (A) Quantile-quantile (Q-Q) plot with MAF categories. (B) The effect of SNPs and B-coefficient (Beta effect size) for UBC risk. Positive values indicate stronger SNP effects. (C) Manhattan plot showing GWAS results for UBC with a red horizontal line indicating the genome-wide significance threshold (p < 5.31 × 10−8). Index variants and significant loci are highlighted in red. The y-axis represents -log10(p-values), and the x-axis represents genomic positions.
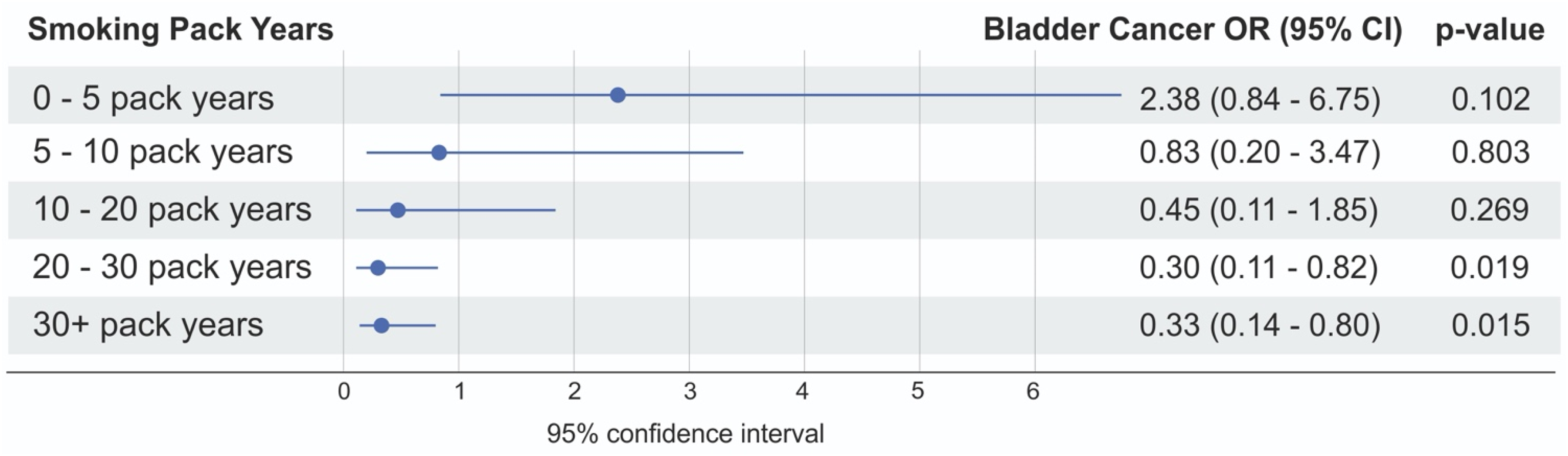
Association between smoking pack-years and bladder cancer risk. The figure shows odds ratios (OR) with 95% confidence intervals (CI) across different smoking categories based on cumulative pack-years. The categories include 0–5(n = 5078), 5–10(n = 5085), 10–20 (n = 4951), 20–30 (n = 4185), and 30 + pack-years (n = 3988). A dose-dependent decrease in OR is observed with increasing pack-years, indicating a protective effect for certain cumulative smoking exposures. Statistical significance is highlighted for categories with p-values < 0.05. UGT1A (variant 2:23369363:G: T) and associated risk of UBC development with smoking duration, highlighting decreased OR with > 20 pack years and UGT1A variant. Results highlighting protective effects of UGT1A variants in heavy smokers.
Locuszoom associations of GWAS
LocusZoom plots were used to visualize the associations of SNPs within significant loci, highlighting local linkage disequilibrium (LD) and recombination patterns. These plots provide a graphical overview of the regional genetic architecture but do not infer causality or augment the robustness of the analysis. We identified 11 SNPs meeting the threshold for GWAS significance (p < 5.31 × 10−8) for association with UBC. These include a single independent signal within the PSCA gene, represented by six SNPs in high LD. To identify potentially causative SNPs, we utilize LocusZoom plots to show the magnitude of the associations for each SNP and the pairwise local LD and recombination patterns/positions in the region of the SNPs of interest.18,19 We focused on 6 individual SNP within the PSCA gene, 2 SNP within TERT, 1 SNP in TACC3, TMEM129 and a sequence variant within UGT1A (UDP glucuronosyltransferase family 1 member A6), (variant 2:233693631: G: T). (Supplementary Figure 2). Measurement of LD provides a measure of non-random association of alleles at two or more loci. Our results indicated high LD of TACC3 with FGFR3, TMEM129, and SLBP, suggesting molecular phenotype of TACC3-associated SNP could be associated with differential expression of more than one gene (FGFR3, SLBP, and TMEM129). This finding has been replicated independently within TCGA data by a separate group. 20 This was contrary to LocusZoom for the UGT1A variant, showing minimal shared association with neighboring SNPs.
