Abstract
In recent, mechanisms behind transition from normal epithelium to premalignancy and finally to oral carcinoma have been linked to genetic and epigenetic multistep process. These epigenetic changes are heritable, potentially reversible, and presented as epigenetic signature normal and carcinoma cells. Such epigenetic alterations have been suggested as DNA methylation, histone modifications, and small noncoding RNAs. These extragenetic changes have been reported to play a role in the modulation of several key signaling, regulatory, and repair routes in oral squamous cell carcinoma (OSCC) and other carcinomas. In this review, we summarize scientific developments in the last decade, substantiating the role of extra- and beyond genetic molecular signatures in OSCC progression, and potential biomarker and implications in chemotherapy intervention.
Introduction
Oral squamous cell carcinoma (OSCC) is an aggressive malignant neoplasm, exhibiting amplitude of etiological, clinical, and molecular heterogeneity. 1 –4 The majority of OSCCs are linked with tobacco, alcohol use, betel quid chewing, human papilloma virus (HPV) infections, and other potential agents, acting to promote carcinogenesis. 1,2,5,6 However, the mechanisms associated with the development of OSCC are known to be in the form of genetic and epigenetic perturbations. 1 –10
In recent time, overwhelming attention is being given to epigenome alterations as “epimutations” specifically centered on aberrant cellular signaling and DNA damage response pathways. 11 –16 There is an accepted notion that aberrant DNA methylation patterns present as major reason behind the epigenome landscape changes in OSCC. However, other epigenetic processes such as posttranslational histone modification, the deposition of histone variants, histone chaperones, and recently noncoding RNA are fueling up the development of OSCC. 11 –13,17
Several epigenetically regulated cellular pathways encompassing DNA damage response, cellular proliferation, and evading cellular death are crucial in OSCC progression. These pathways exhibit a stepwise process, initially altering the normal oral cells to oral epithelial dysplasia and progression into the aggressive and metastatic OSCC. 12 –16,18 In addition, these epigenetically deregulated pathways are considered as a platform to therapeutic interventions to thwart OSCC survival strategy against genotoxic and molecular inhibitor-based approaches. 10,14 –16 In this review, we summarize the recent insights into the epigenetic landscape of OSCC and explore its plausible clinical utility as biomarker and in therapeutic interventions.
Overall genetic landscape in OSCC
The pathophysiology of OSCC is considered as defects at the molecular and cellular level, yet surprisingly there is paucity of knowledge on molecular, diagnosis, and therapeutic intervention. 1 –6 There are several genes that are commonly mutated and/or deregulated in OSCC and therefore may present as important clue to the development and progression of OSCCs in general. 1 –10 In the last decade, there has been resurgence in the interest to analyze several molecular events and proteins involved in OSCC. The several key players/pathways implicated in the molecular settings of OSCC are p53, transforming growth factor beta 1, epidermal growth factor receptor and cyclins, anaplastic lymphoma kinase, wingless homeobox genes (WNT), and mammalian target of rapamycin. 1 –10
Among the class of tumor suppressor genes, phosphatase and tensin homolog (PTEN) is accepted for its crucial role in regulating cell adhesion, cell cycle arrest, migration, apoptosis programming, and differentiation. In a recent gene expression, the study indicates that PTEN expression in OSCC shows loss of heterozygosity (LOH) and diminished level compared to normal mucosa and could be implicated in OSCC carcinogenesis. 7 There is report about association of cornulin (CRNN) gene downregulation with OSCC development. This study suggests that CRNN gene did not show any pathogenic mutation, however, when there was a high frequency of allelic imbalances reported at 1q21.3 region. Further, their data support that downregulation status of CRNN gene could be a good candidate for OSCC prognosis. 6
In a further study to understand the molecular genetics contribution in OSCC, authors indicated for an association between CLPTM1L rs402710C/T and TERT rs2736100A/C single-nucleotide polymorphisms and OSSC risk. Further, their findings suggest that elevated level of tumor relative-threshold level (RTL) values and positive human telomerase reverse transcriptase (hTERT) expression could be used as prognostic markers in OSCC. 8 Inactivation or mutation of the p53 tumor suppressor is a frequent event in tumorigenesis and is regarded as a hallmark of cancer cells. In line with such observation in OSCC, there is evidence on the contribution of mutated p53 and elevated expression of inhibitors of apoptosis proteins in OSCC progression and development. 9,10
The plausible explanation arises from the fact that these carcinogenic settings many unleash genetic and epigenetic disturbances in the normal cellular landscape possibly through chromosomal alteration or mutations in genome as well as genetic materials. 11 –25
The accumulating evidence points out that cohesin complex is responsible for sister chromatid cohesion, where Stromal Antigen 1 (STAG1)/Stromal Antigen 2 (STAG2) is part of the complex, which is regulated by PDS5B. A genetic study suggests about the presence of LOH at STAG1 and STAG2 loci in OSCC. However, both OSCC and normal mucosa showed identical transcript levels of STAG1, STAG2, and PDS5B. 19 The ataxia telangiectasia mutated (ATM) gene is considered to play critical role in DNA damage response, and elevated levels of ATM may be one of the early events in OSCC development. 26 Yap et al. 27 present evidence to show that homeobox only protein homeobox (HOPX) messenger RNA (mRNA) levels are reduced in OSCC, a subset of head and neck squamous cell carcinoma (HNSCC), and may function as a tumor suppressor and substantiate a central role for HOPX in suppressing carcinogenesis. The genetic susceptibility to oral cancer is also suggested due to aberrant rat sarcoma (Ras) pathway, and the prediction that high percentage of Ras mutation in oral carcinoma could be from India. Further, cyclin D1, a downstream member of the Ras pathway, may be overexpressed in the majority of oral cancers and related with poor prognosis. 28
DNA methylation as epigenetic tool in OSCC
