Abstract
Time-lapse (TL) embryo monitoring is the latest technology that is proposed for embryo evaluation and selection for transfer. TL technology enables us to collect significantly more information about the in vitro development of the embryos that can be obtained through the daily-once evaluation under the light microscope. In addition, the embryos do not need to be removed from the culture environment for this. The extra morphokinetic information and the undisturbed culture conditions could both be beneficial for the cultured embryo cohort. Many morphokinetic parameters have been tested in relation to variety of laboratory (e.g. blastocyst development) and clinical (implantation and live-birth rate) outcomes. Most of these studies are retrospective in nature and suffer from methodological problems (heterogeneous patient population, culture conditions not standardized, and small sample size). Several groups attempted to build algorithms, however, have not yet been confirmed externally as attempts so far could not reproduce the expected predictive abilities. Therefore, these algorithms cannot be universally accepted. The latest algorithm proposed for embryo selection was developed based on data from 24 clinics using local stimulation and laboratory procedures. It groups embryos into five categories (KIDScore) based on in and out of range kinetic events. The algorithm was tested in subsets of patients using various fertilization methods or culture conditions and its predictive ability remained the same. The authors, therefore, feel comfortable to recommend it for routine use in any laboratory using TL technology. There is, however, still limited prospective, randomized trial data testing the algorithms. This article reviews TL technology, retrospective and prospective reports on various morphokinetic parameters, and the benefits and shortcomings of currently available algorithms.
Keywords
Introduction
The step limiting the success of in vitro fertilization (IVF) the most is implantation. A healthy embryo, a properly built-up endometrium and appropriate synchronization in between them are required for this step to proceed successfully. In a typical IVF cycle, a cohort of embryos are created in vitro and typically 1 or 2 are selected for transfer. Depending primarily on the age of the woman, on average, 5–40% of them implant and therefore around 1/3 of the cycles result in a pregnancy (European IVF-Monitoring Consortium (EIM), 2016).
It has been realized for some time that the daily-once evaluation of the actual morphology of the developing embryos provides us the limited information about their overall status and therefore does not allow proper selection for transfer. In order to improve our ability to identify the embryo(s) with the highest implantation potential, various methods (extended culture to the blastocyst stage (Glujovsky, 2016), metabolomics, proteomics, granulosa cell gene expression profiling, preimplantation genetic screening (PGS) (Montag et al., 2013)) have been tested. Among them, PGS seemed to be the most effective as the identification of the embryo with a normal, euploid chromosome content has been shown to improve pregnancy rate in younger, good prognosis patients (Dahdouh et al., 2015).
Time-lapse (TL) embryo monitoring is the latest tool that may aid the embryologist’s assessment of the developing embryos. TL technology allows us to continuously monitor embryonic development without the need to remove the embryos from the optimal culture conditions. The extra information obtained through TL monitoring gives us a more detailed knowledge about the kinetic and morphologic changes/abnormalities an embryo undergoes in vitro. The kinetic events can be precisely timed and these timings/intervals can be correlated with various stages of embryonic development, implantation, and live birth (Kovacs, 2014). Ultimately, the kinetic and morphologic parameters obtained through TL could be used to build algorithms that can help to choose the fittest embryo for transfer.
TL technology
There are various TL equipment in use (Primo Vision, Vitrolife AB, Sweden (PV) time-lapse system, embryoscope (ES), early embryo viability assessment (EEVA)) (Kovacs, 2014). They are all built around a similar concept. A TL unit is made up of a camera that takes a picture of the developing embryos at preset (10–20 min) intervals and is connected to a microscope system. This complex either has to be placed into a standard incubator (PV, EEVA) or is part of an incubator already (ES). Embryos in the TL system need to be identified and followed up individually. This can be achieved by culturing the embryos in individual microwells with no connection between the embryos (Embryoslide) or by the use of so called multiwell dishes (Primo Vision Culture Dish, EEVA dish) when each embryo is placed in individual microwells (9–16-well dishes) and they are cultured under a single drop of culture medium. This latter system allows individual embryo tracking but also provides the benefit of group culture via enhanced auto- and paracrine effects (Vajta et al., 2008).
The pictures captured by the camera are then processed by the appropriate software. This way a short film is created by connecting the pictures that can be rewound and fast-forwarded, and in the case of the PV or ES systems, embryos can be evaluated in several focal planes (Kovacs, 2014).
TL parameters
TL monitoring allows us to closely follow the embryos from fertilization up until the transfer. Events of the pronuclear phase, precise timing of cell divisions, duration and synchrony of the cell cycles, the sometimes transient changes of morphology (e.g. fragmentation), timing of compaction, blastocyst formation, and expansion and blastocyst dynamics can be precisely timed (Figures 1 and 2). In addition to the normal events of embryo development, abnormal cellular events can be detected as well that otherwise easily could be missed by relying on the daily-once observation. It has been shown that direct cleavage from 1 to 3 cells, multinucleation, and uneven blastomere size at the 2- to 4-cell stage are strong negative predictors of implantation (Meseguer et al., 2011).
