Abstract
Introduction
Mutations in the filaggrin (FLG) gene are known to cause ichthyosis vulgaris.
Methods
We used whole-genome sequencing (WGS) technology to investigate the genetic causes of rare and complex inherited diseases including rheumatoid arthritis, ichthyosis, and congenital fibrosis of the extraocular muscles type 1 (CFEOM1) in a Chinese family. WGS was performed in four topics, and the identified candidate mutations were further verified through Sanger sequencing.
Results
We identified a mutation in FLG gene (g.152280098 C>A, p.E2422∗) that may be associated with ichthyosis and arthritis. Moreover, a mutation in KIF21A (g.39726207 G>A, p.R954 W) was also determined in affected members as the cause of CFEOM1. The gene interaction network demonstrated an interesting correlation between FLG and genes associated with arthritis and ichthyosis. Functional enrichment analysis of these interacting genes revealed several possible pathways that might be linked to arthritis and ichthyosis.
Conclusion
In general, we confirmed a loss of function mutation in the FLG gene associated with ichthyosis vulgaris and rheumatoid arthritis in this family.
Keywords
Introduction
Whole-genome sequencing (WGS) can provide a comprehensive and detailed investigation of human genetic variations, allowing the investigation of coding and noncoding regions of the genome. Increasingly, WGS studies have provided the strongest evidence that rare genetic variants can have large cumulative effects on human diseases. 1 Family based studies, especially for single gene inherited diseases, often demonstrate the transmission of gene variants from parents to offspring and represent an implicit enrichment strategy for identifying rare gene variants. 2
Rheumatoid arthritis (RA) is a chronic autoimmune inflammatory disease characterized by pannus formation, leukocyte infiltration, and angiogenesis. 3 In clinical praxis, two major types of autoantibodies, which are the rheumatoid factor (RF) and anti-cyclic citrullinated peptide (anti-CCP) antibodies, have been widely used. RA is a common complex genetic disease. The application of high-throughput sequencing to the study of the genetic bases of complex diseases, such as RA and osteoarthritis (OA), has expanded the amount of “omics” data to multiple molecular levels, from the detailed sequencing of the genome to epigenetic markers, such as DNA methylation and microRNAs (miRNAs), transcriptomics and metabolomics.4,5 A larger number of genes, including cytokine gene (interleukin-13 and interleukin-17), are believed to be associated with RA.6,7 Interleukin-13 (IL-13) is a pleiotropic cytokine, secreted by activated T cells, that can affect vessel formation, and an important component of the RA synovial tissue pannus. 8 Interleukin-17(IL-17) may be a key activator of T cell-driven inflammation and thus may contribute to the pathogenesis of RA. 9 Serum IL-13 and IL-17 values increased in RA have been reported in many works.3,10 With the discovery of a number of T cell subsets and their potential plasticity during disease, it became clear that the T cell behavior in autoimmune diseases is much more complex than the first Th1/Th2 concept. 11
Ichthyosis vulgaris (IV; OMIM #146700) and recessive X-linked ichthyosis have been classified as common ichthyoses. 12 IV is one of the most common monogenic skin disorders. The discovery, 13 in 2006, that loss-of-function mutations in the filaggrin gene (FLG, GenBank accession number NM-002016) are the cause of IV and are also powerful genetic risk factors for atopic dermatitis (AD) marked a significant breakthrough in the understanding of eczema pathogenesis. 14 FLG is located on chromosome 1q21 within a cluster of genes that constitute the epidermal differentiation complex and consists of three exons. Exons1 (15 bp) and 2 (159 bp) are small, whereas exon 3 (12,573 bp) encodes nearly the entire profilaggrin protein. 15 Filaggrin, which is processed from profilaggrin, is an abundant protein found in the outer epidermis that is essential for the terminal differentiation of keratinocytes and the formation of an effective barrier against water loss and the invasion of pathogens, allergens, and irritants. 16 Studies of the FLG null mutations have identified a number of significant associations with diseases, including AD, atopic asthma, allergic rhinitis, peanut allergy, and psoriasis vulgaris.17,18 To date, approximately 49 truncating mutations, including the two initially reported mutations (p.R501*and p.2282del4), have been described, and the incidence of these mutations varies considerably among different ethnic groups and populations. 19
Congenital fibrosis of the extraocular muscles type 1 (CFEOM) is a heterogeneous group of disorders that may be associated with other anomalies 20 and is characterized clinically by non-progressive restrictive ophthalmoplegia, with or without ptosis. According to the clinical features and genetic components, CFEOM can currently be classified into three primary categories. CFEOM1, the classic form of CFEOM, is an autosomal dominant trait caused by mutations in the KIF21A gene (OMIM 608283) on chromosome 12q12. 21 CFEOM2 is caused by the mutations in the ARIX gene (OMIM 602753) on chromosome 11q13.4. However, the clinical manifestations of patients in families with CFEOM3 are similar to those with CFEOM1, 22 where CFEOM3 can be diagnosed even if no patients exhibit the classical clinical signs, and KIF21A mutations can also be associated with CFEOM3.