Epigenetic mechanisms are considered as pivotal driver in the complex etiology of various cancer types including OSCC. 11 –16,18,19,29 Among several epigenetic modification processes, DNA methylation is considered as central to achieve the state of gene promoter methylation and also global methylation status. 11 –16,18,19,29 To achieve DNA methylation of target sequence, dedicated enzymes have been reported to carry out such regulatory process such as DNA methyltransferase (DNMTase), a family of enzymes with an ability to transfer methyl group to target DNA. Among the family of DNMTase, DNMT1 is reported to be highly active in cancer to silence tumor suppressor genes via hypermethylation of gene promoter. 11 –16,18,19,29 Several findings converge to common understating that DNA promoter methylation of tumor suppressor genes and global DNA hypomethylation may play pivotal role in OSCC. 14 –16,19 Recently, authors provided data that DNA methylation changes in oral epithelial dysplasia can be used as predictive markers of developing OSCC. DNA methylation alterations such as promoter methylation of angiotensin II receptor type 1, forkhead box I2, and proenkephalin and possibly hypomethylation of long interspersed nuclear elements could be used as a predictive biomarkers in oral epithelial dysplasia. 15 The ability of carcinoma to attach with their surrounding niches is considered critical for their progression and metastasis. In view of approaches to disrupt cell-to-cell interaction, authors support the idea that epigenetic event in the form of cytosine-phospodiester bond-guanine (CpG) methylation promoter may disturb the action of E-cadherin and possibly stop the metastatic tendencies of oral carcinoma. 30 In case of OSCC, the epigenetic impact in metastasis process is still not clear. Recently, Clausen et al. 31 reported that WNT1-inducible-signaling pathway protein 1 (WISP1) expression can be regulated by methylation. They proposed that hypomethylation of WISP1 gene supports metastasis of OSCC and can serve as a predictor marker. Despite progress in cancer biology of OSCC, the 5-year survival rates are stagnant at a poor 50%. In light of potential molecular markers in the premalignant oral tissues, evidence suggests that eye absent (EYA) transcriptional coactivator and phosphatase 4 (EYA4) is considered as a tumor suppressor gene candidate and indicated to be epigenetically controlled via DNA methylation and eventually silenced. Further, authors show that epigenetically repression of EYA4 may elicit enhanced cell proliferation and DNA damage repair capabilities in OSCC. 32
Jithesh et al. 12 performed Illumina GoldenGate high-throughput array comprising 1505 CpG loci on DNA extracted from 44 OSCCs and paired normal mucosa. Their data support the differential methylation levels of a number of genes between tumor tissues and matched normal samples. Their findings corroborate the previous and existing idea to describe OSCC among epigenetically manifested genetic diseases and could serve the purpose toward seeking biomarkers and therapeutic intervention. In one of the epigenetic studies, data from the methylation status of the microRNA-196b (miRNA-196b) gene suggest observation of frequent hypomethylation of the CpG islands located upstream of the miRNA-196b gene in the OSCC tissues than in the adjacent normal tissues. Taken together, their findings remark that the epigenetic mode of alterations in miRNA-196b gene expression contributes toward cell migration and invasion, raising the possibility to be seen as a prognostic marker or therapeutic target for OSCC. 33 In recent effort to investigate the implication of epigenetic mechanisms as hypermethylation of certain target gene promoter, Li et al. 34 reveal that apoptosis protease activating factor-1 (Apaf-1) and death-associated protein kinase (DAPK) gene promoter hypermethylation may exist in OSCC and in turn show low gene expression. Furthermore, they showed that treatment with demethylation drug DAC, APAF-1, and DAPK were demethylated and reexpressed. The evidence also supports the hypermethylation event in cadherin 1 (CDH1) gene, which codes for adhesion protein E-cadherin, may act in OSCC carcinogenesis. 35 The role of p14 alternate reading frame (ARF) as a tumor-suppressor gene is suggested in cell proliferation and division and tumor-induced angiogenesis. The epigenetic control is investigated in p14 (ARF) expression and indicates the hypermethylation of p14 (ARF) produces loss of p53 function and deactivation of p21-induced cell proliferation. There is another example of hypermethylation of tumor suppressor gene cyclin-dependent kinase inhibitor p15 (INK4b), which contributes to promote cell cycle progression during the G1 phase in OSCC. Previously, tissue factor pathway inhibitor-2 (TFPI-2) is considered as potential clinical predictor as well as a key regulator in OSCC. However, authors strongly suggest potential epigenetic route to downregulate such tumor suppressor gene TFPI-2 can play important role in oral tumorigenesis and precisely in metastasis. 36
Based on convincing evidence, hypermethylation in promoter regions of genes can mediate swayed gene functions and lead to malignant cellular transformation. In search of potential markers, Khor et al. 37 identified 33 promoter hypermethylated genes that are probably switched off in OSCC and can play role in oral carcinogenesis. In another recent finding, authors explain the methylation profile of key genes in OSCC progression and survival. In case of Wilm’s tumor suppressor gene, promoter methylation may serve for better prognosis and that mutS homolog 6 and GATA-binding protein 5 (GATA5) gene promoter methylation can help in the prediction of worse prognosis. Besides, they also suggest that GATA5 gene promoter methylation is associated with a shorter survival rate. 38 In line with potential tumor suppressor gene pool, there is understating that the PTEN and p16INK4a (p16) genes are modulated by epigenetic alterations specifically through their promoter hypermethylation. To support earlier views, Sushma et al. 39 provide evidence to conclude that methylation can contribute toward low level of PTEN and p16 expression in OSCC progression and may serve for the purpose of prognostification. Cheng et al. 40 attempted to fortify the notion that DNA methylation exists as epigenetic tools in many cancer. They hypothesized that high-degree DNA methylation to the γ-synuclein may be observed in OSCC. Further, they demonstrated the experimental evidence using methylation inhibitor drug 5-azacitidine (5-aza) and confirmed positive outcome in OSCC clinicopathologic features. In an approach to link epigenetic silencing in OSCC, authors provided proof about the epigenetic changes in keratin 13 gene due to hypermethylation and suggested for the development of diagnostic markers and novel therapeutic approaches. 