Phenome wide association (PheWAS)
A total of 11,544 unique ICD10 and 3109 ICD9 codes were summarized from hospital inpatient, cancer registry, and death registry data of the UKBB cohort. These codes were mapped to 1647 distinct PheCores. We restricted the analysis to PheCodes with at least 20 cases as recommended from previous PheWas guidelines. 21 And grouped into 15 disease categories. Four PheCODEs and two SNP variants were associated with UBC risk at FDR q < 0.05. These PheCODES belonged to prostate cancer (telomerase reverse transcriptase (TERT) variant ID: rs2242652variant ID: rs2242652, OR 1.22, FDR p < 0.0001), and dermatologic condition of the skin including seborrheic keratosis (TERT) variant ID: rs2242652, OR 1.35, p < 0.001). (Figure 3). It is important to distinguish that PheWas does not correlate based on the co-occurrence of ICD9/10 but on UBC predisposition polymorphism and multiple diseases. In other words, co-occurrences of prostate cancer and UBC were seen in patients who carry the variant: rs2242652, subsequently increasing the odds of development of UBC by 22% compared to patients without the variant. These patients are at increased risk for the development of prostate cancer, seborrheic keratosis (SK), and UBC. The interpretation of effect sizes and ORs depends on the reference alleles used in the analysis. For instance, the T allele of rs2294008 in PSCA and the T allele of rs2736098 in TERT were associated with increased UBC susceptibility, while the T allele of rs17863783 in UGT1A showed a protective effect. These findings highlight the importance of specifying reference alleles to ensure biological relevance and reproducibility. All identified SNPs have been previously reported as GWAS signals associated with UBC susceptibility. These include loci within PSCA, TERT, TACC3, TMEM129, and UGT1A, which are consistent with findings from prior GWAS studies.4,22,23
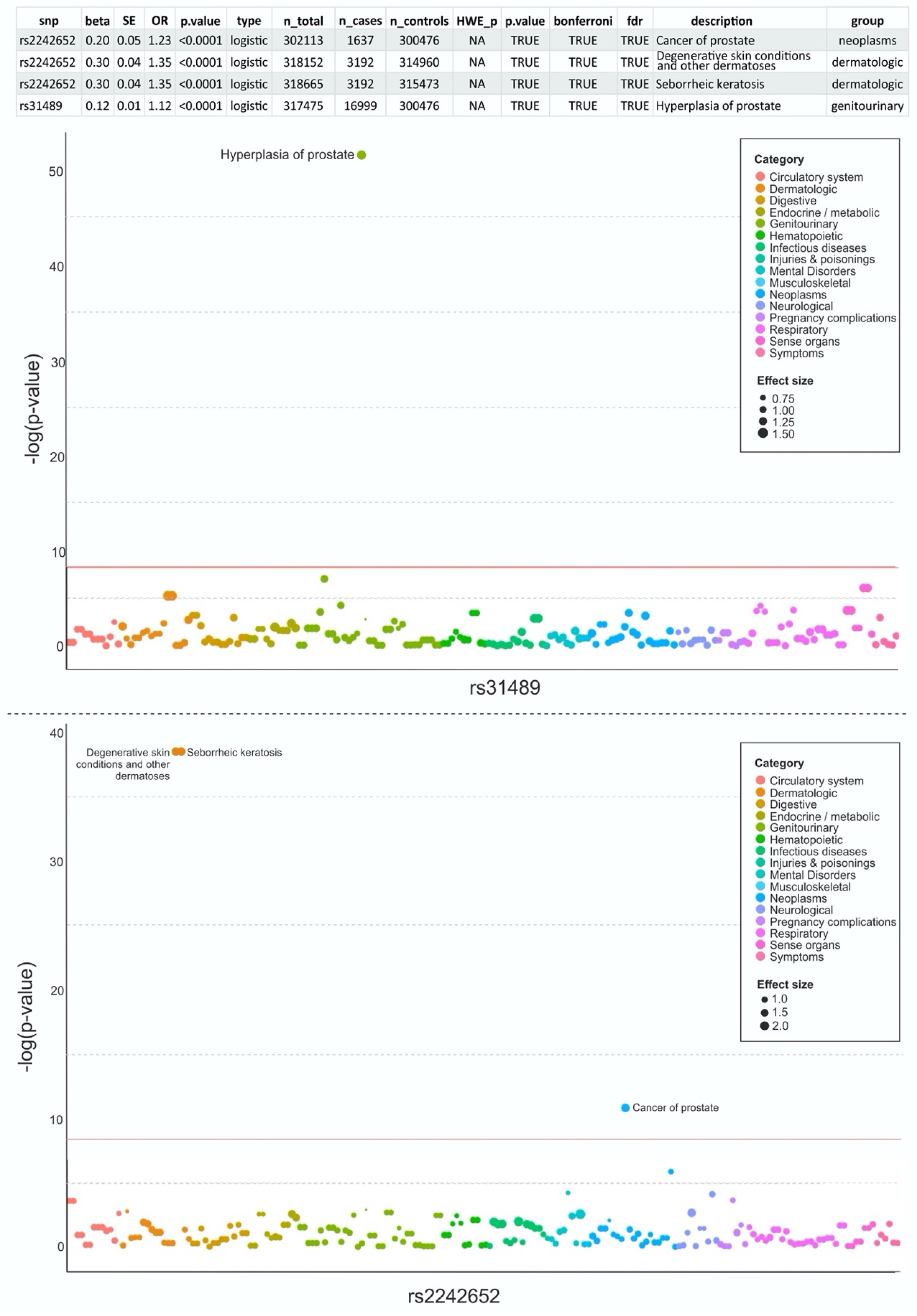
