Abstract
Purpose
Deep convolutional neural networks are favored methods that are widely used in medical image processing due to their demonstrated performance in this area. Recently, the emergence of new lung diseases, such as COVID-19, and the possibility of early detection of their symptoms from chest computerized tomography images has attracted many researchers to classify diseases by training deep convolutional neural networks on lung computerized tomography images. The trained networks are expected to distinguish between different lung indications in various diseases, especially at the early stages. The purpose of this study is to introduce and assess an efficient deep convolutional neural network, called AFEX-Net, that can classify different lung diseases from chest computerized tomography images.
Methods
We designed a lightweight convolutional neural network called AFEX-Net with adaptive feature extraction layers, adaptive pooling layers, and adaptive activation functions. We trained and tested AFEX-Net on a dataset of more than 10,000 chest computerized tomography slices from different lung diseases (CC dataset), using an effective pre-processing method to remove bias. We also applied AFEX-Net to the public COVID-CTset dataset to assess its generalizability. The study was mainly conducted based on data collected over approximately six months during the pandemic outbreak in Afzalipour Hospital, Iran, which is the largest hospital in Southeast Iran.
Results
AFEX-Net achieved high accuracy and fast training on both datasets, outperforming several state-of-the-art convolutional neural networks. It has an accuracy of
Conclusion
The AFEX-Net is a high-performing convolutional neural network for classifying lung diseases from chest CT images. It is efficient, adaptable, and compatible with input data, making it a reliable tool for early detection and diagnosis of lung diseases.
Keywords
Introduction
Nowadays, medical image processing plays an essential role in various medical research fields. Among different types of medical images, computerized tomography (CT) is widely used in medical diagnosis and prognosis due to its inexpensive and popular nature, providing valuable medical supplementary information. CT imaging is an imaging method that uses X-rays to create cross-sectional pictures (slices) of a body organ and its structure. CT imaging has the advantage of eliminating overlapping structures and making the internal anatomy more apparent, making it useful in identifying disease or injury within various regions of the body. For instance, brain CT images can be used to locate injuries, tumors, hemorrhage, and other conditions in the head, while lung CT images reveal the presence of tumors, pulmonary embolisms (blood clots), excess fluid, and other conditions such as emphysema or pneumonia in the lungs.
In this article, we focus on lung CT images to classify different diseases using one of the most powerful tools in deep learning, known as convolutional neural network (CNN). With the advancements in CT and other medical imaging techniques, using convolutional neural networks for medical image classification is an interesting research topic.
On the other hand, the 2019 novel coronavirus was first recognized in Wuhan city of Hubei province of China and rapidly spread beyond the Chinese borders, 1 causing common symptoms such as cough, shortness of breath, chest pain, and diarrhea. 2 Given the current pandemic situation, it is crucial to differentiate early between patients with and without the disease. 3 Currently, reverse-transcription polymerase chain reaction (RT-PCR) is the standard and straightforward test for diagnosing COVID-19 infection, 4 which may take more than 24 hours to achieve a test result in practice. 5
CT is a widely suggested tool to be combined as a clinical test besides polymerase chain reaction test 6 for diagnosing the existence and severity of coronavirus symptoms. 7 Although X-ray images can also be used as the second imaging tool in COVID-19 diagnosis (such as studies done by Ucar and Korkmaz, 8 Pereira et al., 9 Yi et al., 10 Tuncer et al., 11 and Yamac et al. 12 ), using CT besides the RT-PCR test has many advantages. For example, peripheral areas of ground glass, which are a hallmark of early COVID-19, can be detected in CT images, while they can easily be missed in chest X-rays. 13
After CT scanning, the chest CT image should be interpreted by a radiologist to detect the symptoms, which can be a tedious task and may lead to unintentional mistakes when several patients are available. Here, machine learning systems trained based on these images can be of great help in easing medical decisions and saving time. Currently, many successful deep models have been developed to perform classification and segmentation tasks on CT images and can diagnose abnormalities in images without radiologist intervention.
