Abstract
Introduction:
The glucocorticoid receptor (GR) is one of the most widely studied ligand-dependent nuclear receptors. The combination of transcriptional regulatory factors required for the expression of individual genes targeted by GR varies across cell types; however, the mechanisms underlying this cell type–specific regulation of gene expression are not yet clear.
Methods:
Here, we investigated genes regulated by GR in two different cell lines, A549 and ARPE-19, and examined how gene expression varied according to the effect of pioneer factors using RNA-seq and RT-qPCR.
Results:
Our RNA-seq results identified 19 and 63 genes regulated by GR that are ARPE-19-specific and A549-specific, respectively, suggesting that GR induces the expression of different sets of genes in a cell type–specific manner. RT-qPCR confirmed that the epithelial sodium channel (
Conclusion:
These findings indicate that
Introduction
The glucocorticoid receptor (GR) is a well-known ligand-dependent nuclear receptor (NR) capable of activating or repressing thousands of genes in the human body. 1 GR signaling is present in a variety of organ systems, including the nervous, visual, respiratory, reproductive, and epidermal systems, and regulates genes involved in metabolism and immune and inflammatory responses. 2 Similar to other nuclear receptors, the presence of ligands disrupts the binding between the heat shock protein (HSP) and GR due to the higher affinity between GRs and the ligand; the ligand then binds to GR and the complex dimerizes and translocates into the nucleus. 1 Activated GR binds to the glucocorticoid response element to regulate target genes, which then repress the inflammatory, allergic, and immune responses. 3 Synthetic glucocorticoid-like substances, such as dexamethasone (Dex) and prednisolone, are used to treat inflammatory diseases. Such diseases include AMD (age-related macular degeneration), which is caused by an extruded substance known as drusen that accumulates due to retinal pigment epithelium (RPE) cell dysfunction,4–6 and pulmonary diseases, such as asthma. 7 In addition, steroid hormones are used to treat rheumatic arthritis, ulcerative colitis, and organ transplant rejection. 8 Although the GR pathway may have multiple functions, there is a common mechanism of activation in all cells. However, the type of genes regulated by GR differ from cell to cell.9–11 Given that multicellular organisms are composed of many types of cells that are functionally divergent, genes must be differentially activated based on the cell type.
In differentiated cells, the combinations of regulatory transcription factors required for the expression of individual genes vary across cell types. Although the mechanisms of cell type–specific gene expression are not yet clear, early studies have suggested that pioneer factors are important regulators of gene expression.12–14 Pioneer factors are distinguished from activators by their ability to bind to silent chromatin. 15 The recruitment of transcription factors to specific genomic sequences within condensed chromatin is difficult. However, pioneer factors are able to open the binding site and facilitate the binding of proteins to DNA. 13 Pioneer factors are able to regulate epigenetic factors by recruiting histone-modifying enzymes to either activate or repress gene expression, potentially facilitating cellular reprogramming, depending on the external conditions of the cell. 16 The epigenetic effects of pioneer factors can be divided into histone modifications and DNA methylation. Changes to histone modifications affect gene expression by allowing activating or repressive modification enzymes to remain in place. 17 In contrast, DNA methylation involves the binding of transcriptional corepressors to histones, which suppresses gene expression through H3K9 methylation.18,19
Hormone signaling induces chromatin remodeling and converts chromatin to an open state at certain previously inaccessible GR-binding sites.20,21 This suggests that prior to GR binding, pioneer factors remodel the chromatin structure to make it more accessible. FOXA1, a representative pioneer factor that binds to the nuclear receptors, interacts with the estrogen receptor (ER) and retinoic acid receptor (RAR), depending on the cell type.22,23 FOXA1 expression upregulates GR binding at the mouse mammary tumor virus (MMTV) promoter, a chromatin domain regulated by GR.
24
Transcription factors, including GR, are recruited to the MMTV promoter in a FOXA1-dependent manner, leading to increased transcriptional activity.
24
In contrast, recent studies have shown that hormone signal transduction through ER and GR promotes the restructuring of the FOXA1-chromatin complex. Another pioneer factor, C/EBPβ, initially binds GR-binding sites in mouse liver tissue to promote chromatin remodeling and maintain chromatin accessibility.