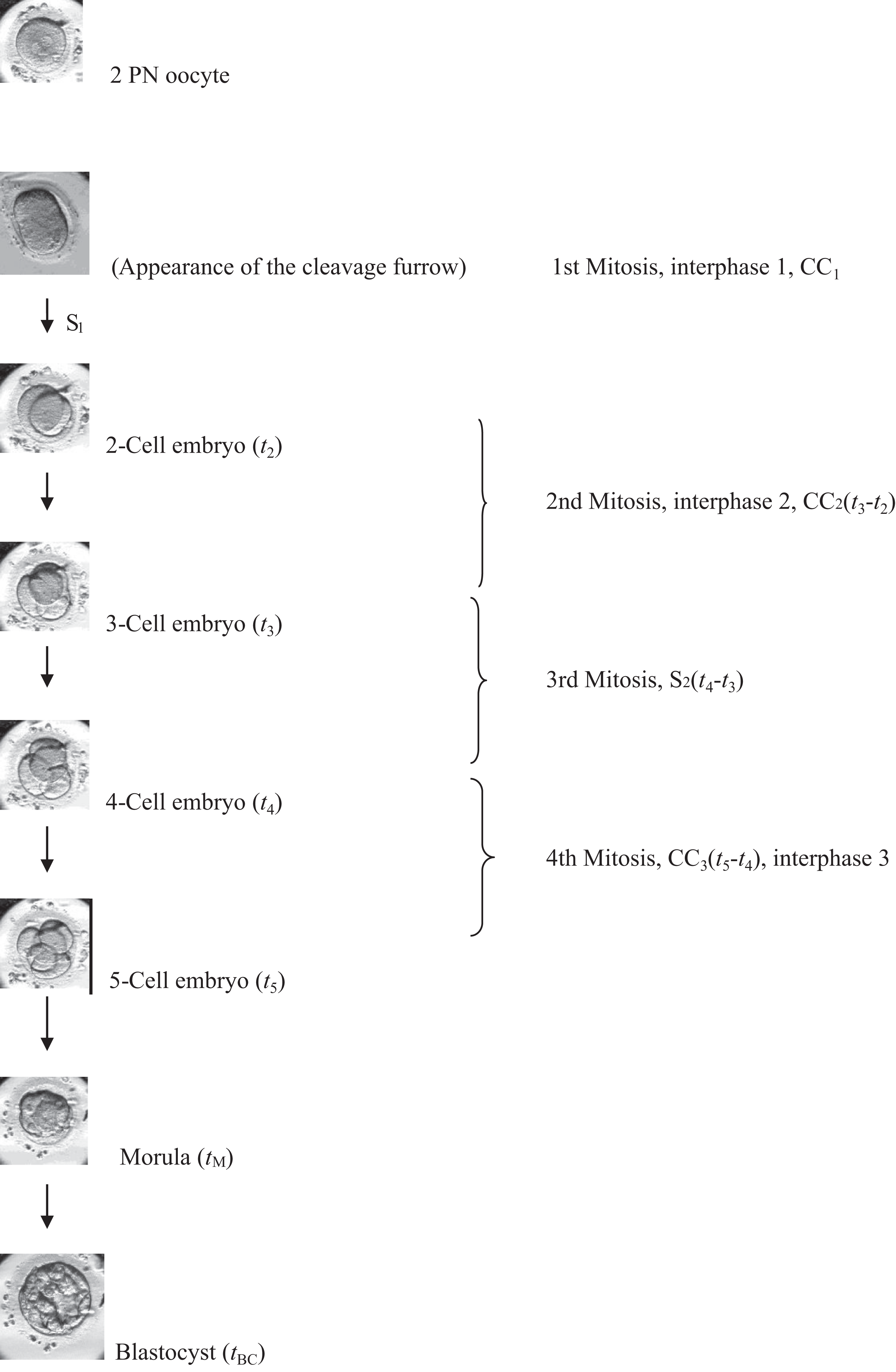
Embryo development from 2PN to blastocyst stage and the various terminology used in the different papers for certain developmental events (with permission from Reproductive Biology and Endocrinology: Kovacs (2014)).
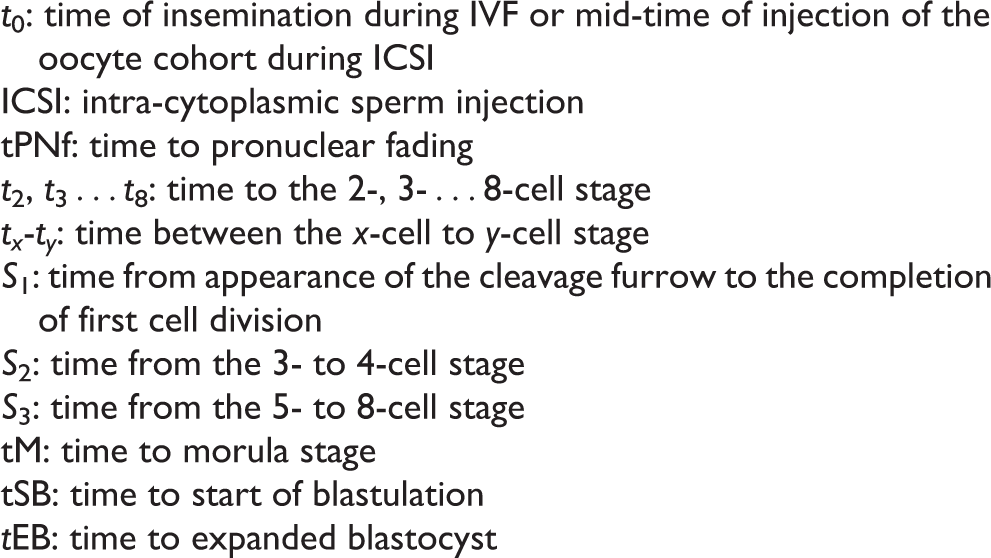
Definitions of kinetic TL parameters.