In our research, we identified a patient with the primary complaints of ankle pain and scaly skin, with varying accompanying symptoms of eczema and ptosis. To date, the presence of three concurrent diseases in a single patient has rarely been described in the literature. We aimed to study the molecular pathogenesis of these three diseases and to explore the correlations among these three diseases within this Chinese pedigree.
Materials and Methods
Clinical subjects
This study is an observational study on human familial-hereditary disease. We selected nine members with this family, which originated from the remote mountainous area of Xinyang, Henan Province, China, as the research objects. Medical histories were obtained, and the pedigree chart is shown in Figures 1(a) and 2(a). This study conformed to the tenets of the Declaration of Helsinki and was approved by the Ethics Committee of the Second Affiliated Hospital of Guizhou Medical University. Written informed consent was obtained from all participating individuals or their guardians. Diagnosis of RA was performed according to criteria of the American College of Rheumatology, the proband’s family was too poor to receive any medical treatment. In addition, any other chronic inflammatory disease was used as the exclusion criteria in this study.

Six family members (labeled as I:2, II:1, II:2, II:3, III:1, and III:3 on the pedigree chart) were examined by an orthopedist, a dermatologist, and an ophthalmologist who performed the necessary clinical and auxiliary examinations, and their medical histories were recorded using a standardized questionnaire. 23 Additionally, six members of the pedigree donated 10 mL samples of whole venous blood per person for testing, which was performed by the Department of Clinical Laboratory. All of the necessary physical and auxiliary examinations were performed at the Second Affiliated Hospital of Guizhou Medical University.
DNA extraction
Six members of the pedigree donated 5 mL samples of whole venous blood per person for genomic DNA isolation. Genomic DNA was extracted using a HiPure Blood DNA-Kit (Magen, Guangzhou, China) according to the manufacturer’s instructions. Genomic DNA samples were stored at −20oC.
Whole genome sequencing and bioinformatics analysis
Whole-genome sequencing was performed on genomic DNA isolated from peripheral blood leukocytes from four cases in the pedigree (II:2, II:3, III:1, and III:3). Genomic DNA (approximately 2 μg) from each sample was used for library construction, according to the standard protocol provided by the manufacturer. Genome libraries were sequenced with an Illumina HiSeq X sequencing system using a 150-bp paired-end run (Illumina, San Diego, CA, USA).
The raw data generated by WGS were processed using the NGS QC Toolkit 24 program to filter out adapters and low-quality reads. SpeedSeq, an ultra-fast personal genome analysis tool, 25 was used to process clean reads by using a series of modules, including reads alignment, duplicate marking, and variant calling. In detail, the alignment of clean reads to a human reference genome (GRCh37/hg19) was performed using BWA-MEM. 26 Duplicate reads were marked using SAMBLASTER, 27 and only uniquely mapped reads were used for the detection of variants. Single-nucleotide variants (SNVs) and indels were identified jointly using FreeBayes (https://github.com/ekg/freebayes) implemented in SpeedSeq. Structural variants were identified by using LUMPY (https://github.com/arq5x/lumpy-sv) and CNVnator (http://sv.gersteinlab.org/).
