Abstract
Occupational health nurses play a key role in evaluating innovative technologies that can aid in providing safe and rapid care and reduce lost work time. A nurse-led employee health clinic participated in a validation study of a novel pathogen detection technique developed by GeneCapture, Inc. Their proposed portable urinary tract infection (UTI) in vitro diagnostic test was challenged with discarded, deidentified urine samples from patients presenting with typical UTI symptoms collected at two university clinics and two multiphysician practices. GeneCapture’s panel for this study was designed to rapidly identify the genetic signature of seven organisms: gram-negative Escherichia coli, Klebsiella pneumoniae, Proteus mirabilis, and Pseudomonas aeruginosa; gram-positive Enterococcus faecalis and Staphylococcus aureus; and fungal Candida species. The results from 40 clinical samples were in 95% agreement (90% specificity, 100% sensitivity) with traditional urine culture results from routine analysis. This successful occupational health nursing collaboration and validation study shows promise for point-of-care diagnoses and earlier treatment for workers with UTIs.
Introduction
Occupational health nurses work in a variety of settings and are one of the largest professional groups providing care to workers (Thompson, 2010). An important aspect of the occupational health nurse role is one of advocacy in promoting environments that ensure safe and quality care while at the same time remaining cognizant of health care costs. Employers view hours not worked because of health-related issues as hours lost (Graeve, McGovern, Nachreiner, & Ayers, 2014). Health care technology can be useful in assuaging those “hours lost” and maintaining productivity.
Occupational health nurses serve a key role in evaluating emerging technologies that can safely and rapidly aid in providing care. Urinary tract infections (UTIs) are one of the most common bacterial infections affecting women of all ages (Arnold, Hehn, & Klein, 2016). UTIs are responsible for nearly 10 million health provider visits each year, with one in five women having at least one UTI in her lifetime (Schappert & Rechtsteiner, 2008). An estimated 30% to 44% of women will have a recurrent UTI within 6 months of an initial infection. Foxman (2014) defines a recurrent UTI as three or more UTIs in 12 months, or two or more infections in 6 months.
Given the frequency and painful discomfort of urinary infections and the need to be seen by a health care provider for diagnosis and treatment, a UTI is a common reason for missing work (Foxman, 2014). Although many women see their health care provider promptly, therapy decisions for UTI often require a lengthy 2 to 3 day culture analysis. On-site employee health clinics are the perfect venue for patients to be diagnosed and treated as rapidly as possible. Care of the employee population at the worksite is basic to occupational health nursing. Employers are always looking for ways to cut costs, such as productivity loss from health-related absences and use of benefits such as sick leave (Graeve et al., 2014).
Work-Site Project
A nurse practitioner-led employee health clinic on the campus of a research intensive university recognized the problem of the frequency of UTI visits, the need for rapid detection, and the importance of returning to work as soon as possible with minimal loss of work productivity. Treatment of UTIs must be performed promptly for symptom relief and patient recovery. It is standard practice for nurse practitioners and other providers to treat UTIs empirically (Colgan & Williams, 2011). Untreated UTIs can lead to complications such as pyelonephritis and/or sepsis, depending on the age of the employee (Arnold et al., 2016). The nurse practitioner (also the director of the clinic) partnered with GeneCapture, Inc., a resident associate company at HudsonAlpha Institute for Biotechnology in Huntsville, Alabama. HudsonAlpha is a bioscience and genomic research institute that includes private companies on their campus. GeneCapture is a start-up company in the medical devices field, building a portable device that identifies the cause of an infection in about an hour. The technology is based on a patented complementary genetic signature hybridization of pathogenic RNA to specially designed synthetic DNA probes (see www.genecapture.com).
Rapid diagnosis of a UTI is essential for proper treatment, antimicrobial stewardship, and prevention of life-threatening complications (Becknell, Schober, Korbel, & Spencer, 2015; Centers for Disease Control and Prevention [CDC], 2012; Marques et al., 2012). Cell culture is the gold standard for diagnosing a UTI, but results generally take 2 to 3 days during which time an unnecessary or wrong antibiotic may have been prescribed (Nicolle, 2014). Cheap, rapid dipstick strips to identify the presence of blood, nitrites, and leukocytes in urine are routinely used by medical staff and are now available for purchase as at-home screeners. However, false positives and negatives are common and elevated levels in the elderly confound their utility (CDC, 2015; Pezzlo, 2014). Amplification-based molecular diagnostics have largely ignored the UTI market due to the misleading problem of amplifying contaminating urogenital flora found in most urine specimens (Beveridge, Davey, Phillips, & McMurdo, 2011). GeneCapture developed an in vitro diagnostic (IVD) platform technology to address this gap.