Heterogeneity in the markers identified as predictors of various outcomes
As TL technology became available, several groups have started to use it and started to correlate certain kinetic events with laboratory and clinical outcomes. These studies are mostly retrospective and rely on building databases of various kinetic markers that can subsequently be correlated with blastocyst formation, implantation, clinical pregnancy or live birth. In order to associate these parameters with a given clinical outcome, the analysis has to be limited to embryos with known implantation data (KID embryos). This is straightforward when a single embryo is transferred (SET); in cycles with a double embryo transfer (DET), only those can be considered for analysis when either both embryos implant or neither of them do.
When the outcomes of these studies are evaluated, their heterogeneity has to be considered as well. Some studies included fresh cycles using only own oocytes; others included cycles with fresh autologous or donated oocytes, while others included frozen oocytes or cryopreserved fertilized oocytes as well. The culture conditions are not uniform; different culture media are used by the various research groups, the amount of out-of-incubator handling is likely to differ, and the oxygen (O2) concentration is not standardized either. The day of transfer varies from day 2 to day 5 and there are reports in which embryos are not transferred just cultured and observed to a certain developmental stage. Obviously, with a shorter culture period, we cannot enjoy the full benefit of the TL technology, as less morphokinetic data is generated and the out-of-incubator handling of embryos is similar to the standard methods (for details of the studies, see Tables 1 –3.).
Results of RCTs using TL technology.
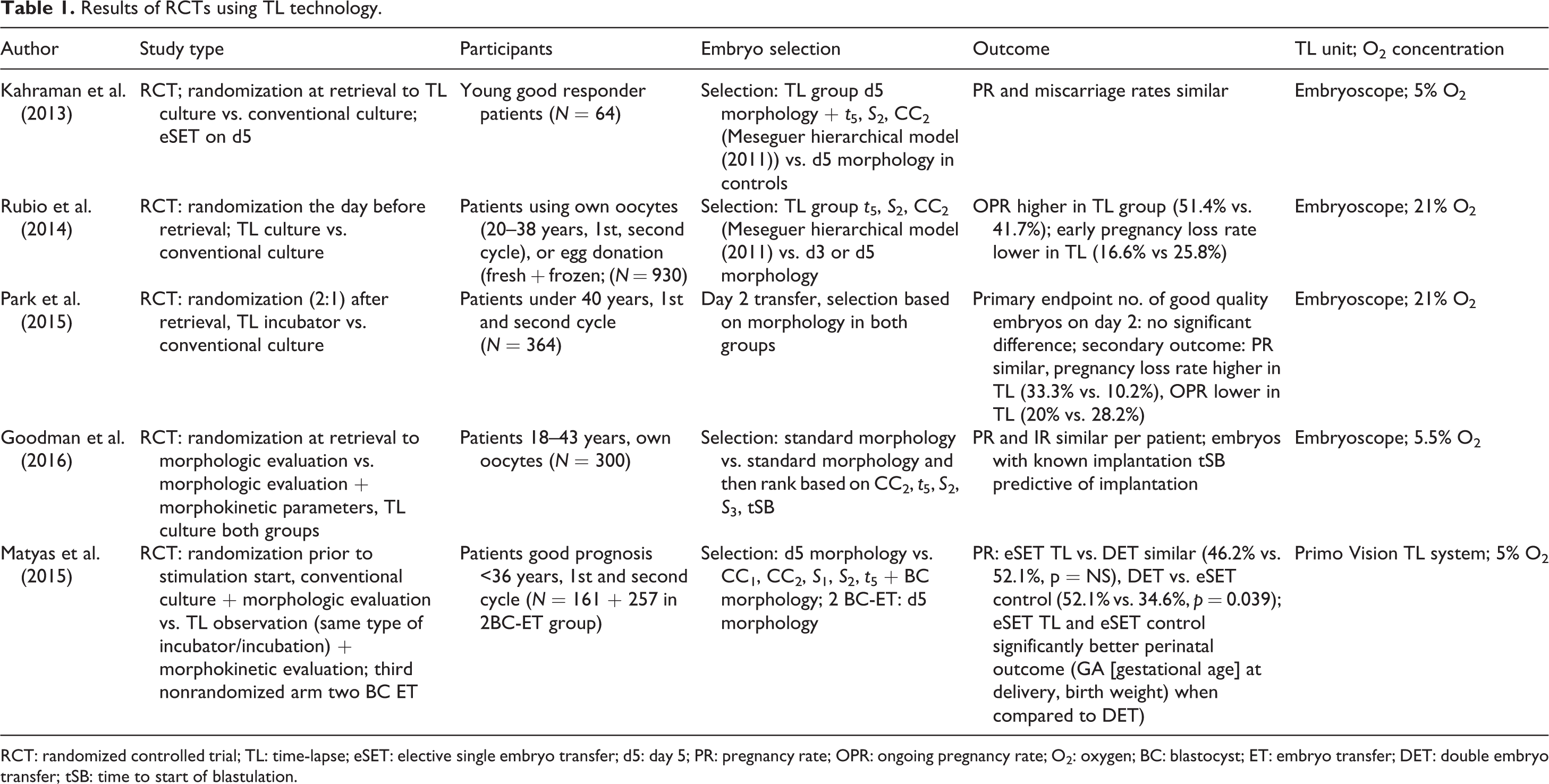
RCT: randomized controlled trial; TL: time-lapse; eSET: elective single embryo transfer; d5: day 5; PR: pregnancy rate; OPR: ongoing pregnancy rate; O2: oxygen; BC: blastocyst; ET: embryo transfer; DET: double embryo transfer; tSB: time to start of blastulation.
Results of prospective and retrospective cohort studies evaluating TL technology.
