Abstract
Introduction
In low-and-middle-income-countries (LMIC), viral suppression is defined as plasma viral load (PVL) below 1000 copies/mL (low-level viremia [LLV]) and threshold for HIV drug resistance (HIVDR) testing. However, there is evidence that drug resistance mutations (DRMs) may emerge at LLV, thus compromising antiretroviral treatment (ART) response
Methods
A cross-sectional study was conducted among individuals with LLV at the Chantal BIYA International Reference Centre, Yaoundé, Cameroon from January 2020 through August 2021. HIV-1 sequencing was performed on protease/reverse-transcriptase, and sequences analysed using Stanford HIVdbv9.5. SSR and HIVDR rates were assessed according to viral-load ranges, with P < .05 considered statistically significant.
Results
In total, 131 individuals were enrolled (median [IQR] age = 41 [30−49] years; 67.9% female; 54.7% at WHO clinical-stage I/II; median ART-duration 7 [4−11] years; median CD4-count 221 [103−402] cells/mm3 and median PVL 222 [96−436] copies/mL). Overall, SSR at LLV was 34.4% (45/131) and increased significantly with decreasing-age (P = .002) and increasing-PVL (P = .017). SSR were doubled at PVL≥150 copies/mL (21.8% at [40−150] vs. 43.3% at [150−1000]; OR = 2.8, P = .01). Of the 45 sequences obtained, 75.6% were recombinant strains (CRF02_AG, CRF09_cpx, CRF11_cpx) and 24.4% pure-subtypes (A1, D, F2, G). Overall, HIVDR prevalence at LLV was 82.2% (37/45), with 74.6% and 15.6% resistance to reverse-transcriptase inhibitors (RTIs) and ritonavir-boosted protease inhibitors (PI/r) respectively. Interestingly, HIVDR rates were similar at PVLs [50−200] versus [200−1000] copies/mL (P = .69). The most frequent DRMs were M184 V (73.3%) and K103N (40.0%) for RTIs and M46I (6.7%) for PIs/r. Overall 55.6% (25/45) of individuals were on suboptimal ART (needing ART-optimisation), with 48.9% (22/45) having suboptimal TLD predictive efficacy. Optimisation need was higher in first-line (81.8%, P = .03), but similar across viral clades and PVL-ranges (P = .6).
Conclusion
In this LMIC context, sequencing for HIVDR is feasible at LLV even with broad HIV-1 diversity, with significantly higher SSR above 150 copies/mL and/or in paediatrics. About 80% of individuals with LLV harbour HIVDR strains, with half of them needing ART optimisations to limit HIVDR emergence and prevent treatment failure. Our findings underscore the clinical benefits of HIVDR during persisting LLV and the need to reconsider the threshold for viral suppression around 200copies/mL in LMICs.
Plain Language Summary
In low- and middle-income countries (LMICs) like Cameroon, virological success is defined as viral loads less than 1000 copies/mL. However, drug-resistant strains can emerge even when viral loads are low and could be challenging for long term management outcomes. This study looked at how well HIV drug resistance (HIVDR) can be detected in people with low viral loads, the rates, and patterns in which they occur and their potential impact on treatment regimens. We found that about one-third of the people could be successfully tested for drug resistance at low viral loads, and this success rate improved significantly with higher viral loads, doubling significantly at a threshold of 150 copies/mL. Up to 80% of those with low viral loads had drug-resistant strains, and about half of them needed their treatment adjusted. This suggests that the current definition of viral suppression (under 1000 copies/ml) might need to be reconsidered and lowered to around 200 copies/ml in LMICs to better control HIV and prevent treatment failure.
Introduction
Though HIV remains a public health priority with an estimated 39 million people living with HIV in 2022, of which about 60% in Sub-Saharan Africa (SSA), the continuous use of effective antiretroviral therapy (ART) has greatly reduced HIV/AIDS related morbi-mortality. 1 In fact improved access to treatment has prevented about 21 million deaths globally, with also great impact in SSA, where AIDS claimed fewer lives than at any point since 1990. 2 The goal of ART is to suppress viral replication, to a level below which acquired drug-resistance mutations cannot emerge. 3 This risk of HIV drug resistance (HIVDR) emergence remains a persistent threat to ART programs, with potential to jeopardize treatment outcomes. 4 Globally in 2022, 86% of PLHIV knew their status, 76% were receiving ART and 71% were virally supressed. 2 This latter population (virologically suppressed) have improved and near normal quality of life, with zero risks of horizontal transmission in both high and low-income settings.5–8
There is significant heterogeneity in the definitions of viral suppression (VS) among persons receiving ART. Specifically, European recommendations require repeating any viral load (VL) > 50 copies/mL after 1−2 months. A second VL > 50 copies/mL is defined as virological failure (VF) and this requires performing HIVDR and tropism testing, among other measures. Meanwhile in the USA, a VL > 50 copies/mL requires continuing the current regimen with 3-monthly VL monitoring without HIVDR testing if VL remains below 200 copies/mL. Thus, in both European and USA recommendations, two consecutive VL ≥ 200 copies/mL (VF) require performing HIVDR testing. 2 In South Africa, a VL > 50 copies/mL requires a second VL monitoring within 3 months maximum, while HIVDR testing is only required for at least two consecutive VL > 1000 copies/mL, applicable after 2 years on dolutegravir- or PI-based regimens. For several low- and middle-income countries (LMICs), WHO guidelines are being implemented, recommending enhanced adherence counselling (EAC) and VL monitoring after 3 months following a first VL > 50 copies/mL. When this second VL falls between 50 and 1000 copies/mL, EAC is continued and VL monitored quarterly. HIVDR testing is recommended only to optimize the third-line regimen for people failing second-line ART with two consecutive VL > 1000 copies/mL within 3−6 months interval. CD4+ T-cell count monitoring is only recommended in case VL is not accessible.9,10 These raise issues on delineating a suitable threshold for VS, especially for translational application in LMIC. Subsequent VF and HIVDR selection are more likely to occur at VL ≥ 200 copies/mL,11–13 and even for some cases at 50−200 copies/mL.14–16 These conflicting data, coupled to scarcity of evidence in LMICs, suggest a contextual gap that remains questionable.
In Cameroon for instance, the VS threshold is <1000 copies/mL,17,18 with VLs between 50−1000 copies/mL termed low level viremia (LLV). Nationwide in 2022, 94.1% of people living with HIV knew their status, of whom 82.7% were on ART and 94.1% of these had VS, reflecting somewhat good programmatic performance progress towards achieving the 95/95/95 targets.19,20 However, considering the threshold for VS (1000 copies/mL), persistent viral replication at this LLV may correlate with HIVDR emergence and subsequent VF, therefore jeopardizing long-term clinical outcomes.21–25 In an era where sustained viral control remains challenging, determining optimal VS thresholds in a context of wide viral diversity is crucial. Likewise, feasibility of HIVDR testing at LLV has not been sufficiently evaluated in the routine clinical practice of LMICs like Cameroon. Due to the low VLs, amplification performance may be significantly lower in people with LLV as compared to treatment failing individuals (VL > 1000 copies/mL), which in turn may call for optimization of in-house procedures for an improved sequencing success rate.26–28 Therefore, to ensure ART durability in LMICs, we sought to evaluate the sequencing success rates (SSR) at LLV and to determine the genotypic resistance patterns at LLV and their adequacy with therapeutic regimens.