As an example, Castiglione et al. 14 designed an optimized CNN to classify patients into the infected and non-infected categories based on the CT images. In another attempt, Singh et al. 15 proposed a novel deep convolutional algorithm based on multi-objective differential evolution (MODE) to classify COVID-19 infected patients from non-infected ones. Additionally, Yang et al. 16 proposed an architecture based on the DenseNet model to classify lung CT images into COVID-19 and health classes, and misclassification cases in the test dataset were assessed by a radiologist. Despite the excellent performance of these models in classifying lung CT images and detecting COVID-19 symptoms in patients, they provide no information about other abnormalities that can be diagnosed using these images.
In a different research conducted by, 17 a fully automatic deep learning system with a transfer learning approach was proposed for the diagnosis and prognosis of COVID-19 disease. This deep model was first trained using CT images with lung cancer, and then COVID-19 patients were enrolled to re-train the network. The trained network was externally validated on four large multi-regional COVID-19 datasets. Although this research has achieved interesting results, its training was not done using COVID-19 CT images.
There are some successful research studies that have attempted to distinguish between COVID-19 and other lung diseases. For example, the work done by Pu et al. 18 used patients with positive RT-PCR tests as samples from the COVID-19 class and patients with community-acquired pneumonia (CAP) as samples of the other class, and proposed a three-dimensional (3D) convolutional model to classify these two classes. Additionally, Xu et al. 19 proposed a deep learning system to screen CT images in three steps. First, abnormal regions were segmented using a 3D deep learning model, then the segmented images were categorized into the three classes of COVID-19, Influenza-A viral pneumonia, and healthy by the ResNet model. Finally, the infection type and total confidence score of each CT image were calculated with the Noisy-or Bayesian function.
Heidari et al. 20 proposed the use of preprocessing algorithms to enhance the performance of CNN in predicting the likelihood of COVID-19 from chest X-ray images. Similarly, Wang et al. 21 developed a tailored deep CNN design, called Covid-Net, specifically for the detection of COVID-19 cases from chest X-ray images. Their study demonstrated the potential of CNN in accurately identifying COVID-19 cases. Gulakala et al.22,23 utilized generative adversarial networks (GANs) for data augmentation and progressively growing GAN for CNN optimization, respectively, to achieve rapid diagnosis of COVID-19 infections. These studies highlighted the importance of data augmentation techniques in improving the performance of CNN for COVID-19 detection. Liang et al. 24 and Uddin et al. 25 also explored the use of CNN for automated detection of COVID-19 from medical images, further supporting the potential of CNN in this application. Waheed et al. 26 proposed the use of auxiliary classifier GAN for data augmentation to improve COVID-19 detection, demonstrating the effectiveness of GAN-based techniques in enhancing CNN performance for this task.
In a more general and interesting research, Ardakani et al.
27
used 10 different deep CNNs to classify chest CT slides of patients into three classes. Their research aimed to compare the capabilities of different deep networks in confronting pulmonary infections. Their study showed that the ResNet-101 and Xception networks achieved the best performance; both had an area under the curve (AUC) of
It is worth mentioning that although the diagnosis of COVID-19 is critical, the importance of identifying other lung diseases, such as lung cancer, should not be ignored. According to WHO reporting, lung cancer has been the leading cause of death worldwide in recent years 28 . Although the risk of other infections in patients with lung cancer is still to be determined, 29 these patients are more vulnerable to other infections than others due to inherent characteristics and care needs. Additionally, ground-glass opacities or patchy consolidations, which are the most frequent indications of COVID-19 in chest CT images, 30 may also appear in the early stages of radiotherapy pneumonitis (RP) in patients with lung cancer and are not easily distinguishable from each other.
The research conducted by Ibrahim et al. 31 was the first study to utilize a multi-classification deep learning model for diagnosing COVID-19, pneumonia, and lung cancer diseases. In this study, the authors did not have access to a dataset that included images from these classes, and CT images of different classes were collected from various public sources. They evaluated five deep architectures to classify these public chest X-ray and CT datasets, and based on their experiments, the VGG19+CNN model achieved the best result. Although their work was pioneering and valuable, it is uncertain whether images collected from different sources and imaging devices can form a compatible and reliable dataset together. In other words, images collected from each medical imaging device encompass certain characteristics related to that device that might interfere with learning algorithms.