Variant filtering was applied to exclude variant sites where the samples had a <5 × read depth. Individual variant calls with read depths <5 × or with quality scores <30 were also excluded. The data were analyzed for variants in genes previously associated with arthritis, ichthyosis, and CFEOM disease and for variants in novel candidate genes. The association of genes with Mendelian diseases/phenotypes was annotated based on the NCBI ClinVar (https://www.ncbi.nlm.nih.gov/clinvar/) and Online Mendelian Inheritance in Man (OMIM, http://www.omim.org/) databases. To identify novel recessive or dominant/de novo genetic causes, genes with variants that passed the filtering strategy were further prioritized by using VarElect implemented in TGex (http://tgex.genecards.cn/). Remarkably, variants were finally identified according to different diseases/phenotypes. For CFEOM, only variants present in both cases X and Y were retained. For arthritis and ichthyosis, to reflect the different possible inheritance patterns of the underlying mutations, variants were filtered based on autosomal or sex-linked recessive and dominant/de novo inheritance models.
Functional gene interactions were determined using the STRING database (https://string-db.org/), with a confidence threshold of >0.4. Candidate genes associated with arthritis and ichthyosis were collected using Phenolyzer (http://phenolyzer.wglab.org/). The interaction networks were visualized using the Cytoscape software. 28 Enrichment analyses of the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway were conducted using the clusterProfiler package in R language. 29 The hypergeometric test statistical method was applied, and the whole human genes were used as background genes. The pathways with q-value thresholds <0.05 were considered significantly enriched functional categories, and only the pathways potentially involved in the process of arthritis were shown as dot plot.
Variant validation
PCR primers and conditions for FLG and KIF21A mutations.

Abbreviations: F=forward primers, R=reverse primers.
ELISA assay and flow cytometric analysis
Serum concentrations of IL-13 and IL-17 and cytokine secretions of IL-13 and IL-17A of CD3+T cells from four cases in the pedigree (II:2, II:3, III:1, and III:3) were measured using ELISA-kit (Invitrogen, USA) and FCM (BD, USA), respectively. The analysis of serum parameters was based on a quantitative sandwich ELISA, according to the manufacturer’s instructions. Peripheral blood mononuclear cells (PBMCs) were cultured with 1 × cell stimulation cocktail plus protein transport inhibitors (Invitrogen, USA) for 5 hours. Cultures were harvested and fixed and permeabilized with fixation buffer and permeabilization buffer (Invitrogen, USA). The extracellular and intracellular staining was performed by following the manufacturer’s instructions.
Results
Clinical findings
According to this family history, the genetic inheritance pattern of ichthyosis is autosomal dominant. However, the proband is presented with palmar dry skin accompanied by a mild hyperlinearity and eczema skin and epidermal blisters on the buttocks and back. The proband also presented with an abnormal epidermal barrier function of the skin, the primary finding of skin that was dry and rough, similar to scales, and taupe scaly stripes on the lower legs (Figure 1(b)–(d)), according to this family history and the results of NGS, the phenotypic characteristics of ichthyosis is IV.
Laboratory data for four samples.