Materials and Methods
Study Design and Setting
We carried out a cross-sectional and analytical study at the Chantal Biya International Reference Centre for Research on HIV/AIDS prevention and management (CIRCB), between January 2020 and August 2021. Included were virally suppressed ART experienced individuals, with low level viremia (plasma viral load ≥ 50 copies/mL to < 1000 copies/mL). Excluded were participants on treatment interruption at the time of genotypic resistance test, defined as a the stop of ART medication within a given period (herein greater than one month) according to national guidelines. 29 We added a control group, of treatment failing individuals (VL ≥ 1000copies/mL), to permit comparison of mutational profiles with the virally the suppressed group.
Procedure for Genotypic Resistance Testing at Viral Loads <1000 Copies/mL
Viral RNA was extracted from 1 mL of plasma using the QIAGEN kit, following the manufacturer's instructions. For the amplification of the pol region, a previously validated in-house genotyping assay was used. 30 Failed samples were repeated using the ThermoFisher commercial genotyping kit (see supplementary Figure S1). Briefly for the in-house assay, the target sequence was first amplified with Reverse Transcription PCR using BS primers (5′-GAC AGG ATT ATT TTT TAG GG-3′) and FRA S1 (5′-TT CCC CAT ATT ACT ATG CTT-3′) in 25 μL of reaction mixture for 40 cycles. The PCR conditions were: 50°C for 30 min, 94°C for 2 min, 95°C for 30 s, 51°C for 30 s, 72°C for 2 min, followed by extension at 72°C for 10 min. A semi-Nested PCR was subsequently performed using BS primers (5′-GAC AGG ATT ATT TTT TAG GG-3′) and TAK3 (5′-GGC TCT TGA TAA TGA TAT TAT GT-3′) in 50 μL of reaction mixture for 30 cycles. The PCR conditions were 93°C for 12 min, 94°C for 30 s, 53°C for 45 s, 72°C for 2 min, followed by extension at 72°C for 10 min. The PCR products were revealed using 1% agarose gel electrophoresis. The successfully amplified samples were purified and then sequenced by the following primers: B (5′-AGC AGA CCA GAG CCA ACA GC-3′), F (5′-CCA TCC ATT CCT GGC TTT AAT-3′), SEQ1 (5′-GAA TGG ATG GCC CAA AA-3′), SEQ2 (5′-TTG AGA TAC AAT GGA AAA GGA AGG-3′), SEQ3 (5′-CCC TGT GGA AAG CAC ATT GTA-3′), SEQ4 (5′-GCT TCC ACA GGG ATG GAA-3′), SEQ5 (5′-CTA TTA AGT CTT TTG ATG GGT CA-3′), TAK3 (5′-CCT TGT TTC TGT ATT TCT GCT-3′). The products of the sequencing reaction obtained were purified by exclusion chromatography on SEPHADEX G50 resin and then sequenced using the genetic analyzer (ABI 3500®).
Bioinformatic Analysis of Resistance Mutations
The resulting sequences encompassing the pol region were assembled and edited using Recall _v2.28. Sequence interpretation was done using Stanford HIVdb.v.9.5 (http://www.hivdb.stanford.edu/ accessed on 26 June 2023). Subtyping was done first using rapid subtyping tools (COMET, REGA and BLAST), with confirmations done using molecular phylogeny on MEGA 11.
Data Entry and Statistical Analysis
Data was entered and analyzed using Epi Info version 7.2.2.6. Categorical variables were described in terms of frequency and percentage. The chi-square or Fischer's exact test were used to compare proportions of the qualitative variables. The values were expressed with their 95% confidence interval. All P-values < .2 underwent multivariate analysis to adjust for potential bias and to determine predictors of HIV drug resistance. The threshold of statistical significance was set at 5%.
Results
Study Population
We retained a total 131 participants, majority being female (67.9%), median age [IQR]; 41 [30−49] years, with majority of participants aged between 20−49 years (61.8%). Majority of participants were on second-line PI/r-based regimens (n = 81, 61.8%), while 28.2% were on first line NNRTI-based regimens and 4.6% on third line regimens (DRV/r based). Overall, the median [IQR] duration on treatment was 7 [4−11] years, with 44.3% of participants reporting good adherence levels (defined as at least 95% of ART). This was self-reported, with the participant reporting the number of doses missed in the month, and adherence calculated as [(X-Y)/X] × 100%, where X = number of days in the month and Y = number of missed pills/days in the month). Table 1 shows a summary of study population characteristics.
Study Participant Characteristics.
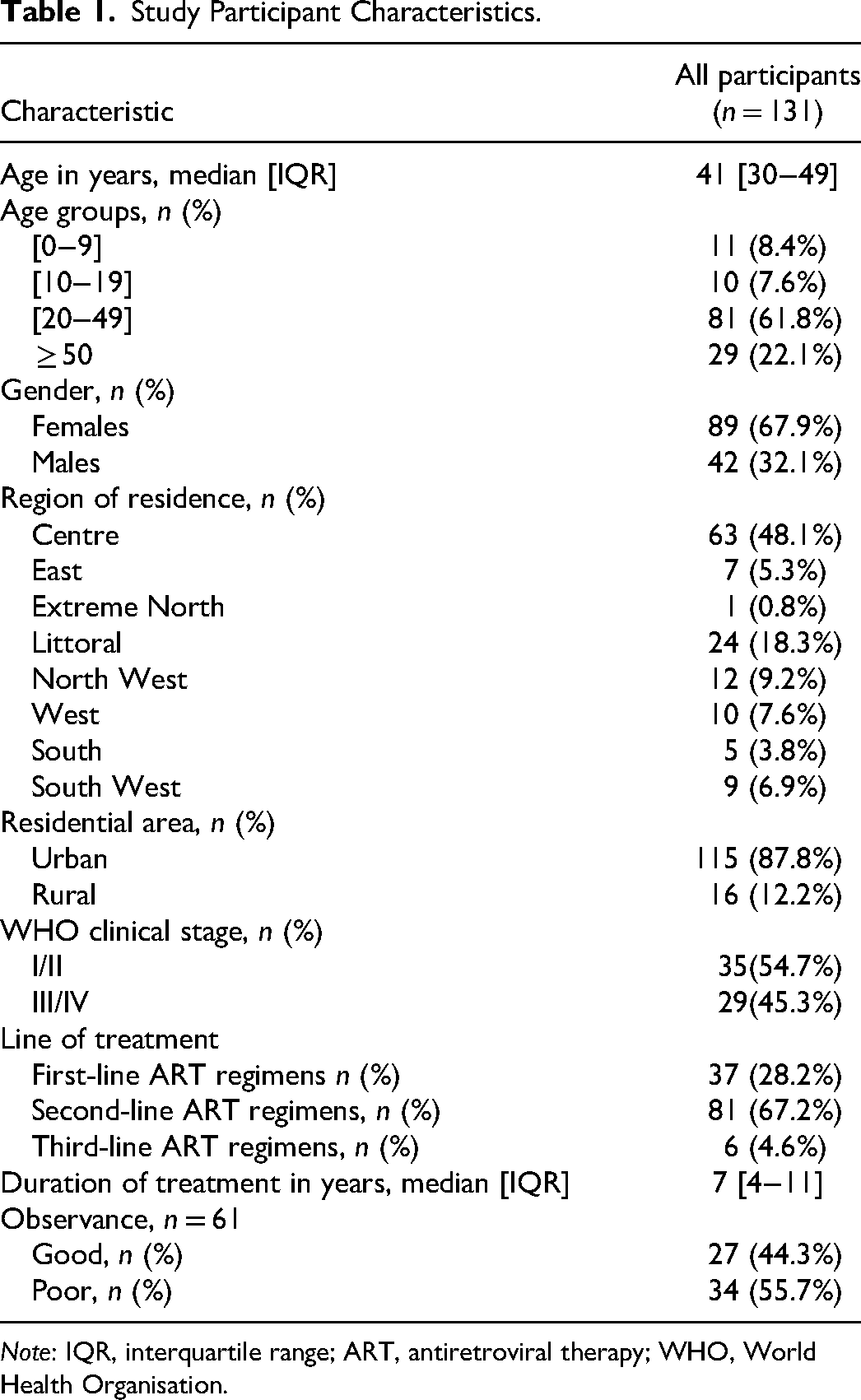

Note: IQR, interquartile range; ART, antiretroviral therapy; WHO, World Health Organisation.