Based on experiments conducted by Torralba and Efros, 32 even a single-source dataset appears to have inherent bias. Additionally, according to research by López-Cabrera et al., 33 if there is any bias in the collected images, such as corner labels and typical characteristics of a medical device, the classification model might easily learn to recognize these biases in different classes rather than focusing on the main features separating the different classes.
Therefore, in this article, we aim to address the global challenge of classifying chest CT images using a lightweight and fast deep CNN-based architecture trained on an unbiased dataset. The proposed CNN architecture is an adaptive version of the one developed by Gao et al. 34 for classifying CT brain images. To the best of our knowledge, our proposed method (illustrated generally in Figure 1) is the second supervised deep learning model in the area of multi-class CT chest images classification (after the work done by Ibrahim et al. 31 ), where COVID-19, lung cancer, and normal images are included (see samples in Figure 2). However, an important aspect of our work is that all the 10,715 chest CT images were collected from one imaging device with the same device configuration to limit the risk of bias in images and achieve a reliable dataset.

Overview of the framework of the proposed method. It has been investigated on two different datasets, the first one (which is a collected dataset by authors) has three COVID-19, cancer and normal classes while the second one has only two COVID-19 and normal classes.
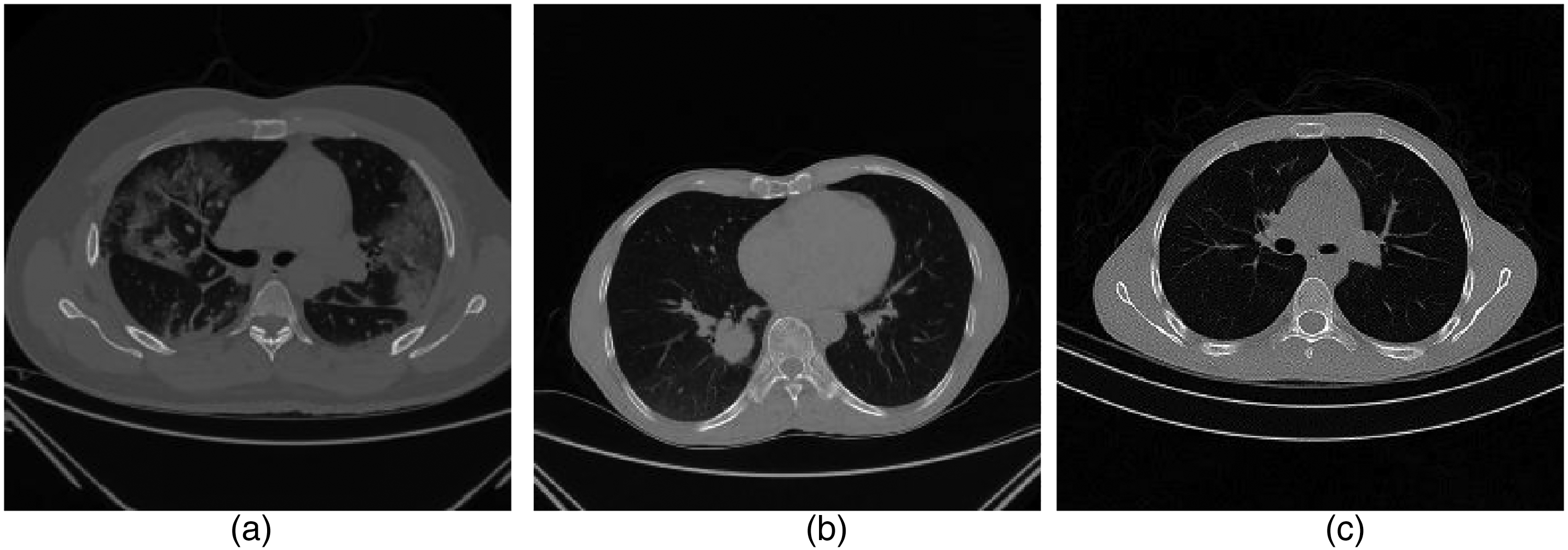
The example of computerized tomography (CT) chest images from subjects showing: (a) COVID-19, (b) cancer, and (c) normal images.