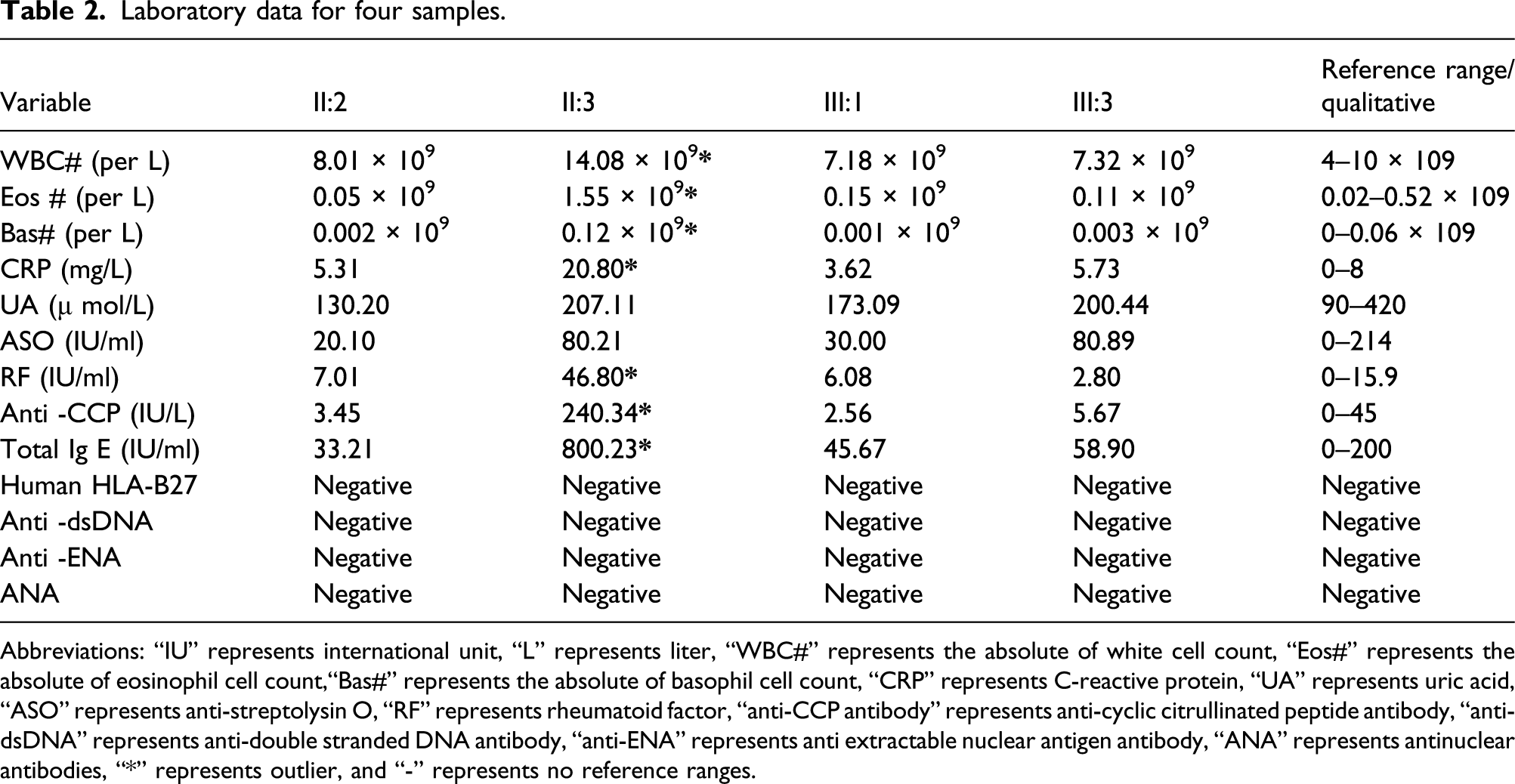
Abbreviations: “IU” represents international unit, “L” represents liter, “WBC#” represents the absolute of white cell count, “Eos#” represents the absolute of eosinophil cell count,“Bas#” represents the absolute of basophil cell count, “CRP” represents C-reactive protein, “UA” represents uric acid, “ASO” represents anti-streptolysin O, “RF” represents rheumatoid factor, “anti-CCP antibody” represents anti-cyclic citrullinated peptide antibody, “anti-dsDNA” represents anti-double stranded DNA antibody, “anti-ENA” represents anti extractable nuclear antigen antibody, “ANA” represents antinuclear antibodies, “*” represents outlier, and “-” represents no reference ranges.