Each point of the Manhattan plot represents an SNP, with the y-axis displaying the negative log-transformed p-value (-log(p-value)) for the association between each SNP and the condition. The SNPs are color-coded by their category (e.g., genitourinary, dermatologic, neoplasms), as shown in the legend, and the size of the points corresponds to the effect size, where larger points indicate a greater effect size (0.75, 1.00, 1.25, and 1.50). The horizontal red lines represent the threshold for statistical significance (p < 0.05). Data from the UK Biobank were used for this analysis, and the results are based on logistic regression models adjusting for relevant covariates, including age, sex, body mass index, and smoking status.
Gene-level functional analysis
We conducted FUMA analysis to map and annotate the genetic associations to understand better the genetic mechanism underlying UBC. 8 Using the GWAS UKB data, FUMA analysis identified eight genomic risk loci (Figure 4), with nine lead SNPs from 578 candidate SNPs and 308 mapped genes. GSEA was undertaken to test the possible biological mechanism of the eight candidate genes implicated in UBC, with adjusted p < 0.05 used for further analysis and based on prioritized genes, putative biological mechanisms revealed that most overlapping genes of significance are associated with flavonoid and xenobiotic glucuronidation in addition to metabolic processing of xenobiotics with the overlapping genes of interest, including UGT1A1, 6–10. SNP within UGT1A8 represented the strongest association with UBC with enrichment P-value (−10 log10). The SNP within UGT1A8 that represented the strongest association with UBC is rs17863783 (G > T), as identified through GWAS and functional annotation analyses. This again highlights the importance of UGT1A polymorphism and the association of UBC, which is highlighted in GWAS (Figure 4).
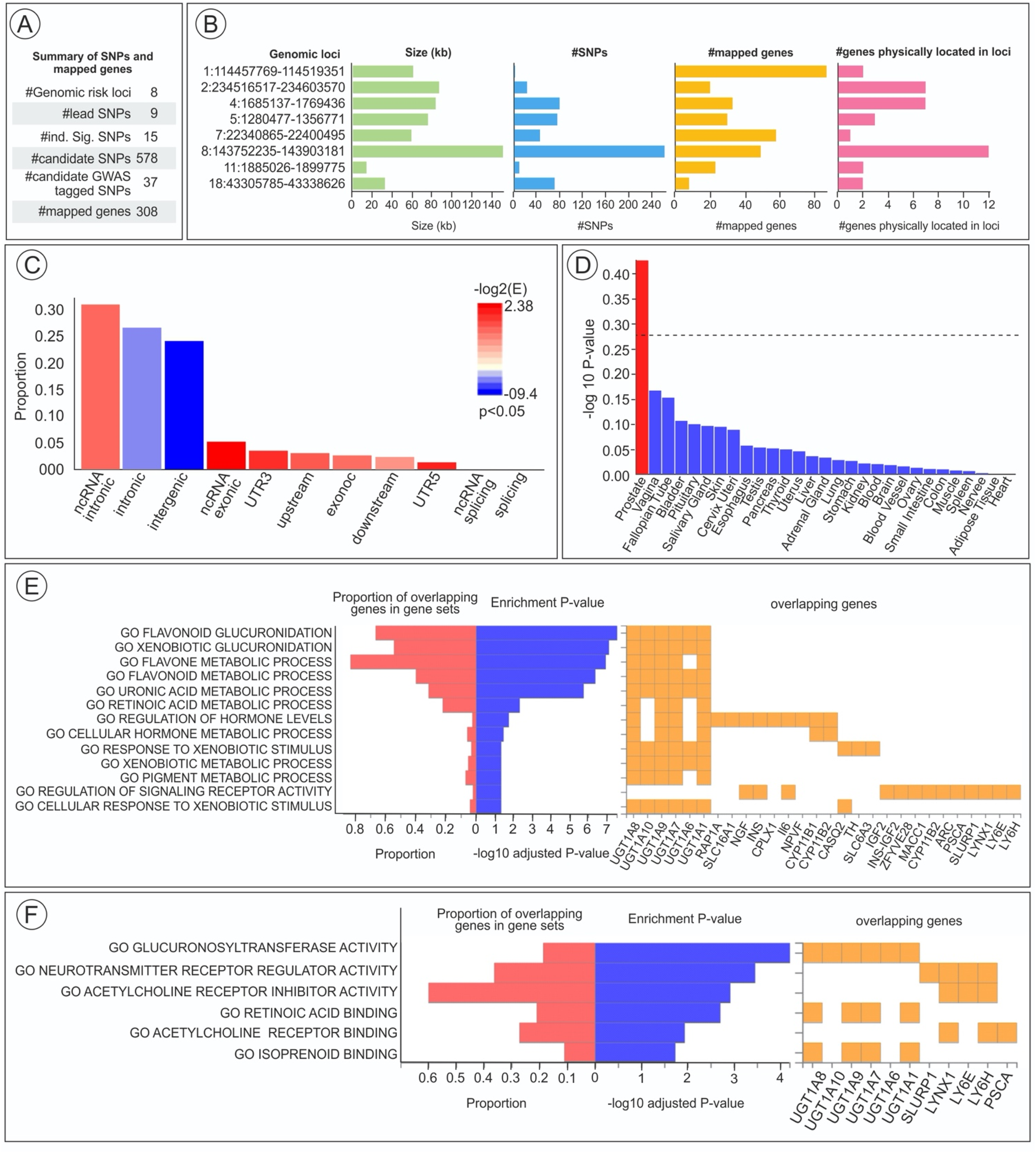
FUMA analysis showing A. Number of Identified SNPs and genomic risk loci with representative B. Size(kb), number of SNPs, and mapped genes. C. Proportion of UBC-associated genes in GWAS SNPs in different genomic annotation categories. D. Tissue-specific gene expression and shared biological function across multiple tissue types. Significant enrichment at Bonferroni corrected p-value ≤ colored in red. (E-F) Gene set enrichment analysis represented the biological function of prioritized genes of interest.