In short, the main contributions of this work are as follows:
A novel lightweight CNN-based model with Adaptive Feature EXtraction layers (AFEX-Net) is suggested for the classification of chest CT images into three classes. The proposed network has an adaptive pooling strategy with adaptive activation functions, increasing model robustness. The proposed network has few parameters compared to other CNN models used in this area (e.g. ResNet50 and VGG16) with faster training while preserving the accuracy. The low computational time of the proposed model makes it highly attractive in the clinic. The proposed model has been evaluated on collected chest CT images from one origin to limit the learning risk of bias.
Hence, three significant properties of the proposed network (low number of parameters, adaptivity, and robustness) make it an easy-to-train and efficient network to be used with clinical data.
The rest of this article is organized as follows: In the “Methodology of AFEX-Net, the proposed deep model” section, we have explained (a) the preprocessing step used to make an unbiased dataset (“Image preparation” section), and (b) the proposed deep CNN in details (“Proposed deep CNN for chest CT images classification” section). The “Evaluation” section provides the obtained results from the proposed network and discusses its performance compared to other efficient networks. Finally, conclusion is presented in the “Conclusion” section.
Methodology of AFEX-Net, the proposed deep model
This section will focus on the proposed model and its characteristics. The proposed model is developed based on lung CT images collected from the radiology section of Afzalipour Hospital—the biggest hospital in Southeast of Iran—in Kerman, Iran, during 6 months. This model is the result of a cooperation between Kerman University of Medical Sciences and Shahid Bahonar University of Kerman.
Based on Figure 1, there are two major steps in the proposed approach: (1) preparing the chest CT images by removing the background and probable biases and enhancing the dataset reliability, and (2) applying and evaluating the proposed AFEX-Net on the dataset. These two steps are explained in “Image preparation” and “Proposed deep CNN for chest CT images classification” sections meticulously.
Image preparation
Deep CNNs consist of many layers with a vast number of neurons that can automatically extract valuable features from images. These extracted features are then utilized for abnormality detection or the classification of medical images by the network. 35 Although there is a belief that CNNs are powerful feature extractors, the bias in the input images can influence their outcome by diverting the network’s attention to irrelevant features. According to Figure 2, each slice of the CT image contains extraneous curves and noises outside the chest region, which can lead to biases during the learning process. Therefore, this study aims to perform background subtraction by eliminating the out-of-chest region to prevent undesirable biases.
An interesting work conducted by Moldovanu et al. 36 suggests a robust algorithm for removing non-brain tissues in MR images. Here, the idea of the mentioned algorithm is utilized and modified to be suitable for preparing the chest CT images (see Figure 3).
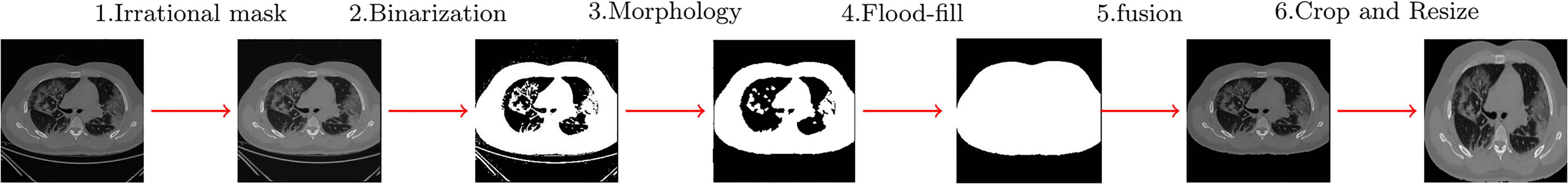
Preprocessing steps done on computerized tomography (CT) images.