They developed a method called “CAPTURE” for Confirming Active Pathogens Through Unamplified RNA Expression. The system uses a combination of specific nucleic acid probes (captors) to quickly identify the pathogens in a patient sample. The portable instrument and disposable UTI panels were designed to enable a UTI diagnosis in physician’s offices, clinics, long-term care facilities, and remote or underserved locations rather than remote, diagnostic labs. Enabling home health aides to carry and use a portable IVD to test symptomatic patients during regular home visits could reduce the delay in obtaining needed treatments and would avoid the inconvenience and cost of a visit to a medical facility, especially as more Americans choose to age at home.
Innovation
DNA that is complementary to each selected pathogenic nucleotide sequence is synthesized into the loop of a structured captor probe (Figure 1a). All captors have unique loop regions, but share the same universal stem sequences. When pathogenic nucleic acids pass over a microarray of such captors, they will bind to their complementary loop regions, and this new, more thermodynamically favorable binding forces those captors’ stems to open (Figure 1b). Next, a fluorescently labeled universal detector is allowed to bind to the common tail of any open captor (Figure 1c). Thus, the binding of the detector simply and quickly reveals the presence of the pathogen in the sample. The instrument actuates the movement of the sample over the array and reads the fluorescent signals that indicate which probe was activated.
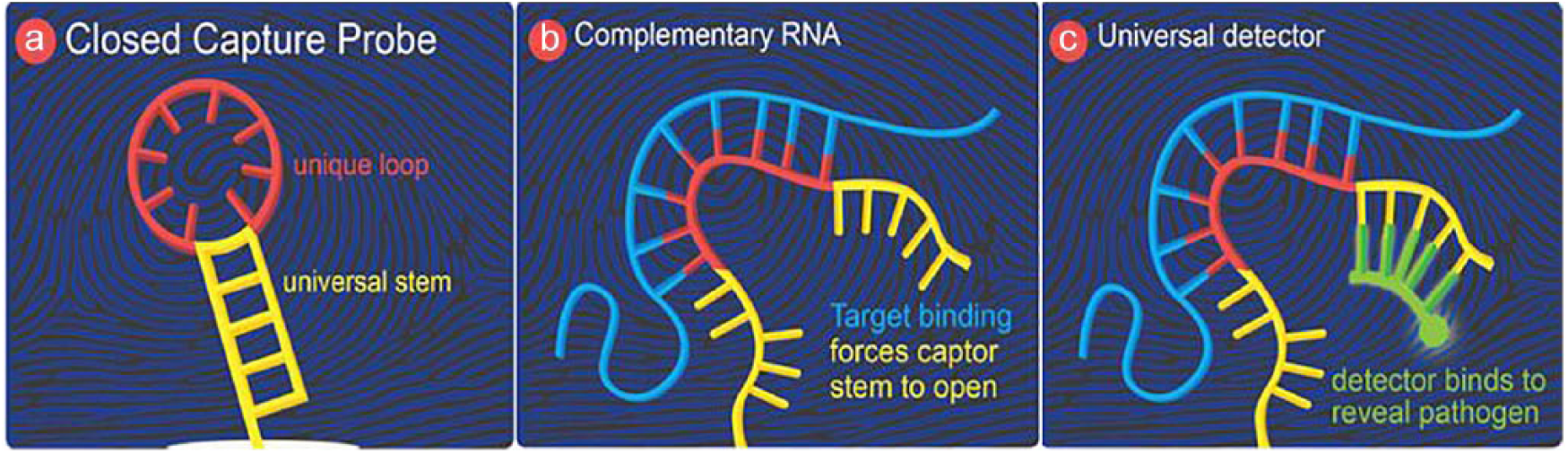
CAPTURE assay principle.
A desktop prototype at the GeneCapture facility was used for this study. This molecular approach with a fixed microarray allowed the diagnostic to screen for many pathogens at once.