Polygenic risk score
At the p-value threshold of 0.001 for SNP selection, the PRS model fit corresponded to poor PRS association with UBC, with improved predictive capability at the SNP p-value threshold of 0.05 and 0.1 with subsequent PRS model fit of 0.018 and 0.025. In other words, the PRS model with input of SNP with present p < 0.05 and p < 0.1 from GWAS could explain 1.8% and 2.5% of the variance in UBC.
Discussion
Our study focuses on performing a GWAS to identify genetic loci associated with UBC risk, supported by functional analyses and phenotypic evaluations. This investigation aimed to elucidate specific genotypes contributing to a genetic predisposition for UBC within the UKB cohort, focusing particularly on environmental exposure contexts and functional pleiotropy. Concordant with existing literature, our GWAS identified significant associations between elevated UBC risk and particular genetic variants localized in PSCA, TERT, TACC3, and TMEM129. The genetic loci within the PSCA gene on chromosome 8q24, initially reported by Wu et al., 5 and subsequently corroborated by Rothman et al., consistently demonstrate an association with susceptibility to UBC, with an OR closely mirroring our findings (OR 1.13 in prior work versus OR 1.18 in this study).22,23 Moreover, our analytical framework uncovered supplementary distinct variants within the PSCA gene (rs2976393, rs1045531, rs2978982, rs2976394, rs2976391), augmenting the well-established rs2294008 variant previously confirmed as strongly associated with UBC risk. Variants in the TERT and FGFR3 genes represent the most frequently occurring somatic alterations in urothelial carcinoma of the bladder. Our investigation corroborated an association between germline TERT variants, specifically rs13167280(G > A) and rs2736098(C > T), and elevated risk of UBC. Notably, the rs2736098(C > T) polymorphism, situated in the second exon of the TERT gene, is a synonymous mutation that does not alter the amino acid sequence (Asn305Asn). Nonetheless, the presence of the A allele has been linked to shortened telomere lengths.