25
C/EBPβ is required for the recruitment of GR and for additional chromatin remodeling at multiple GR-controlled
Here, we examined the differential patterns of GR-mediated gene expression in two different cell lines. Furthermore, we suggested that the mechanism underlying this cell type–specific gene expression might depend on the pioneer factor FOXA1. Based on this, we identified genes regulated by GR following treatment of two different cell lines, A549 and ARPE-19, with Dex, and examined how gene expression varied depending on FOXA1. Finally, we identified the factors that regulate cell type–specific gene regulation in A549 and ARPE-19 cells.
Materials and methods
Cell culture
Human RPE cells (ARPE-19) purchased from the American Type Culture Collection (Manassas, VA, USA) were grown in Dulbecco’s modified Eagle’s medium F-12 obtained from Welgene (Daegu, Korea) at 37°C under 5% CO2 conditions. A549 cells were obtained from the Korean Cell Line Bank (KCLB, Seoul, Korea) and maintained in Roswell Park Memorial Institute (RPMI) 1640 medium with 2 mM L-glutamine and 10% fetal bovine serum (FBS) at 37°C under 5% CO2 conditions.
mRNA sequencing
Total RNA was extracted and an mRNA sequencing (mRNA-seq) library was prepared using the TruSeq Stranded mRNA kit (Illumina, San Diego, CA, USA). Polyadenylated mRNAs were purified using poly-T oligo-coupled magnetic beads. mRNAs were then fragmented using divalent cations under elevated temperature conditions. The fragmented RNA was subsequently used for first- and second-strand cDNA synthesis using reverse transcriptase with random primers and DNA polymerase I. These cDNA fragments were purified and enriched by polymerase chain reaction (PCR) to create a cDNA library. Each constructed library was sequenced using an Illumina NextSeq500 instrument (Illumina). The original image data were converted into sequence data and stored in the FASTQ format. Genes showing an absolute fold change of at least 2 and with a false discovery rate (FDR) < 0.05 between the groups were considered to be differentially expressed. Gene set enrichment was evaluated using EnrichR. 28
RNA interference
Small interfering RNA (siRNA) experiments were performed according to previously published methods. 29 Transfection of APRE-19 cells was performed with Oligofectamine (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. The following siRNA sequences were used: siFOXA1(1): 5′-GAGAGAAAAAAUCAACAGCdTdT- 3′ (sense) and 5′-GCUGUUGAUUUUUUCUCUCdTdT-3′ (anti-sense); siFOXA1(2): 5′-GCGAAGUUUAAUGAUCCACdTdT-3′ (sense) and 5′-GUGGAUCAUUAAACUUCGCdTdT-3′ (anti-sense); siAP-1(1): 5′-CGGACCUUAUGGCUACAGUdTdT-3′ (sense) and 5′-ACUGUAGCCAUAAGGUCCGdTdT-3′ (anti-sense); siAP-1(2): 5′-GGCAUGUGCUGUGAUCAUUdTdT-3′ (sense) and 5′-AAUGAUCACAGCACAUGCCdTdT-3′ (anti-sense); and siNS: 5′- UUCUCCGAACGUGUCACGUdTdT-3′ (sense) and 5′-ACGUGACACGUUCGGAGAAdTdT-3′ (anti-sense).
Quantitative RT-PCR (RT-qPCR)
ARPE-19 and A549 cells were treated with Dex (100 nM) for 16 or 24 h; then, the total RNA was extracted using Trizol (Invitrogen, Carlsbad, CA, USA) and reverse transcribed (RT) using an iScript cDNA synthesis kit (Bio-Rad Laboratories, Hercules, CA, USA) in a total volume of 20 μL. The RT product was used for qPCR analysis with specific primers. The RT-qPCR primer sequences are listed in Table 1.
Primers used for RT-qPCR experiments.

Statistical analysis
The
Results
Differential gene expression mediated by GR in ARPE-19 and A549 cells
To explore the differing patterns of GR-mediated gene expression in two GR-responsive cell lines, ARPE-19 and A549, transcriptional analysis was performed on these cells following their treatment with Dex (Figure 1(a)). Eighty-six genes were differentially expressed between the two cell lines (Figure 1(b)). There were four genes that showed altered expression following Dex treatment, when compared to the control, in both cell lines. In ARPE-19 cells, the expression of 15 and 8 genes was increased and decreased, respectively, compared to the control (Figure 1(c), left panel). In A549 cells, 50 and 17 genes were upregulated and downregulated, respectively, compared to the control (Figure 1(c), right panel). RNA-seq analysis revealed that there were 19 ARPE-specific Dex-regulated genes. Moreover, while 63 genes were found to be A549-specific, four genes showed significantly different expression in both cell lines (Figure 1(d)). Based on the RNA-seq analysis, we confirmed the presence of cell-specific genes regulated by GR in ARPE-19 and A549 cells. These results suggest that the GR induced gene expression is cell type-specific.