Study
To test the CAPTURE method in the occupational setting, an advanced practice occupational health nurse partnered with GeneCapture, Inc., researchers. During the development phase of CAPTURE, the occupational health nurse visited the GeneCapture, Inc., laboratory at different stages to serve as a clinical consultant. The study was approved by the institutional review board from the University of Alabama in Huntsville, and a memorandum of understanding was obtained between the two institutions.
GeneCapture’s UTI panel consists of a set of proprietary surface bound captors printed in a microarray included in each assay cartridge. The captors were designed to target organisms causing more than 90% of UTIs. Reusable prototype cartridges, with interchangeable arrays, were used in this study for cost purposes. The method to clean and decontaminate the cartridges has been verified. Two configurations of the microarray were used in this study including a preliminary panel in Cohort 1 and an intermediate panel in Cohort 2.
Samples spiked into clean urine were made from multiple clinical isolates of each organism targeted on the panel. These spiked samples were processed through the CAPTURE assay, and the resulting fluorescent signals from the entire panel of captors were used to generate pathogen-specific “fingerprints” that subsequently allowed for the identification of organisms in the clinical samples.
Study Sample
Cohort 1 samples were collected from the university clinic in the fall of 2016 and Cohort 2 samples were collected primarily in the summer of 2017 when patients with a suspected UTI signed the consent form allowing GeneCapture to study their deidentified, discarded urine. An initial set of four samples was processed to ensure that the manual sample prep protocol used during this study was sufficient for actual clinical samples. A total of eight patient samples were included in Cohort 1. For Cohort 2, 10 additional patient samples from the university clinic and 10 samples from a family practice clinic and 22 samples from a urologist practice were collected for a total of 42 specimens. All samples were picked up daily (with two clinic pick-ups per day) or immediately upon notification and were processed within 1 hour of receipt. All samples sent to GeneCapture were allocated a unique identification number. The primary samples were sent for routine culture analysis to the laboratory of choice for each practice and those results were later made available to GeneCapture by sample number for validation.
Method and Materials
The sample preps were performed manually in the GeneCapture BSL2 lab. The purpose of the sample prep is to release the pathogenic RNAs. The sample prep process is quite simple when compared with other molecular diagnostics because the CAPTURE technique does not require any enzymatic steps. The prep is accomplished in three steps: concentration (centrifugation), lysis (heat and beads), and fragmentation (zinc hydrolysis). The volume of the crude lysate was diluted to 1.1 ml in buffer and transferred to an assay cartridge or frozen at −80°C for later use.
Before being assayed, the clinical samples were assigned numbers to remove provider practice information from the subsequent analysis. All assays were run in GeneCapture’s desktop prototype device. The arrays were scanned for fluorescent signals through the back of the glass slide while still immersed in the final rinse buffer using a custom laser scanner.
Results
The identification method used in this study is based on the change in normalized intensity (ΔNI) profile of the entire panel to identify discreet patterns of signal in the presence of each pathogen. To fully implement this approach, assay runs of spiked known clinical isolates were used to create reference “fingerprints” of each species’ profile across the panel. The fingerprint profiles for Klebsiella pneumoniae and Pseudomonas aeruginosa are shown in Figure 2 where the trace for each bacterium is the average of four runs of multiple strains.