41 The communication between carcinoma and their adherence surfaces is crucial for progression and development. The authors suggested that differential regulation of E-cadherin by epigenetic mechanisms may be linked with OSCC pathophysiology. 42 One of the approaches to show the epigenetic contribution in OSCC, expression of mucosa-associated lymphoid tissue 1 (MALT1) was studied. MALT1 is expressed in the nucleus of oral epithelial cells and can be epigenetically silenced, suggesting that the detection of MALT1 expression is a useful predictive and prognostic determinant in the clinical management of oral carcinoma. 43 The silencing of several genes due to epigenetic regulation is considered as key features in cancer. Koike et al. 44 provide data to suggest the downregulation of four and a half Lin11, Isl-1 & Mec-3 (LIM) domains 1 (FHL1) gene as a frequent event during oral carcinogenesis. Further, they also propose that FHL1 downregulation in OSCCs could be through DNA methylation of the promoter region instead of histone deacetylation or mutation. In view of therapeutic hindrances, radio-resistance poses a problem for radiotherapy of oral cancer. To understand the role of deregulated epigenetic changes in radio-resistance, Lin et al. 45 suggest that transcriptional repression of fragile histidine triad could be through hypermethylation, H3K27me3, and overexpression of H3K27 methyltransferase zeste homolog 2 (EZH2) in OSCC.
Epigenetic role of histone signature in OSCC
The cellular process as posttranslational modifications is pivotal in normal as well as carcinoma to achieve certain regulations and responses. 13 –16 The presence of histone posttranslational modifications is a highly observed phenomenon in OSCC. 13 –16 The scientific insights describe that epigenetic alterations are observed at the N-terminal tails, within each of the four histone complexes (H3, H4, H2A, and H2B). 13 –16 To achieve unique histone code signature within the target cells, several processes are employed such as methylation, acetylation, adenosine diphosphate (ADP)-ribosylation, phosphorylation, ubiquitination, and sumoylation with the precision of amino acid residues incorporated inside the histone tails. 13 –16 Both acetylation and methylation of lysines can activate or inactivate chromatin and transcriptional status of genes. 13 –16 A number of methylated residues and their position within the histone tail denote the status of genes as silenced, inactivated, and switched on. Taken together, the histone signature in the form of potential modifications may correlate with the developmental process of OSCC and other cancer types such as breast, cervical, and hepatocellular carcinoma. 13 –16,46 In line with the plethora of epigenetic process such as DNA methylation, posttranslation modification of histones, contribution of chromatin remodelers are indicated as mechanisms to influences gene expression. 13 –16,47 Recently, wide appreciations are received for chromatin remodeling, which is illustrated as dynamic changes in chromatin architecture to control gene expression at the transcriptional level. It is widely accepted that such remodeling is mostly carried out by players such as covalent histone modifications by specific enzymes, that is, histone acetyltransferases, deacetylases, methyltransferases, and kinases, and adenosine triphosphate (ATP)-dependent chromatin remodeling complexes such as SNF2H and SNF2L. Targeting chromatin remodeling pathways are currently evolving as a major therapeutic strategy in the treatment of several cancers. 13 –16,47
Histone deacetylation catalyzed by various histone decetylases (HDACs) including HDAC1, HDAC2, and HDAC6 plays a significant role during oral carcinogenesis. 13 –16 There are findings to support the overexpression of HDAC2 and HDAC6 in OSCC tissue and its association with advanced tumor stage, tumor size, and metastasis. 48 In one of the studies, a DNA repair protein poly (ADP-ribose) polymerase 1 is indicated to influence chromatin structure modification and can facilitate DNA repair through highly abundant chaperon molecule chromatin assembly factor-1/p60 in OSCC. 19
One of the players in the histone epigenetic alterations is noted as HDAC2 and its expression is highly elevated in OSCC. Further, evidence leads to the idea that HDAC2 can be crucial for migration/invasion potential of OSCC. The authors extend the evidence to support that HDAC2 maintains hypoxia-inducing factor-1α (HIF-1α) stability and can lead to the enhanced cell invasion/migration ability in OSCC. 49 The potential of inhibitor of trichostatin A as HDACs inhibitors is tested on proliferation and apoptosis of OSCC and data confirmed that PTEN/AKT pathway maybe involved observed response. 50 Methylation of H3K4, H3K36, and H3K79 may play a role in activating histone marks that engaged in gene transcription activation. Furthermore, findings support that epigenetic alterations as HDAC carried out by various HDACs are crucial during oral carcinogenesis OSCC and possibly by stabilizing the HIF-1α. 18
Besides epigenetic phenomena as DNA methylation, evidence is available to demonstrate the contribution of deregulated posttranslational modifications as histone methylation to regulate gene expression in OSCC development. 46,48 –53 In the prospects of potential epigenetic players presented as prognostic factor, authors suggest that the histone modification-related proteins, such as aurora kinase B-a serine/threonine-protein kinase of H3S10, G9a, EZH2, and suppressor of variegation 3–9 homolog 1 (H3K9 methyltransferase), may be useful in OSCC early detection excluding suppressor of variegation 3–9 homolog 2. 46 The interest in the study of molecular inhibitors to epigenetic process such as histone modification is appreciated. One group carried out research on the use of aurora kinase inhibitor VX-680 toward histone H3 phosphorylation at Ser10 and indicated inhibition of histone modification in OSCC and subsequent apoptosis. 51 The epigenetic mechanism modulates the chromatin landscape, and one of the possible ways involved methylation of specific lysine residues in histone tails. In particular, findings led to the fact that H3 lysine 4 histone methylation is engaged in transcriptional activation in OSCC. 52 In another study, authors have attempted to establish link between homeobox (HOX) transcript antisense RNA (HOTAIR), a long intergenic noncoding RNA and histone H3K27 methylation and H3K4 demethylation for epigenetic gene silencing to promote cancer metastasis. Their findings indicate that HOTAIR expression in OSCC is enhanced compared with nontumor tissue and may be associated with metastasis. Further, authors delineate the mechanisms of actions of HOTAIR by regulating E-cadherin through binding to EZH2 and H3K27me3 with the E-cadherin promoter. 53
