Abstract
Background and Aims:
Porphyromonas gingivalis (P. gingivalis) is considered as an important pathogen responsible for periodontal disease which is characterized by inflammation of gingiva and destruction of periodontal ligament and alveolar bone leading to loss of tooth. Along with clinical investigations, suitable microbiological analysis needs to be performed which could provide more insight into the disease severity. We aim to quantify P. gingivalis by real-time PCR (RT-PCR) and analyze its association with demographic data including clinical parameters.
Materials and Methods:
The study consisted of chronic periodontitis patients (CP group) and healthy subjects (H group) with 120 samples in each group. RT-PCR was carried out by the SYBR Green assay to target 16S ribosomal ribonucleic acid species-specific region of P. gingivalis. Standard strain of P. gingivalis ATCC 33277 was used as a control.
Results:
In the CP group, 79.16% samples were found positive for P. gingivalis, whereas 29.17% samples were positive in the H group. A significant difference was found when the prevalence was compared within males and females (P < .001 for both). In the older age groups, we found a higher rate of detection of P. gingivalis. As analyzed by Spearman’s correlation test, the number of cells of P. gingivalis was significantly associated with probing depth (P = .02) and clinical attachment level (P = .01) in the CP group. The mean cell number of P. gingivalis was found to be increasing with increasing levels of probing depth and clinical attachment level (P < .001 and P = .01, respectively).
Conclusion:
The present study reaffirms that the P. gingivalis microbe is significantly associated with the chronic periodontitis and that its level varies with the severity of the disease. Colonization of the bacterium is significantly associated with severe forms of the disease.
Introduction
Chronic periodontitis is a common disease of the oral cavity characterized by inflammation of gingiva and destruction of periodontal ligament and alveolar bone leading to loss of tooth. 1 The interactions between microbes present on the tooth biofilm and the host tissues lead to host inflammatory/immune response, ultimately resulting in clinical signs and symptoms of periodontitis. 2 The microbial flora of the oral cavity is highly complex and diverse, and is composed of more than 700 bacterial species and phylotypes. Several bacteria, especially the red complex which form a part of the flora in the subgingival niche, are known to trigger the initiation of periodontal disease. 3 Among them, Porphyromonas gingivalis (P. gingivalis) is known to play an important role in pathogenesis of periodontal disease and is considered a keystone pathogen. 4
P. gingivalis is a gram negative, nonmotile, nonspore-forming anaerobic bacterium. The microbe possesses a number of virulent factors such as fimbriae, cysteine proteases, collagenases, hemagglutinins, and lipopolysaccharides. 5 These factors help the microbe to form a biofilm on the calcified hard tissues and epithelial cells of the gingiva. The bacteria invade the host gingival tissue and colonize in the deep periodontal pockets that provide an ideal envi-ronment for its growth and survival. 1
The initial diagnosis and treatment of the disease are based on the clinical investigations such as probing depth (PD) and clinical attachment level (CAL) measurements are deeply invaded by bacterial colonization. However, with the development of different molecular and microbiological techniques, it has also been advised to perform the microbiological analysis along with the clinical investigations to provide more insight into the disease severity. 6 Anaerobic culturing is the most commonly used technique for the detection of gram negative anaerobes, but it has several drawbacks. 7 Also, due to the slow growth rate of anaerobes, it imposes a challenge on the fast cultivation of this microbe. Many other alternate techniques have been tested for the detection of P. gingivalis in the oral cavity. These include molecular methods such as hybridization, immunoassays, and PCR.8-10 However, each technique has its own advantages and drawbacks, and it is important to choose a method of detection based upon the objectives of the proposed study. Quantitative examination is of real significance because the abundance of bacteria is more imperative than mere occurrence of bacteria which may assist in understanding the bacterial etiology in periodontitis all the more unequivocally. The real-time PCR (RT-PCR) method is a rapid, sensitive, and reliable technique for the detection and quantification of individual microbial species.11,12
The role of P. gingivalis in oral health and disease has been explored by many researchers around the globe using different laboratory techniques. However, different results have been obtained because of various environmental factors, lifestyles, and techniques used for analyses. In India, the occurrence of P. gingivalis in patients with periodontal disease was analyzed by conventional methods such as microbial culture and qualitative PCR.10,13,14 The aim of the present study is to quantitatively analyze the occurrence of P. gingivalis in patients with chronic periodontitis and healthy individuals by RT-PCR and to correlate its association with demographic data and clinical parameters in these subjects.
Materials and Methods
The current study consisted of 120 chronic periodontitis patients (CP group) and 120 healthy subjects (H group). Ethical clearance for the study was obtained from the Institutional Review Board (Certificate No. 819). All the participants included in the study were screened to meet the inclusion and exclusion criteria.
