Abstract
Intrathecally administered RNase H1-active gapmer antisense oligonucleotides (ASOs) are promising therapeutics for brain diseases where lowering the expression of one target gene is expected to be therapeutically beneficial. Such ASOs are active, to varying degrees, across most or all cell types in the cortex and cerebellum of mouse and non-human primate (NHP) brain regions with substantial drug accumulation. Intrathecally delivered ASOs, however, exhibit a gradient of exposure across the brain, with more limited drug accumulation and weaker target engagement in deep brain regions of NHP. Here, we profiled the activity of a tool, ASO, against Malat1 in three deep brain regions of NHP: thalamus, caudate, and putamen. All neuronal subtypes exhibited knockdown similar to, or deeper than, the bulk tissue. Among non-neuronal cells, knockdown was deepest in microglia and weakest in endothelial stalk. Overall, we observed broad target engagement across all cell types detected, supporting the relevance of intrathecal ASOs to diseases with deep brain involvement.
Introduction
Lowering the expression of a specific target gene in the central nervous system (CNS) is expected to be therapeutically beneficial across a range of diseases. 1 Antisense oligonucleotides (ASOs) with gapmer design, where the central region is unmodified DNA with 5' and 3' wing residues containing 2' sugar modifications, can trigger RNase H1 degradation of a target RNA, lowering its expression. 1 More than 10 such gapmer ASOs administered intrathecally have entered clinical development for brain diseases, and one—tofersen, targeting SOD1—has been approved.
Available autopsy data from humans who received intrathecally administered ASOs support relatively broad brain distribution,2,3 but as such data remain limited, our understanding of intrathecal ASO pharmacology comes primarily from pharmacology and toxicology studies in non-human primates (NHP) such as cynomolgus macaques, whose brain is ∼5% the mass of a human brain and ∼200 times larger than a mouse brain. 4 An intrathecal ASO against Malat1 distributed broadly throughout the brain of cynomolgus macaques, albeit with drug accumulation more than an order of magnitude lower in deep brain regions than in many areas of the cortex and spinal cord. 5 Accordingly, target engagement in deep brain regions was also more modest. An intrathecal ASO against PRNP achieved broad knockdown across various cell types in the cynomolgus macaque cortex and cerebellum, with a pattern of activity similar to that observed in mouse. 6 Nevertheless, the cortex and cerebellum exhibited strong drug accumulation in that study, with 21 and 16 µg/g of ASO respectively. The cell-type specificity of gapmer ASO, activity in deep brain regions with more limited drug concentration has not yet been evaluated.
Here, we use single nucleus sequencing to analyze tissue from an NHP model, cynomolgus macaques, treated intrathecally with a Malat1 ASO in order to characterize target engagement across cell types in each brain region.
Methods
ASO treatment of NHP
The Malat1 ASO used here has been described previously.5,6 Its sequence is
Single-nucleus sequencing
We utilized the same wet lab protocol 7 and computational pipeline 8 described previously. 6 Briefly: frozen tissue was triturated in Kollidon VA64, Triton X-100, bovine serum albumin, and RNase inhibitor, passed through a 26-gauge needle, washed and pelleted, passed through a cell strainer, and flow sorted for DAPI signal. After cell counting, a volume estimated to contain 17,000 nuclei was submitted to the Broad Institute’s Genomics Platform for 10X library construction (3′ V3.1 NextGEM with Dual Indexing), which was performed according to manufacturer instructions and sequenced (100 cycles) on an Illumina Novaseq 6000 S2. Data were analyzed on Terra.bio using Cumulus Cell Ranger 7.2.0 (Dockstore workflow https://dockstore.org/workflows/github.com/lilab-bcb/cumulus/Cellranger:2.5.0) with flags—include introns and—secondary. Because MALAT1 is not annotated in the Ensembl Macaca fascicularis 6.0 transcriptome, we created our own custom reference. We downloaded the Macaca_fascicularis_6.0 GTF file, used cellranger mkgtf to filter for protein_coding, lncRNA, and antisense transcripts, manually added lines for MALAT1 on the minus (−) strand spanning positions chr14:8898261-8905725, and finally compiled with cellranger mkref 8.0.1. Count matrices were then aggregated using Cell Ranger 7.1.0 (aggr with the—normalize flag set to none) to yield one UMI count matrix per species and brain region. Statistical analyses and data visualization were conducted using custom scripts in R 4.2.0.