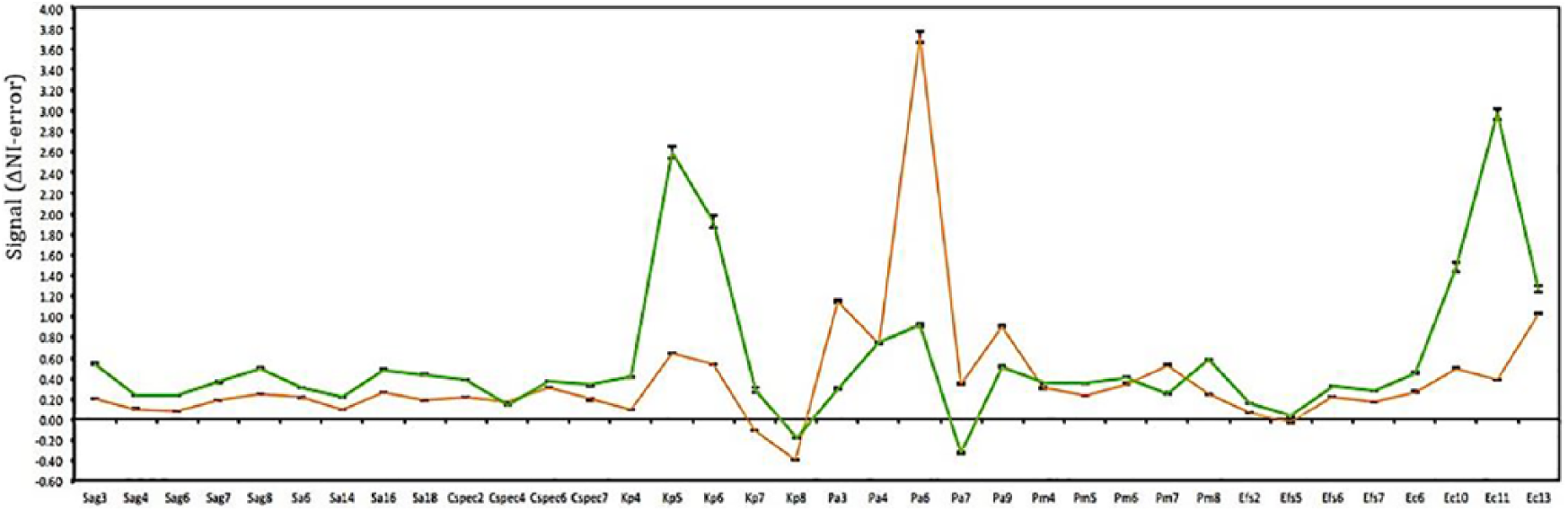
Consensus profiles of two genetically distinct uropathogenic bacteria: Klebsiella pneumoniae (green) and Pseudomonas aeruginosa (orange).
The key to this method’s success is the determination of clearly distinct patterns in the panel signal that is organism-dependent. The y values in Figure 2 represent signal intensity and the x axis represents specific captor probes.
The captors identifying Candida were designed to distinguish infections from this genus against bacterial infections. Five isolates were obtained from the CDC’s Candida bank or Huntsville Hospital (Huntsville, AL). The five organisms tested gave a definitive identification of Candida. In other words, the captors were able to identify the organism correctly.
As illustrated in Figure 3, the Candida specific captors present variable profiles: Candida albicans, Candida glabrata, and Candida tropicalis being the most similar and Candida parapsilosis and Candida lusitaniae more unique. These results do suggest the ability of the CAPTURE assay to differentiate various species of Candida as well as to distinguish either Candida or bacteria as a causative infectious agent.
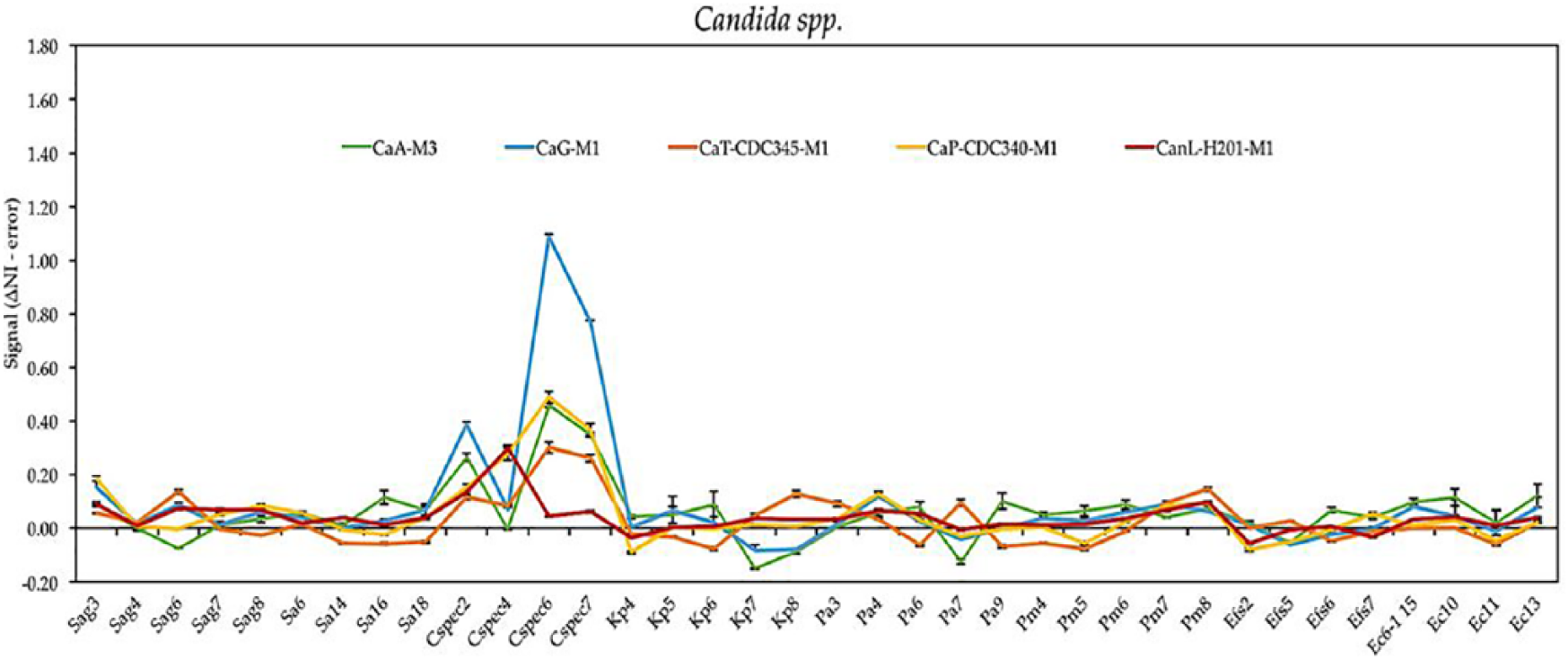
Panel signal when exposed to five different species of Candida; Candida albicans (green), Candida glabrata (blue), Candida tropicalis (orange), Candida parapsilosis (yellow), and Candida lusitaniae (red).
