Abstract
Endometrial carcinoma (EC) is one of the most common gynecological cancers in many developing countries. Although tremendous advances have been made in the diagnosis and treatment of EC, there is still no adequate biomarker currently available for predicting the prognosis of this cancer. In this study, we found that miR-103 expression was significantly upregulated in EC tissues than their paired non-carcinoma tissues. Overexpression of miR-103 significantly promoted EC cell proliferation, while downregulation of miR-103 significantly suppressed EC cell proliferation. In addition, ZO-1 expression was significantly downregulated in the EC tissues than their paired non-carcinoma tissues. We also found an inverse correlation between ZO-1 and miR-103. Moreover, ZO-1 was validated as the direct target of miR-103. The downregulation of ZO-1 significantly enhanced EC cell proliferation. In conclusion, miR-103 could regulate EC cell proliferation through directly targeting ZO-1. Our results provide a potential development of microRNA-based targeted approaches for the treatment of EC.
Introduction
Endometrial carcinoma (EC) is a common malignancy of the female genital tract and one of the most common gynecological cancers in many developing countries. 1 Several risk factors have been reported to be involved in EC, such as hypertension, postmenopausal status, infertility, family history of EC, and long-term used of estrogen. 2 In spite of tremendous advances made in the diagnosis and treatment of EC in recent years, there is still no adequate biomarker currently available for predicting the prognosis of this cancer. Therefore, it is important to elucidate the molecular mechanisms underlying the development and progression of EC for the novel diagnostic, prognostic, and/or treatment strategies for this disease.
MicroRNAs (miRNAs) are a class of endogenous, 20–25 nucleotides in length, non-coding single-stranded RNAs that repress the expression of target genes by degradation or inhibiting translation of the targeted mRNA. 3 MiRNAs have been found to have important roles in regulating diverse basic cellular functions, including proliferation, migration, invasion, and epithelial–mesenchymal transition (EMT) process. 4 Numerous miRNAs have been found to be aberrantly altered in a variety of cancers including EC. 5 Currently, miRNAs are attractive candidates for therapeutic targets in cancer. Thus, the identification of miRNAs and their targets is essential for cancer research.
In this study, we identified the expression of miR-103 in EC tissues and their adjacent non-carcinoma tissues. We also investigated the role of miR-103 in EC cell proliferation by overexpression and downregulation of miR-103. Subsequently, we discussed the molecular mechanism underlying its effects, and ZO-1 was identified as a directly functional target for miR-103. Moreover, we detected the function of ZO-1 in EC cell proliferation by downregulation of ZO-1. Our results provide the first evidence of regulatory mechanisms of miR103/ZO-1 axis in EC. Therefore, this study provides a potential novel candidate therapeutic target in EC.
Materials and methods
Ethical approval
The study was approved by the Institutional Ethics Committee of our hospitals, and written informed consent was obtained from all participants.
Tissue samples and cell lines
The human endometrial cancer cell lines, HHUA and JEC, were obtained from American Type Culture Collection (Rockville, MD, USA) and maintained in Minimum Essential Medium Eagle (Sigma-Aldrich, St Louis, MO, USA) supplemented with 15% of fetal bovine serum (FBS; Gibco, Rockville, MD, USA), 100 U/mL of penicillin, and 100 μg/mL of streptomycin (Gibco). The cells were incubated at 37°C in a humidified atmosphere containing 5% CO2 and 95% air.
Human EC tissues were obtained from 14 patients treated at the Oncology Division in the Department of Gynecology and Obstetrics of Liaocheng People’s Hospital (Shandong, China), between March 2016 and May 2018. This study was approved by the ethics committee, and informed consent was obtained from each patient. All pathology specimens were reviewed in our institution, and histological classification was performed according to the World Health Organization (WHO) criteria, whereas pathological stage was determined in accordance with the International Federation of Gynecology and Obstetrics (FIGO) guidelines. None of the patients had received preoperative chemotherapy or radiation. Age, histological type, stage, grade, additional disease (i.e. diabetes, obesity, and blood pressure), and treatment information were recorded in all cases. Biopsies were snap frozen in liquid nitrogen and stored at −80°C until subsequent analysis.