The exclusion criteria included subjects diagnosed with diabetes, HIV infection, and pregnancy; those using immunosuppressant medications, tobacco in any form; and those needing prophylactic antibiotics, on antibiotic therapy or oral prophylaxis within the last 3 months. All subjects needed to have a minimum of 20 teeth in dentition. The presence of gingival inflammation, presence of bleeding on probing, probing depth ≥5 mm, and clinical attachment loss ≥ 3 mm were the criteria used to select the patients for chronic periodontitis group. Healthy subjects were selected on the basis of no signs of gingival inflammation, absence of bleeding on probing, probing depth ≤5 mm, and clinical attachment loss ≤3 mm. An informed consent was obtained from each participant before enrollment in the study.
Subgingival plaque samples were collected with the help of a universal curette. Selected area was isolated, air dried and supragingival plaque was removed using cotton rolls. Subgingival plaque was collected and transferred into vial containing Tris EDTA buffer. Clinical parameters such as gingival index (GI), plaque index (PI), probing depth (PD), and clinical attachments level (CAL) were recorded during each sample collection.
Standard strain of P. gingivalis ATCC No 33277 was used in the study. The initial colony count of the bacterial broth was adjusted to 109 colony forming units/ml (CFU/ml) by using 0.5McFarlands turbidity standard. DNA extraction was carried out by the “modified proteinase K method” as described previously. 15 Briefly, the samples were vortexed, washed, and then bacterial cells were lysed by using lysis buffers containing inorganic reagents. Protein was degraded by the addition of proteinase K (10 mg/ml). The samples were then centrifuged and the supeant containing DNA was collected and stored at –20℃ till fuher processing.
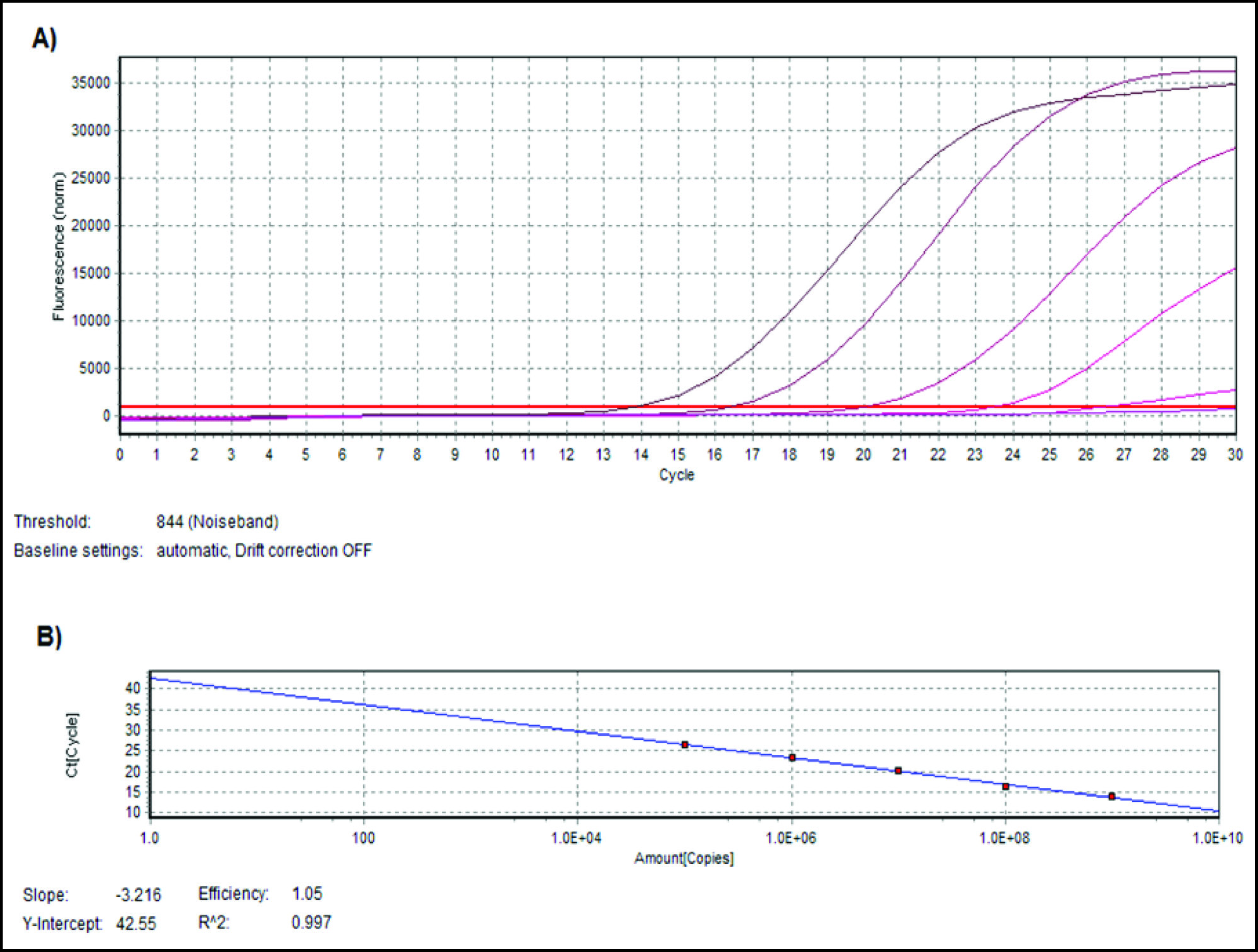
DNA extracted from the standard strain of P. gingivalis was serially diluted to obtain concentrations ranging from 109 to 105 CFU/ml to generate a standard curve. 16S ribosomal ribonucleic acid species specific gene of P. gingivalis was amplified by the primers; Forward primer 5ʹ-AGG CAG CTT GCC ATA CTG CG-3ʹ and reverse primer being 5ʹ-ACT GTT AGC AAC TAC CGA TGT-3ʹ. 9 RT-PCR was performed in 20µl total volume with FastStart Universal SYBR Green Master, 2× concentrated master mix (Roche, Switzerland) that contained 2.5 mM MgCl2, FastStart Taq DNA Polymerase, Reaction Buffer, Nucleotides (dATP, dCTP, dGTP, dUTP), and SYBR Green I dye. Primers at a concentration of 8 pmole/µl and DNA template of about 100 ng concentration were added to the reaction mixture. The tubes were kept in Realplex mastercycler (Eppendorf, Germany) and the thermal cycling conditions included the following: initial denaturation at 95℃ for 5 minutes followed by 35 cycles of denaturation at 95℃ for 30 