The robustness of our GWAS was augmented by incorporating LocusZoom, a bioinformatic tool facilitating the detailed examination of relationships between individual SNPs and their adjacent genetic loci. LocusZoom is a standard visualization tool in GWAS that summarizes SNP associations within loci and their LD relationships. While useful for interpreting regional genetic architecture, it does not infer causality or enhance analytical robustness. This methodology aims to unveil potential interlocus interactions. Specifically, the SNP in TACC3, designated as rs199500838 GGGGT: G, representing an intronic variant deletion, demonstrated high linkage disequilibrium with the FGFR3 and TMEM129 genes. This observation may indicate a shared genetic architecture or co-regulatory mechanisms between these loci. Given FGFR3's established involvement in UBC cellular proliferation and differentiation and the nascent understanding of TMEM129's functionalities, such associations may yield valuable insights into co-dependent mechanisms and predispositions to UBC. Subsequent functional analyses are imperative to clarify the biological significance of these genetic interrelationships. The presence of the UGT1A polymorphism (rs17863783 G > T) was found to confer a protective effect against the development of UBC. 24 The implicated T allele of rs17863783 is a coding synonymous variant (Val209Val) that impacts the mRNA expression of the functional isoform UGT1A6.1. Although synonymous variants do not inherently alter the amino acid sequence, this specific variant in UGT1A has been demonstrated to affect splice sites, thereby upregulating the mRNA expression of UDP-glucuronosyltransferase.
Exposure to aromatic amines, commonly found in industrial chemicals and tobacco smoke, is robustly associated with elevated UBC risk. 2 UGTs function to conjugate UDP-glucuronic acid with the N-hydroxylated derivatives of various substrates, including aromatic hydrocarbons, thereby rendering them water-soluble and facilitating their excretion through feces and urine. While the initial conjugation is principally carried out by hepatic Phase II metabolism, the resultant water-soluble glucuronides may become unstable in urine, especially at acidic pH levels (<6). This can result in the reformation of oncogenic forms of aromatic amines, creating DNA adducts and initiating carcinogenesis within the bladder epithelium.
UGT1A, which exhibits substantial expressions in bladder tissue, plays a role in re-conjugating aromatic amines to facilitate their excretion. 24 Our observation of the UGT1A variant (rs17863783 G > T) as conferring protective effects against UBC development, notably in individuals with high tobacco smoke exposure, aligns logically with its functional role. This variant enhances UGT1A expression within bladder epithelium via alternative mRNA splicing, thereby providing protection selectively in individuals exposed to specific environmental carcinogens, such as tobacco smoke and industrial chemicals while exerting a neutral effect in other contexts. Additional observational studies corroborate these findings 24 (Figure 5).
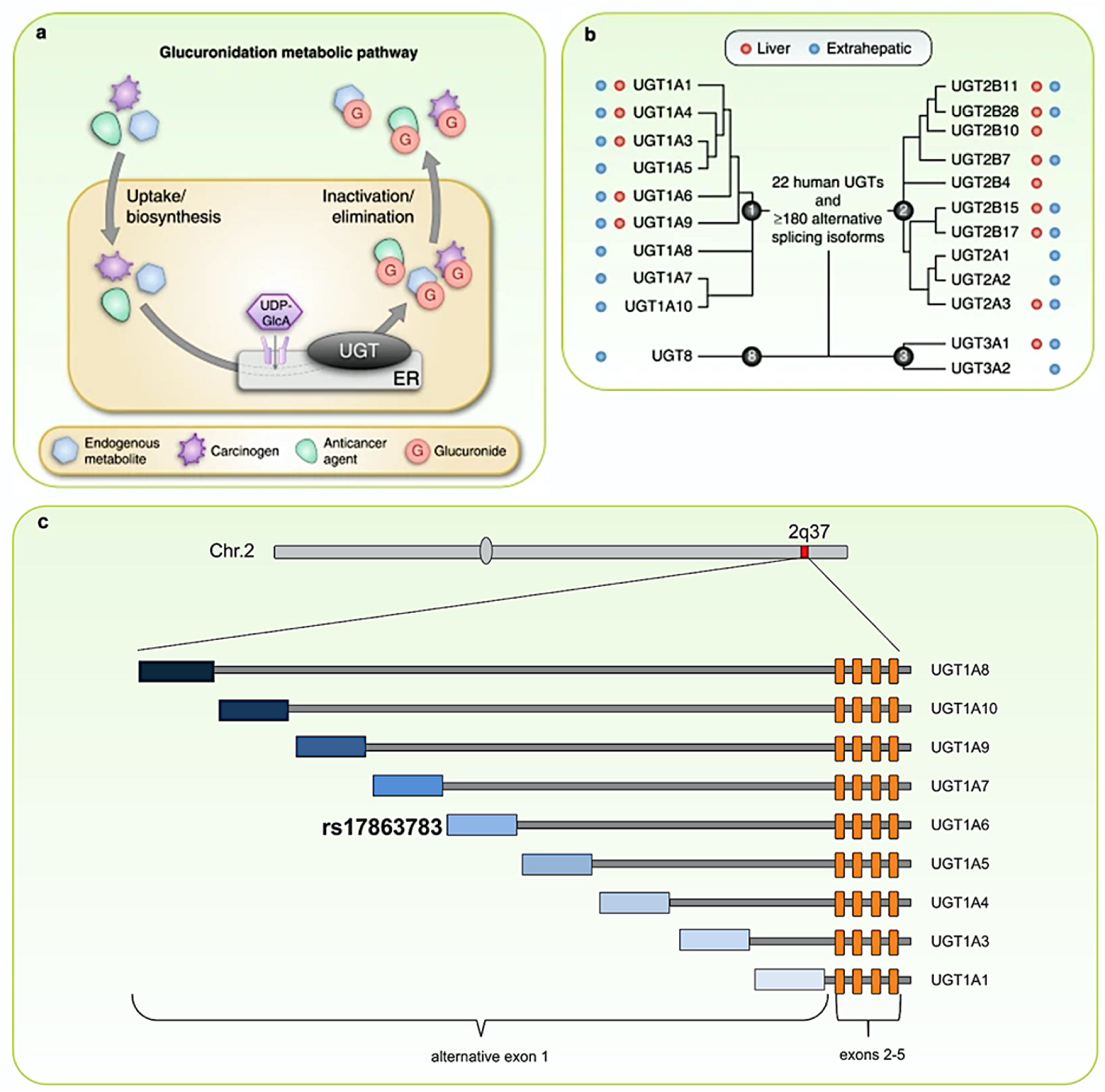
Glucuronidation metabolic pathway (a) and organ location (b), as well as associated documented polymorphisms (c) in UGT1A1-10 including rs17863783 (UGT1A6), synonymous variant (Val209Val), which impacts the mRNA expression of the functional isoform. (Figure modified from Dudec et al. 25 and Allain et al. 26 )