Detailed ophthalmologic examinations of the clinical features of the extraocular muscle of the six individuals were performed. We clinically characterized this Chinese family with autosomal dominant CFEOM1. The three affected members (I:2, II:3, and III:1) shared the typical clinical characteristics of CFEOM1, including congenital, non-progressive ptosis, infraducted globe position in the primary gaze, and upgaze and horizontal gaze palsy in both eyes. Aberrant movements, chin-up head position, and forced duction testing of the superior rectus muscles and medial rectus muscles were positive (Figure 2(b)). The other members (II:1, II:2, and III:3) showed the normal control sequence.
Molecular genetics
Analyses for variants within known CFEOM1, RA, and ichthyosis disease genes revealed two pathogenic gene mutations (KIF21A and FLG).
A total of four samples (II:2, II:3, III:1, and III:3) were included in this WGS study. WGS produced an average of 139.4 GB of clean data per sample, after the removal of adapters and low-quality bases (Supplementary Table S1). In general, each sample had at least 98.82% of reads aligned to the reference genome, and the average coverage of regions was 48.9-fold per sample. A minimum of 99.2% of the target regions were covered at least 1-fold, 99.1% of the target regions were covered at least 5-fold, and 99% of the target regions were covered at least 10-fold. Above all, the quality of the sequencing data conforms with the follow-up analysis standard.
Gene mutation sites in the studied pedigree.

Abbreviations: D: damage, A: disease causing automatic, AD: autosomal dominant.
The profilaggrin/FLG resides on chromosome 1q21 and consists of three exons and two introns, exon 1 (15 bp) consists only of a 5’ untranslated (UTR) sequence, exon 2 (159 bp) contains the translation initiation codon, and exon 3 is extremely large (>12 kb) and encodes most of the profilaggrin polypeptides with almost completely homologous 10, 11, or 12 repeats. There exist polymorphic variations in the number of filaggrin repeats. In the repeat 6, the nonsense heterozygous mutation FLG (g.152280098 C>A, p.E2422*) was detected in the proband. The mutation (p.E2422*) resulted from a single nucleotide base substitution of C → A at position (g.152280098) (Table 3).
A missense heterozygous mutation in KIF21A (g.39726207 G>A, p.R954 W) has also been identified in other two members (I:2 and III:1) (Table 3). However, the nonsense heterozygous mutation FLG (g.152280098 C>A, p.E2422*) was not detected in any of the other five members (I:2, II:1, II:2, III:1, and III:3).
The mutation sites were verified by gene sequence analyses
The PCR products of the FLG and KIF21A genes were directly analyzed by Sanger sequencing. The sequence from the FLG repeat 6, corresponding to codons 2420–2,424, shows a heterozygous transversion g.152280098 C → A, resulting in the nonsense mutation p.E2422* in the proband; however, this FLG mutation was not found in the other five individuals (I:2, II:1, II:2, III:1, and III:3) (Figure 3). The KIF21A gene shows the same heterozygous base substitution g.39726207 G → A in three members (I:2, II:3, and III:1), which resulted in the substitution of an arginine residue at codon 954 with a tryptophan residue (p. Arg954Trp). This KIF21A mutation was not found in the other individuals (II:1, II:2, and III:3). The Sanger sequence results can be seen in Figure 4. All of the Sanger sequencing results for the FLG and KIF21A genes were consistent with the WGS analyses.

Interaction network and functional enrichment analysis for the FLG gene
To gain insight into the functional associations between FLG and other genes, especially those related to arthritis and ichthyosis, a gene-interaction network was constructed based on the interaction dataset collected from the STRING database. As a result, varying degrees of interactions were observed between FLG and 17 genes, including three candidate arthritis genes (IL-13, PIK3R1, and PADI4
The downstream effects of the putative protein damage from the disease-associated variants were also investigated by exploring the affected functional pathways. As a result, these genes were found to be significantly enriched in a total of 80 pathways (Supplementary Table S2). Notably, several enriched pathways associated with arthritis or autoimmune diseases, such as the Jak-STAT signaling pathway
Immunological investigations
We also found that the concentrations of IL-13 and IL-17 in the serum and the frequency of CD3+T cells producing IL-13 and IL-17A in PBMCs of the proband suffering from RA were higher than those in the other three members (II:2, III:1, and III:3) (Figure 6). (A&B) The cytokine secretion of IL-13 and IL-17A of CD3+T cells and the serum concentrations of IL-13 and IL-17 from four cases in the pedigree (II:2, II:3, III:1, and III:3) were measured using FCM and ELISA-kit, respectively.