Tumor suppressor and oncogenic miRNA in epigentic contribution
There is an emerging trend in harnessing the highlighted potential of miRNAs in cellular gene regulation steps in both normal and cancer cells. 11,52 –55 miRNAs are elaborated as small noncoding RNAs that posttranscriptionally control gene expression through mRNA degradation or limiting the translation of target mRNAs. 11,52 –55 It is important to note that plethora of miRNAs is reported to become notorious in the sense of acting as onco-miRNAs in OSCC and other carcinoma types. 11,52 –55 It has been suggested that epigenetic modifications and miRNAs are interlinked with each other in the expression pattern of genes such as Ras, p53, and B-cell lymphoma 2. 11,52 –55 Their inherent regulatory role is commonly manipulated in several types of cancer. It is believed that epigenetic factors may be able to dictate miRNome (defined as the full spectrum of miRNAs for a specific genome) profile witnessed in several cancer types such as breast cancer, OSCC, and prostrate cancer. 11,55 On the other hand, players of the epigenetic signature such as DNMTases and HDACs may be placed under the guidance from a dedicated group of miRNAs (defined as epi-miRNAs). Besides, certain miRNAs can indirectly manipulate the expression of tumor suppressor genes associated with epigenetic alterations. The inclusive investigations see the cross talk between miRNAs and epigenetic in the form of Dicer and DGCR8 expression regulation via epigenetic pathway in human carcinogenesis and prospective drugs/inhibitor cocktails. 11 To dissect the molecular-level alterations in OSCC, several miRNAs are reported to contribute to its progression and development, and the findings indicate that miRNA-375, miRNA-127, miRNA-137, the miRNA-200 family, and miRNA-205 are overactivated in OSCC; and their silencing may pave options for therapeutic intervention. 56 In a recent study, the action of several miRNAs including tumor suppressor miRNA-126, an endothelial-specific miRNA is described to implicate in various types of human cancer. 11,52 –55 However, its role in OSCC remains unclear. Yang et al. 57 established that miRNA-126 acts as a tumor suppressor in OSCC cells partially may be by the downregulation of epidermal growth factor-like domain 7, which is critical for progression and angiogenesis.
In search of potential noncoding RNA player, authors suggest the potential of miRNA-17/20a presenting as a tumor migration suppressor and prognostic marker of OSCC patient. 58 Emerging knowledge substantiate that miRNAs deregulation can manage the level of key molecular proteins, which may be involved in the oral cancer progression. Possibly, few oncogenic miRNAs may have an important contribution toward chemosensitivity of OSCC. The work carried out in implication of miRNA-221 to the resistance of human OSCC cells to Adriamycin suggests that sensitivity of OSCC can be accentuated via increased TIMP3 expression. 59 In line with the potential miRNAs contribution OSCC progression and chemoresistance, Zhang et al. 60 suggest that miRNA-32 may act as tumor suppressor agent due to the interference of EZH2-mediated epigenetic regulation in OSCC. Recent study unveiled that miRNA-101 is downexpressed in OSCC and may correlate with zinc finger E-box binding homeobox 1 (ZEB1) expression, lymph-node metastasis, and poor prognosis in OSCC patients. 61,62
Evidence is emerging to show the role of miRNA-125a in the OSCC progression. The process is understood as tumor suppressor role of miRNA-125a by repressing the expression level of estrogen-related receptor α functions as a transcription factor and finally disrupting the prosurvival signaling pathway. 63 Expression of miRNA-216a is reported to be reduced in OSCC. However, overexpression of miRNA-216a can significantly suppress the proliferation, colony formation, migration and invasion of the OSCC, and potential pinpointing of eukaryotic translation initiation factor 4B, which was observed to be upregulated in the OSCC. 64 The prediction about the two tumor suppressor miRNAs, miRNA-329 and miRNA-410, led to experimental evidence, supporting their action to quench Wnt-7b, an activator of the Wnt/β-catenin pathway, and logically dampening Wnt/β-catenin signaling pathway in OSCC. Furthermore, authors extend their study to claim that a demethylation agent and an HDACs inhibitor can reexpress miRNA-329 and miRNA-410 in OSCC to achieve tumor suppressive role. 65 There are data showing higher differential methylation pattern of miRNA-9-1 and miRNA-9-3 in oral and oropharyngeal cancer. Further, they suggest that demethylation treatment using 5-aza-deoxycytidine can restore its expression. 66
In a recent approach, role of tumor suppressor miRNA-143 is reported in clinicopathological factors and prognosis of OSCC patients and suggested to be downregulated. 67 To enrich the evidence to provide the existence of differential expression of miRNAs in OSCC and associated clinical implications, findings such as upregulation of miRNA-1275 and miRNA-144 have been associated with the nodal invasion. 68,69 Besides role of miRNAs in OSCC progression and development, noncoding regulatory sequences can be a part of chemotherapy drug-resistance obstacles. These noncoding regulator sequences may participate in drug-resistance obstacles by overexpressing certain DNA damage response genes responsible for the repair of genomic instability created by drugs. In line with such an observation, Ghosh et al. 70 bring the evidence of resistant-designated miRNA expression signature in OSCC and facilitate the development of cisplatin resistance specifically modulating stem-cell-like and epithelial–mesenchymal transition-type properties. It is a well-accepted view that miRNAs act as regulator and predictor in OSCC. In search of biomarkers in OSCC, Xu et al. 71 provide insights on the importance of miRNA-483-5p abundance in serum of patients to serve as prognostic and diagnostic purpose.