Cell type assignment
Preliminary cell types were assigned using scType, 9 and final cell type determinations were made after manual curation in Loupe Browser. In order to identify cell types, we used reported markers from projects SCP2706 and SCP2719 in the Single Cell Portal 10 as well as previous publications on mouse, human, and NHP deep brain regions.9,11–13
Statistical modeling
To estimate knockdown in each cell type, we utilized the same negative binomial model described previously. The model was specified as glm.nb (target umi ∼ celltype + celltype:treatment +offset(log(total umi))). The coefficient for each cell type-treatment interaction term (in natural logarithm space) was then exponentiated to yield the mean estimate of the residual target RNA in that cell type, and the 95% confidence interval was that mean estimate ±1.96 of the model’s standard error. Individual animal point estimates were obtained by adding the model’s residuals to each coefficient before exponentiating.
Data and source code availability
An analytical dataset and R source code sufficient to reproduce these analyses will be made available at https://github.com/ericminikel/nhp_deepbrain_aso and the full sequencing dataset will be posted to the Single Cell Portal at https://singlecell.broadinstitute.org/
Results
Selection of brain regions for evaluation
We re-analyzed the previously reported 5 PK and pharmacodynamic (PD) parameters of a Malat1 ASO. In a dose-response study in mouse and rat, the median inhibitory concentration (IC50) in tissue varied by brain region from 0.1 to 4 µg/g (Supplementary Fig. S1), being highest in the cerebellum. We examined the PK versus PD across brain regions in NHPs (Fig. 1A) that each received three doses of 25 mg on study days 1, 14, and 28, and tissue was collected on day 42. While most CNS tissues exhibited <20% residual target RNA, five regions exhibited weaker target engagement (Fig. 1A–B). The three regions of the the basal ganglia—putamen, globus pallidus, and caudate—exhibited the lowest drug accumulation, with 3/4 animals having drug accumulation at the lower limit of quantification in caudate and in putamen (Fig. 1C). The thalamus exhibited somewhat higher drug accumulation and deeper target engagement, intermediate between basal ganglia and cortex. The cerebellum was unique in having modest target engagement despite higher drug accumulation, likely due to the predominance of cerebellar granule neurons, which exhibit comparatively weak ASO uptake 5 and activity. 6 Because the cerebellum was already examined for a PRNP ASO, 6 we chose not to revisit it here. We selected the thalamus, caudate, and putamen for further evaluation by single nucleus RNA sequencing (snRNA-seq).