Immunological and Virological Profiles
At LLV, the median CD4 count was 221 [103−402] cells/mm3 with 54.7% having CD4 counts between 0−250 cells/mm3 and 45.3% with CD4 counts ≥250 cells/mm3. Median CD4 count significantly decreased with increasing age; that is median of 814 cells/mm3 in participants aged 0−9 years; 399 cells/mm3 in those 10−19 years, 191 cells/mm3 in those aged 20−49 years and 210 cells/mm3 in those aged ≥50 years (P < .0001). CD4 count did not vary by sex (P = 1), treatment regimen (P = .8), WHO clinical status (P = .5), adherence (P = .3), nor viral load (P = .6). With respect to virological status, the median viral load was 222 [96−36] copies/mL, with 48.1% (n = 63) of participants having viral load in the range [50−200], 33.6% (n = 44) in the range [200−500] and 18.3% (n = 24) in the range [500−1000] copies/mL. On bivariate analyses (Table 2), residual LLV was significantly higher in females (P = .004), WHO clinical stages III/IV (P = .03) and poorly adherent participants (P = .04). After multivariate analysis adjusting for age, sex, WHO clinical stage and adherence; female sex, advanced disease stage at diagnosis (WHO III/IV) and poor adherence, remained independent determinants of residual viremia (adjusted P values; .003, .02 and .03 respectively).
Median Viral Load by Age, Sex, Clinical Stage, Treatment Line, CD4 Count, Adherence, and Residential Area.
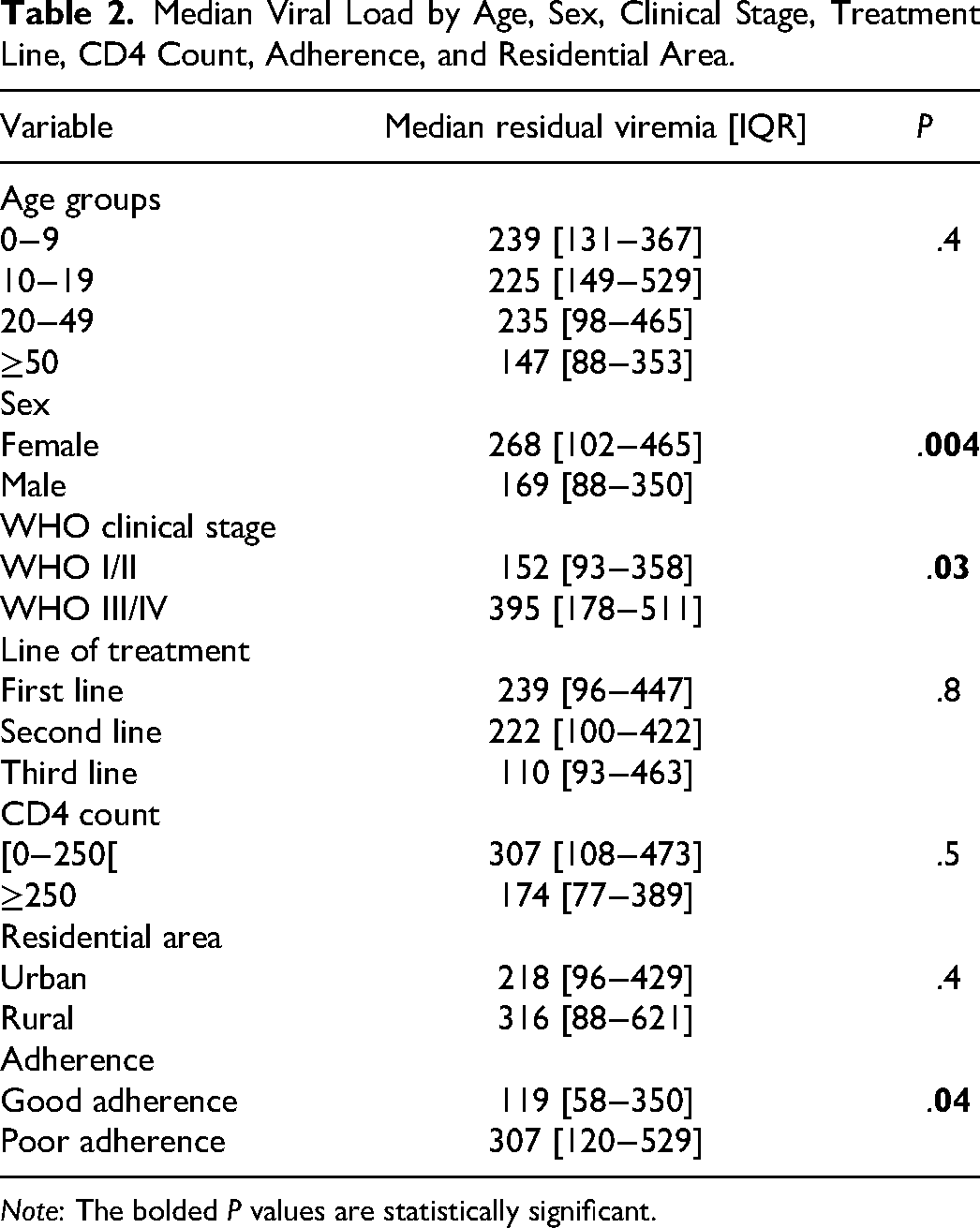
Note: The bolded P values are statistically significant.
Sequencing Success Rates
Of the 131 participants included, genotypic resistance testing was successful in 45, giving us an overall sequencing success rate of 34.4% (95% CI: 26.3%−43.2%). Sequencing success rates did not by sex (P = .6), residential area (P = .4), WHO clinical stage (P = .8), nor line of treatment (P = 1). The median viral load was higher in those with successful sequencing (301 [150−518] copies/mL), as compared to those with unsuccessful sequencing (151 [87−405] copies/mL), P = .03. Though not statistically significant, individuals with viral load 200−1000 copies had two times higher chances of sequencing success as compared to those with viral loads <200 copies/mL (41.8% vs. 27.8%; OR = 1.9, P = .099). Considering the 150 copies/mL as threshold, sequencing success rates were significantly higher at viral load [150−1000] (43.3%) as compared to viral loads [50−150] copies/mL (21.8%), OR = 2.8, P = .01. Elsewhere, sequencing success rates significantly varied by age (P = .005), with success rates of 72.7% in participants aged 0−9 years; 60.0% in participants aged 10−19 years; 30.9% in those aged between 20−49 years and 20.7% in those aged 50 years and more (see Figure 1). After multivariate analysis adjusting age, sex, WHO clinical stage and viral load, age and viremia remained independent determinants of sequencing success (P = .002 and .017 respectively).