The proposed algorithm has four basic steps as follows:
Applying an irrational mask based on the Gregory-Leibniz infinite series to the input image. Applying an image binarization method on the previously filtered image using a proper and adapted binarization threshold. Eliminating the undesired tissues from the binary image using morphological operations (using proper and adaptive disc size) and applying the flood-fill algorithm to fill holes in the image. Cropping the final images (removing the black background) and resizing the resulted images to a unique size containing only the chest area.
Figure 3 depicts a chest CT image with the background removed. These modifications ensure that all chest CT images are placed in an unbiased condition for use by the proposed CNN. This approach allows for network training without concern for deviation caused by focusing on unnecessary elements in the background.
Proposed deep CNN for chest CT images classification
Deep CNNs are well-known supervised methods that have proven their efficiency in many areas, including classification and segmentation tasks on medical images. Here, we have proposed an adaptive deep CNN, whose architecture is inspired by the one developed by Gao et al. 34 Our proposed network is adaptive and more general, bringing adaptation to the network to make it applicable in different situations.
Obviously, hyperparameter tuning is essential to adapt a previously developed network for a different purpose and to get the most out of it. CNNs have many hyperparameters that can affect the network performance, such as the number of convolution layers,37,38 pooling layer,39,40 size of mini-batches, 41 neurons’ activation function,19,42 etc. There are many researches devoted to investigating the influence of these parameters. This article focuses on the role of pooling operation and activation functions in a deep CNN to make them adaptive, as they introduce non-linearity in the network and boost its performance in confronting complex data.
There are some traditional activation functions used in networks, such as sigmoid, tanh, and the ReLU family, with the latter being the most popular. Nonetheless, using adaptive activation functions seems a better idea because they have higher learning capabilities than the traditional ones and can adapt themselves to the training data. Also, they improve the convergence rate, especially at early stages in training, as well as the network accuracy. 43 Using adaptive activation functions, it is possible to have smaller networks with fewer parameters while positively affecting the efficiency and accuracy of the network, 44 which seems very interesting.
Based on the explanations mentioned above, in this article, we have developed a CNN with an “Adaptive Feature EXtraction” box called AFEX that consists of adaptive activation functions and an adaptive pooling layer.
The adaptation parameters should be learned; therefore, they participate in the back-propagation algorithm and get tuned during the learning process.
The proposed AFEX-Net consists of 21 different layers, including the input layer, feature extraction layers (convolution layers (Conv), adaptive activation function (AAF), and adaptive pooling layers (A-Pool)), and classification layers (flatten layer (F), fully connected layer (FC), and softmax function (S)), arranged as illustrated in Figure 4.

The framework of AFEX-Net for the classification of COVID-19, cancer, and normal CT lung images includes the following components: BN (batch normalization), Conv (convolution layer), A-Pool (adaptive pooling layer), AAF (adaptive activation function), D (dropout), F (flatten), FC (fully connected layer), and S (softmax classifier). AFEX-Net: adaptive feature representation enhancement network; CT: computed tomography.
In this figure, batches of input images of size
The essential feature extraction stage is associated with the convolution layer that extracts the most related features of each image. The proposed network is based on six convolution layers; the first four are followed by three adaptive activation function layers, while the last two utilize rectified linear unit (RELU) activation functions. Each adaptive activation function layer is followed by an adaptive pooling layer itself.
The classification part of the proposed network starts with a flatten layer (which converts the last two-dimensional (2D) convolution layer into a one-dimensional vector), then a fully connected layer and softmax layer are used. The softmax function calculates the likelihood of each class for a given input image and classifies the input image into the three alternative classes. Furthermore, we used two dropout layers as a regularization method after the two last convolution layers to prevent the overfitting problem. Details of all the layers used in the proposed CNN and related parameters are summarized in Table 1.
CNN architecture used in AFEX-Net.

BN: batch normalization, Conv: convolution layer, A-Pool: adaptive pooling layer, AAF: adaptive activation function, FC: fully connected layer. AFEX-Net: Adaptive Feature EXtraction network; CNN: convolutional neural network.
Total params: 4,453,127.
Trainable params: 4,451,909.
Non-trainable params: 1218.