Of the 51 samples collected, one was not sent for culture and was excluded. Although the CAPTURE assay was sufficiently robust in that it only required simple crude cell lysates, one sample (from a patient noted as having prostate cancer) could not be processed through the GeneCapture manual protocol and was also excluded. Three samples encountered human or machine anomalies during the assay process (one broken slide, two positive control errors) and were also excluded.
The volumes of the 46 remaining samples ranged from 7.5 ml to 70 ml (Mdn = 35 ml; M = 32 ml). Of the samples with patient data, nine were confirmed to be from males and 19 from females with an age range from 22 to 83 years (Mdn age = 64 years; M age = 66 years). Two of the samples were confirmed to be from non-White ethnicities: one African American and one Hispanic.
Six patients self-reported to be on medicines at the time of the sample collection (Table 1). Significantly, none of these medicines appeared to have interfered with the successful outcome of the CAPTURE assay. Six of the 46 samples (including one on antibiotics) were reported by CAPTURE as negative because the causative organisms were not targeted on this study’s intermediate UTI panel. These nontarget organisms were identified by culture as Enterobacter cloacae, Enterobacter aerogenes, Streptococcus viridans, Acenitobacter radioresistens, Staphylococcus saprophyticus, and Citrobacter freundii. Four of these samples came from the urology practice, which generally had an older, less healthy patient population.