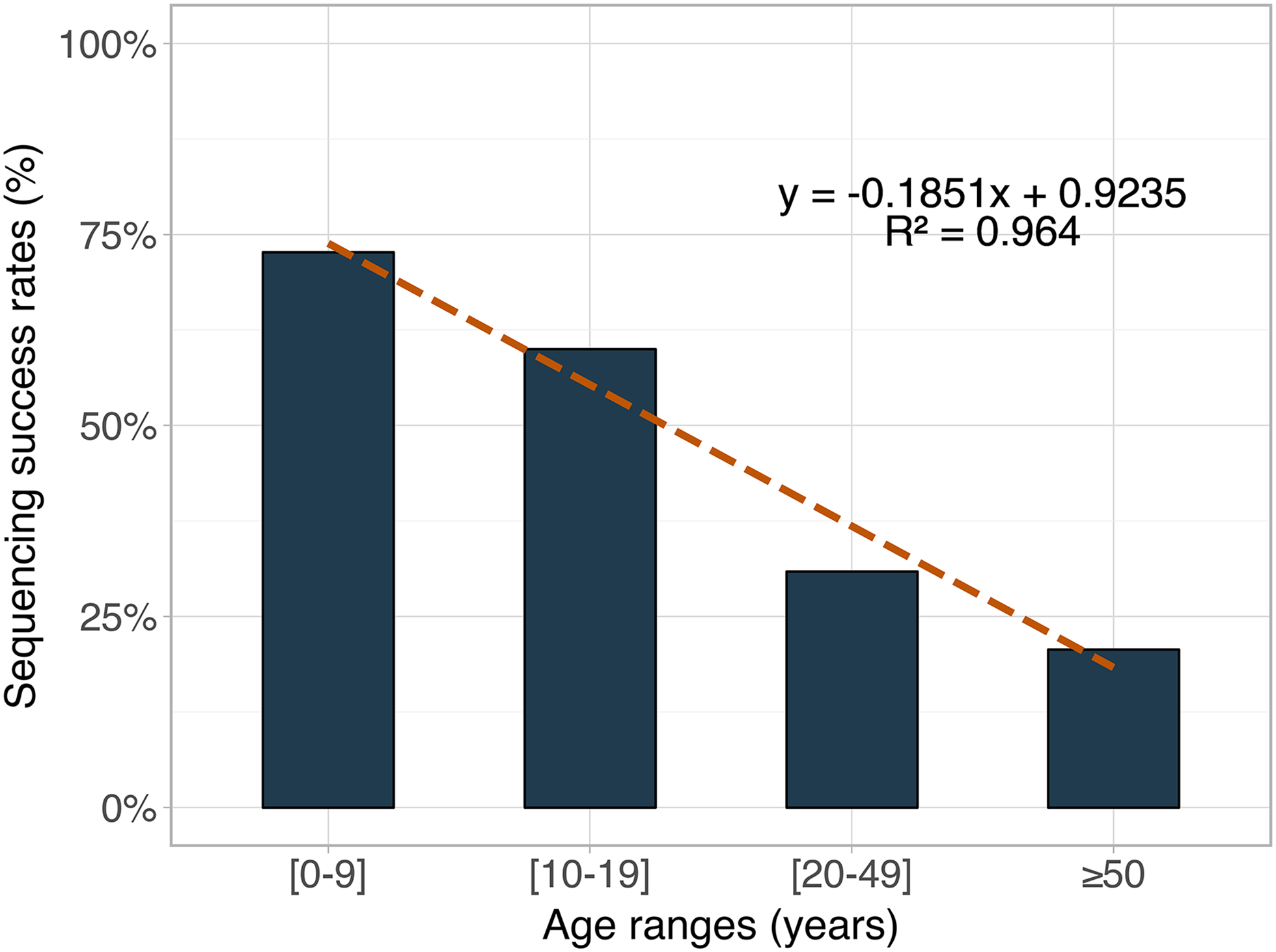
Sequencing success rates by age.
Genotypic and Resistance Profile
Majority of participants harboured a recombinant viral clade (34/45; 75.6%), notably CRF02_AG (32/45; 71.1%), CRF09_cpx (1/45; 2.2%), and CRF11_cpx (1/45; 2.25). Only 11/45 (24.4%) of participants harboured a pure viral clade, notably A1 (6/45; 13/3%), D (2/45; 4/4%) F2 (2/45; 4.4%) and G (1/45; 2.2%). Overall, 37/45 participants harboured a virus with at least one major drug resistance mutation (DRM), giving us an overall rate of HIVDR of 82.2%, (95% CI: 68.0%−92.0%]). By drug class, NRTI and NNRTI resistance were each 74.6% (95% CI: 60.5−87.1%), while overall PI/r resistance was 15.6% (95% CI: 6.5%−29.5%). NRTI/NNRTI dual class resistance was 68.9% (95% CI: 53.4%−81.8%) while NRTI/NNRTI/PI/r triple class resistance was 13.3% (95% CI: 5.1%−26.8%). Overall, the most prevalent NRTI mutation was the M184V mutation (73.3%) followed by the K65R (25.6%) and lastly the T215Y mutation (20.0%). The most frequent NNRTI mutation was K103N (40.0%) followed by A98G (20.0%) and lastly Y181C and G190A (13.3%). M46I and L76 V were the most prevalent PI/r resistance mutations (6.7%), followed by the I84V mutation (4.4%). The following important PI/r accessory mutations were present: M36I (88.9%), L89M (77.8%), K20I (62.2%), L63P (26.7%), L10I and K20R (13.3%) and L89I (6.7%). Figure 2 shows the major drug resistance mutations (DRMs) by drug class.