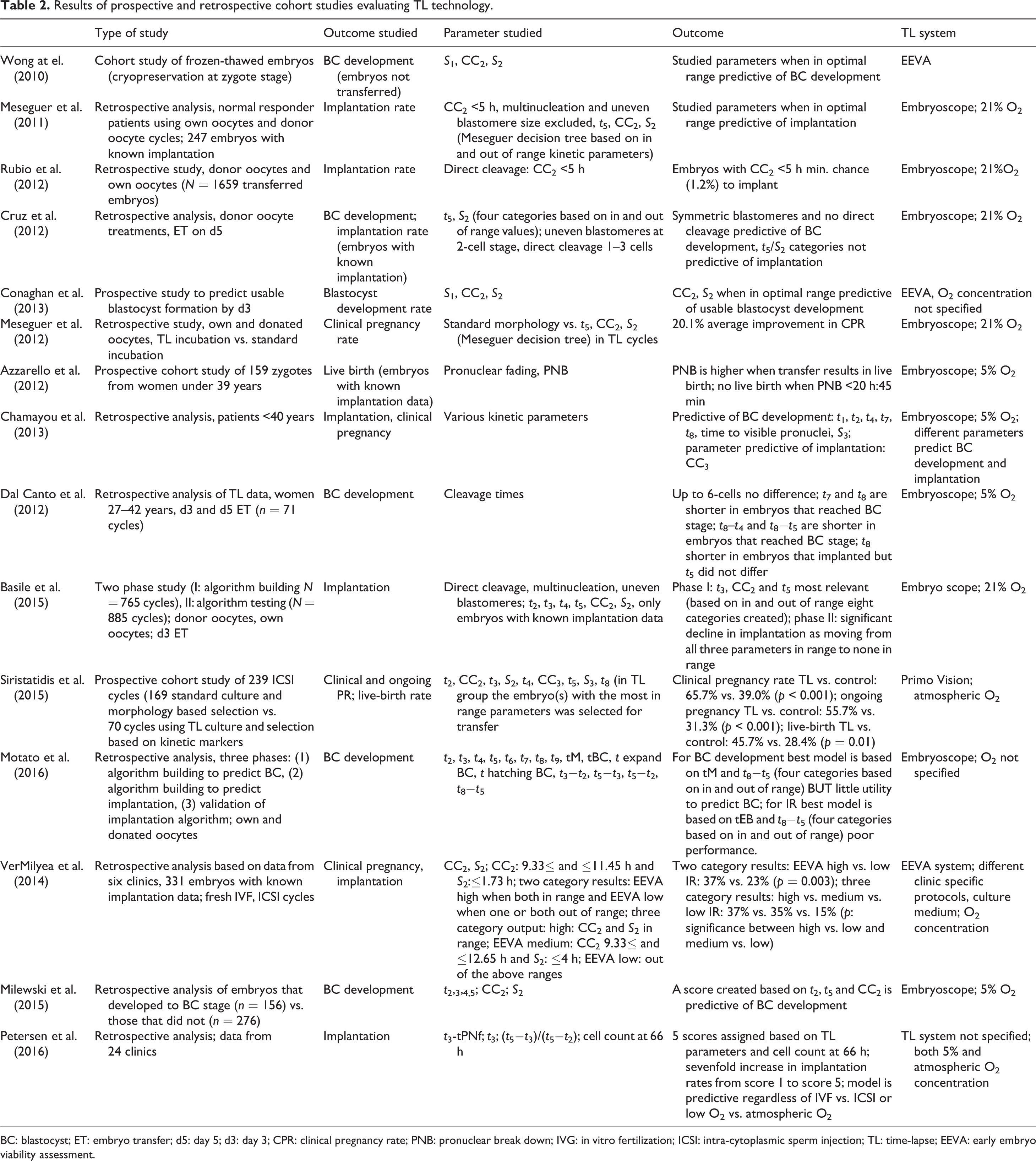
BC: blastocyst; ET: embryo transfer; d5: day 5; d3: day 3; CPR: clinical pregnancy rate; PNB: pronuclear break down; IVG: in vitro fertilization; ICSI: intra-cytoplasmic sperm injection; TL: time-lapse; EEVA: early embryo viability assessment.
Results of studies evaluation aneuploidy and TL technology.
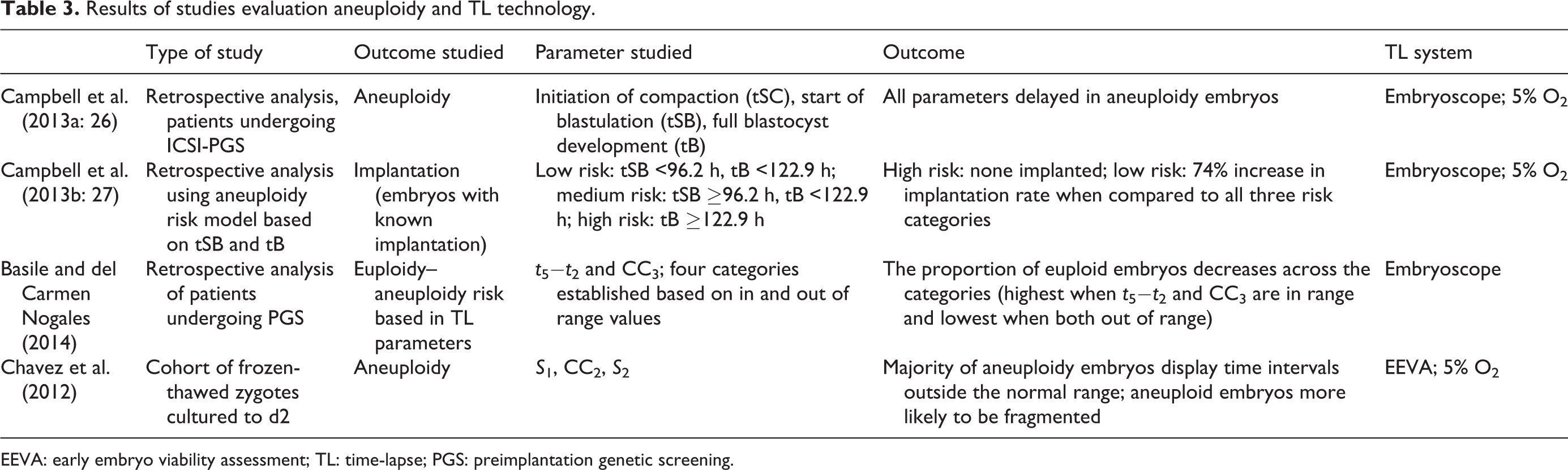
EEVA: early embryo viability assessment; TL: time-lapse; PGS: preimplantation genetic screening.
