Abstract
Chronic inflammation in asthmatics is initiated/exacerbated by many environmental factors, such as bacterial lipopolysaccharide and allergens. Phospholipase A2 and histone acetyltransferase/deacetylases are enzymes involved in inflammatory process, particularly in lipid inflammatory mediators production and control of transcription of many inflammatory genes, respectively. The aim of the study was to identify differences in the inflammatory process in patients with severe and non-severe asthma, taking as a criterion expression of two groups of enzymes: phospholipases A2 and histone acetyltransferases/deacetylases. Thirty-two patients with severe, non-severe atopic to house dust mite asthmatics and 14 healthy volunteers were recruited. Peripheral blood mononuclear cells were stimulated with
Background
Phospholipases A2 (PLA2), present in cells involved in inflammation, participate in the production of lipid inflammatory mediators by releasing arachidonic acid from cell membranes.3,4 Both in vitro and in vivo studies indicate secretory phospholipase A2 from group IIA, V, X, XII, and cytosolic PLA2: α, β, γ are involved in asthma pathogenesis.5,6 Moreover, our previous studies showed that cytosolic phospholipase A2 is involved in the pathogenesis of asthma.7,8 We have previously shown that microsatellite fragments (T)n and (CA)n in the promoter region of the cPLA2α gene (
Bacteria and allergens are the factors that participate most in asthma pathogenesis and exacerbation.9–14 Proteins of house dust mite, and among them, Der p1 and Der p2, are major indoor allergens. Der p1is potent immunomodulator, able to stimulate production of many cytokines in many cells types and might influence the cells phenotype.15,16
Lipopolysaccharide (LPS) is a strong modulator of immunologic reactions – it induces the release of many pro-inflammatory cytokines and mediators responsible for the enhancement of inflammation. 17 LPS also regulates the phospholipases expression and enzymatic activity and is also used as an adjuvant.18–20
Gene expression regulation undergoes through control, involving many transcription factors but also by epigenetic mechanisms DNA methylation and histone acetylation. The acetylation of histones involves lysine residue modification in the tails of histone proteins. Histone acetyltransferases (HAT) are the enzymes that participate in the acetylation process In turn, histone deactylases (HDAC), act by removing acetyl residues, resulting in chromatin conformational change and inhibition of transcription.
21
An imbalance between HAT and HDAC activities was observed in patients with asthma.22,23 In relation to phospholipases, studies show that p300 acetyltransferase is involved in the regulation of
In bronchial biopsies from patients with asthma, an increased level of HAT is observed. Therefore, even a small reduction in HDAC activity, in comparison to normal airways, favors accelerated inflammatory gene expression.26,27 HDAC/HAT activities may contribute to glucocorticoid insensitivity which appears in patients with severe steroid–insensitive asthma.
There is evidence that the different HDACs act on different patterns of acetylation, thus regulating the expression of different types of genes. In biopsies from asthmatic patients, an increase in HAT activity and a reduction in HDAC activity occurred, favoring increased inflammatory gene expressions.27,28
The aim of this study was to identify differences among severe, non-severe asthmatic and healthy subjects in the expression of two groups of enzymes – phospholipases A2 and histone acetyltransferases and deacetylases.
Phospholipases A2 play a crucial role in the generation of inflammatory lipid mediators, and a group of enzymes with acetyltransferase and deacetylase activity that regulate gene expression by modifying chromatin structure.
Materials and methods
Reagents
The LoTox natural
Patients
Thirty-two patients with severe and non-severe asthma of atopic origin, as well as 14 healthy volunteers, were enrolled into the study. The project, conducted from May 2014 to February 2015, was approved by the

Scheme of subject enrollment procedure.
To exclude atopy, the following skin tests were executed to healthy volunteers: mites:
Detailed characteristics of the patients and controls are shown in Table 1.
Parameters and characteristics of patients and controls.

FEV1: forced expiratory volume; WBC: white blood cells; NE: neutrophils; LIMF: lymphocytes; MO: monocytes; EOS: eosinophils; BASO: basophils; HGB: hemoglobin; PLT: platelets; CRP: C reactive protein.