Real-time quantitative reverse transcription polymerase chain reaction
Total RNA was isolated from frozen tissues or cells with mirVanaTM miRNA isolation kit (Thermo Fisher Scientific, Boston, MA, USA). After removal of residual DNA, the RNA was reversely transcribed into complementary DNA (cDNA) with a RNA-to-cDNA Kit (Thermo Fisher Scientific) according to the manufacturer’s protocol. Quantitative real-time polymerase chain reaction (qRT-PCR) was performed on an ABI 7500 system (Applied Biosystems, Foster City, CA, USA) using SYBR Green I detection and gene-specific primers. The sequences of the primers used for PCR amplification were as follows: ZO-1, 5′-CCCTCAAGGAGCCATTC-3′ (forward) and 5′-GAGTTTGCTCCAACGAGA-3′ (reverse); β-actin, 5′-TAGTTGCGTTACACCCTTTCTTG-3′ (forward) and 5′-GCTGTCACCTTCACCGTTCC-3′ (reverse). 6
For quantification of miRNAs, 1 μg of total RNA was reverse-transcribed using the Taqman advanced miRNA cDNA synthesis kit according to manufacturer’s recommended protocols (Applied Biosystems). U6 small nuclear RNA (snRNA) was used as the internal control and reverse-transcribed by TaqmanTM microRNA reverse transcription kit by following manufacturer’s protocol (Applied Biosystems).
The relative expression levels were normalized to β-actin or U6 using the 2–ΔΔCt method. 7 All the reactions were run in triplicate.
Western blotting analysis
Cells extracts were prepared using lysis buffer (Thermo Fisher Scientific), and protein concentrations were determined using a DC Protein Assay kit (Bio-Rad, Richmond, CA, USA). Total protein (15 μg) was separated on 4%–15% precast gels (Bio-Rad) and transferred onto nitrocellulose membranes. After blocking in 5% non-fat milk for 2 h at room temperature, the membranes were incubated with primary antibodies (anti-ZO-1, 1:500; Abcam, Cambridge, MA, USA) overnight at 4°C. After being washed with tris buffered saline with Tween-20 (TBST) three times, the membranes were incubated with horse radish peroxidase-conjugated secondary antibodies (1:1000; Thermo Fisher Scientific). The bands were visualized using an enhanced chemiluminescence kit (Pierce, Minneapolis, MN, USA). β-Actin (1:500; Sigma-Aldrich) was used as a loading control for analysis.
MiR-103 mimic and miR-103 inhibitor transfection
Transfection of miR-103 mimic, miR-103 inhibitor, and the corresponding mimic/inhibitor negative control (miR-Ctl) was performed using the RNAiMAX Reagent (Invitrogen, Carlsbad, CA, USA) by following the manufacturer’s instructions. MiR-103 mimic, miR-103 inhibitor, and miR-Ctl oligos were commercially available from Thermo Fisher Scientific. Cells were collected for subsequent analysis after 48 h of transfection.
Luciferase reporter assay
The 3′-untranslated region (3′-UTR) luciferase assay is a common tool to study the expressed miRNA and target 3′-UTR interactions. For ZO-1 target gene verification, ZO-1 3′-UTR luciferase assay was performed as previously described. 6 Briefly, HeLa cells were seeded into a 24-well plate at a density of 2 × 105 cells per well. The cells were cotransfected with 40 nM of either miR-103 or miR-Ctl, 20 ng of either wild-type ZO-1-3′-UTR-WT or ZO-1-3′-UTR-mutant (WT: UGU UAC AUU UUU AAG UGC UGC A or mutant: UGU UAC AUU AUA AUG ACC ACG A), 6 and 2 ng pRT-TK (Promega, Madison, WI, USA) with lipofectamine 2000 (Invitrogen). After 24 h of transfection, cells were harvested, and dual-luciferase activities were measured with the Dual Luciferase Reporter Assay System (Promega). Results are represented by normalizing firefly luciferase activity to Renilla luciferase activity.
ZO-1 silencing
The siRNA for ZO-1 (ZO-1-KD, forward, 5′-CTGATCAAGAACTAGATGA; reverse, 5′-TCATCTAGTTCTTGATCAG) were transfected into HHUA and JEC cells using lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. siRNA negative control (Ctl-KD) was obtained from Sigma-Aldrich. The cells were cultured in a six-well plate for 48 h, and the ZO-1 expression level was detected by Western blot and qRT-PCR to determine the interference efficiency.