Our results were further substantiated through functional analysis using FUMA, identifying genomic loci 2:234516517-234603570 as significant in UBC incidence. The highest enrichment was observed in Glucuronosyltransferase activity along the biological pathway. Published research has correlated UGT1A immunostaining with normal bladder tissue, showing diminished immunoreactivity in tumor tissues. A robust correlation was noted between lower UGT1A expression and recurrence in patients with high-grade non-muscle-invasive UBC (NMIUBC). Additionally, UGT1A expression is positively regulated by 17β-estradiol, which may elucidate the sex disparities and underlying mechanisms contributing to the incidence of UBC. 27 While our findings are primarily hypothesis-generating, it is crucial to comprehend the potential ramifications of UGT1A variant activity on the risk of cancer development. Understanding this could pave the way for the creation of therapies aimed at prevention rather than merely curative interventions post-cancer onset. Similarly, screening for this specific UGT1A variant could aid in the risk stratification of patients undergoing surveillance as well as those in the diagnostic workup for hematuria.
Several well-established UBC susceptibility genes, including GSTM1 and NAT2, did not emerge in our analysis. The GSTM1 null genotype, characterized by a large deletion, is a known risk factor for BC, particularly in smokers, but may not be adequately tagged by the SNP genotyping array used in the UKB. Similarly, NAT2 acetylation polymorphisms, which influence the metabolism of aromatic amines, are typically analyzed as functional haplotypes rather than individual SNPs, which may explain their absence in our results. Recent studies, such as those by Hein et al. 28 and Garte et al., 29 suggest that SNPs can effectively tag these variants in certain populations. Future studies employing more targeted approaches or sequencing-based methods are needed to fully explore the contributions of GSTM1 and NAT2 variations to UBC risk.
Our findings regarding the protective role of the UGT1A6 variant (rs17863783:G > T) in UBC risk among heavy smokers are consistent with prior analyses of UGT1A variants and their interaction with smoking. 30 These studies highlight the role of genetic variation in modulating the effects of environmental exposures, such as tobacco smoke, on UBC susceptibility. The potential protective effect of the UGT1A rs17863783:G > T variant in heavy smokers is consistent with the role of UGT1A enzymes in detoxifying tobacco-related carcinogens. Previous studies have identified associations between UGT1A variants and bladder cancer risk, highlighting the importance of genetic factors in modulating susceptibility to environmental exposures . Further research is needed to quantify this interaction and to explore the underlying biological mechanisms. This study identifies several significant loci associated with UBC risk, including PSCA (rs2294008), TERT (rs2736098), TACC3 (rs199500838), TMEM129 and UGT1A, many of which align with previously reported associations from UKB- based genome-wide association studies. Although a formal comparison with publicly available UKB GWAS summary statistics was not conducted, the consistency of these findings underscores the robustness of our approach. Differences in study design, population adjustments, and analytic methods may limit direct comparability. Future research may explore formal comparative analyses with these summary statistics to further validate and refine these findings. It should be noted that, UKB is a cohort study with longitudinal follow-up data, offering the opportunity to assess time-dependent risk for exposures that may vary dynamically with age. While this approach could provide additional insights, particularly for non-genetic risk factors, our study focused on genetic susceptibility, where exposure timing is less relevant. Future investigations may leverage the cohort design to explore temporal interactions between genetic and environmental risk factors.