PBMC isolation and stimulation
PBMCs were isolated by centrifugation on Histopaque 1077, a density gradient cell separation medium according to the manufacturer’s instructions. Cells were plated on a 24-well plate and cultured in AIM-V medium overnight. The next day, 6 × 106/ml PBMCs were stimulated in vitro with LoTox nDer p1 (1 µg/ml) or LPS from
RNA extraction and reverse transcription
Total RNA was isolated from PBMCs using the RNeasy Cell Mini Kit with QIA shredder according to the manufacturer’s instructions. RNA was DNase treated, purified, eluted in 30 μl of RNase-free water and stored at −80°C for further analysis. RNA was then reverse transcribed to cDNA using High Capacity cDNA kit.
TLDA cards
Each card of the low-density array contained eight separate loading ports that fed into 48 separate wells for a total of 384 wells per card. Each 2-μl well contained a lyophilized TaqMan assay to enable a single gene to be detected. In this study, the TLDA card was configured into eight identical gene sets (
Real time PCR
qPCR was performed as described previously.
10
Briefly, 300 ng of cDNA was used for each port of TLDA. The array was centrifuged for 1 min twice at 306 ×
The card was then sealed and quantitative PCR (qPCR) was performed using an ABI Prism 7900HT (Applied Biosystems) sequence detection system. RQ Manager 1.2.1 software was used to analyze raw qPCR data. The results were analyzed in comparison to the expression of the reference gene, using Livak’s method, and represented as the relative expression of mRNA in the form of RQ = 2–∆∆Cq. The untreated sample was used as a calibrator.
The results are presented as relative expressions to the untreated control.
Immunoblotting
Total proteins from the PBMCs of patients with asthma and healthy controls were extracted in RIPA protein extraction buffer supplemented with protease inhibitor, and the protein concentrations were determined by the BCA protein assay kit. NuPAGE Bis-Tris Gels (Life Technologies) were used to separate proteins (200V, from 110 mA – start to 70 mA – end). Subsequently, the samples were transferred onto a nitrocellulose membrane using the eBlot Protein Transfer System. To block non-specific binding, 5% non-fat dry milk diluted in TBST (0.01% Tween 20 in TBS) was used. Then, membranes were washed three times with TBST and incubated for 12 h at 4°C with primary antibodies and subsequently for 1 h with secondary antibodies at room temperature. After incubation with secondary antibodies and triple washing in TBST once again, colorimetric detection of bands by means of BCIP/NBT Alkaline Phosphatase Substrate (Millipore) was performed following the manufacturer’s instructions.
Densitometric analysis of bands was performed with ImageJ 1.34s software (Wayne Rasband, National Institutes of Health, Bethesda, Md., USA) and the results are presented as fold change in optical density.
Statistical analyses
Data were analyzed using Statistica software (v. 10.0; StatSoft, Tulsa, OK). The distribution of data and the equality of variances were checked by Shapiro-Wilk and Levene’s tests, respectively. Significant changes were determined by ANOVA with the appropriate post-hoc tests as multiple comparison procedure. Values of
Principal component analysis (PCA) was performed using the SigmaPlot 13.0. according to its technical manual, with using protein data. PCA is a statistical technique that analyzes a data set in which observations are described by several inter-correlated quantitative dependent variables. In this study, PCA analysis shows chosen gene expressions in relation to asthma severity.
Results
Blood and spirometry parameters
Complete blood count, spirometry and CRP analyses revealed significantly higher forced expiratory volume baseline in severe asthmatics, comparing to non-severe asthmatics healthy volunteers. Other parameters showed no significant differences (Table 1). Cell viability, assessed by trypan blue, was over 90% before stimulation and over 84% – after LPS stimulation.