The rapid emergence of data pertaining to the role of various miRNAs is highly witnessed. In line with such attempt, Yu et al. 72 support the oncogenic action of miRNA-9 in OSCC being overexpressed and intercept the Wnt/β-catenin signaling pathway. In addition, authors bring evidence to confirm that C-X-C motif chemokine receptor 4 (CXCR4) gene can be a direct target of miRNA-9 and participate to downplay the activation of Wnt/β-catenin through CXCR4 activation. The clue lies in the observation of differential expression profiles of miRNAs expression in OSCC. To support on such theory, there are reports on the downregulation of miRNA-125b and miRNA-100 in OSCC and possibly linked to the development, progression, and sensitivity to ionizing radiation. 73
In a recent expression profile study of miRNAs in OSCC revealed the contribution of miRNA-155 and miRNA-221 in carcinogenesis and also confirmed the overexpression of miRNA-221. 74,75 Accumulating data on oncogenic miRNAs, there are evidence to link the role of miRNA-29b in OSCC suppression and may be achieved through the downregulation of C-X3-C motif chemokine ligand 1, a cell–cell adhesion regulator. 76 Another prominent research carried out by Deng and Liu 77 showed that miRNA-506 acts as tumor suppressor in OSCC, controlling the expression of GATA-binding protein 6 transcriptional factor, which is indicated to drive growth and metastasis as pro-cancer gene. To delineate the role of miRNAs in OSCC progression and development, Wu et al. 61 observed that miRNA-101 downregulation can inversely impact the ZEB1 expression. In turn, they suggested that elevated expression of miRNA-101 significantly inhibits OSCC cell proliferation, apoptosis resistance, migration, and invasion. Further example of epigenetic contribution is manifested by downregulation of miRNA-99a in OSCC tissues, promoting tumor development, growth, and survival. 78 The role of oncogenic miRNA-17/20a is also described as a prognostic predictor of OSCC patients’ outcome and a tumor migration suppressor miRNA. 58 In a recent study, the action of several miRNAs has been reported for various cellular-signaling pathways. The authors suggest that increased expression of miRNA-146a enhance the oncogenicity of OSCC via modulation of toll-like receptors and cytokine signaling. 79
The right approach to utilize the differential epigenetic signature in OSCC is widely found. One example describes the loss of miRNA-146a expression in high-grade OSCC. The authors dissected out that the transcription factor sex determining region (SRY)-box 2 can be a putative target of miRNA-146a. Finally, they suggest that the loss or decrease in miRNA-146a may be the added characteristics associated with aggressiveness in OSCC. 80 Presently, Xu et al. 81 suggest that downregulated tumor suppressor miRNA-138 may unleash molecular mechanisms through interaction with the 3′-untranslated region (3′-UTR) of Yes-associated protein 1 (YAP1). In addition, they showed that overexpression of miRNA-138 in OSCC reduces the expression of YAP1. Concomitantly, the downregulation of miRNA-138 can inhibit YAP1 expression level.