Our PheWAS analysis identified associations between specific genetic variants and prostate conditions, consistent with studies reporting a high frequency of double primary cancers involving the bladder and prostate, suggesting shared genetic or environmental risk factors. 31 Additionally, the association with seborrheic keratosis is notable, as these benign skin lesions. 32 often harbor FGFR3 and PIK3CA mutations similar to those found in low-grade papillary bladder tumors. 33 This genetic overlap indicates potential shared pathways in the pathogenesis of these conditions, warranting further investigation into their clinical significance.
Additional evidence on the risk of UBC development and exposure revealed no association between residential exposure to fine particles and nitrogen dioxide and UBC after adjusting for smoking status. Other studies have previously reported no significant association between PM2.5 and UBC risk. 34 Similarly, the association between exposure to diesel fumes, pesticides, and nocturnal shift work displayed a limited correlation with the development of UBC after adjustment for tobacco smoking. Although the relationship between diesel exposure and UBC has remained equivocal, constraints within the UKB methodology and the absence of metrics for assessing cumulative duration and dosage of exposure further complicate this analysis. Specifically, the questionnaire employed in the study merely solicited participants to categorize their exposure as frequent, occasional, nonexistent, or unknown. As such, the ability to accurately quantify the risk associated with UBC is confounded by variables such as temporal exposure and cumulative dose.
The odds ratio for smoking observed in this study is lower than previously reported in the literature, where ORs are often in the range of 3.35,36 This discrepancy likely reflects differences in how smoking was modeled. The observed effect size may be diluted by grouping all ever-smokers into a single category, regardless of intensity or duration. Analyses stratified by smoking intensity in this study showed stronger associations, consistent with prior research emphasizing the dose-dependent relationship between smoking and BC risk. Differences in study design and population characteristics may also contribute to the variation in effect sizes.
The absence of certain previously reported GWAS hits in our study may be attributed to differences in study design, population characteristics, and statistical power. Our study utilized the UK Biobank cohort, which differs from other bladder cancer GWAS in population structure, linkage disequilibrium patterns, and allele frequencies. Byun et al. discussed similar heterogeneity in UBC GWAS results. 37 Additionally, smaller effect sizes or lower minor allele frequencies of these SNPs in this cohort may have contributed to their lack of significance in our analysis.
Limitations
Acknowledging certain limitations inherent in utilizing the UKB data for this manuscript is important. First, the self-reported nature of exposures such as diesel fumes, pesticides, and nocturnal shift work may introduce reporting bias. Second, the UKB dataset lacks granularity in capturing the cumulative duration and dosage of such exposures, limiting the depth of our risk assessment. Specifically, the simplistic categorization options provided in the questionnaire hinder the precise quantification of exposure levels. Additionally, the UK Biobank population may not be entirely representative of broader demographics, potentially limiting the generalizability of our findings. Lastly, the cross-sectional design of the UK Biobank constrains our ability to make causal inferences or to understand longitudinal changes related to UBC susceptibility. Therefore, while the UK Biobank offers a valuable resource for hypothesis-generating studies, these limitations need to be carefully considered when interpreting the results.
Another limitation of our study is the inability to stratify UBC cases by tumor stage due to the use of ICD codes that do not specify this information. This limitation restricts our capacity to analyze associations between genetic and environmental risk factors and specific tumor stages. Future research should incorporate detailed staging data to enhance the understanding of these associations. Additionally, while this study included unmatched controls to leverage the full diversity and statistical power of the UKB cohort, we recognize that a matched analysis could offer additional insights, particularly for age- and sex-specific risk factors. Future studies may consider this design further to refine genetic and environmental associations with UBC risk.
Closas et al. 36 demonstrated that certain genetic variants exhibit differential associations with NMIBC and MIBC, underscoring the heterogeneity in bladder cancer pathogenesis. However, due to the lack of detailed clinical staging information in the UKB dataset, our study could not assess such subtype-specific associations. Future research utilizing datasets with comprehensive clinical staging data is necessary to determine the genetic factors uniquely associated with NMIBC and MIBC. Diseases such as UBC exhibit considerable phenotypic heterogeneity, and the granularity of clinical data within the UKB may be insufficient for stratifying cases into more uniform subgroups. This introduces a potential confounder that could undermine the validity of GWAS outcomes. Notably, the UKB lacks parameters for clinical staging, relegating our analyses to a dichotomous categorization based on the mere presence or absence of UBC, without the ability to assess disease aggressiveness.
Furthermore, the UKB may manifest healthy volunteer bias, similar to many epidemiological databases. This is particularly relevant as individuals with aggressive disease phenotypes who are undergoing intensive medical interventions are less likely to participate in longitudinal studies and contribute biospecimens. This discrepancy in participation introduces an additional layer of selection bias, thereby confounding the interpretability of GWAS findings derived from this dataset.