Self-Reported Patient Medications Did Not Interfere With the CAPTURE Assay (Diagnostic Test for Possible Urinary Tract Infection)
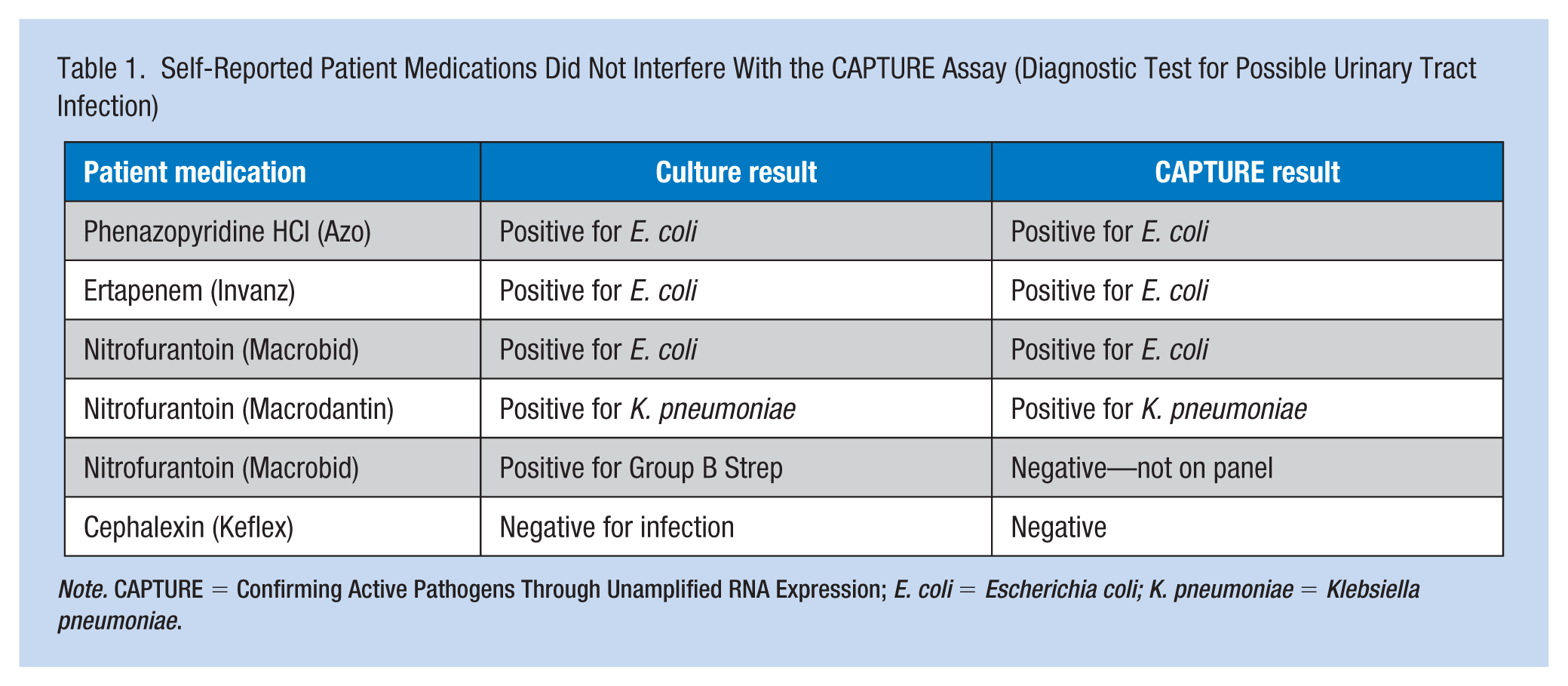
Note. CAPTURE = Confirming Active Pathogens Through Unamplified RNA Expression; E. coli = Escherichia coli; K. pneumoniae = Klebsiella pneumoniae.
For the remaining 40 samples, cell culture results determined that 21 were positive for UTI and 19 were negative. The CAPTURE assay successfully confirmed all of the culture-negative samples as negative. For the culture-positive samples, the CAPTURE assay readily confirmed 17 positive for Escherichia coli and two positive for K. pneumoniae. There were two culture-positive E. coli samples that the CAPTURE assay did not identify. One of these had no signal and we believe the RNA was degraded during the manual sample prep process. The second had such a low signal across the array that its identification was not achievable with the described comparison to the profiles. In summary, the overall sensitivity of the CAPTURE assay is 19/21 or 90%, and the specificity is 19/19 or 100%, with a culture agreement of 95% (Table 2).