Gene expression
House dust mite allergen and LPS modulated the expression of selected phospholipase A2 genes. An increased expression of
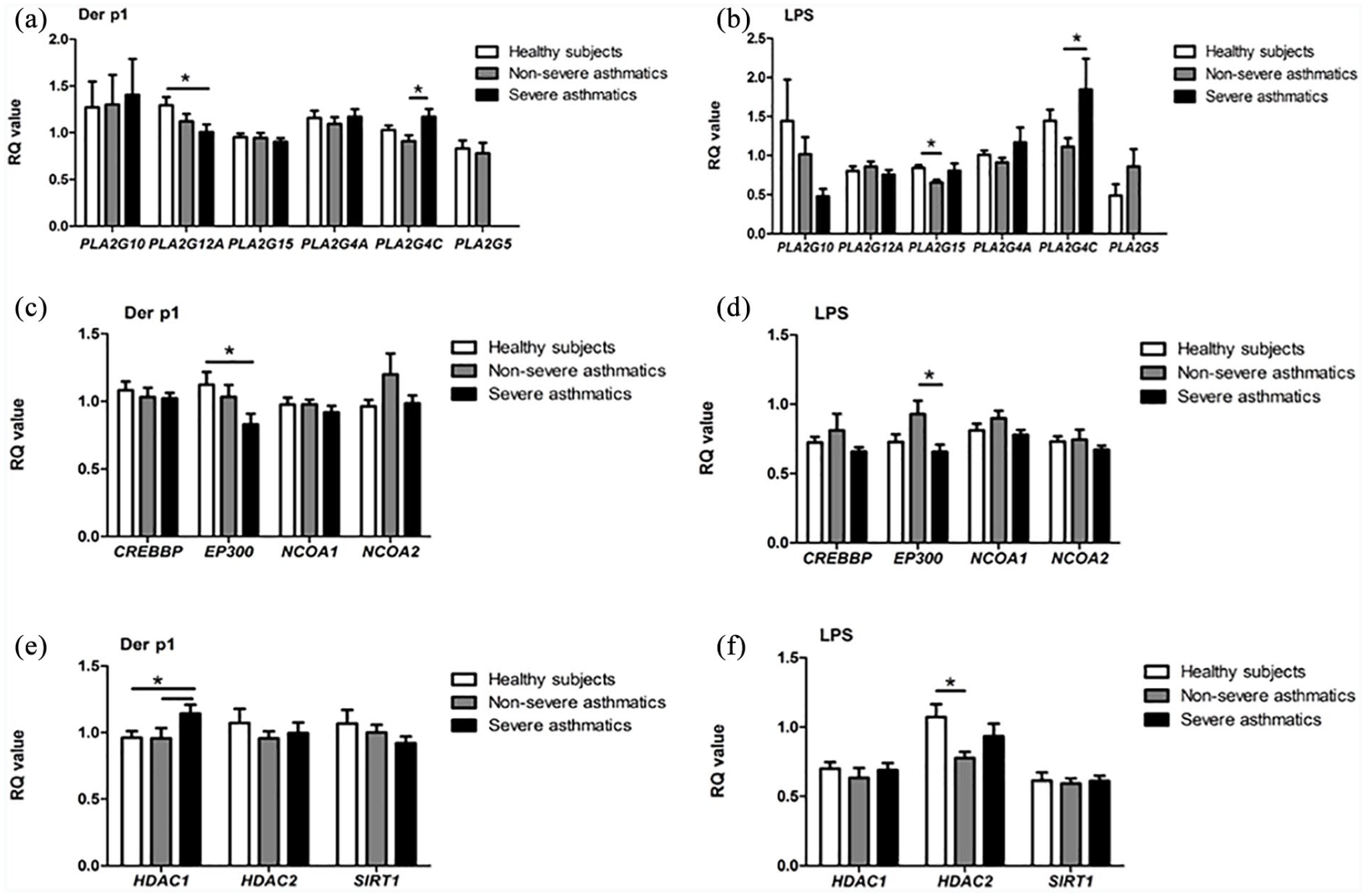
Results of qPCR. Expression of phospholipase A2 (a, b) histone acetylase (c, d), histone deacetylase (e, f) genes in the PBMCs of healthy subjects (white bar,
Four histone acetyltransferase gene expressions was assessed in our study. We observed that the stimulation of PBMC in severe asthmatics with nDer p1 – comparing to healthy subjects, and LPS – comaring to non-severe asthmatics, resulted in a decreased expression of
Among histone deacetylase genes, a difference in
Protein expression
Based on the qPCR results, we evaluated the concentrations of the selected proteins. The immunoblotting analysis showed that among the phospholipases A2, cPLA2γ expression differed between the studied groups and reflected the changes observed on mRNA level (Figure 3a).