Finally, the patient populations studied are rather heterogeneous as well. The significance of this may be limited though. Ultimately, we wish to differentiate healthy, ready to implant embryos from the unhealthy cohort. Healthy embryos (when cultured under similar conditions) should be more likely to follow a strict, normal kinetic developmental pattern regardless of parental characteristics.
Due to the heterogeneity of the studies, it should not be surprising that a wide variety of kinetic markers have been associated with the various clinical outcomes (Tables 1–3). Sometimes, even the same group identified various markers as “predictive” when a different outcome was studied (e.g. blastocyst development vs. implantation). One of the earliest studies in which an algorithm was proposed to identify the embryos with a better chance to implant incorporated t 5, CC2, and S 2 into the hierarchical model and showed that those embryos that had kinetic parameters in the optimal ranges were more likely to implant (Meseguer et al., 2011). In a later study, Cruz et al. (2012) categorized embryos based on in or out of range t 5 and S 2 values but found no significant difference in the implantation rates across the four kinetic categories of embryos. Basile et al. (2015) proposed a different hierarchical model in which S 2 was replaced by time to the 3-cell stage (t 3). This final hierarchical model was built using in and out of range t 3, CC2, and t 5 parameters to predict implantation. These studies were published by the same group and were carried out in the same chain of clinics. The patient population involved is rather heterogeneous as fresh autologous and donated oocytes as well as the use of frozen oocytes were included to build the database. The sample size of the studies (from a few hundred to the thousands) is likely to influence the predictive value of the various studied markers. Very importantly though, these authors have identified markers that are associated with minimal chance of success and these parameters are therefore proposed as deselection markers. Rubio et al. (2012) have shown that embryos with very short interval between the 1- and 3-cell stage (<5 h) have a minimal chance to implant. Furthermore, it has been shown that uneven blastomere size at the 2-cell stage and multinucletaion at the 4-cell stage are strong negative predictors of implantation (Meseguer et al., 2011). These deselection markers should be part of an algorithm built.
The parameters identified as significant are influenced by the technology itself. The EEVA system using dark field technology and an automated software analysis can reliably follow embryos up to the second and third cell cycle and therefore their algorithm depends on early markers (Conaghan et al., 2013; Wong et al., 2010). Wong et al. (2010), using frozen-thawed fertilized oocytes, found S 1, CC2, and S 2 to be predictive of blastocyst formation. More relevant clinical outcomes could not be studied, as the embryos were not transferred. Later on, Conaghan et al. (2013) using the EEVA system have confirmed the improved ability to predict development to the blastocyst stage using the same early kinetic markers. Clinical outcome was not discussed in this study either.
There are groups that evaluated morphokinetic markers to predict embryonic aneuploidy. Campbell et al. (2013a, 2013b) published two papers on this topic. In their first study, they showed that late kinetic events (time to start of compaction, time to start of blastocyst formation, and time to blastocyst formation) were all delayed in aneuploid embryos. Early markers, however, failed to correlate with genetic health (Campbell et al., 2013a). In a subsequent study, they tested the predictive ability of time intervals (time to start of blastulation (96.2 h) and time to full blastocyst formation (122.9 h)) and proposed low-, medium-, and high-risk categories for aneuploidy based on them (Campbell et al., 2013b). Chavez et al. (2012) studied embryos obtained from frozen-thawed fertilized oocytes and found that euploid embryos followed a much tighter early development pattern as S 1, S 2, and CC2 parameters were more homogeneous when compared to aneuploid embryos. Basile and del Carmen Nogales (2014) found t 5–t 2 and t 5–t 3 as the most reliable parameters to predict euploidy and built a model using in and out of ranges of these parameters to predict healthy chromosome content.
Algorithms predicting implantation
Once those morphokinetic markers that correlate with a given clinical outcome have been identified they need to be built into an algorithm that could improve our ability to identify the embryos with a higher chance to implant and result in a pregnancy/live birth (Table 4).
TL algorithms predicting clinical outcome (implantation or pregnancy rate).