Discussion
The whole genomes were sequenced to elucidate the key disease-causing genes in this Chinese family, and a heterozygote nonsense mutation in FLG (g.152280098 C>A, p.E2422*) and a heterozygote missense mutation in KIF21A (g.39726207 G>A, p.R954 W) were present in the proband, who suffered from the three diseases (RA, ichthyosis, and CFEOM1). Three patients with CFEOM1 in this pedigree shared a common mutation in KIF21A (g.39726207 G>A, p.R954 W) (Figures 3 and 4).
Ichthyoses are a heterogeneous group of inherited scaling disorders caused by abnormal cornification. 31 Ichthyosis syndrome often involves multiple organs with varying degrees of ichthyosis lesions, is caused by mutations in genes that are primarily involved in skin barrier formation, and is a manifestation of gene pleiotropism. Currently, ichthyosis is associated with many diseases. One-third to one-half of patients with IV show concurrent AD. 32 The development of hypothyroidism and juvenile rheumatoid arthritis in a patient with harlequin ichthyosis (HI) has been reported. 33 An unusual association between RA with vasculitis myositis and ichthyosis has been reported in France. 34 A case of juvenile idiopathic arthritis (JIA) associated with HI has been reported in the UK 35 A case of a patient with non-pitting edema, ichthyosis, arthritis, and presenting a manifestation of leprosy has been reported in India. 36 All of the reports imply that ichthyosis results from the FLG mutation, which may also be significantly associated with the development of RA.
Filaggrin is a multifunctional protein that can alter the skin barrier function and the inflammatory response. 37 Loss-of-function mutations in the (FLG; OMIM #135940) have been reported to cause semi-dominant keratinizing disorders, such as IV and AD.17,38 Autoantibody formation to citrullinated (pro) filaggrin has proven to be a highly specific serological marker for RA, the laboratory data from the proband displayed abnormal values for anti-CCP (Table 2). In the 2008 study by Huffmeier et al., there was no difference in the frequency of FLG loss-of-function mutations in patients with RA compared to controls, that is to say the investigated FLG variants do not confer an overall risk for the development of RA. However, loss-of-function mutations in the FLG gene may contribute to the development of humoral autoimmunity, targeting citrullinated determinants in early RA. 38 However, in the proband, a concurrence of a heterozygote missense mutation in KIF21A (p.R954 W) and a heterozygote nonsense mutation in FLG (p.E2422*) had been identified. The FLG heterozygous mutation (p.E2422*) is a recurrent and rare mutation in both European and Asian populations. 39 Our results suggest significant contribution of the FLG heterozygous mutations (p.E2422*) in these Chinese pedigrees to the development of autoantibodies that recognize citrullinated peptide determinants.
Understanding the genetic bases of complex diseases, such as arthritis, has been the focus of many researchers over the last decade. A key goal of this study is to identify biomarkers that can predict disease onset.
4
The field of complex trait genetics is progressing rapidly, but one specific challenge is the dissection of the multiple gene–gene interactions that are likely to contribute to the heritability that remains to be explained in arthritis. Based on the interaction dataset collected from the STRING database, some gene (IL-13, PIK3R1
The diagnosis of CFEOM1 in the studied family was based upon the observations of autosomal dominant inheritance, bilateral congenital ptosis, bilateral infraducted globe position in primary gaze, and severely restricted upgaze. Genetic analyses demonstrated that the phenotype was linked to the CFEOM1 locus. A missense heterozygous KIF21A mutation (g.39726207 G>A, p.R954 W) has subsequently been observed in this family. This mutation is commonly observed in previously reported pedigrees and in sporadic cases.