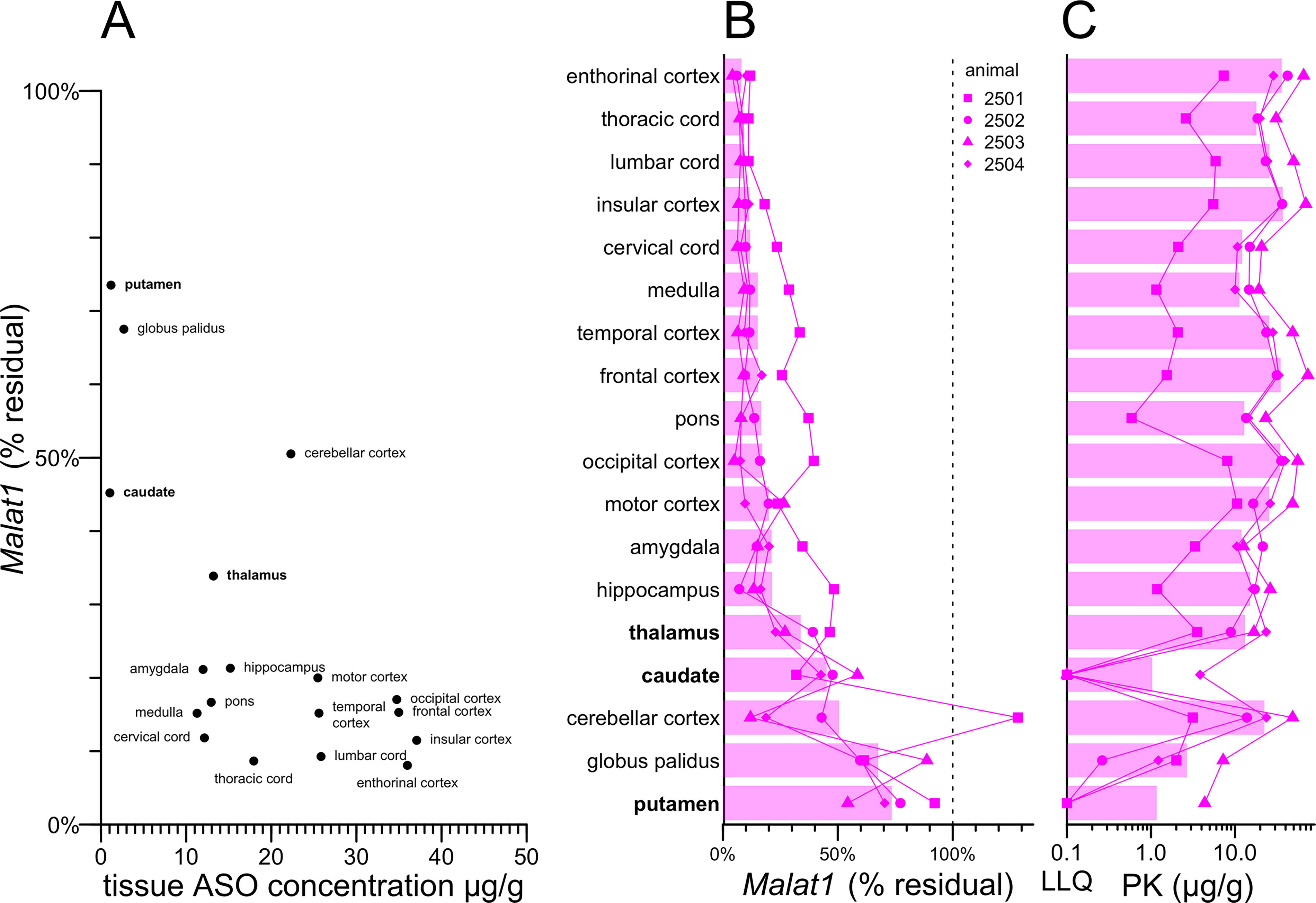
Pharmacokinetic and pharmacodynamic measurements across NHP CNS regions. All data in this figure are reproduced from Jafar-nejad & Powers et al.
Although all animals received the same dose of active drug, one animal (2501) exhibited more limited drug exposure and weaker knockdown across all CNS regions examined (Fig. 1B–C), while another (2503) exhibited the highest drug exposure and deepest knockdown across most regions (Fig. 1B–C). When the four animals were compared to one another across each of the CNS regions, higher drug exposure was significantly correlated with deeper target engagement (Supplementary Fig. S2).
Identification of cell types in NHP deep brain
We performed snRNA-seq according to our previously published protocol 6 on three brain regions each from two control NHP treated with aCSF and four NHP treated with Malat1 ASO (18 samples total). We obtained transcriptomes from 130,553 single nuclei (Supplementary Table S1). The 18 samples averaged 675 million reads each, mapping to 7,253 nuclei, with each nucleus having a median of 6,724 unique molecular identifiers (UMIs) and 2,846 unique genes (Supplementary Table S1). We used cell type-specific markers reported for mouse and marmoset 10 to identify cell types (Fig. 2 and Supplementary Fig. S3). Across all three brain regions, we identified fast-spiking interneurons (IN) expressing parvalbumin (PVALB) and midbrain (MB)-derived IN expressing OTX2, GATA3, and KIT. In the thalamus and caudate we also identified IN expressing somatostatin (SST). In thalamus, we found excitatory neurons expressing RXFP1, and in the caudate and putamen, DRD1- and DRD2-positive spiny projection neurons (SPNs), formerly known as medium spiny neurons. In addition to oligodendrocyte progenitor cells (OPCs) and mature oligodendrocytes, in each brain region we also identified committed oligodendrocyte precursors/newly formed oligodendrocytes (COP/NFOL), typified by expression of IFI44L and a higher level of ENPP6 expression than seen elsewhere. In the caudate, we also identified radial glia by their particularly high expression of VIM.