Cell proliferation assay
CyQuant assay (Thermo Fisher Scientific) was used to monitor cellular proliferation. Cells were seeded at a density of 5000 cells per well in the 96-well plates. After 1, 2, 3, 4, or 5 days of incubation, the media was removed and 100 µL of CyQuant solution was added to the wells at the same indicated time. Then, the plates were incubated in the dark for 45 min at room temperature. The CyQuant solution was prepared immediately before use. Subsequently, plates were read at excitation at 497 nm and emission at 520 nm.
Colony formation assay
The cells were seeded in the six-well plates at a density of 1000 cells in each well after 24 h of transfection of miR-103 or miR-Ctl and maintained in the cell culture media for 2 weeks. Colonies were fixed with methanol and stained with 2% Giemsa solution (Merck, Darmstadt, Germany) for 10 min after culture for 2 weeks.
Statistical analyses
SPSS software (version 16.0.1; SPSS Inc., Chicago, IL, USA) for Windows was used for the statistical analyses. All data were expressed as mean ± standard error (SE; n = 3 independent experiments). The two-tailed Student’s t-test was used to compare two groups. The Spearman’s rank correlation analysis was performed to analyze the correlation. P (probability) < 0.05 was considered to indicate a statistically significant difference.
Results
miR-103 is overexpressed in the EC tissues
In our preliminary bioinformatic analysis data, among the top miRNAs overexpressed in EC, we found that miR-103 was not previously associated with EC. In addition, previous studies reported that miR-103 was upregulated in colorectal cancer tissues. 6 In this study, we first determined the expression level of miR-103 in 14 pairs of EC tissues and adjacent non-carcinoma tissues by qRT-PCR. Figure 1 shows that miR-103 was significantly upregulated in the human EC tissues in comparison with the non-carcinoma tissues, which was consistent with previous study. U6 snRNA was used as the internal control.

miR-103 is overexpressed in endometrial carcinoma (EC). miR-103 level is determined by qRT-PCR in 14 paired EC tissues and adjacent non-carcinoma tissues. Values represent mean ± SE, *P < 0.05 versus non-carcinoma tissues.
These results suggest that overexpressed miR-103 may involve in the tumor development and progression.
miR103 overexpression promotes the proliferation of EC cells
The effects of miR-103 on the human EC cell proliferation were determined by Cyquant assay and colony formation assay. Figure 2(a) shows that miR-103 was significantly upregulated or downregulated after miR-103 mimic or miR-103 inhibitor transfection in both EC cells. Next, we found that miR-103 overexpression remarkably enhanced the proliferation of endometrial cells than miR-Ctl and untransfected cells (Figure 2(b) and (c)). In addition, both EC cells transfected with miR-103 exhibited significant increase in colony numbers compared with miR-Ctl (Figure 2(d)). Conversely, cells transfected with miR-103 inhibitor had significantly suppressed cell proliferation and reduced cell colony formation compared with miR-Ctl and untransfected cells (Figure 2(b)–(d)).

Effects of miR-103 on human EC cells in vitro: (a) miR-103 level was analyzed by qRT-PCR after cell transfection with miR-103 mimics, miR-103 inhibitor, or miR-Ctl. (b and c) Relative cell proliferation rate of miR-103, miR-103 inhibitor, or miR-Ctl transfected cells and untransfected cells in vitro was monitored by Cyquant assay. (d) Representative colony formation assays of EC cells transfected with miR-103, miR-103 inhibitor, or miR-Ctl. Values represent mean ± SE (n = 3), *P < 0.05 versus miR-Ctl; ^P < 0.05 versus untransfected cells.
ZO-1 is directly repressed by miR103
In the previous study, miR-103 has been evaluated to be directly targeted by ZO-1 in colorectal cancer. 6 To investigate the underlying mechanism of miR-103 in EC, we further detected the expression of ZO-1 in EC tissues and their paired non-carcinoma tissues by qRT-PCR. As shown in Figure 3(a), ZO-1 mRNA levels were downregulated in the human EC tissues compared to the non-carcinoma tissues. To further explore the relationship between miR-103 and ZO-1 in endometrial cancer, correlation analysis between the expression of ZO-1 was performed. The expression of ZO-1 was negatively correlated with miR-103 (Figure 3(b)).