Differential gene expression induced by GR in ARPE-19 and A549 cells: (a) experimental setup for the identification of cell type–specific genes. (b) Heat map generated from RNA-seq analysis of ARPE-19 and A549 cells treated with Dex (100 nM) for 16 h, showing the differential expression of GR-responsive genes between the two groups. Values represent the log2 fold change (FC) relative to the vehicle-treated control (CTR). GR target genes are divided into three groups: ARPE-19-specific genes, A549-specific genes, and genes regulated by GR in both cell lines (non-specific genes). (c) The number of genes that were upregulated or downregulated by GR in ARPE-19 or A549 cells. (d) Venn diagram showing the number of genes that are ARPE-19-specific, A549-specific, and non-specific.
Activation of cell type–specific signaling pathways by GR
To confirm the expression pattern of the cell type–specific genes, we performed pathway analysis using genes regulated by GR in A549 and ARPE-19 cells. Pathway analysis (KEGG) was performed using EnrichR and revealed that GR was associated with different pathways in ARPE-19 and A549 cells. Specifically, genes that were differentially regulated by GR in ARPE-19 cells were enriched for the PI3K-Akt signaling pathway (

Activation of cell type–specific signaling pathways by GR. (a, b) Among the GR-responsive genes, ARPE-19-specific and A549-specific genes were categorized according to gene function via KEGG pathway analysis. Statistically significant pathways (
Identification of cell type–specific genes regulated by GR
After identifying the presence of cell type–specific genes using transcriptome analysis, RT-qPCR was used to validate the genes selectively expressed in A549 and ARPE-19 cells, along with the genes regulated by GR in both cell types. After 16 and 24 h of Dex treatment in ARPE-19 cells, epithelial sodium channel (

Identification of cell type–specific genes regulated by GR. (a, c) Expression of GR target genes (
FOXA1 acts as a pioneer factor for ARPE-19-specific ENACα expression
Among the genes identified by RT-qPCR,
ARPE-19 cells deficient in FOXA1 showed significantly reduced
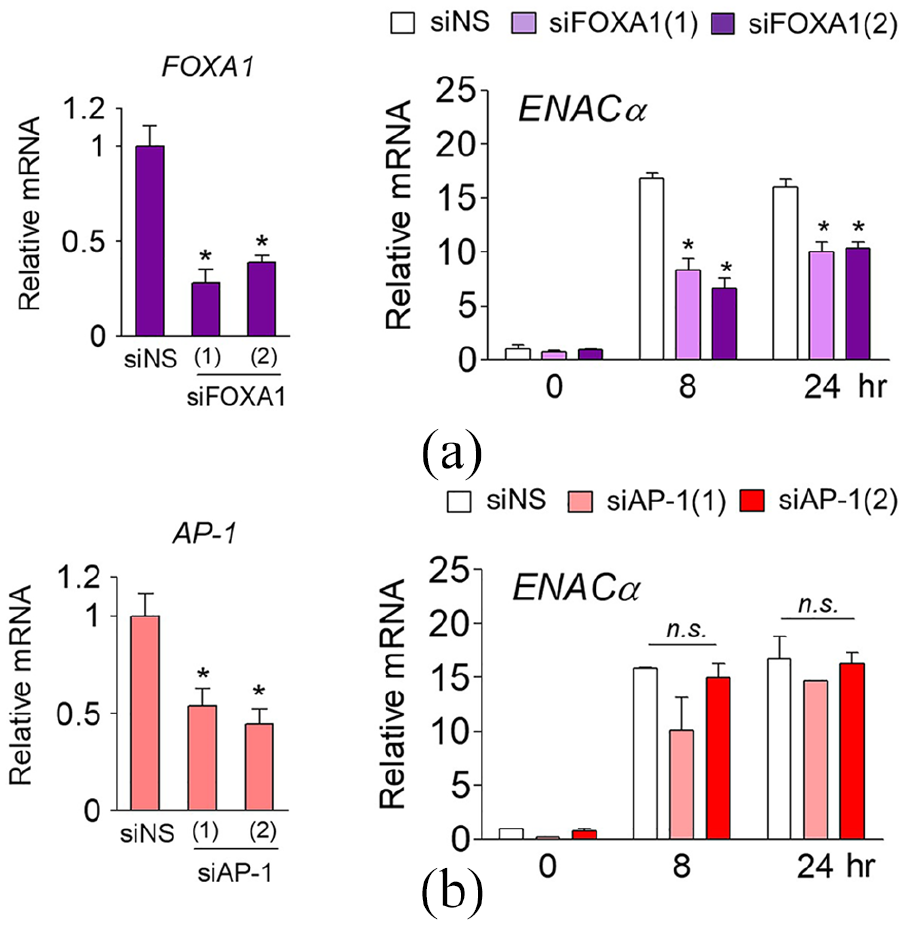
FOXA1 is a pioneer factor mediating ARPE-19-specific expression of
Discussion
Most GR-binding sites are constitutively open and accessible. However, the binding of pioneer factors to specific sites on genomic loci causes local chromatin accessibility and promotes the recruitment of other transcription factors. FOXA1 has been studied as a pioneer factor that interacts with steroid receptors to bind to specific genomic loci. In particular, FOXA1 is the main factor interacting with ER during the development of breast cancer.23,30–32 FOXA1-mediated regulation of GR has been identified from the analysis of the MMTV promoter region, a chromatin domain regulated by GR. FOXA1 alters the chromatin structure, regardless of whether GR binds to MMTV.
24
The binding of GR is enhanced by FOXA1, which optimizes the chromatin structure and mediates the binding of additional transcription factors associated with GR. In a FOXA1-dependent manner, FOXA1 and GR recruit related transcription factors, including NF1 and Oct1, leading to in an increase in transcriptional activity of the genes.