Preliminary Validation of CAPTURE Versus Gold-Standard Culture Testing for Urinary Tract Infection Diagnosis
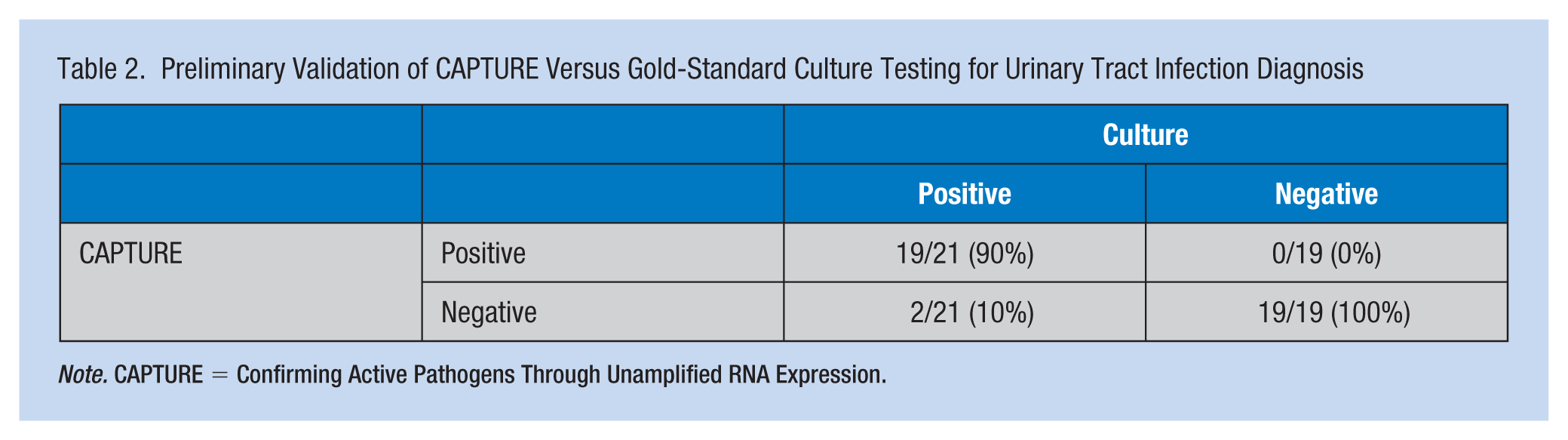

Note. CAPTURE = Confirming Active Pathogens Through Unamplified RNA Expression.
The dynamic range of the organisms identified in this study was from 2E4 CFU/ml to 1E8 CFU/ml, with the lowest CAPTURE-positive sample having a total of 1E6 CFU. The quantitative nature of the direct CAPTURE assay will allow reporting of the number of organisms in a sample in the ranges currently used for UTI reporting. The determination of such values was not a goal of this preliminary validation study.
The average time for culture results to be returned to the clinics or physicians’ offices was 3.2 days; the shortest time was 23 hours and the longest was 9 days. However, 14 of the culture results were faxed to the clinics or physicians’ offices over the weekend when they were closed, further delaying the ability to treat 10 of these patients who had positive cultures in a timely manner.
Discussion
The preliminary clinical validation of the CAPTURE UTI IVD resulted in higher than anticipated values for agreement, sensitivity, and specificity. The results suggest that the CAPTURE assay’s clinical effectiveness for rapidly detecting UTI in workers is promising.
The percentage of negative urine cultures reported in this study (48%) is consistent with typical ambulatory culture results, where roughly 65% of all urine specimens sent for culture are negative (Pezzlo, 2014). The rapid determination of those negative samples is important to prevent the unnecessary use of antibiotics. Likewise, the rapid determination of the true causative agent of a UTI allows the correct medicine to be rapidly prescribed to prevent extended patient suffering while waiting for culture results. The delays observed in this study due to weekend reporting (14 out of 51, or 27%) were larger than expected and further highlight the need for same day, point-of-care testing.
Quantitation of the organisms causing a UTI is a critical part of UTI diagnostics. Culture results report the causative pathogen along with its concentration range (from 10,000 to >100,000 CFU/ml). Samples of certain uropathogenic bacteria are considered infectious at levels down to 1,000 CFU/ml. GeneCapture’s scanner creates a sensitive, low-noise image of the final wet array that is able to detect small changes in fluorescence. These fluorescence values are quantitative and will be used to determine the presence of pathogenically relevant bacterial loads.
Three of the six samples that were culture-positive but whose infectious agent was not yet part of the CAPTURE assay (the two Enterobacter species and S. viridans) had enough homology to existing captors on the Cohort 2 panel that they gave sufficient signal to indicate the presence of bacteria. However, without a comparative fingerprint, these organisms could not be specifically identified.
There were five samples that were reported by culture as mixed urogenital flora. The CAPTURE results for these samples were also called mixed flora because each sample showed low signals from various captors that target both gram-positive and gram-negative organisms. The CAPTURE assay should be able to identify the profiles of mixed infections as well as mixed flora, but an additional learning set will be needed.
This study has many implications for occupational health nurse practice and the medical and nursing field in general. This form of rapid diagnostic test allows for the determination of the infecting pathogen in an employee clinic for timely and accurate treatment. In the future, this technology has the potential to be used in areas with little or no access to laboratories or medical facilities. The potential for use in pandemic emergence or for public health tracking is also important. GeneCapture’s rapid test can get therapy started 30 hours sooner than the current laboratory culture results.