seconds, annealing at 60℃ for 30 seconds and extension at 72℃ for 30 seconds. Melting curve analysis (dissociation curve) was performed in the range of 60℃ to 95℃ to check the specificity of the amplification reaction. Cycle threshold (Ct) values for all standard samples were obtained and a standard curve was plotted (Ct values against quantity). Regression line with slope of –3.2 and R2 value close to 1.0 is considered as optimum. Cycle threshold for unknown samples were obtained and plotted onto the standard curve and corresponding quantity was obtained.
Statistical Analysis
The statistical analysis was performed using graph pad prism software version 5.1 (GraphPad Software Inc., USA). The results of the H group and the CP group were compared by using Fisher’s exact test. The relationship between prevalence of P. gingivalis with clinical parameters was analyzed by using U Mann-Whitney test. The correlation of the clinical parameters with cell numbers of P. gingivalis was performed by using Spearman’s correlation coefficient test. The comparison of occurrence and number of cells of P. gingivalis with different categories of PD and CAL was done by one-way ANOVA. The value of P < .05 was considered statistically significant.
Results
The amplification plot and standard curve were obtained as shown in Figures 1A and 1B, respectively. Out of the total 120 samples from the CP group, 95 samples were found positive for P. gingivalis (79.16%), whereas 35 samples from the H group out of 120 were positive for P. gingivalis (29.17%). The CP group had significantly higher prevalence of P. gingivalis compared to the H group as statistically analyzed by Fisher’s exact test (P < .001).
The occurrence of P. gingivalis was compared for males and females of the CP group and the H group. Among 59 males of the CP group, 48 samples were found positive (81.35%) whereas in the H group of 75 males, 20 were found positive for P. gingivalis (26.66%). The difference between the groups was statistically significant (P < .001). Among females, the CP group had higher prevalence of P. gingivalis (77.04%) compared to the H group (33.33%) which was statistically significant (P < .001). When prevalence was compared between the gender irrespective of the CP or H group, more number of females (58.49%) showed presence of P. gingivalis compared to males (50.74%), but the difference was not statistically significant (P = .39).
The occurrence of P. gingivalis was analyzed in different age groups as shown in Table 1. There was significantly higher prevalence of P. gingivalis detected in 41 to 50 and 51 to 62 age group categories. The P value was statistically significant only within these old age group categories when compared by Fisher’s exact test (P < .001). For the age group of 21 to 30 and 31 to 40 years, the difference was not statistically significant (P = .20 and P = .08).