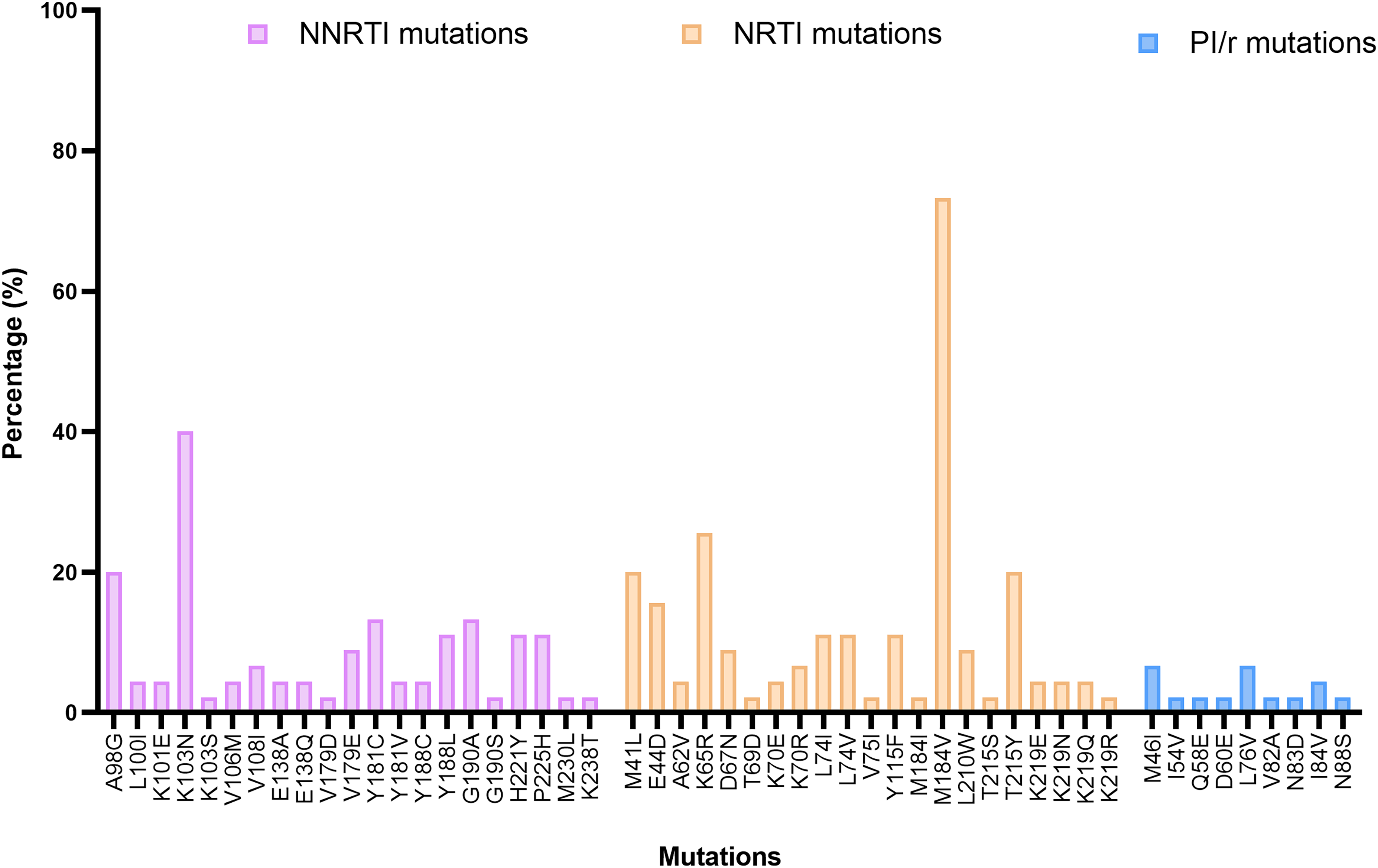
Major drug resistance mutations by drug class.
HIVDR Potential Determinants
Neither sex (P = 1), residential area (P = 1), WHO clinical stage (P = 1), age (0.1), viremia (P = .3), line of treatment (0.4), nor viral subtype (0.2) significantly affected the presence of major DRMs at LLV. Interestingly, all children aged 0−9 years (n = 8, 100%), harboured a virus with at least one drug resistance mutation. At LLV, overall HIVDR rates were similar at viral loads between [50−200] copies/mL (88.2%) and [200−1000] copies/mL (78.6%), P = .69. Comparing drug resistance levels in LLV individuals with virologically failing individuals (VL ≥ 1000 copies/mL), those on virological failure had significantly higher rates of overall HIVDR (P = 0.006), of NNRTI resistance (P = .002) and of NRTI/NNRTI dual class resistance (P = .01). Rate of NRTI resistance was on the threshold of significance (P = .06), while PI/r and triple class resistance were similar (P = .7 and .9 respectively). See Table 3.
Comparison of Resistance Profiles at Low Level Viremia and Virological Failure.
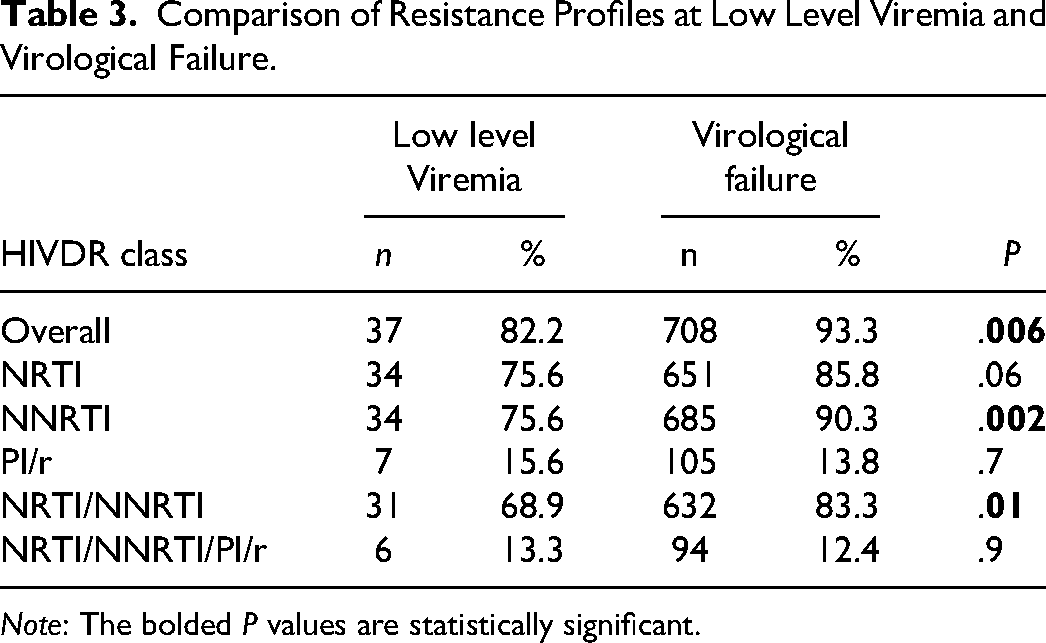

Note: The bolded P values are statistically significant.
Potentially Active ARV Drugs
According to viral susceptibility scores as per the Stanford algorithm, drug susceptibilities are shown on Figure 3. Notably, DRV/r showed highest levels of efficacy (93.3%) followed by ATV/r (88.9%) and LPV/r (88.4%). As concerns NRTI, AZT preserved efficacy in 64.4% of participants while TDF in 51.2%. First generation NNRTI showed low levels of efficacy, 20% for EFV and 15.6% for NVP. Taking into consideration ongoing therapies of individuals, treatment optimisation was proposed for 25/45 participants (55.6%; 95% CI: 40.0%−70.4%). Need for treatment optimization was significantly higher in participants on first line regimens (81.8%), as compared to second (50.0%) and third (0%) line participants (P = .03). Seven out of eight (87.5%) children between 0−9 years needed treatment optimization as opposed to 4/6 (66.7%) adolescents (10−19 years) and 14/29 (48.3%) adults (≥20 years) though this difference did not appear statistically significant (P = .2). Neither sex (P = .2), residential area (P = .7), WHO baseline clinical stage (P = .7), nor viral load range (P = .6) significantly influenced the need for treatment optimization. With respect to viral load ranges, optimization was necessary for 11/17 (64.7%) at viral load range [50−200] copies/mL, 8/16 (50%) at [200−500] copies/mL, and 6/12 (50%) at [500−1000] copies/mL (P = .6).