A detailed differential expression study of miRNAs is reported by Ganci et al. 82 and their findings corroborate the notion of deregulated miRNAs may contribute to OSCC development and predict risk factors. On the other hand, several molecular protein players are produced by the carcinoma for their own growth and survival. In light of such molecular cross talk and regulatory miRNAs and molecular protein actors, Lu et al. 83 show that miRNA-29a can directly target 3′-UTR of matrix metallopeptidase 2 gene and point toward negative regulation. Further, they provide evidence suggesting that ectopic overexpression of miRNA-29a can inhibit OSCC cell invasion. Exploring the diverse regulatory role of miRNAs in OSCC, authors suggest that miRNA-27a can participate in thermal-sensitivity response by targeting the Hsp110 and Hsp90 expression. 84 Recently, Baba et al. revealed added function of mRNA-115-5p, which help in the OSCC metastasis and also suggested its potential as therapeutic intervention molecules. 85 In a similar kind of effort, Wu et al. provide data to claim that miRNA-17-5p can increase or decrease apoptosis-related proteins in irradiated oral carcinoma 3 (OC3) cells and modulate p53 protein expression contributes toward radiosensitivity of the OC3 cells. 86
In a recent study, authors suggest downregulation of miRNA-200b and miRNA-203, and upregulation of miRNA-138 and miRNA-183 are associated with clinical and pathological features of oral cavity and oropharynx SCC tumors. 87 In addition, Xu et al. 88 show that miRNAs are central in energy metabolism and strongly emphasize the role of miRNA-340 to create metabolic perturbations in OSCC via controlling the glucose transporter 1 (Glut1) expression. The deregulation of several regulatory miRNAs is frequent cellular events in carcinoma, and the role of miRNA-92b is also suggested in OSCC via promotion of nuclear factor-κB (NF-κB) signaling axis. 89 Recently, the role of miRNA-188 is elucidated as tumor suppressor in OSSC because its expression is frequently downregulated. The authors suggest that miRNA-188 may regulate the key proteins in OSCC progression, namely, cyclin D1 and matrix metallopeptidase 9. 23
HPV-related epigenomics in OSCC
HNSCC and cervical cancer are the principal malignancies wherein the epigenetic alterations related to HPV have been implicated. As indicated by multicentric studies, the accumulation of aberrant DNA methylation of genes is a characteristic feature of “epigenetic field for cancerization.” Such alterations may be directly induced by viral agents, such as HPV, and their components, or alternatively by chronic inflammation in response to the infections. 90
During malignant transition, integration of high-risk HPV genomes often occurs at fragile locations of the host genome. This integration can happen with a single copy (type I) or a concatemer (type II) of HPV genome. 91 It was shown by Kalantari et al. 92 that the methylation status of locus control region (LCR) in type II integration was relatively more, consistent with hypothesis of an epigenetic silencing of the internal copies of the viral genome. In brief, the epigenetic alterations in the expression pattern of the viral onco-proteins E6 and E7 are noted to cause substantial effects. The primary effects include modulation of DNA methylation, upregulation of the DNMTases, induction of the histone lysine demethylases, and altered expression of miRNAs. 93 Further, evidence suggestive of chromatin modifications has also been put forth, such as increase in expression of EZH2 enhancer and a global loss of H3K27me3 levels. 94
In HNSCC context, several studies have provided significant data regarding the epigenetic changes noted in HPV-positive cases, such as hypermethylated genetic profile and various unique methylated CpG loci. 20,95,96 In a recent study, individual genes/loci [p16 (cell cycle), DAPK, Ras association domain family member 1 (RASSF1; apoptosis), breast cancer 1 (BRCA1), MutL homolog 1 (MLH1; DNA repair), epithelial cadherin (ECAD; cell to cell adhesion), glutathione S-transferase Pi 1 (GSTP1; carcinogen metabolizing), methylated in tumor1 (MINT1), methylated in tumor2 (MINT2), and methylated in tumor3 (MINT3)] related to various pathways of carcinogenesis were investigated for hypermethylation in HPV-positive as well as -negative cases of HNSCC. They observed that promotor methylation of p16, DAPK, RASSF1, and MINT31 was significant with the positive cases. The viral integration seemed to cause CpG island methylation of the tumor suppressor genes and a relatively favorable survival rate. 97 In another study, epigenetic variations in the cyclin-dependent kinase Inhibitor 2A (CDKN2A) locus with changes in its protein expression were noted in HPV-positive oropharyngeal cancer. This is an established tumor suppressor gene in HNSCC and often affected by epigenetic silencing or deletion. 98 Another study on Indian population showed a significant correlation between promoter hypermethylation of PTEN and p16 genes in HPV-positive OSCC tissue samples. 39 Concerning the HPV-negative HNSCC cases, differential expression of GLI1 gene of the Hedgehog-signaling pathway was noted. 99
In addition to methylation, HPV-positive HNSCC has been noted to have distinct miRNA signatures. Some of these miRNAs are also expressed in cervical carcinoma suggestive of common altered pathways for HPV, irrespective of the anatomic site. Few examples of these common miRNAs include miRNA-125a-5p, miRNA-218, miRNA-145, miRNA-143, miRNA-31, and miRNA-21. 21,99 The viral onco-proteins E6 and E7 have significant effects on the expression of miRNAs associated with genes playing pivotal role in malignant transformations. For example, miRNA-200c and miRNA-34a are upregulated, whereas miRNA-574-3p is downregulated with respect to p53 gene; miRNA-15a and miRNA-16 are upregulated with respect to cyclin D1; miRNA-195 and miRNA-497 are downregulated in relation to cyclin E; miRNA-143 and miRNA-145 are downregulated in relation to Ras gene. 21 Specific to HNSCC, there is report on overexpression of has-miRNA-363_st and has-miRNA-21_st and decreased expression of hsa-miRNA-31_st and has-miRNA-193b-star_st was observed in their study. 100 A study by Wald et al. 101 on HPV-positive cells of HNSCC showed relative upregulation of miRNA-363, miRNA-33, and miRNA-497, whereas miRNA-155, miRNA-181a, miRNA-181b, miRNA-29a, miRNA-218, miRNA-222, miRNA-221, and miRNA-142-5p were downregulated. Another study on HPV-positive oropharyngeal squamous cell carcinoma revealed strong expression of miRNA-9, which has a role in E-cadherin modulation and subsequent metastatic potential. 102 The miRNA expression in HNSCC is still under extensive research. In future, developing a validate miRNA panel for HPV-positive squamous cell carcinoma can aid in easy screening tests, diagnostics, and targeted therapeutic strategies. Recently, Zhang et al. 103 show genetic study data to suggest that an insertion (Ins)/deletion (Del) polymorphism (rs3783553) in the 3′-UTR of interleukin 1α (IL-1α) could result in functional and influential susceptibility to HPV16-associated OSCC, particularly for squamous cell carcinoma of the oropharynx. 103 In support of the link between miRNAs regulation and HPV infection, Lajer et al. 104 report that downregulation of miR-375 and upregulation of miR-31 in OSCC are linked to the clinical manifestations due to HPV infection. Recently, Yuan et al. 105 have provided insights in the involvement of E2F1rs3213180 polymorphism toward susceptibility to HPV-associated OSCC.