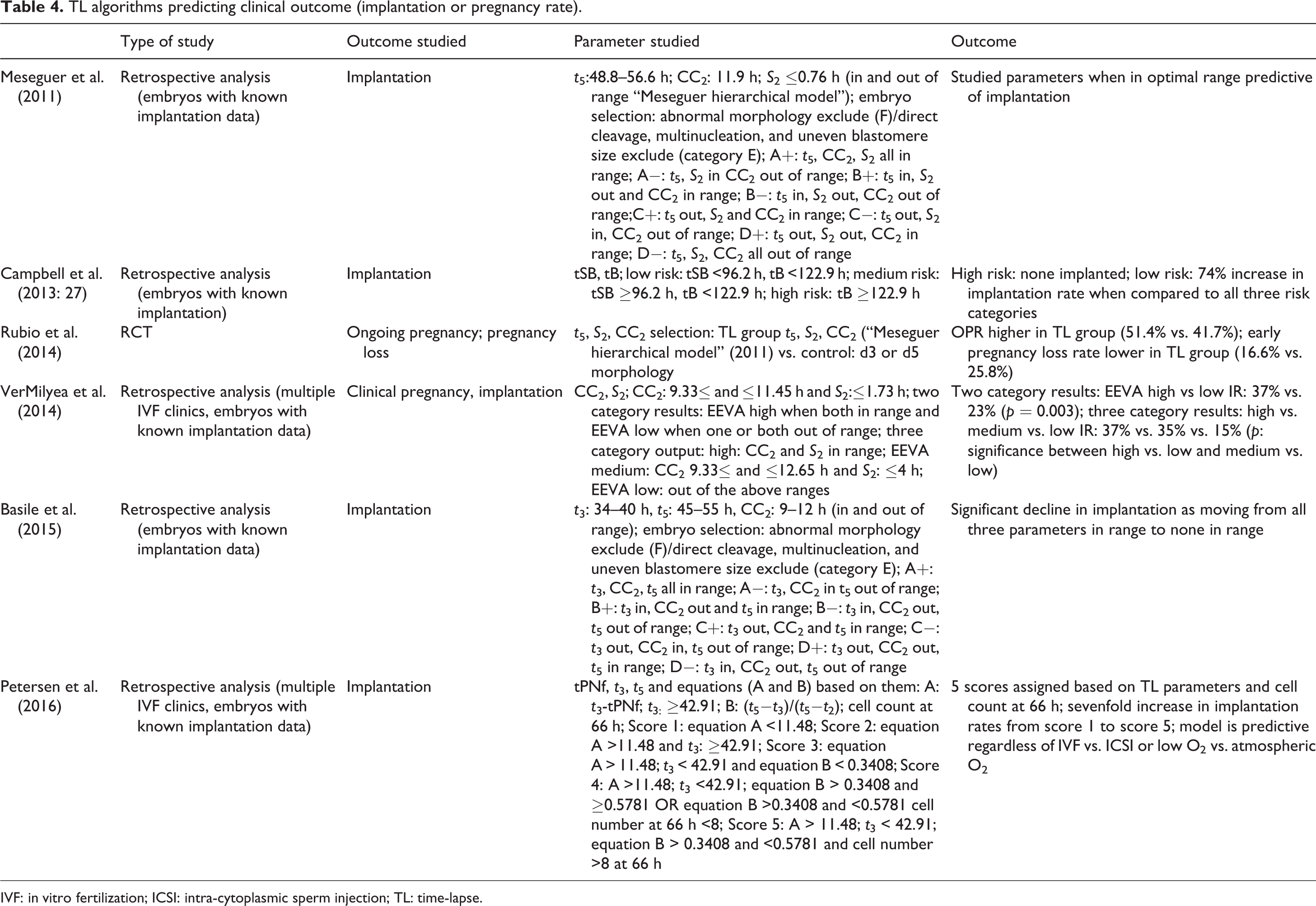
IVF: in vitro fertilization; ICSI: intra-cytoplasmic sperm injection; TL: time-lapse.
The first algorithm to predict implantation was published by Meseguer et al. (2011). 247 embryos with KID were available to build the algorithm. According to their description, those embryos with clear morphologic abnormalities should be discarded and should not be considered for transfer. Those embryos with direct cleavage 1–3 cells, multinucleation, and uneven blastomere size at the 2- to 4-cell stage are not recommended for transfer either (deselection markers). The remaining embryos were split into eight kinetic categories (A+ (highest chance to implant), A−, B+, B−, C+, C−, D+, and D− (lowest chance to implant)) first based on in and out of range of t 5 (48.8–56.6 h), then in and out of range of S 2 (≤0.76 h), and finally based on in and out of range of CC2 (≤11.9 h). About 66% of the embryos in the best, A+ category implanted, while only 15% of those in the lowest D− category implanted successfully. Only 8% of those embryos that were deselected based on exclusion criteria implanted. As mentioned earlier, Campbell et al. published two papers assessing late TL kinetic markers and aneuploidy and implantation (Campbell et al., 2013a, 2013b). In their first study, time to start of blastulation and time to full blastocyst development emerged as predictive for euploidy. In a subsequent study, a retrospective analysis based on 69 cycles, they showed that none of those embryos considered high risk for aneuploidy based on time to full blastocyst formation ≥122.9 h implanted, while 72.7% of those identified as low risk (time to start of blastulation <96.2 h and time to full blastocyst formation <122.9 h) implanted (Campbell et al., 2013b). Basile et al. (2015) proposed a different algorithm based on retrospective analysis of morphokinetic data of 1137 embryos with known implantation outcome. Embryos considered nonviable based on morphology were discarded (n = 55) and embryos showing any of the deselection criteria according to the Meseguer algorithm (n = 197) were suggested to be excluded too. The remaining 885 embryos were first split based on in and out of range of t 3 (34–40 h), then based on in and out of range of CC2 (9–12 h), and finally based on in and out of range of t 5 (45–55 h) (A+ (highest implantation potential, A−, B+, B−, C+, C−, D+, D− (lowest implantation potential)). About 32% of A+ embryos implanted, while only 19% of those categorized as D−. The implantation rate was 17% among those embryos identified based on deselection criteria (Basile et al., 2015). Motato et al. (2016) proposed a hierarchical model to predict blastocyst implantation based on retrospective analysis of 832 blastocysts with KID. Two markers were included in the algorithm; time to expanded blastocyst ≤112.9 h versus >113 h and t 8−5: ≤5.67 versus ≥5.68 h (categories A, B, C, and D). Implantation rates decreased across the four groups from 72.7% in category A to 39.7% in category D. It is interesting to point out that this study had two parts and the other part in which markers to predict blastocyst formation were searched identified different markers as predictive of blastocyst formation. Those markers, however, were not able to predict implantation (Motato et al., 2016).