Relative cPLA2γ (a), HDAC1 (b), HDAC2 (c), p300 (d) protein expression. PBMC from severe (black bar), non-severe asthmatics (gray bar) and healthy subjects (white bar) were in vitro stimulated with nDer p1 or LPS. Each sample of all participants was analyzed by Western Blot six times showing similar results, and the blots presented are representatives. The
The immunoblotting analysis showed a similar trend in the p300 protein concentration (Figure 3d). However, histone deacetylase (HDAC1 and HDAC2) protein concentrations did not differ significantly between the studied groups (Figures 2c and 3b).
Principal component analysis
Results of the PCA of the PBMCs after stimulation with nDer p1 for all subjects revealed that

Results of PCA in the PBMCs after stimulation with nDER p1 for all subjects. A biplot of individual subjects according to their scores of the first (PC1) and second (PC2) principal component (with prediction ellipse 95%). The HA group (healthy subjects; 1–13) is marked in turquoise; the NSA group (non-serve asthmatics; 14–28) is marked in navy blue; and the SA group (serve asthmatics; 29–39) is marked in red.
The results of PCA in the PBMCs after stimulation with LPS for all subjects showed that

Results of PCA in the PBMCs after stimulation with LPS for all subjects. A biplot of individual subjects according to their scores of the first (PC1) and second (PC2) principal component (with prediction ellipse 95%). The HA group (healthy subjects; 1–13) is marked in turquoise; the NSA group (non-serve asthmatics; 14–28) is marked in navy blue; and the SA group (serve asthmatics; 29–40) is marked in red.
Discussion
In present study we assessed how bacterial lipopolysaccharide and house dust mite allergen influence expression of selected phospholipase A2 and histone acetyltransferase/deacetylase genes and proteins in studied groups. PBMC from severe asthmatics turned out to be more sensitive to stimuli – in this group, allergen treatment resulted in more explicit modulation, comparing to control group. This might be explained by the fact that on the surface of PBMC, there are PAR receptors, which are involved in Der p1-signal transduction. 35 We therefore hypothesized that observed changes are specific to this allergen action. However further studies with specific PAR inhibitors or siRNA may be needed to understand the mechanism of observed changes.
LPS and allergen-stimulated PBMC of patients with phenotype of severe asthma characterized increased cPLA2γ gene and protein expression in comparison to non-severe asthmatics. Human cPLA2γ shares 30% homology and the same amino acids in the catalytic center with cPLA2α. cPLA2γ, unlike cPLA2α, lacks the regulatory phosphorylation sites, thus, the C2 domain is Ca2+ –independent and is constantly bound to the cell membrane. 36 The enzyme possesses phospholipase A2 and lysophospholipase/transacylase activities. 37 The mechanism of this phenomenon might be connected to phospholipids remodeling, which cPLA2γ contributes. 38 Bickford et al. 39 proved that TNF-α induces the cPLA2γ mRNA and protein expression. This mechanism is important – it highlights the significant role of TNF-α in the pathophysiology of severe asthma and its increased presence in PBMCs and the airway of severe asthmatics.39,40 They also 39 revealed an increased expression of cPLA2γ in allergic inflammation in a mouse model of asthma. In our study, the increased expression of cPLA2γ in severe asthmatics may appear by a mechanism of pro-inflammatory cytokines such as TNF or NF-κB, which may be induced by Der p1/LPS and involved in further cellular mechanism. 41
House dust mite allergen increased expression of the
Both LPS and Der p1 decreased the expression of the
The PCA revealed that the
In our study, some data from gene and protein expression is contradictory. The first study indicating the possibility of a correlation between the concentration of mRNA (easy to determine) and protein concentration (difficult to determine) appeared in 2004. However, only approx. 40% of the protein concentration corresponded to the amount of mRNA. 48 The lack of correlation between mRNA and protein expression may be a result of the time difference of transcription and translation. Furthermore, the process of gene expression is a multistep process, and each of the stages might be responsible for the observed differences in the expression of mRNA and protein (control of transport of mRNA, mRNA stability, translation process control, control of protein degradation). Our study is limited because it did not separately analyze protein concentration in the nucleus and cytoplasm. As HAT and HDAC proteins are more abundant in the nucleus, this type of analysis may be more valuable in the assessment of mRNA and protein correlation. Another limitation is lack of the statistical power calculation, however the results obtained by statistical analyzes performed, allow us to consider that the size of the groups were sufficient.