Just for the purpose of comparison and to demonstrate the efficiency of the proposed model, the well-known VGG16 47 and ResNet50 48 deep models were selected, as they are recognized for their powerful feature extraction capabilities. The VGG16 is a CNN with 23 layers and a total of 107,008,707 trainable parameters. The ResNet50 is also a CNN that utilizes 50 layers for feature extraction from images, with 23,587,587 parameters, including 23,534,467 trainable and 53,120 non-trainable parameters. Therefore, the proposed AFEX-Net, VGG16, and ResNet50 models will be trained using the chest CT images. Table 2 provides a brief comparison of these three CNN models based on the number of network parameters and the time required for training these networks.
Comparing the three AFEX-Net, ResNet50, and VGG16 convolutional networks based on the number of parameters and training time.

AFEX-Net: Adaptive Feature EXtraction network.
As shown in the table, AFEX-Net has almost 5 times fewer parameters than ResNet and 24 times fewer parameters than the VGG16 network. Comparing these three networks, it is evident that AFEX-Net is a fast and lightweight network that can be tailored for various medical image classification purposes. These networks will be further compared in detail in the next section.
Evaluation
In this section, we will describe the collected chest CT images used as the dataset, the conducted experiments on the dataset using the three AFEX-Net, VGG16, and ResNet50 models, and discuss the obtained results.
Dataset
COVID-Cancer-set (CC)
The CT images were collected from the radiology section of Afzalipour Hospital in Kerman, Iran. This hospital is the largest in southeast Iran and has been the primary academic and health department for COVID-19 since the pandemic outbreak in the area. The collected dataset, which we refer to as COVID-Cancer-set (CCs), comprises 10,715 2D chest CT images from 46 patients. These are divided into 4883 slices of COVID-19, 3047 slices of lung cancer, and 2785 slices of normal images. It is important to note that images with closed-lung mode were excluded from the dataset, as well as COVID-19 slices without any lung infections. All processes were monitored by a medical expert.
The images were randomly divided into training and test sets, with 6719 2D slices used for training, and the remaining 3996 slices (including 1954, 1067, and 975 2D slices of COVID-19, lung cancer, and normal, respectively) reserved for the test set.
COVID-CTset
COVID-CTset is a publicly available dataset obtained from.
49
It contains 2282 COVID-19 images from 95 patients and 9776 normal images from 282 persons, all in TIFF format and with a resolution of 512
Experimental setup
To use the adaptive activation function mentioned in relation 1,
Models’ training parameters for CC dataset.

AFEX-Net: Adaptive Feature EXtraction network; CC: COVID-Cancer-set.
The initial value for neuron weights was sampled from a truncated normal distribution
The three models were implemented using Python 3 and Keras 51 with TensorFlow 52 backend as the deep learning framework and were run on Google Colab 53 with 358.27 GB storage, 12 GB RAM, and an NVIDIA Tesla K80 GPU processor.
Performance metrics
The performance of the three models was evaluated based on accuracy, sensitivity, specificity, and precision metrics, formulated as follows:

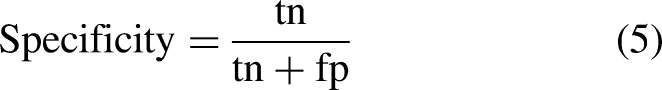

Sensitivity or recall evaluates a model’s ability to correctly predict positive samples for each available category of data or determine images with a specific disease. In contrast, specificity evaluates a model’s ability to correctly exclude negative samples that do not have a given disease. Additionally, precision is used to be more confident about the predicted positive samples by preventing the occurrence of false negatives.
To complete the evaluation criteria, the categorical cross-entropy loss function was also calculated for the three mentioned models, as given by: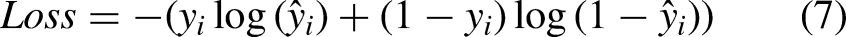
Moreover, a confusion matrix is presented for each model to better visualize the performance of that model in confronting chest images from the three available classes.