Inflammation and epigenetic changes in oral cancer
Cancer-associated inflammatory response has been documented a century ago by Rudolf Virchow. 106 This persistent unresolved state of chronic inflammation is splendidly observed in the tumor microenvironment. It comprises of varied components of the immune system modulating the initiation and progression of carcinogenesis. 107 Major mediators of inflammation such as cytokines, cyclooxygenase-2 (COX-2), and NF-κB are known biomarkers detected in several cancers. 108,109 Interestingly, inflammation-associated carcinogenesis provides great therapeutic opportunity in targeted therapy. 110,111 In addition, when stimulated, the immune cells generate mediators such as reactive oxygen and nitrogen species, which subsequently lead to oxidative DNA damage, activation of oncogenes, inactivation of repair enzymes, and so on, aiding in the initiation of carcinogenesis. Ironically, these reactive species are known mediators of apoptotic and necrotic cell death. 112
Besides direct genetic alterations, current research on tumor-associated inflammation has revealed its substantial role in epigenetic control of cancer. CpG and promotor hypermethylation have been observed in inflammatory conditions such as Helicobacter pylori infection, ulcerative colitis, and Barrett’s esophagus, which are noteworthy factors associated with gastric carcinogenesis. Further, inflammation-induced epigenetic alterations may occur via histone modifications, contributing considerably to genetic instability. 109
Oral mucosa is a vulnerable location for various kinds of inflammatory stress including carcinogenesis. 112 At present, limited studies have been conducted to prove the role of inflammatory components in epigenetic alterations associated with oral cancer. One study on oral cancer cell line by Arif et al. 113 discovered a histone modification (hyperacetylation of histone H3K14) which was nitric oxide dependent. As described earlier, reactive nitrogen species are produced by activated immune cells, hence it can be hypothesized that inflammation contributes to such genetic alterations. 18 Another study showed that inflammatory mediator like IL-6 can induce global hypomethylation as well as altered DNA methylation patterns in OSCC cell lines. 114 A study by Shaw et al. 115 showed positive correlation between CpG island methylation phenotype positive oral cancer cases and an increased host inflammatory response. Such cases also exhibited a favorable clinical behavior. To conclude, further valid data are essentially required for determining the pleiotropic role of inflammation in the epigenetics of oral cancer.
Epigenetic intervention in OSCC
The recent findings suggest that the precedence of changes in OSCC, which may not be initially resistant against genotoxic drug treatment, however in course of time due to epigenetic heterogeneity accumulation, these cells eventually will acquire drug resistance capability. 11,13 Therefore, appreciable in vitro, preclinical and clinical attempts are reported to harness the aberrant epigenetic settings in OSCC (Figure 1). 11 –13,116 –126
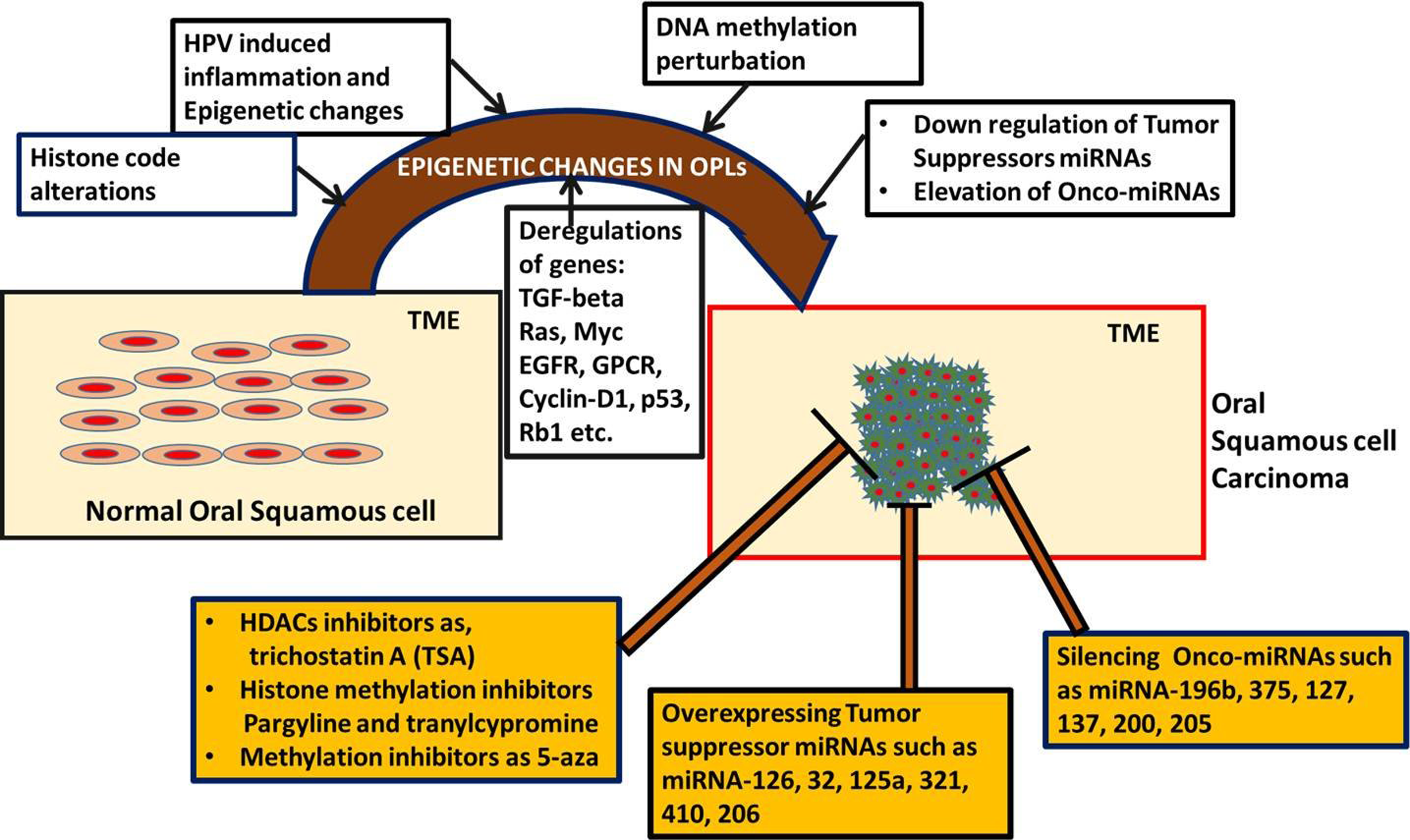
The molecular landscape in oral squamous cell carcinoma progression and development, involving epigenetic regulation, aberrant cell signaling pathways, human papilloma virus infection, and inflammation. Besides aberrant situation of molecular, genetic, and epigenetic settings, several therapeutic options are also reported in the form of inhibitors to epigenetic players and microRNAs as potential agent.