24
The interaction between NRs and transcription factors can be either dependent or independent of DNA binding. Mechanisms independent of DNA binding include the binding between GR and AP-1, a pro-inflammatory transcription factor, which is mediated through protein–protein interactions in the absence of GR responsive element (GRE). AP-1 also maintains the accessible chromatin structure, allowing the selective access of GR to specific sites. The loss of AP-1 significantly reduces the binding of GR to the DNA.33,34 This could be due to the fact that GR must bind with AP-1 to bind the DNA or because of the increased chromatin accessibility established by the activation of AP-1. In addition, AP-1 is recruited to the TR responsive element (TRE) inflammatory gene to induce transcription and serves as a pioneer for the subsequent recruitment of GR.33,34 Regulators that involve in GR binding may be cell type-specific. The AP-1 motif was identified at the GR-binding site in hepatocytes, and the enrichment of motifs for other proteins (e.g. SP1 and forkhead motifs) was confirmed.
35
These results suggest that the role of this factor in the binding of GR to the chromatin is to provide cell type–associated specificity. In this study, we demonstrated that different genes are expressed in different cell types in response to a common activator (GR), resulting in the activation of different biological pathways. In addition, various transcriptional mechanisms are involved in different cell types to control the expression of the same gene, suggesting that pioneer factors play an important role in these processes. One limitation of our study was that we used only two cell lines in our RNA-seq experiment. Follow-up studies can also aim to identify if there are additional patterns of GR-mediated gene expression in various tissue-specific cell types. Another limitation of this study is that the existence of intracellular signaling pathways coupled with epigenetic regulators (e.g. coactivators or corepressors) of cell type–specific gene expression was not considered. For example, receptor-mediated intracellular signaling cascades such as SGK1, ERK, and PKA have been reported as a mechanism regulating the expression of ENAC.36,37 Most recently, GR has been reported to activate transcription via the direct binding of a 1.3-kilobase portion of the
Conclusion
Our findings indicate that the expression of
Supplemental Material
2020_05_18_A549_ARPE19_Supple_Figures_R1 – Supplemental material for Cell-specific expression of ENACα gene by FOXA1 in the glucocorticoid receptor pathway
Supplemental material, 2020_05_18_A549_ARPE19_Supple_Figures_R1 for Cell-specific expression of ENACα gene by FOXA1 in the glucocorticoid receptor pathway by Young Sun Chung, Hong Lan Jin and Kwang Won Jeong in International Journal of Immunopathology and Pharmacology
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Basic Science Research Program through the National Research Foundation (NRF) of Korea funded by the Ministry of Education (2014R1A1A2056066) and by the Gachon University research fund of 2018 (GCU-2018-0665).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