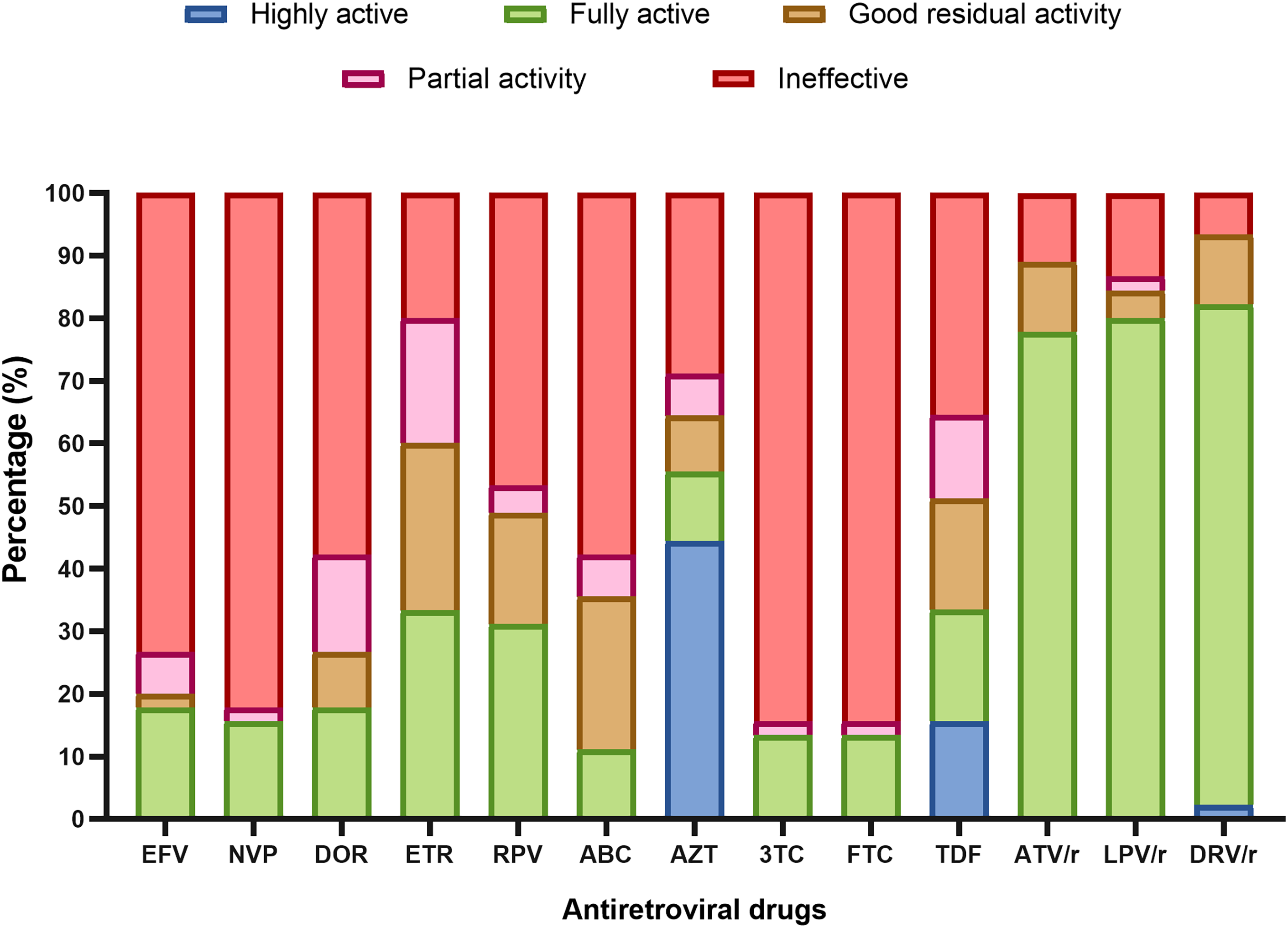
Potentially active drugs. The figure shows potentially active drugs (EFV: efavirenz, NVP: nevirapine, DOR: doravirine, ETR: etravirine, RPV: rilpivirine, ABC: abacavir, AZT: zidovudine, 3TC: lamivudine, FTC: emtricitabine, TDF: tenofovir, ATV/r: atazanavir boosted by ritonavir, LPV/r: lopinavir boosted by ritonavir, DRV/r (darunavir boosted by ritonavir) according to genotypic susceptibility scores (GSS) obtained from the Stanford algorithm. In red are ineffective drugs (GSS ≥ 60); in pink are drugs with potential activity (GSS = 30−59); in orange with good residual activity (GSS = 10−29) in green fully active drugs (GSS = 0−9) and in blue, highly active drugs (GSS < 0).
Potentially Efficacy of TDF-3TC-DTG (TLD) at low Level Viremia
Considering the non-exposure to integrase inhibitors in our study population, and thus expected low level of INSTI resistance in integrase-naïve individuals in this population (<1% overall; 0% in ART-naïve and 0.8% in ART-experienced) 31 we estimated TLD predictive activity by considering full potential activity of DTG in this population. Therefore, considering overall TDF potential activity of 53.1% (considering a GSS < 30), overall TLD predictive activity was 51.1% at LLV. This predictive efficacy was similar with respect to the line of treatment (P = .9), viral load ranges (P = .2) among other factors (Figure 4).
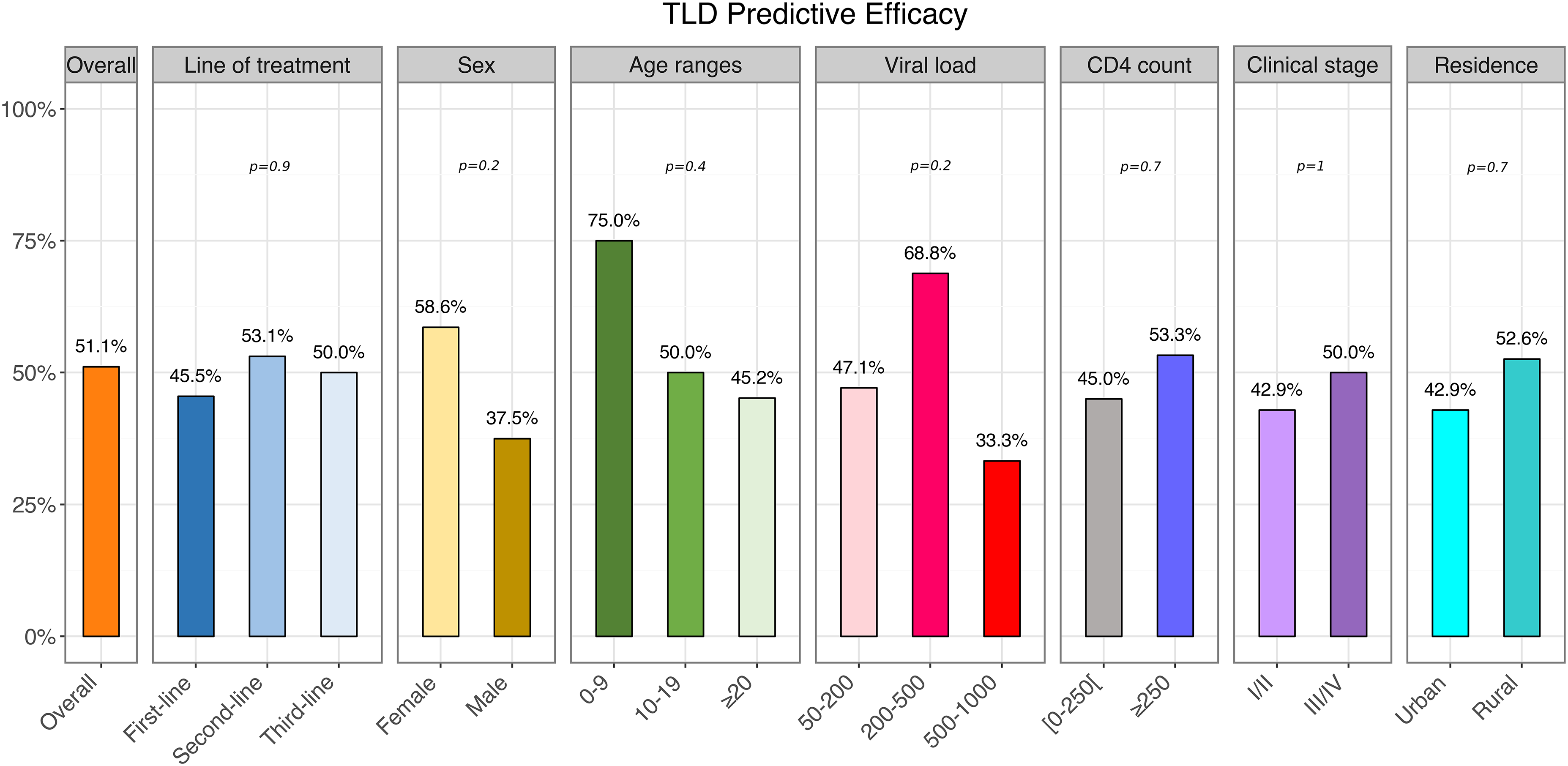
Estimated TDF-3TC-DTG (TLD) predictive activity in the overall population and with respect to study variables.