Despite these methodological limitations, it is reassuring to note that our findings are corroborated by extant observational research in scientific literature. This study provides both confirmatory and novel insights into BC genetics and its association with environmental risk factors. Significant associations in well-established BC susceptibility loci, including PSCA, TERT, TACC3, and TMEM129, validate previous findings and reinforce their role in BC risk. Additionally, the PheWAS analysis revealed links with prostate conditions and seborrheic keratosis, suggesting shared genetic pathways and emphasizing the utility of large-scale biobank data in uncovering emerging risk factors for UBC. This lends credence to the robustness of our functional GWAS analyses within the framework of environmental exposures and UBC susceptibility. Consequently, we are highly motivated to further investigate the mechanistic relationships between the genetic loci identified in our study and the oncogenesis of UBC.
Supplemental Material
sj-docx-1-blc-10.1177_23523735251370863 - Supplemental material for Genetic susceptibility and environmental risk factors in bladder cancer: Evidence from the UK biobank
Supplemental material, sj-docx-1-blc-10.1177_23523735251370863 for Genetic susceptibility and environmental risk factors in bladder cancer: Evidence from the UK biobank by Laura Bukavina, Ilaha Isali, Sneha Parekh, Sarah Psutka, Nicole Uzzo, Steven Leonard, Adam Calaway, Sunil Patel, Petros Grivas, Angela Jia, Andres Correa, Jason R Brown, Alexander Kutikov, Lee Ponsky, Robert Uzzo, Mohit Sindhani, James Catto, Chen-Han Wilfred Wu and Philip H Abbosh in Bladder Cancer
Supplemental Material
sj-xlsx-2-blc-10.1177_23523735251370863 - Supplemental material for Genetic susceptibility and environmental risk factors in bladder cancer: Evidence from the UK biobank
Supplemental material, sj-xlsx-2-blc-10.1177_23523735251370863 for Genetic susceptibility and environmental risk factors in bladder cancer: Evidence from the UK biobank by Laura Bukavina, Ilaha Isali, Sneha Parekh, Sarah Psutka, Nicole Uzzo, Steven Leonard, Adam Calaway, Sunil Patel, Petros Grivas, Angela Jia, Andres Correa, Jason R Brown, Alexander Kutikov, Lee Ponsky, Robert Uzzo, Mohit Sindhani, James Catto, Chen-Han Wilfred Wu and Philip H Abbosh in Bladder Cancer
Supplemental Material
sj-xlsx-3-blc-10.1177_23523735251370863 - Supplemental material for Genetic susceptibility and environmental risk factors in bladder cancer: Evidence from the UK biobank
Supplemental material, sj-xlsx-3-blc-10.1177_23523735251370863 for Genetic susceptibility and environmental risk factors in bladder cancer: Evidence from the UK biobank by Laura Bukavina, Ilaha Isali, Sneha Parekh, Sarah Psutka, Nicole Uzzo, Steven Leonard, Adam Calaway, Sunil Patel, Petros Grivas, Angela Jia, Andres Correa, Jason R Brown, Alexander Kutikov, Lee Ponsky, Robert Uzzo, Mohit Sindhani, James Catto, Chen-Han Wilfred Wu and Philip H Abbosh in Bladder Cancer
Supplemental Material
sj-docx-4-blc-10.1177_23523735251370863 - Supplemental material for Genetic susceptibility and environmental risk factors in bladder cancer: Evidence from the UK biobank
Supplemental material, sj-docx-4-blc-10.1177_23523735251370863 for Genetic susceptibility and environmental risk factors in bladder cancer: Evidence from the UK biobank by Laura Bukavina, Ilaha Isali, Sneha Parekh, Sarah Psutka, Nicole Uzzo, Steven Leonard, Adam Calaway, Sunil Patel, Petros Grivas, Angela Jia, Andres Correa, Jason R Brown, Alexander Kutikov, Lee Ponsky, Robert Uzzo, Mohit Sindhani, James Catto, Chen-Han Wilfred Wu and Philip H Abbosh in Bladder Cancer
Footnotes
Abbreviations
Ethics statement
This study was exempt by the Institutional Review Board of University Hospitals Cleveland Medical Center.
Author contributions
1.
2.
3.
4.
5.
6.
7.
8.
9.
10.
11. Jason R. Brown: Review
12.
13.
14.
15.
16.
17. Chen-Han Wilfred Wu: Validation
18.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/orpublication of this article:
Ilaha Isali, Sneha Parekh, Steven Leonard, Adam Calaway, Sunil Patel, Angela Jia, Andres Correa, Jason R. Brown, Lee Ponsky, Mohit Sindhani, Chen-Han Wilfred Wu, Philip H. Abbosh: No conflicts of interest to declare.
Data availability statement
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