There are some limitations on this study. First, the specific molecular consequences of the mutations in FLG are not clear in this study, since early inflammatory arthritis is a clinically heterogeneous disease, cytokine networks are known to play a critical role in the pathogenesis of rheumatoid arthritis. Based on the results of this study, IL-13 and IL-17A might be of better usefulness in the prediction of RA activity status than RF and anti-CCP. 3 However, treating with cDMARD or biologic therapy might affect the frequencies of infections, the levels of cholesterol, and the distribution frequency of lymphocytes or neutrophils (IL-13 and IL-17A). 44 Second, we did not provide the results of auto-antibodies against filaggrin/citrullinated filaggrin in the proband sera, the correlation between the level of auto-antibodies against filaggrin/citrullinated filaggrin and the development of RA will be determined in our subsequent investigations. Indeed, it will be more convincing if we provide information about control of RA patient with ichthyosis and healthy control for the flow cytometry analysis. Finally, the calculation and justification of the sample size selected in this study was not done, and although we provided the questionnaire(s) used in this study, we did not pretest in the Methods, which we believe is worth investigating and will be characterized in our future studies.
Conclusions
To our knowledge, this is the novel reported case of a patient with RA, IV, and CFEOM1. A loss-of-function mutation in the FLG (p.E2422*) was associated with ichthyosis vulgaris and rheumatoid arthritis in these Chinese pedigrees, and there could be larger causal relationship between the FLG heterozygous mutations (p.E2422*) and IL-13 gene or anti-CCP antibodies. The occurrence of three rare comorbid conditions raises the possibility that they may be associated, although the mechanism is still quite unclear. Additionally, we identified a common, recurrent, missense KIF21A (p.R954 W) mutation in this Chinese family with CFEOM1 phenotypes. These reports will be helpful for guiding genetic study and the clinical diagnosis of polygenetic hereditary diseases.
Supplemental Material
sj-xlsx-1-eji-10.1177_20587392211032805 – Supplemental Material for A loss of function mutation in the filaggrin gene associated with ichthyosis vulgaris and rheumatoid arthritis
Supplemental Material, sj-xlsx-1-eji-10.1177_20587392211032805 for A loss of function mutation in the filaggrin gene associated with ichthyosis vulgaris and rheumatoid arthritis by Xinxin Xu, Qingqing Ma, Mu Lin, Mubo Liu, Chaolin Huang, Jianchao Ying and Jun Ye in European Journal of Inflammation
Supplemental Material
sj-xlsx-2-eji-10.1177_20587392211032805 – Supplemental Material for A loss of function mutation in the filaggrin gene associated with ichthyosis vulgaris and rheumatoid arthritis
Supplemental Material, sj-xlsx-2-eji-10.1177_20587392211032805 for A loss of function mutation in the filaggrin gene associated with ichthyosis vulgaris and rheumatoid arthritis by Xinxin Xu, Qingqing Ma, Mu Lin, Mubo Liu, Chaolin Huang, Jianchao Ying and Jun Ye in European Journal of Inflammation
Footnotes
Acknowledgmnts
We thank the doctors and the patients and their families for their participation in this study. We also thank Shanghai Sunny Biotechnology Co., Ltd. for their help with this study.
Authors contributions
X. Xu and Q. Ma performed clinical investigations, X. Xu, M. Lin, M. Liu, and C. Huang performed genetic studies and bioinformatics analysis, and J. Ying contributed to bioinformatics analysis. J. Ye and J. Ying contributed to conceptualization and supervision and wrote and revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the start-up funds from the First Affiliated Hospital of Wenzhou Medical University (Grant Number: 2018QD014).
Ethical approval
Ethical approval for this study was obtain from Ethical Committee of the Second Affiliated Hospital of Guizhou Medical University (ID:20180015).
Informed consent
Written informed consent was obtained from all subjects before the study.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