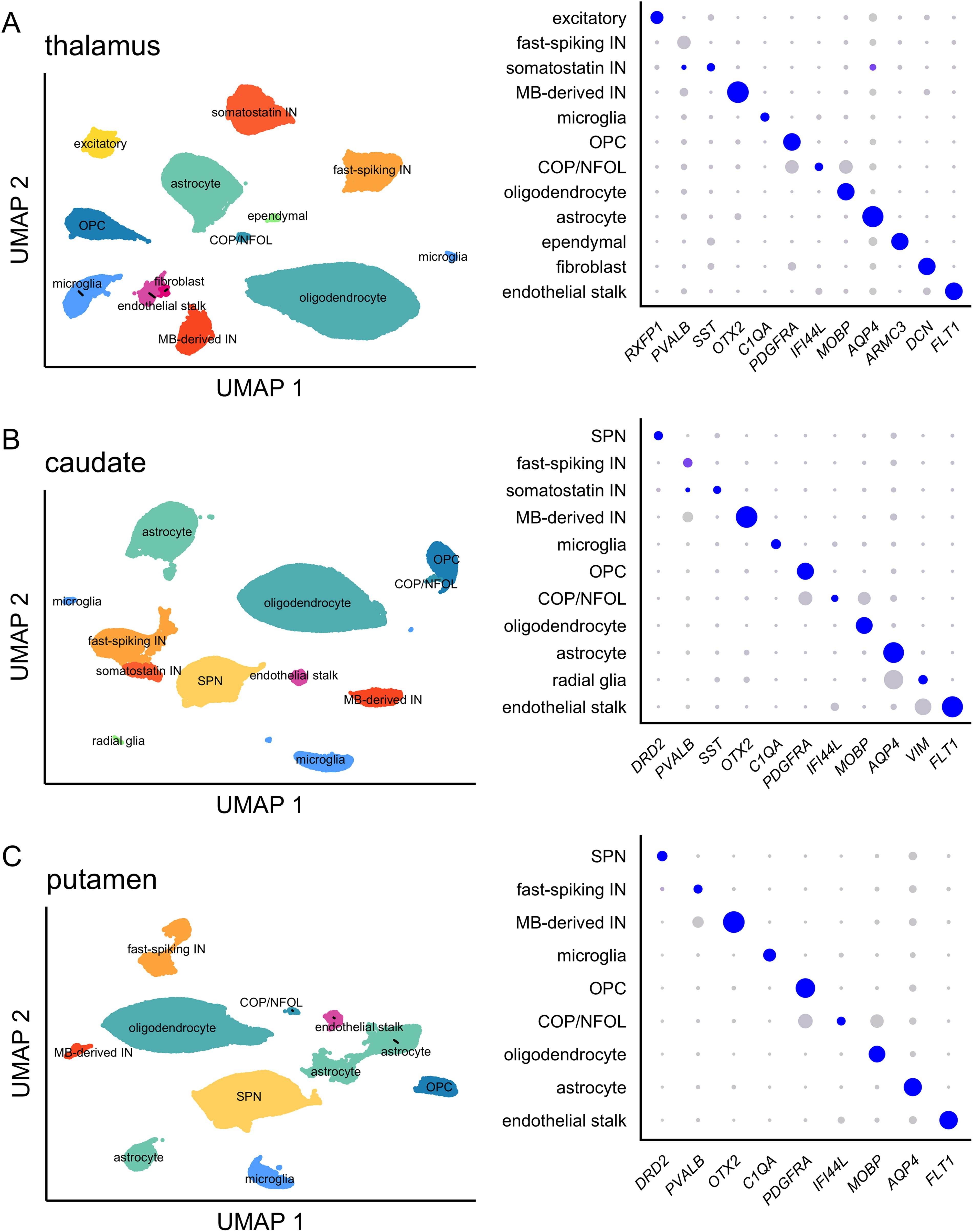
Cell types identified in NHP deep brain. Uniform manifold approximation and projection (UMAP) plots (left) and dotplots (right) for
While broadly distributed cell types such as oligodendrocytes and astrocytes were seen in roughly equal proportion across all animals, more regionally specific cell types were minimal or absent from some animals (Supplementary Fig. S4). For instance, in the thalamus, excitatory neurons were identified almost exclusively in 1/6 animals. Few or no SPNs were recovered in 2/6 animals in the caudate and 1/6 in the putamen. Dissection differences between animals likely account for these differences, as well as for the imperfect agreement between overall knockdown in single-cell sequencing (Fig. 3). For modeling the activity of the Malat1 ASO, we only included animal + brain region + cell type combinations with ≥10 cells observed, and we only fit models for brain region + cell type combinations where at least 1 control and 2 treated animals met this criterion.