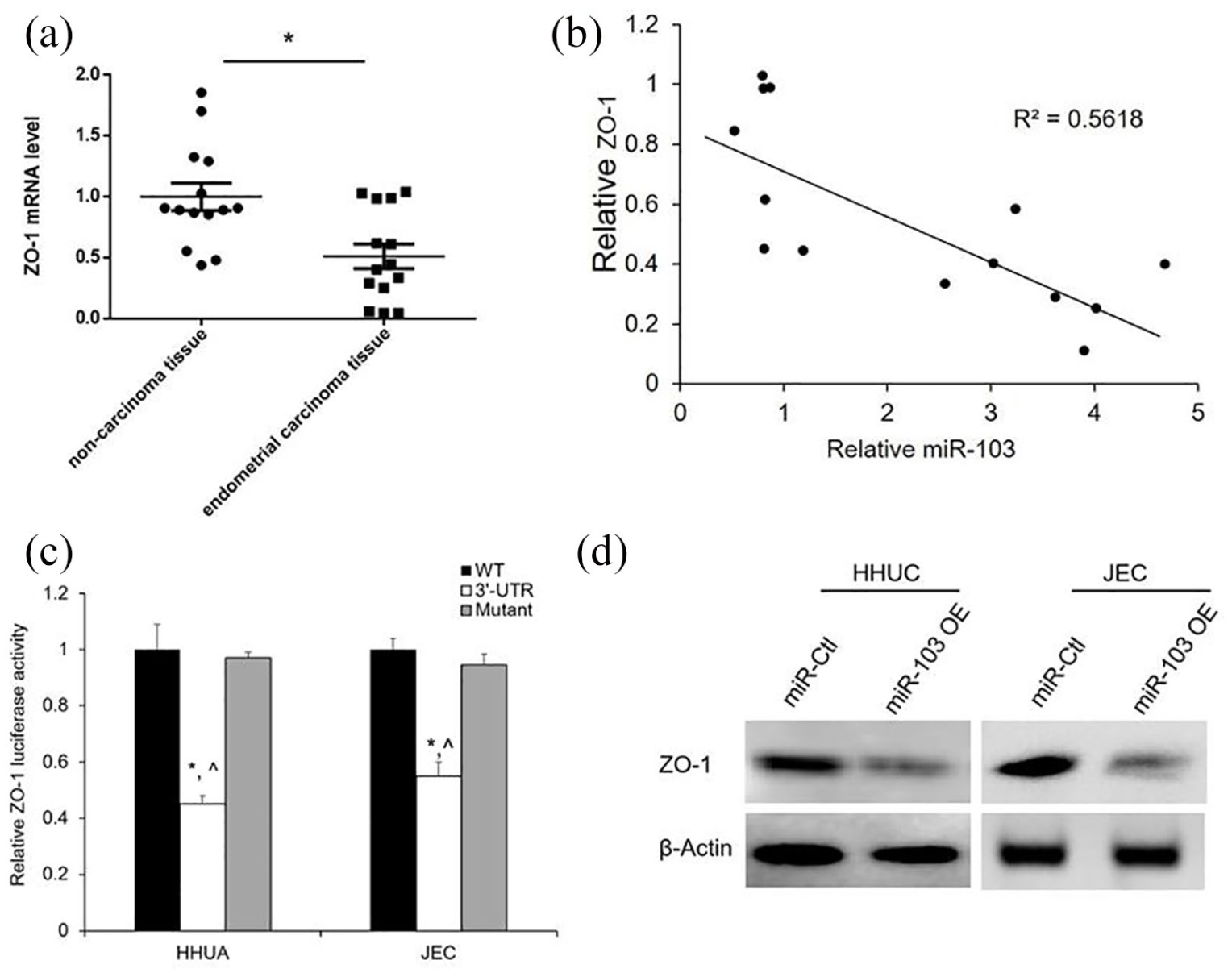
miR-103 directly represses ZO-1 in human EC cells: (a) relative ZO-1 mRNA expression level was identified by qRT-PCR in 14 matched EC tissues and adjacent non-carcinoma tissues. Values represent mean ± SE, *P < 0.05 versus non-carcinoma tissues. (b) Correlation analysis revealed a negative correlation between miR-103 and ZO-1 expression in EC tissues, P < 0.05. (c) The 3′-UTR luciferase assay demonstrated a direct binding of miR-103 and ZO-1. Values represent mean ± SE (n = 3), *P < 0.05 versus WT; ^P < 0.05 versus mutant. (d) miR-103 overexpression inhibited the ZO-1 protein expression was revealed by Western blotting.