(A) Amplification plot and (B) standard curve generated for standard DNA extracts of P. gingivalis ATCC No. 33277 with concentrations ranging from 109 to 105 colony forming units/ml (CFU/ml).
Prevalence of P. gingivalis, Compared Within Each Age Category
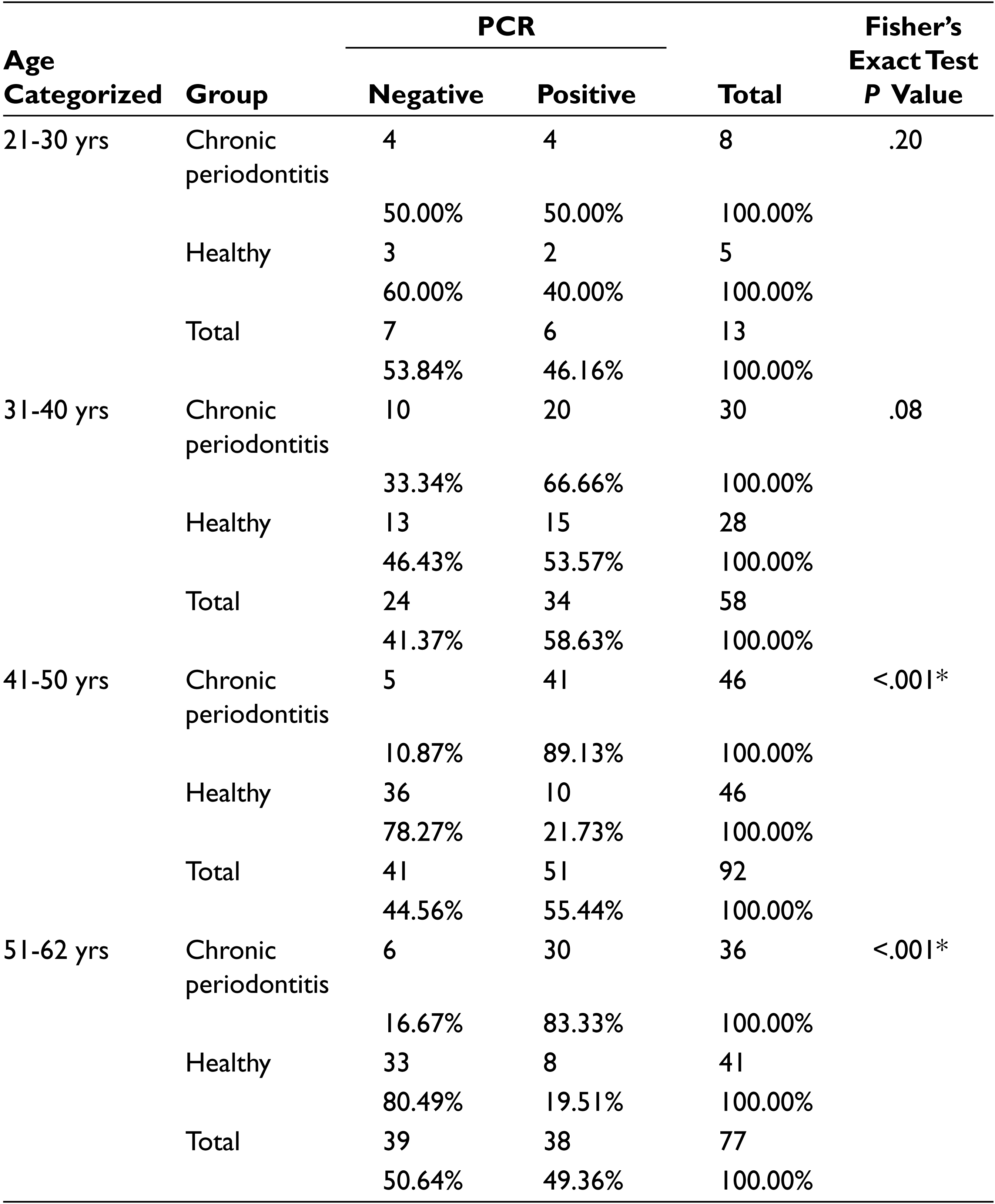
*P value is significant if < .05.
The comparison of the presence and absence of P. gingivalis in relation to CAL in CP subjects showed that the difference was statistically significant (P = .007). There was no significant association found among PD, GI, and PI when analyzed in 120 CP subjects as shown in Table 2.