Discussion
At the moment, VS (VL < 1000 copies/mL) is still considered as a threshold for therapeutic success and for ineligibility to HIVDR testing in several LMICs like Cameroon. In this frame, the present study serves as a proof of concept for a suitable threshold for effectiveness of sequencing in real life and considerations for HIVDR in individuals with LLV in Cameroon and similar LMICs where HIV molecular epidemiology may be either comparable, or less diverse, leaving the possibility for better performance with optimally designed protocols. Overall genotyping success rate was low (34.4%) which is different from other studies carried out in different contexts which report higher rates of success at LLV.32,33 One of the main explanations to this difference is HIV genetic diversity. 22 Previous studies have described high HIV-1 genetic diversity in Cameroon with both pure and recombinant subtypes.34–37 In this study, we successfully sequenced just seven viral subtypes (three recombinant and four pure), suggesting that many others may have been missed. Though the overall low success rate may justify the public health policy which currently recommends genotypic resistance testing only at viral load values greater than or equal to 1000 copies/mL, it however highlights the feasibility of genotypic testing even in our context, despite the diverse HIV-1 genetic diversity as compared to other settings. Furthermore, optimisation of currently used assays taking into consideration local molecular epidemiology may also be necessary to improve this sequencing results. 38 Interestingly, sequencing success rate did not vary by sex, residential area, clinical stage of disease, nor CD4 count, but increased significantly with decreasing age and increasing viral load. In fact, participants aged 0−9 years had a sequencing success rate of 72.7%, which decreased with increasing age with those aged greater than 50 years having success rates of just 20.7%. To add, sequencing success rates increased significantly with increasing viral load, with two times higher rates of success at viral loads above 200 copies/mL. This therefore implies that, at these low-level viremias (50−1000 copies/mL), as expected, the high HIV-1 genetic variability and residual viral load could significantly affect sequencing success rates, but also multiple changes in the viral diversity due to longer viral evolution appears to be a substantial determinant. In effect several studies have shown that earlier ART initiation limits viral diversity, which will be the case for the paediatric population, with relatively early ART initiation as compared to the older population.39–42 In a context where paediatric HIV management still remains a major challenge with high rates of virological failure, HIVDR and mortality, using HIVDR for treatment optimisations is therefore very feasible and should be considered to improve outcomes in this special population.43–45 Interestingly, we observed a decrease in absolute CD4 count with increasing age, with significantly higher CD4 counts in children and adolescents as compared to adults. Indeed, it is known that children usually have higher CD4 counts than adults, reason why the CD4% is usually preferably used, although the CD4 count can be used as from 5 years, with good correlation with adult/adolescent values. 46 Immunosenescence has been shown to affect T lymphocyte subsets, with a general reduction of most subsets with aging. However, a study by Li et al in healthy subjects showed that although CD3 and CD8 subsets linearly decreased with aging, CD4 subsets appeared more stable. 47 This probably explains the similar levels of mean CD4 count observed in our results, in adults 19−49 years and those 50 years and above. Castro et al showed that children starting ART at an early age and those with a higher CD4 nadir, have higher mean CD4 counts (as compared to those starting after 10 years, or those with lower CD4 nadir). 48 This would possibly also explain in the virally suppressed population in our study, the higher CD4 count levels observed in the children and adolescents, who generally start ART treatment at a much earlier age (known time of exposure), as compared to adults where diagnosis is usually much later when significant immune depletion may have already occurred.
At this LLV, rates of HIVDR were high, with 82.2% of participants harbouring at least one drug resistance mutation. It should be noted that though significant rates of HIVDR have been reported at LLV, our observed rates appear higher than those observed in some studies and similar to those observed in others.33,49,50 The high rates observed in our study would have been favoured by the higher treatment duration in our context (averagely 7 years), the use of low genetic barrier drugs, poor adherence (about half of participants with poor adherence), the high viral genetic diversity. It should be noted that these mutations are mostly due to the continuous viral replication despite low level viremia, but may also be the regression of some already existing variants to minority populations following treatment changes. 51 All individuals aged 0−9 years harboured at least one drug resistance mutation. Indeed, paediatric HIV/AIDS remains a major bottleneck to eliminating HIV/AIDS by 2030. This population boosts of higher HIV associated morbi-mortality as compared to adult populations in LMICs like Cameroon. Rates of HIVDR are also high in this population, favoured by factors such as higher risks of pre-treatment drug resistance, limited treatment options and formulations, poor adherence and the use of low genetic barrier drugs in LMICs for paediatric management and prevention of mother to child transmission in LMICs. 52 Therefore, the high rates of drug resistance observed in this population are not surprising, but rather further highlight the HIV management challenges in the paediatric population. Unfortunately, the true contribution of pre-treatment drug resistance to the profiles observed in our study (both the paediatric and adult population) cannot be fully ascertained, as HIV genotyping is not systematically recommended before ART initiation in Cameroon and similar resource limited settings. 29 However, the significant treatment exposure, as well as the compatibility of resistance profiles with acquired drug resistance patterns in these virally suppressed individuals, will suggest a negligible contribution of PDR in the observed profiles. To add, in this real-world clinical scenario, it would be almost impossible to ascertain the exact contribution of pre-treatment drug resistance to the resistance profiles in these individuals, also highlighting the programmatic and clinical importance of these findings. While accessing HIVDR rates with respect to viremia, we found similar rates of resistance at viral loads between [50−200] copies/mL and [200−1000] copies/mL. Though some studies have shown greater risks of HIVDR and subsequent virological failure at viral load greater than 200 copies/mL, our studies remain concordant with many other studies which showed similar rates of HIVDR and subsequent risks of virological failure at viral loads between 50−1000 copies/mL.4,14,53 Although the overall high rates of HIVDR observed at LLV are slightly lower (P = .006), that in treatment failing individuals VL ≥1000 copies/mL), these results still call for prompt measures at all low-level viremia levels, to ensure viral undetectability (VL < 50 copies/mL), which appears to remain the optimal virological threshold. Indeed, data remains conflicting with respect to the optimal threshold for VS and resistance testing, with some authors recommending 50 copies/mL and others 200 copies/mL but importantly not 1000 copies/mL. Although the WHO has for years recommended the 1000 copies/mL threshold which remains important also in the “Undetectable = Untransmissible” concept, 8 it is worth noting that these guidelines would usually suggest the minimal requirements for good performance in a public health approach, but should not be a limitation for clinicians as viral replication at low level viremia, should be an indication for further investigation, for possible causes such as suboptimal adherence, pharmacological interactions, HIVDR or virus production from clonally expanded HIV-infected cells. 