It should be mentioned here that while several groups consider multinucleation as a negative predictor of implantation/pregnancy, there are reports that disagree with this. Balakier et al. (2016), in a retrospective analysis, evaluated the impact of multinucleation on embryonic euploidy and implantation/pregnancy rates. They found that multinucleation was more frequent at the 2-cell stage (43.2%) when compared to the 4-cell stage (15%), suggesting that there are mechanisms that can correct this abnormality. In addition, they reported similar rates of multinucleation in euploid versus aneuploid embryos (40.8% vs. 46.7%). Finally, of those embryos showing multinucleation at the 2-cell stage, 61 were transferred and a 45.9% of clinical pregnancy rate obtained with them.
VerMilyea et al. (2014) used the EEVA prediction model developed by Wong et al. (2010) and Conaghan et al. (2013) to study its predictive ability for implantation and clinical pregnancy. TL recordings were obtained from six different clinics that used clinic specific stimulation and culture protocols. Embryos were selected based on morphology for transfer and the TL recordings of those embryos where implantation was known (n = 331) were analyzed. A two- (high and low implantation potential) and a three-category (high, medium, and low implantation potential) evaluation was tested based on CC2 and S 2 (CC2: 9.33≤ and ≤11.45 h and S 2:≤1.73 h; 2-category results: EEVA high when both in range and EEVA low when one or both out of range vs. 3-category results: EEVA high: CC2 and S 2 in range; EEVA medium: CC2 9.33≤ and ≤12.65 h and s 2: ≤4 h; EEVA low: out of the above ranges). Both the two- and three-category models were predictive of implantation and clinical pregnancy.
The latest model was published by Petersen et al. (2016). This algorithm is based on five kinetic and one morphological event. The algorithm was built based on retrospectively collected data from 24 clinics using local embryology practices. The algorithm assigns five scores to the embryos and a sevenfold increase in implantation rate can be seen starting from the lowest score to the highest. This score, the KIDScore, is described in detail in the next section.
External validation
There are many potential confounders when morphokinetic parameters and clinical outcome are correlated. Patient characteristics, stimulation methods, culture conditions, O2 concentration, and so on can all influence embryo development and as a result the kinetic markers. Therefore, when one group recommends an algorithm as a model to predict implantation or pregnancy, a different group in a different patient population using different culture conditions may not find the results helpful. In order to accept and introduce an algorithm in routine practice, it has to be tested in a different clinical setting.
Freour et al. (2015) studied the Meseguer hierarchical model (Meseguer et al., 2011) in a retrospective analysis. About 2240 embryos obtained from 450 couples were considered for the analysis. There were no exclusion criteria, any couple undergoing intra-cytoplasmic sperm injection (ICSI) treatment was considered eligible. Embryos were cultured in ES under a reduced (5%) O2 environment and transfers were performed at cleavage as well as blastocyst stages. Embryos were selected primarily based on morphology and then based on kinetic analysis. 528 embryos with KID were analyzed. The authors observed a heterogeneous distribution of implantation rates across the kinetic categories. The correlation coefficients were lower than in the original study by Meseguer et al. (2011), were modest only, and were not significant. Based on their results, the authors did not recommend routine application of the original model but proposed each clinic to develop their own predictive algorithm.
Another study, conducted by Kirkegaard et al. (2014), tested the Conaghan model. The Conaghan model (Conaghan et al., 2013) predicts blastocyst formation using two early kinetic markers, CC2 and S 2. Those cleavage stage embryos that follow in range kinetics are more likely to turn into usable blastocysts. Kirkegaard et al. tested whether the same model is able to predict implantation. TL kinetic data was obtained from seven clinics. Embryos were cultured in ES up until the transfer on day 2 or 3. Selection for transfer was based on morphology. The model was applied to those 1519 embryos that had KID. In the overall cohort, the implantation rate was 17.4%. Among embryos that were considered usable by the model the implantation rate was 22.7%, while among those embryos that were considered nonusable the implantation rate was 14.2%. More importantly, half of the embryos that eventually resulted in a pregnancy were identified as nonusable by the model and potentially would have been discarded.
A different approach for “external validation” is to pull data from different clinics using different daily practices.
This approach was used by Petersen et al. (2016) and they proposed a different algorithm to predict implantation using day 3 embryos. The model was built on retrospective analysis of TL imaging of 3275 embryos with known implantation data. Data was obtained from 24 clinics using heterogeneous patient populations, culture conditions, and embryo handling. Five kinetic events and one morphological event are included in the algorithm. The first parameter is t 3-tPNf (pronuclear fading). Optimally, this would be over 11.48 h (if shorter the embryo is given score 1). When it is >11.48 h, we proceed to the second checkpoint, t 3 which ideally would be less than 42.91 h (if longer the embryo is given score 2). When t 3 is <42.91 h, a third parameter is calculated by the following formula: (t 5 − t 3)/(t 5 − t 2). If the equation is <0.3408, the embryo is given score 3, if it is ≥0.5781, it is given a score of 4. If it is between 0.3408 and 0.5781 and the embryo fails to reach the 8-cell stage by 66 h and it is also given a score of 4. For those embryos that reach the 8 or more cell stage by 66 h a score of 5 is assigned. Across the five score groups, the implantation rate increased sevenfold from 5.18% (score 1) to 36.17% (score 5). The strengths of this scoring system (KIDScore) over the other algorithms have to be emphasized. The algorithm was built based on TL recordings over 3000 embryos with known implantation outcome. The data was obtained from 24 clinics treating patients with a wide range of problems and using different culture conditions. The model predicts a clinically relevant outcome rather than surrogate markers of success. The algorithm was tested in subsets of cases based on method of fertilization and O2 concentration and very similar values were obtained when compared to the entire data set. This suggests that the algorithm works well both in low and ambient O2 concentration and both with IVF and ICSI fertilization.