Clinical Outcomes in Patients with Chronic Periodontitis Based on the Presence or Absence of P. gingivalis
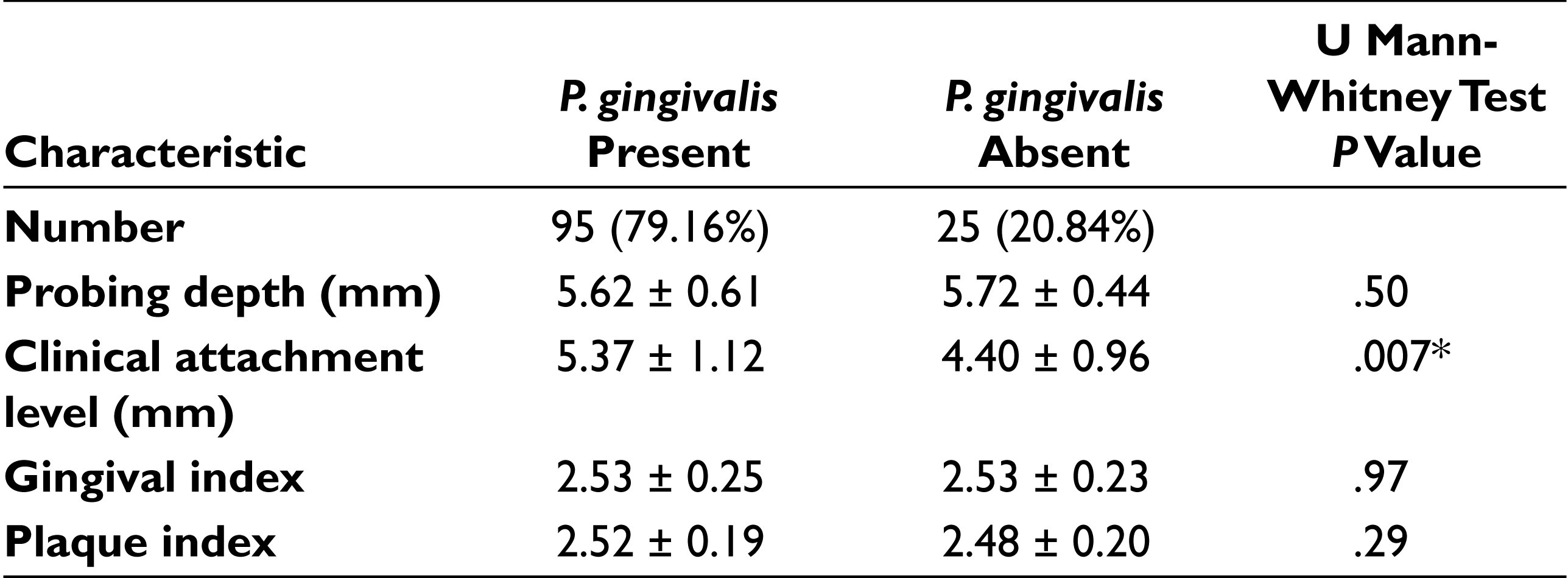
*P value is significant if < .05.
The number of cells of P. gingivalis present in each sample was obtained from RT-PCR. Our samples showed that a higher level of P. gingivalis was detected in the CP group (mean 1.47 × 108 cells) in contrast to the H group (mean 6.99 × 105 cells). The lowest detection limit by RT-PCR was found to be 1.24 × 102 cells. Spearman’s correlation test was applied to know the relationship between clinical parameters and number of cells of P. gingivalis. There was no significant association found between the level of P. gingivalis with GI and PI in both the CP and H groups. However, the association was significant with PD and CAL in the CP group (P = .02 and P = .01, respectively). In the H group, there was no significant association between PD and CAL with copy numbers, as shown in Table 3.