To verify whether ZO-1 is directly repressed by miR-103, HHUA and JEC cells were cotransfected with the ZO-1 3′-UTR and miR-103 mimic or miR-Ctl. On the contrary, the ZO-1 3′-UTR mutant was cotransfected with miR-103 mimic or miR-Ctl. Luciferase results showed that miR-103 significantly reduced the luciferase activity of ZO-1 3′-UTR (Figure 3(c)), but had no effect on the luciferase activity of the mutant (Figure 3(c)).
Subsequently, we detected the ZO-1 expression level after overexpression of miR-103 by Western blotting in HHUA and JEC cells. Results revealed that the ZO-1 protein levels were significantly decreased by miR-103 mimic than the miR-Ctl (Figure 3(d)).
Taken together, these results indicate that miR-103 directly targets ZO-1 in EC cells. miR-103 overexpression contributes to the tumor development and progression.
ZO-1 silencing enhances the proliferation of EC cells
To examine the molecular mechanism underlying miR-103, we silenced ZO-1 via transfecting siRNAs in HHUA and JEC cells. ZO-1 siRNA significantly reduced the mRNA and protein levels in both HHUA and JEC cells (Figure 4(a) and (b)). Similar to miR-103 overexpression, ZO-1 silencing markedly enhanced the proliferation than Ctl-KD and untransfected cells (Figure 4(c) and (d)). Moreover, the ZO-1 silencing also significantly increased the colony formation compared with Ctl-KD (Figure 4(e)).
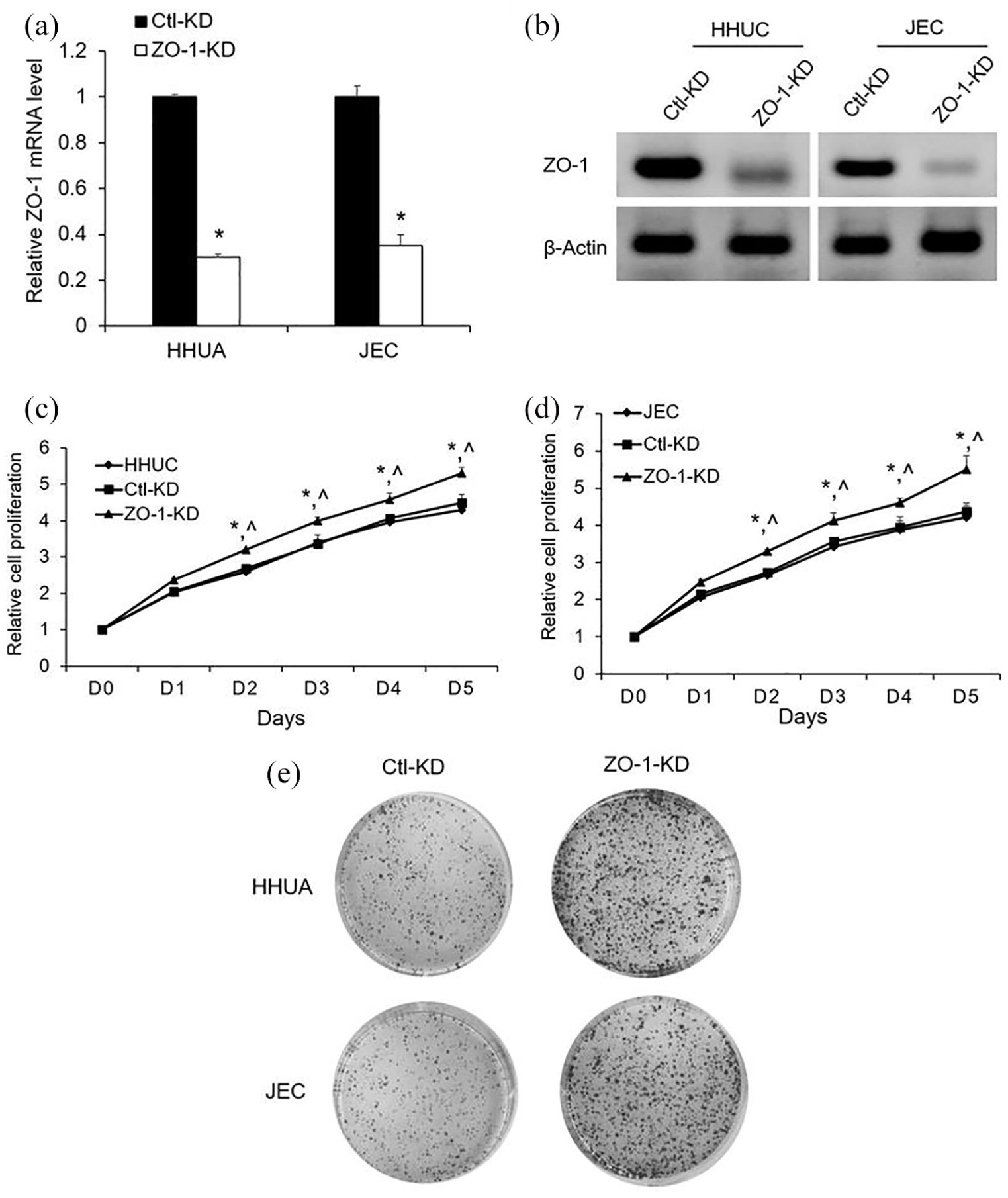
Effects of ZO-1 on human EC cells in vitro: (a) ZO-1 mRNA levels were analyzed by qRT-PCR after cell transfection with ZO-1 siRNA (ZO-1-KD) or control siRNA (Ctl-KD). (b) ZO-1 siRNA inhibited the ZO-1 protein expression was revealed by Western blotting. (c and d) Relative cell proliferation rate of ZO-1-KD, Ctl-KD, and untransfected cells in vitro was monitored by Cyquant assay. (e) Representative colony formation assays of EC cells transfected with Ctl-KD or ZO-1-KD. Values represent mean ± SE (n = 3). *P < 0.05 versus Ctl-KD; ^P < 0.05 versus untransfected cells.
Discussion
MiRNAs, small non-coding RNAs, were identified in 1993. 8 They can silence a broad set of target genes and play important roles in the post-transcriptional level of modulating gene expression. Their expression and functions are cell and tissue specific. Currently, miRNAs have been revealed to be aberrantly expressed in cancer cells and attractive in research as oncogenes or tumor suppressor genes in the process of tumorigenesis. 