Randomized controlled trials using algorithm-based embryo selection
Ultimately, the true predictive value of the various markers or their combinations (algorithms) would be tested prospectively, ideally in a randomized controlled trial (RCT) when the algorithm is compared to the current standard, daily-once morphological assessment. Unfortunately, there is still limited randomized control trial data that assesses the discriminating ability of the various TL markers/algorithms prospectively. In order to test the benefit of the TL parameters alone, test and control embryos should be cultured under similar conditions. Otherwise, if a difference is noted between the randomized groups, we will not know what to contribute the benefit to if the culture conditions also differ. It is also true that if the culture conditions are identical, then the selection of the embryos in the TL group has to rely solely on predefined TL parameters/algorithm to prove the superiority of TL algorithm-based selection over standard morphological evaluation. We have to keep this in mind when RCT data is analyzed.
Kahraman et al. (2013) tested the Meseguer hierarchical model (based on t 5, S 2, CC2) in a small RCT among good prognosis patients. In the TL group, embryos were cultured in ES, while in the control group, embryos were cultured in conventional incubators. The authors did not find a difference in clinical outcome between the groups. In a much larger study, Rubio et al. (2014) randomly assigned patients to conventional culture and embryo selection for transfer based on day 3 and/or day 5 morphology versus culture in ES and selection for transfer based on t 5, S 2, and CC2 (Meseguer hierarchical model). Patients aged 20–38 years were eligible to participate. The use of autologous and fresh or frozen donated oocytes was allowed. The day of transfer was not standardized as both day 3 and day 5 transfers were allowed. Unfortunately, some of the patients decided to follow the other protocol (not the assigned) after randomization. The authors found lower miscarriage rate and significantly improved ongoing pregnancy rate in the TL arm. Park et al. (2015) randomly assigned patients to embryo culture in a TL system (ES) versus in a conventional incubator. They selected embryos for transfer based on day 2 morphology. Their aim was to see whether the proportion of good quality embryos was higher if embryos were cultured in the TL unit. They found no difference in the proportion of the top quality day 2 embryos with the two incubation methods. This should not necessarily be surprising as the culture conditions differed and the duration of culture was very short. If we accept that by limiting the out-of-incubator handling of the embryos we can improve their growth potential, then in this study, this benefit was not utilized as the embryos had to be removed for fertilization check and day 2 assessment in both groups. Goodman et al. (2016) randomly assigned patients to morphologic assessment versus assessment based on morphology plus TL parameters. Embryos in both groups were cultured in TL units. In the TL group, embryos were primarily selected for the transfer based on morphology and TL parameters were only considered when similar quality embryos were available for transfer. Clinical outcome was similar in the two groups. In the study by Matyas et al. (2015), young and good prognosis patients were randomly assigned to single blastocyst transfer (1) selected based on day 5 morphology versus (2) based on a predefined TL score (algorithm) comprising both kinetic and morphologic parameters. A third group of nonrandomized patients undergoing double blastocyst transfer (DET) was included as well. Clinical and perinatal outcomes were compared. Embryos in all groups were cultured under similar conditions and the amount of out-of-incubator handling was identical too. The pregnancy rate was higher in the DET group when compared to the SET group evaluated using standard morphology; however, it was similar in the TL-selected SET group. The perinatal outcome was significantly better in the SET groups when compared to DET. Based on the results, it was concluded that one can successfully compensate for the fewer embryos transferred if the selection is based on a complex morphokinetic TL score and with this approach the perinatal outcome can be improved (Matyas et al., 2015).
Conclusions
TL technology is relatively new but it is being introduced in everyday embryology laboratory use for multiple reasons (undisturbed culture, detailed morphokinetic data, quality control, ease of workload, and improved documentation). As data has been accumulated, numerous papers were published primarily using retrospective data analysis to test the predictive abilities of different morphokinetic events. As a logical next step, algorithms built based on multiple morphokinetic parameters were proposed. The initial optimism regarding the benefits of these hierarchical models was overshadowed by the lack of ability to externally validate them. It was realized that there are multiple factors that could affect a model’s ability to predict a clinical outcome. Most recently, a new algorithm was introduced that was built based on data collected from 24 clinics using different laboratory technologies in a heterogeneous patient population. The model seems to predict implantation well as from the lowest to the highest score a sevenfold increase in implantation rate can be seen. For the first time, it seems that we have a model that is ready for introduction into routine daily care. This is particularly helpful for smaller clinics performing fewer cycles where it could take a very long time to collect enough data to build their own model especially if different algorithms have to be built for different clinical settings. Eventually, the model by Petersen et al. will also need to be tested prospectively.
It seems that there are competing technologies for the selection of the embryo with the highest chance to implant. It may be time for a paradigm shift, and rather than testing the methods against each other, we should combine them to explore their full benefit. The noninvasive TL technology could be used to “prescreen” embryos and the more expensive, invasive PGS could be applied to a smaller set of embryos to reduce costs and intervention. They together could improve IVF success in a patient-friendly way.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