Spearman’s Correlation Test; Relationship Between Clinical Parameters and the Number of Cells of P. gingivalis by RT-PCR
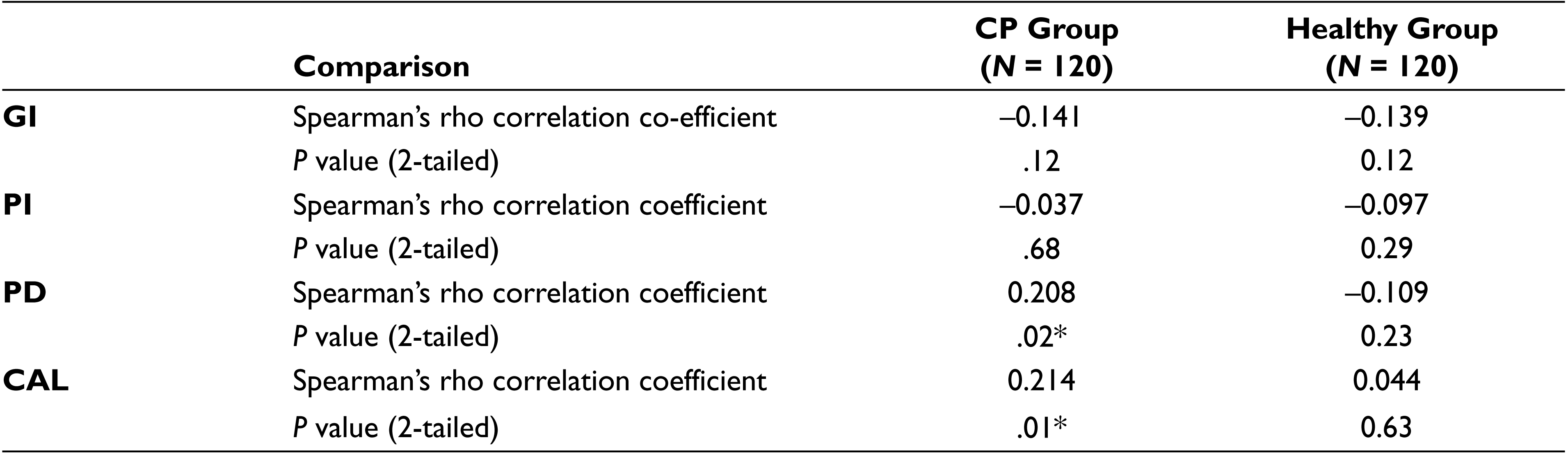
*P value is significant if < .05.
The PD was categorized into three groups: mild (5-6 mm), moderate (6-7 mm) and severe (≥7 mm). P. gingivalis was detected in 72 samples out of 97 samples (74.22%) in mild group. Prevalence of P. gingivalis was 100% in both moderate and severe group. The prevalence of P. gingivalis increased significantly with deeper PD (P = .02). The presence of P. gingivalis to severity of the disease was also analyzed quantitatively. The mean cell numbers of the bacterium increased with deeper PD. The difference between the groups is found to be significant with P < .001 (Table 4). The shallow PD was detected with lower cell numbers, whereas the deep PD had highest cell numbers. There was no significant difference in the quantity of P. gingivalis found between mild and moderate groups. However, there was a highly significant difference found between mild to severe and moderate to severe groups (P < .001 in both cases) (Figure 2A).
Association of Occurrence and Number of Cells of P. gingivalis in CP Patients With Different Categories of Probing Depth and Clinical Attachment Level
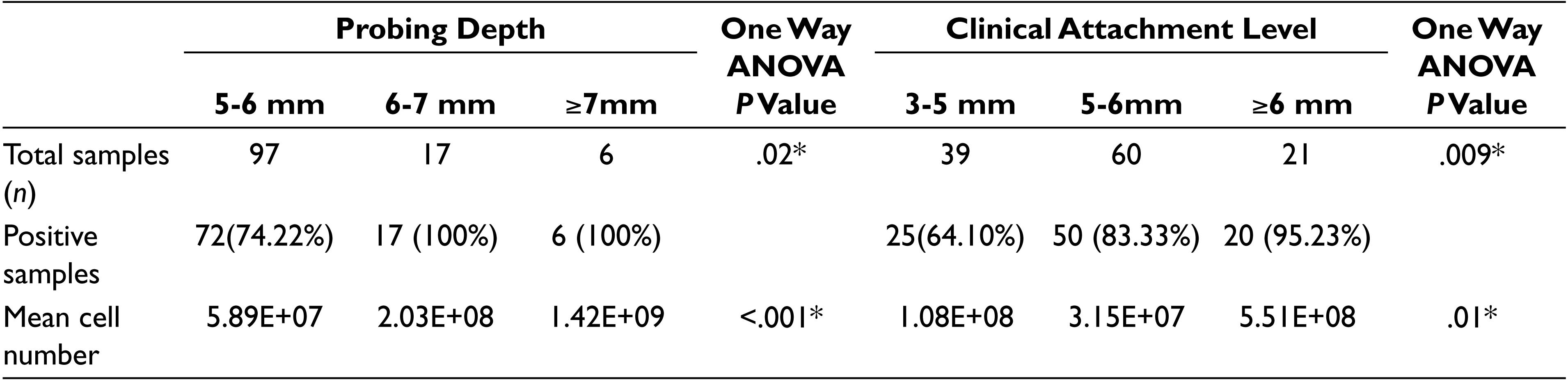
*P value is significant if < .05.