The emphasis on epigenetic therapy is acknowledged by focusing on the HDAC inhibitor, such as phenylbutyrate, to seek their influences to DNA damage repair and survival. The authors gathered evidence to suggest that phenylbutyrate can accelerate DNA repair and survival in normal cells after radiation. Further, they suggested that phenylbutyrate may demonstrate reduction in the oral tumor incidence, burden, and progression and can be interrelated with suppression of onco-miRNAs and Rad51 overexpression. 116
The problem of drug resistance toward genotoxic drugs including cisplatin and dose-related toxicity are two major obstacles in OSCC clinical outcome. To achieve success in chemotherapy, the exploitation of epigenetic deregulation mechanism in OSCC is highlighted to look for combinative treatment regimen. The authors suggest that concomitant treatment with suberoylanilide hydroxamic acid (one of the most potent inhibitors of HDAC) aggravate tumor cell sensitivity to less toxic doses of cisplatin. 117 Recently, Langevin et al. 118 conducted a study to explore the potential biomarkers in the form of epigenetic signature. They identified seven CpG loci appear as novel prognostic biomarkers by noninvasive oral rinse collection technique in OSCC patients. In another effort to support the idea of DNA methylation could be a promising cancer biomarker in OSCC, Huang et al. 119 proposed the detection of paired box gene 1 (PAX1) DNA methylation using oral swabs. This gene is reported to be silenced by methylation in cervical and oral cancers and possibly acts as tumor suppressor gene. 119,127 Recently, findings of Wang et al. 120 support the cancer causing roles of aberrant and high level of lysine (K)-specific demethylase 1A (LSD1) in OSCC. LSD1 is considered to be the first histone demethylase to be identified that demethylates histone H3K4 and H3K9. 10 –16 Further, they indicate that LSD1 abridgment in OSCC cells can result in impaired cell proliferation, migration/invasion, tumorsphere formation, and reduced xenograft growth while inducing cell apoptosis and enhancing chemosensitivity to 5-fluorouracil. Besides the use of LSD1, chemical inhibitors (pargyline and tranylcypromine) induce protein reduction probably via enhanced protein and establish as viable therapeutic inhibitors in OSCC cells. 120 Recently, some clinical trials in the form of biomarker studies and therapeutic interventions are documented (Tables 1 and 2).
List of clinical trials as epigenetic biomarkers and therapeutics.

OSCC: oral squamous cell carcinoma; HPV: human papilloma virus; HNSCC: head and neck squamous cell carcinoma.
List of genes and their physiological functions.

Conclusion
In conclusion, the epigenetic phenomena within normal oral epithelial cells or OSCC are interpreted as the opportunity of turning a gene on and off in perspectives of prognosis and therapeutic intervention, encompassing both molecular inhibitors and genomic insults. During the extensive review, to explore ways to mend the epigenetic landscape of OSCC, new hopes and promises for better drugs/inhibitors combination along with the current regimen of drug therapy were revealed. The prospects are further strengthened by the ongoing interventions at the epigenetic level and the corresponding clinical trials.
Footnotes
Acknowledgments
We would like to thank the Cancer and Translational Research Lab for their constant support and motivation and our host institute Dr D.Y. Patil Biotechnology and Bioinformatics Institute, Dr D.Y. Patil Vidyapeeth, Pune, for providing resources. The authors acknowledge that Cancer and Translational Research Lab are supported by DST-SERB, New Delhi, Government of India, India (SERB/LS-1028/2013) and DPU Pune, India (DPU/05/01/2016).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