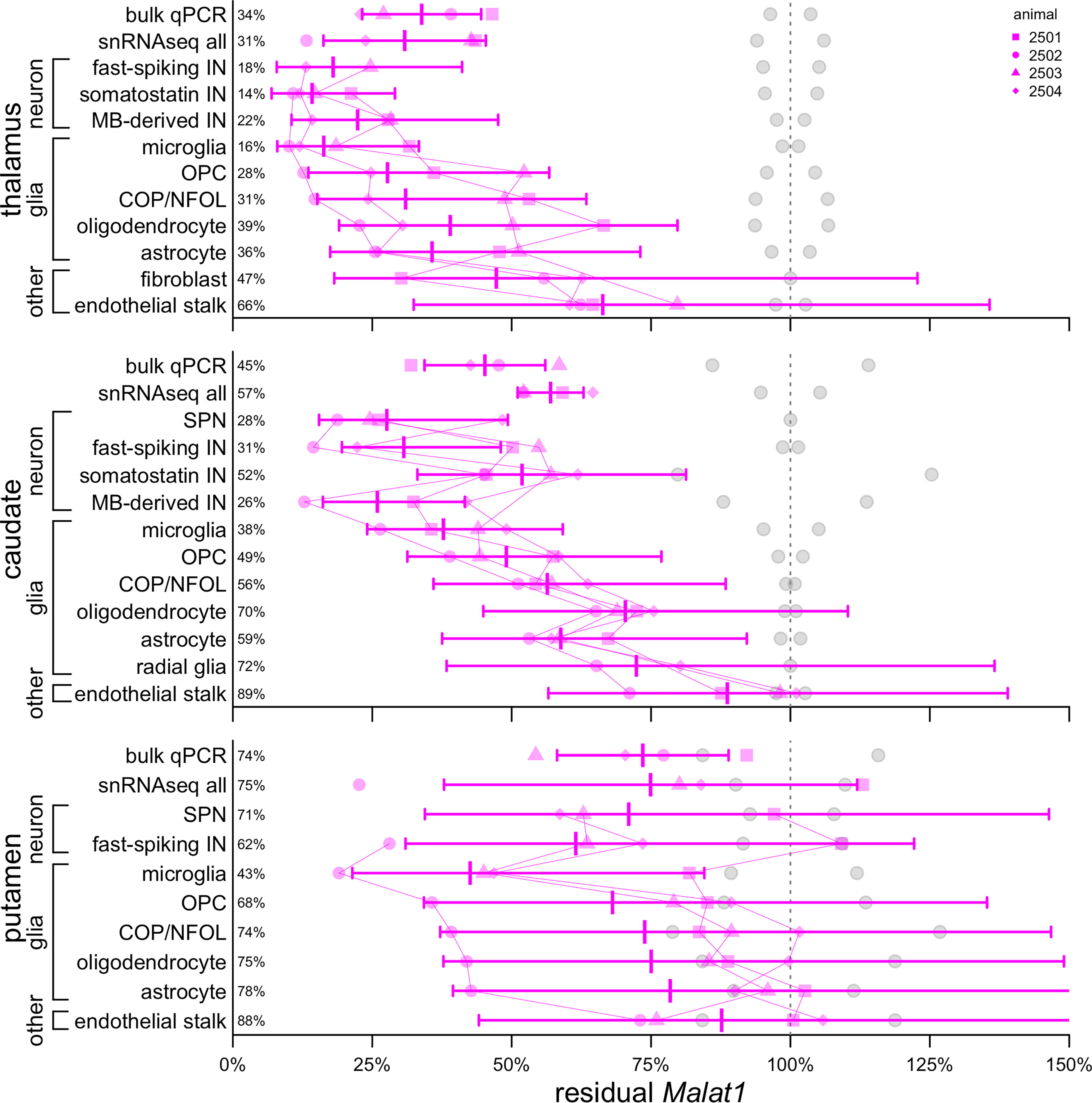
Target engagement by cell type in each NHP deep brain region. Gray points represent control animals (N = 2), magenta points represent ASO-treated animals (N = 4). Each animal’s data are represented by a unique shape and connected by thin lines between cell types. Bars represent means and 95% confidence intervals. Mean residual Malat1 is shown as a percentage at left. ASO, antisense oligonucleotides.