9 In addition, miR-103 was observed to regulate tumorigenesis in colorectal cancer by targeting ZO-1. 6 However, the roles of miR-103 in the progression of human EC are still unclear. In this study, miR-103 was found to be upregulated in EC tissues compared to the adjacent non-carcinoma tissues. In addition, the overexpression of miR-103 can promote the proliferation of EC cells. Conversely, the downregulation of miR-103 significantly suppressed the proliferation of EC cells.
Decreased expression of ZO-1 has been observed to correlate with increased invasiveness in breast, colorectal, and digestive tract cancers. 10 ZO-1 has also been revealed to be involved in tumor invasion-associated EMT, implicating its role in tumor growth process. 11 Downregulation of ZO-1 expression was found to link with disruption of cell–cell interactions leading to carcinogenesis and deep myometrial invasion in malignant endometrium. 12 Since ZO-1 has been reported to be directly regulated with miR-103 in colorectal cancer and miR-103 was investigated to be upregulated in EC tissues, we subsequently studied whether mi-103 could potentially play a role in human EC through ZO-1. In this study, ZO-1 was found to be directly targeted by miR-103 by 3′-UTR luciferase assay and Western blotting. In addition, this result was also supported by the negative correlation between miR-103 and ZO-1 in EC tissues. Furthermore, ZO-1 silencing was found to significantly enhance the proliferation of EC cells. Although our in vitro results provide the potential therapeutic target, the future efforts will be directed toward developing in vivo model to verify the in vitro results.
In summary, we conclude that miR-103 is upregulated in human EC tissues. Overexpressed miR-103 enhances the proliferation of EC cells, while the downregulated miR-103 suppresses the proliferation of EC cells. Moreover, ZO-1 is directly repressed through miR-103. Furthermore, ZO-1 silencing significantly promotes the proliferation of EC cells. These results may help to further elucidate the complex underlying molecular mechanisms that regulate development and progression of EC. Besides, this newly identified miR103/ZO-1 axis suggests the potential to be a prognostic marker or therapeutic target for patients with EC in the future.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