Collaboration of Clinical Occupational Health Nurses With Researchers
Occupational health nurse involvement in this research was initially participation in the validation study and then generation of new knowledge and front line diagnostic involvement. By virtue of their workplaces, occupational health nurses are uniquely placed to design, conduct studies, and evaluate effectiveness of programs. Occupational health nursing research investigates health and safety issues affecting workers and this is a solid example (Salazar, 2002). This was not just an intellectual exercise, but rather a project that is applicable to practice at the point of care delivery. Ensuring safe, competent, and ethical care is the cornerstone of nursing and with occupational health nurses especially, ensuring that employees return to work healthy and as rapidly as possible is paramount. This innovative method of diagnosing and treating UTI allows for faster and accurate therapy. It will help productivity through prevention of health-related absences and less use of benefits such as sick leave and short-term disability.
This study has also provided an example of important collaboration between product developers and front line clinicians. Occupational health nurses should be open to opportunities for collaborations that could enhance the services offered to workers. In this instance, the opportunity to work with scientists from GeneCapture came about when the Chief Executive Officer of the company reached out to the clinic. GeneCapture was looking for clinical partners. This led to participation in the validation study, which in turn led to opportunities for students to work with bench scientists, and the development of a product that can help with rapid diagnosis and treatment of a common infection. Occupational health nurses are encouraged to meet their goal of worker health and safety and provide cost-effective, quality services to their population. To help advance the field, occupational health nurses are a valuable source for validation of new products, and their operational input can have significant impact on the quality outcomes of new product designs. In this case, the successful validation of rapid on-site diagnosis of UTIs leads to a promising improvement in patient care.
Footnotes
Conflict of Interest
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Carter Wright reports being an employee from GeneCapture both during the conduct of the study and outside the submitted work. Dr. Chittur reports other from GeneCapture, outside the submitted work. In addition, Dr. Chittur has a patent “Compositions, Methods and Devices Comprising Stem-Loop Captor Molecules” pending. L. Savannah Dewberry reports being an employee of GeneCapture, Inc., during the conduct of the study and outside the submitted work. Paula Koelle reports being an employee of GeneCapture, Inc., during the conduct of the study and outside the submitted work. In addition, Koelle has a patent “Compositions, Methods and Devices Comprising Stem-Loop Captor Molecules” pending. Elizabeth Gates reports being an intern at GeneCapture, Inc., during the conduct of the study. J. Ethan Stallings reports other from GeneCapture, Inc., during the conduct of the study; other from GeneCapture, Inc., outside the submitted work. Zachary McGee reports being employed by GeneCapture, Inc., during the conduct of the study and outside the submitted work. In addition, McGee has a patent “Compositions, Methods and Devices Comprising Stem-Loop Captor Molecules” pending.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Author Biographies
Louise C. O’Keefe is assistant professor, College of Nursing and Director of the Faculty and Staff Clinic at The University of Alabama in Huntsville.
Paula Koelle is a graduate of Rhodes College with a BS in chemistry and of Harvard University with a MA in biochemistry and molecular biology. She is currently the chief scientist with GeneCapture.
Zac McGee is a graduate of the University Of Alabama in Huntsville with a BS in chemical engineering, chemistry, and mathematics who is currently employed as a research specialist with GeneCapture.
L. Savannah Dewberry is a graduate of the Science and Technology Honors College at the University of Alabama, Birmingham with a BS in biomedical engineering who is currently pursuing her PhD at the University of Florida.
Carter Wright is a graduate of the University of Tennessee at Chattanooga with a BS in biology who is currently pursuing her PhD at the University of Alabama in Huntsville.
J. Ethan Stallings is a graduate of the University of Alabama in Huntsville with a BS in chemical engineering and chemistry who is currently employed as a research associate with GeneCapture.
Elizabeth Gates is a graduate of the University of Alabama in Huntsville with a BSN in nursing who was selected for a BioTrain Internship with GeneCapture. She is currently practicing as a RN and pursuing her DNP at the University of Alabama in Huntsville.
Krishnan Chittur, is a Professor Emeritus of Chemical Engineering at the University of Alabama in Huntsville, and a founder and the CTO of GeneCapture, Inc