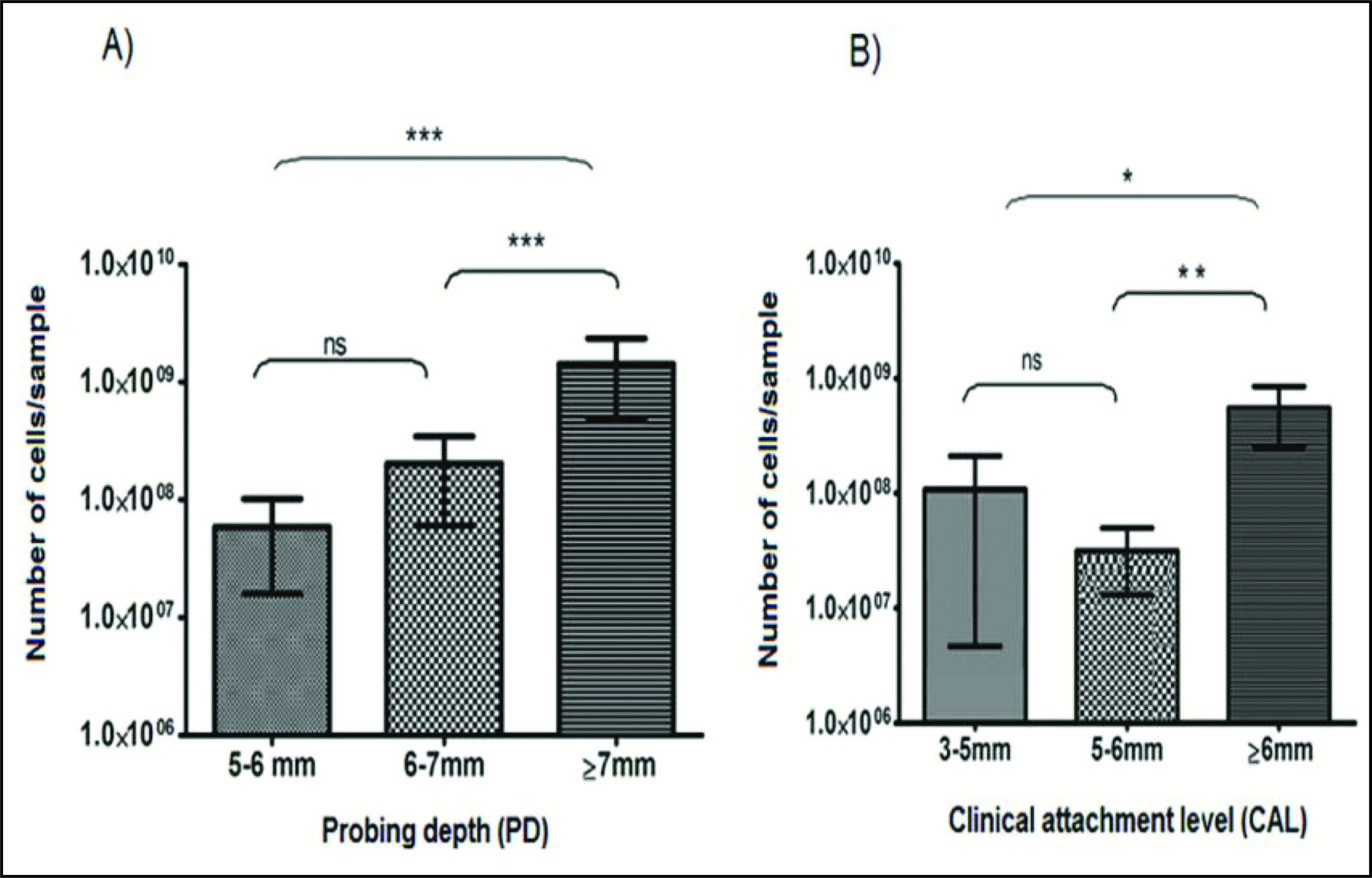
The CAL was also categorized into three groups. The prevalence in mild (3-5 mm), moderate (5-6 mm), and severe groups (>6 mm) were 64.10%, 83.33%, and 95.23%, respectively. The difference between the groups was significant with P = .009. The quantity of P. gingivalis in the mild group was more compared to the moderate group, with the highest quantity being present in the severe group. The difference between the groups was significant (P = .01) (Table 4). On comparing the mean cell numbers, there was no significant increase found between mild and moderate groups. However, there was a significant difference in the cell numbers between mild and severe groups and also between moderate and severe groups (P < .05, P < .01, respectively) (Figure 2B).