Asthma is characterized by the presence of persistent inflammation (higher CRP level), hence the inflammatory cells undergo natural priming. The patients undertake specific therapy to manage this. Since we cannot exclude these aspects during study design, they should be taken into consideration in critical analysis of results from molecular studies. Three strategies were used to decrease the negative impact of these factors in our study: first, we carefully selected patients (without exacerbation, without high doses of GC, drug deprivation); second, after the isolation of PBMCs, we incubated them overnight before stimulation with Der p1; and third, we decided to validate the results from gene expression on the protein level. Only for cPLA2γ, we reported both the elevated gene and protein expression in severe asthmatics. Current literature does not indicate other diseases with cPLA2γ involvement, except for breast and colon cancer. However, more sophisticated studies are needed to confirm specificity of observed changes to severe asthma phenotype.
In the present study, we used PBMC as an experimental model. Though these cells might only partially reflect the changes occurring in targeted organs, the presence of systemic inflammation in chronic diseases also influences the function of PBMC. Additionally, the changes in the PBMC are easily observable, and they might be easily employed for possible therapeutic solutions.
Conclusion
In conclusion, this project provided differences observed in gene expression of selected phospholipases A2 in PBMCs of patients with severe and non-severe asthma, compared to a control of healthy volunteers. Enzymes responsible for epigenetic control of gene expression in the pathogenesis of asthma were also evaluated. This project constituted a complex analysis of LPS/Der p1 and phospholipases A2/histone acetyltransferases/deacetylases interactions and helps to verify existing literature data. This study aimed to explore the molecular responses of bacterial factor (lipopolysaccharide) and house dust mite allergen exposure in those with varying degrees of asthma. The results of the study show that cytosolic phospholipase A2 gamma (cPLA2γ) may be the molecule that differentiates the responses seen between different asthmatics.
Supplemental Material
sj-pdf-1-iji-10.1177_2058738421990952 – Supplemental material for Expression of cPLA2γ mRNA and protein differs the response of PBMC from severe and non-severe asthmatics to bacterial lipopolysaccharide and house dust mite allergen
Supplemental material, sj-pdf-1-iji-10.1177_2058738421990952 for Expression of cPLA2γ mRNA and protein differs the response of PBMC from severe and non-severe asthmatics to bacterial lipopolysaccharide and house dust mite allergen by Ewa Pniewska-Dawidczyk, Izabela Kupryś-Lipińska, Gabriela Turek, Dorota Kacprzak, Joanna Wieczfinska, Paulina Kleniewska, Piotr Kuna and Rafal Pawliczak in International Journal of Immunopathology and Pharmacology
Supplemental Material
sj-pdf-2-iji-10.1177_2058738421990952 – Supplemental material for Expression of cPLA2γ mRNA and protein differs the response of PBMC from severe and non-severe asthmatics to bacterial lipopolysaccharide and house dust mite allergen
Supplemental material, sj-pdf-2-iji-10.1177_2058738421990952 for Expression of cPLA2γ mRNA and protein differs the response of PBMC from severe and non-severe asthmatics to bacterial lipopolysaccharide and house dust mite allergen by Ewa Pniewska-Dawidczyk, Izabela Kupryś-Lipińska, Gabriela Turek, Dorota Kacprzak, Joanna Wieczfinska, Paulina Kleniewska, Piotr Kuna and Rafal Pawliczak in International Journal of Immunopathology and Pharmacology
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Science Centre [Dec-2012/05/N/NZ5/02630] and Medical University of Lodz [503/0-149-03/503-01-001-19-00].
Ethics approval
Ethical approval for this study was obtained from Bioethical Committee for Research Studies of the Medical University of Lodz (ethics approval number: RNN/102/11/KE).
Informed consent
Written informed consent was obtained from all subjects before the study.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