Residual Malat1 in bulk tissue averaged 31% in the thalamus, 57% in the caudate, and 75% in the putamen by snRNA-seq, and 34%, 45%, and 75%, respectively, by qPCR (Fig. 3). When broken down by cell types, confidence intervals were large and generally overlapping, as expected based on the small sample size. Just as observed previously in the cortex and cerebellum, 6 all neuronal subtypes identified here exhibited residual target RNA at least as low as the bulk tissue, with putamen SPNs (71% residual) being nearest to the bulk (75%), while some neuronal subtypes were far lower than bulk, for instance, 14% and 18% residual for somatostatin and fast-spiking IN in the thalamus, versus 31% in bulk tissue. Also consistent with previous results, 6 microglia exhibited relatively strong knockdown, deeper than any other glial cells, and in every brain region, endothelial stalk exhibited the weakest knockdown of any cell type. Overall, while some confidence intervals overlapped 100%, results were consistent with at least some knockdown in every detectable cell type in every brain region (Fig. 3).
Histograms of Malat1 UMIs for each region and cell type generally indicated that the entire distribution shifted left and remained unimodal (Supplementary Fig. S5). Only a few cell types in the putamen exhibited bimodal distributions of Malat1 UMIs after ASO treatment, but this appeared to be due to different levels of knockdown in different animals. Distributions were consistent with knockdown in every individual cell, again as expected based on prior work. 6
Discussion
We previously reported a single nucleus characterization of ASO activity in the mouse brain as well as in the cortex and cerebellum of NHP. 6 Here we extend these results by examining three NHP deep brain regions in animals that received an intrathecal ASO against Malat1. Despite more limited drug exposure in these regions and weaker bulk knockdown, all our key results closely mirror the previous findings in the cortex and cerebellum: 6 we observed ASO target engagement in every cell type detected; all neuronal subtypes have target engagement at least as deep as the bulk tissue; microglia exhibit especially deep knockdown, while endothelial stalk cells exhibit the weakest knockdown; and distributions of target RNA counts across individual nuclei are consistent with ASO activity in every single cell. This last property of ASOs is in contrast to a viral-vectored zinc finger repressor, for which we observed nearly complete elimination of the target transcript only in transduced cells. 14 Given the number of ASO molecules administered in a dose, it is expected that every cell should exhibit some activity. 6 An ASO shuttled across the blood–brain barrier using the transferrin receptor (TfR) exhibited particularly strong knockdown in vascular endothelial cells, 15 while this ASO, delivered intrathecally, exhibited the weakest activity in endothelial cells. It is possible that ASOs intrinsically have little uptake into endothelial cells and that activity for the shuttled ASO is due to escape during the process of transcytosis across the blood–brain barrier. For diseases where endothelial target engagement is critical, intrathecal delivery may be less preferred.
Our study has several limitations. We examined just 18 samples from 6 animals treated with 1 ASO. Due to our small sample sizes, many confidence intervals overlap each other or the 100% mark. The animals in this study received three doses of 25 mg ASO at 2-week intervals, which likely corresponds to greater drug exposure than achieved in clinical dosing regimens. The cynomolgus macaque brain is just 5% the mass of a human brain. We were unable to quantify knockdown in certain cell types because too few such cells were observed in some animals, likely due to dissection differences; the brain tissue from these animals has already been used for multiple prior studies, leaving imperfectly matched tissue punches for this work. We lack a single-cell pharmacokinetic method for quantifying drug accumulation by cell type, so we are unable to discern whether differences in target engagement are attributable to differences in drug uptake. For instance, in the cerebellum, IC50 is higher than in other brain regions (Supplementary Fig. S1), which may relate to poor drug uptake in granule cells, but we have not been able to assess this for deep brain regions.
The finding of broad target engagement across cell types even in deep brain regions with more limited drug exposure may support the relevance of intrathecal dosing to treatment of whole brain diseases or diseases with particularly prominent deep brain involvement. Ultimately, the hypothesis that intrathecal ASO delivery can adequately address such diseases in the larger human brain can only be tested clinically.
Footnotes
Author Disclosure Statement
P.J., J.C., and H.Z. are employees and shareholders of Ionis Pharmaceuticals. S.M.V. acknowledges speaking fees from Abbvie, Biogen, Eli Lilly, Illumina, Ultragenyx, and Voyager; consulting fees from Alnylam, Invitae, and Regeneron; research support from Eli Lilly, Gate Bio, Ionis, and Sangamo Therapeutics. E.V.M. has received speaking fees from Abbvie, Eli Lilly, Vertex, and Voyager; consulting fees from Alnylam, Deerfield, and Regeneron; and research support from Eli Lilly, Gate Bio, Ionis, and Sangamo Therapeutics. The other authors declare no conflicts of interest.
Funding Information
This work was funded by Ionis Pharmaceuticals. The authors thank Briana Nobel for technical assistance with monkey tissue collection.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