Discussion
Periodontal disease is an inflammatory response to polymicrobial colonization of oral pathogens leading to tissue destruction. P. gingivalis has been a major pathogenic colonizer in periodontal cases as found out by previous studies. 16 Severity of disease condition has to be correlated with the presence of this bacterium to establish its role in the progression of the disease.
The prevalence of P. gingivalis in the CP and H groups (79.16% and 29.17%) in the present study is similar to the findings of Griffin et al and Lopez NJ et al.17,18 Our results are also in agreement with study by Krishnan Mahalakshmi et al, which was carried out on Indian subjects by PCR. 13 Joshi et al found 66% prevalence of P. gingivalis in the CP group which was also carried out in Indian subjects. 10 There are researchers who also found more than 80% prevalence of P. gingivalis in the CP group.19,20 In few other studies, we found less prevalence of P. gingivalis in the CP group which was ranging from 45% to 53%.21,22 The difference in the prevalence of bacteria in the CP group could be due to different geographical locations, inclusion criteria, techique used for detection, and variation in sample size between other previous studies. The prevalence of P. gingivalis in the H group was found to be 29.17% which is well in accordnce with other studies.17,23 This suggests that P. gingivalis is present not only in chronic periodontitis group but to a lesser extent it can also get colonize in healthy individuals. This could also establish the fact that presence of P. gingivalis could become a potential risk factor to develop a diseased condition in healthy individuals. This leads us to believe that the non-virulent strains of P. gingivalis could acquire the pathogenic ability and transform into virulent strains responsible for disease progression. The present study showed a strong association of P. gingivalis with chronic periodontitis. Highly significant statistical difference was observed between the CP group and the H group (P < .001). These findings were in agreement with previous studies.18,22
Both males and females of the CP group were detected with more number of positives compared to healthy group. Whereas when prevalence was compared within males and females in both the groups together, the statistical significance was not observed. This suggests that P. gingivalis could infect any adult individual irrespective of the gender. The older age groups were detected with more number of positive cases compared to younger age groups, which could mean the older age group is at higher risk to get infected with P. gingivalis. The pathogenicity of P. gingivalis is attributed to different virulent factors which directly or indirectly destabilize the immunogenic responses from host and help the bacterium to invade the host tissue.
In our study we have quantitatively studied the colonization of P. gingivalis in relation to the severity of the disease. The highest cell numbers of P. gingivalis was obtained in the CP group compared to the H group. The colonization of the bacterium is more in the CP group than in the H group. The lowest detection limit by RT-PCR was found to be 1.24 × 102 bacterial cells which suggest that this technique can be used with high sensitivity for the detection of a range of pathogens responsible for periodontal disease. There is significant association of cell numbers with PD and CAL in the CP group, whereas in the H group there was no significant association found. Torrungruang et al reported significant association between higher prevalence of P. gingivalis with probing depth of >5 mm. 24 Similarly, in the current study we found that sites with deeper PD showed higher levels of P. gingivalis than in sites with shallow PD. As analyzed by Spearman’s correlation test, the levels of P. gingivalis was significantly associated with PD (P = .02) and CAL (P = .01) in the CP group. Abdulaziz et al also found positive correlation between P. gingivalis and increasing probing depth. He also reported the higher rate of detection of P. gingivalis in samples with CAL >6 mm. 25 Klein et al found 90% prevalence in samples with probing depth >5 mm whereas with probing depth ≤5 mm, the prevalence was 40%. 26 Similar results were obtained by other researchers where they found P. gingivalis cell count increase significantly with increased probing depth.27,28 Higher prevalence of P. gingivalis in deeper pockets reveals that deep pockets provided ideal environment for the microbe to grow and survive.
As stated earlier, the virulence plays an important role in the bacterial colonization. This implies the need for the quantitative evaluation of this periodontal pathogen with respect to the occurrence of different virulence genotypes to be analyzed by a suitable molecular technique in order to get clear insight into the disease progression. Although microbiological culture technique is still considered as a gold standard for the detection of P. gingivalis, RT-PCR provided several advantages over the conventional methods. In addition to quantitative evaluation, RT-PCR offers the advantage of eliminating false positives which could otherwise play abysmal role in conventional detection techniques.
Conclusion
The present study reaffirms that P. gingivalis is significantly associated with the chronic periodontitis group and that its level varies with the severity of the disease. The colonization of the microbe is significantly associated with severe forms of the disease. RT-PCR technique can be effectively used further to correlate the pathogenicity of the bacterium taking into consideration the different virulent factors of P. gingivalis.
Footnotes
Declaration of Conflicting Interests
The authors declare that there are no conflicts of interests regarding this study.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