54 Importantly, the adoption of a lower threshold in LMICs should also take into consideration diagnostic capacity and sensitivities which may be a major challenge. Nonetheless, in recent years, there has been scale-up of viral load testing in LMICs with lower detection limits giving thus a chance for a revision of the suppression thresholds.55,56 Although sequencing success rates at LLV in this context may be low as in our findings, this performance could be ameliorated with regular updates in genotyping primers and protocols which is a major perspective for us following this study. Meanwhile, at viral loads above 200 copies/mL, genotyping was successful in about half of these individuals and even much higher in paediatrics, which is significant on a public health level, suggesting that this could be a suitable threshold in our context. It is also worth considering that, this 200 copies/mL would be more optimal in preventing transmission.5,6 Taking into consideration the mutational profiles, the most prevalent mutation (NRTI) was the M184 V mutation (73.3%). Selected by 3TC/FTC, this mutation serves and an adherence marker and increases viral susceptibility to AZT/TDF, while reducing viral replicative fitness.57–59 NNRTI resistance was also high, favoured by their low genetic barrier and long half-life, with the most frequent mutation being the K103N mutation, causing high levels of resistance to first generation NNRTI. We also note the presence of other NNRTI mutations causing cross-resistance to second generation NNRTI (DOR, ETR, and RPV). This supports transition to the current DTG based regimens with higher genetic barrier. PI/r drug resistance was relatively low (15.6%), with the most frequent mutation being the M46I and L76 V mutations. The overall low PI/r resistance reflects the higher genetic barrier of PI/r but was also favoured by the low PI/r exposure in our study population. 49
Taking into consideration the resistance profiles and the genotypic susceptibility scores, all PI/r maintained potential good levels of efficacy, with 93.3% for DRV/r, 88.9% for ATV/r and 88.4% for LPV/r. This further illustrates their higher genetic barrier and justifies their use in second- (ATV/r and LPV/r) and third- (DRV/r) line regimens in our context. As concerns NRTI, AZT and TDF had overall potential good efficacy in 64.4% and 51.2% of participants respectively. In the NNRTI class, EFV and NVP had low levels of efficacy (20% and 15.6% respectively), again supporting transition DTG based regimens. 60 Meanwhile, overall full predictive efficacy of TLD was estimated in just half of these individuals (53.1%). This alerts the fact that, suboptimal transitions on TLD are still possible even a LLV with possible risks of failure in the long run. This calls thus for careful TLD transitions (taking into consideration factors such as therapeutic histories, clinical status, and genotypic profiles) even at LLV. Elsewhere, there were also considerate levels of cross-resistance to second generation NNRTI, though second generation NNRTI efficacy appeared higher. Taking into consideration the participant treatment histories, genotypic and susceptibility profiles, treatment optimisation was needed for 55.6% of the participants. Therefore, similar to other studies, treatment optimisation appears necessary even at low level viremia, in order to attain viral undetectability and prevent subsequent treatment failure. 61 By treatment regimen, need for optimisation was higher in participants receiving NNRT-based first regimes (81.8%) as compared to those of second line (50.0%) and third line (0.0%). This again supports the phasing out of low genetic barrier drugs in first line, and use of higher genetic barrier drugs. Elsewhere, though not statistically significant (P = .2), treatment optimisation was needed for seven out of eight (87.5%) children between 0−9 years, 4/6 (66.7%) adolescents (10−19 years) and 14/29 (48.3%) adults (≥20 years) showing again the need for targeted efforts to the pediatric population. Lastly, need of optimisation was similar in all low-level viremia ranges (50−200; 200−500 and 500−1000, P = .6) again suggesting the threshold of 50 copies/mL as the most optimal in our context. In effect, apart from the virological consequences of LLV (HIVDR and virological rebound), persistent LLV may be responsible for many other consequences such as residual immune activation and inflammation, which contribute to disease. 21 In our study, the median residual LLV was significantly higher as expected in those with advanced clinical stage (WHO III/IV) at diagnosis and poor adherence but unexpectedly also higher in females. Though most studies have reported no significant difference in median residual viremia between males and females, Oliviera et al. described higher tissue HIV-DNA levels in females suggesting thus a possible sex related mechanism in viral persistence, needing further investigation.62,63 Increased attention to these above-mentioned groups may be necessary to for optimal therapeutic outcomes.
Conclusion
In this LMIC harbouring a broad HIV-1 genetic diversity, sequencing appears to be efficient in people with LLV, especially in paediatrics and increasing viral load at LLV, with a doubled performance at VL >200 copies/mL. Of clinical relevance, eight out of ten virally suppressed individuals (<1000 copies/mL) in this setting may harbour HIVDR with up to half of them needing treatment optimisations to attain viral undetectability and prevent subsequent virological failure and HIVDR emergence. Thus, a revision of the current 1000 copies threshold down to 200 copies/mL might be convenient to define viral suppression in LMIC, as well as for HIVDR testing as an efficient approach for treatment optimisation that would contribute in the control of the HIV pandemic by 2030 at global level.
Supplemental Material
sj-docx-1-jia-10.1177_23259582241306484 - Supplemental material for Plasma Viral Load of 200 Copies/mL is a Suitable Threshold to Define Viral Suppression and HIV Drug Resistance Testing in Low- and Middle-Income Countries: Evidence From a Facility-Based Study in Cameroon
Supplemental material, sj-docx-1-jia-10.1177_23259582241306484 for Plasma Viral Load of 200 Copies/mL is a Suitable Threshold to Define Viral Suppression and HIV Drug Resistance Testing in Low- and Middle-Income Countries: Evidence From a Facility-Based Study in Cameroon by Collins Ambe Chenwi, Rachel Audrey Nayang Mundo, Alex Durand Nka, Ezechiel Ngoufack Jagni Semengue, Grâce Angong Beloumou, Aude Christelle Ka’e, Willy Leroi Togna Pabo, Désiré Takou, Aissatou Abba, Sandrine Claire Djupsa, Evariste Molimbou, Naomi-karell Etame, Aurelie Minelle Kengni Ngueko, David Kob Same, Jolle Nounouce Bouba Pamen, Aristide Stephane Abah Abah, Serge Clotaire Billong, Rogers Ajeh Awoh, Gregory Edie Halle-Ekane, Giulia Cappelli, Anne-Esther Njom-Nlend, Anne-Cecile ZK Bissek, Elvis Temfack, Maria Mercedes Santoro, Francesca Ceccherini-Silberstein, Vittorio Colizzi, Jean Kaseya, Nicaise Ndembi, Alexis Ndjolo, Carlo Federico Perno and Joseph Fokam in Journal of the International Association of Providers of AIDS Care (JIAPAC)
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported Chantal Biya International Reference Center for Research on HIV and AIDS prevention and management (grant number 2020/2021 Budget Plan).
Ethical Considerations
An ethical clearance was obtained from the regional ethics committee of the Centre region of Cameroon (Reference: CE N°-0135-/CRERSHC/2021), and a research authorisation obtained from the Director General of the CIRCB (Reference N°:1625/021). Informed consent was provided by each study participant, blood samples were collected by trained phlebotomists, and data were processed by using unique identifiers for purpose of confidentiality and privacy of the study participants’ information.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