Results
CCs dataset
Using the evaluation metrics mentioned in the “Performance metrics” section, this section provides the obtained results from VGG16, ResNet50, and the proposed AFEX-Net models. However, to show the efficiency of the proposed network architecture, we also use the AFEX-Net* model in comparisons, which uses the AFEX-Net architecture without applying the adaptive layers. This allows for a fairer comparison among the architecture of these models, where the lightweight AFEX-Net competes for the big networks such as VGG16 and ResNet50, and no adaptation was used.
All these networks have been trained and evaluated several times, using the random division of images as described in the “Dataset” section. The results reported in the following figures and tables are based on the average of each network performance in these runs.
Figure 5 illustrates the accuracy and loss charts obtained during the network training of the four AFEX-Net, AFEX-Net*, VGG16, and ResNet50 models. As can be seen in this figure, the models seem similar in training, and there is no meaningful difference among their final results; all get converged in 100 epochs to very near accuracy and loss values. However, taking a closer look at this figure, AFEX-Net has performed better than other networks in terms of accuracy, as was expected from its adaptive characteristics. Also, ResNet50 has achieved higher accuracy and less loss than other models in lower epochs, which might be because of using residual blocks in its heavy architecture. Nevertheless, it is worth mentioning again that based on Table 2, the training time for the proposed architecture in AFEX-Net and AFEX-Net* is much lower than the training time for the two other models. This should be considered a privilege for the proposed architecture, making this architecture suitable to be used in systems with limited hardware facilities or an embedded network in the medical platforms.

The training accuracy (left) and training loss (right) are compared in the four utilized models on the COVID-Cancer-set (CCs) dataset. The horizontal axis shows the number of epochs, while the vertical axis displays the model accuracy and loss in the left and right plots, respectively. (a) Training accuracy; and (b) training loss
Although network behavior in the training phase is essential, the network efficiency in confronting unseen samples plays a prominent role in judging its capabilities. Table 4 presents the average results obtained from evaluating the mentioned four models on the test data. For a visualized comparison of the obtained results, refer to Figure 6. In this table, sensitivity, specificity, and precision measures for the COVID-19, cancer, and normal classes were calculated for each model. Additionally, the overall loss and accuracy for each model were assessed by averaging the three classes.

Visual comparison among the average performance of four models on the test set of COVID-Cancer-set (CCs) dataset.
Average of the obtained results from the four models on test data of CCs dataset (best obtained results are in bold).

AFEX-Net: Adaptive Feature EXtraction network; CC: COVID-Cancer-set.
* AFEX-Net without adaptive layers.
As can be seen in Table 4, the proposed AFEX-Net averagely has the highest accuracy and the lowest loss on test data compared to the other three networks. AFEX-Net* and VGG16 networks are very similar to AFEX-Net in terms of overall accuracy, while they have higher losses than that. Also, this table reveals a fantastic finding on the performance of the ResNet50 network; although the behavior of this network was hopeful during the training phase (see Figure 5), its performance on test data is not remarkable. In fact, it has lower accuracy and much higher loss than the other three evaluated networks; its accuracy is about
Figure 6 clearly visualizes how the performance of AFEX-Net architecture in terms of sensitivity, specificity, precision, and accuracy was the highest compared to the other networks. Again, this figure emphasizes that the ResNet50 network is not successful in distinguishing the COVID-19 and cancer images because the precision of this network is much lower than its specificity, especially in cancer images (see the ResNet50 row in Table 4 for details). Totally, based on the obtained results from CCs dataset during training and test of these models, it is evident that AFEX-Net, with its lightweight architecture and fewer parameters, was able to provide the most remarkable results compared to already existing and successful models. The proposed AFEX-Net performance in all terms of the evaluation metrics was effective and superior. It achieved
COVID-CTset
As a secondary analysis, we applied our proposed model to the COVID-CTset dataset and compared it against other state-of-the-art models. Figure 7 provides a clear comparison between the accuracy and loss curves of AFEX-Net and AFEX-Net* in the COVID-CTset dataset. Interestingly, the adaptive layers yield better results, particularly in the initial epochs when there is limited information for learning. This demonstrates that although our proposed models achieve similar convergence results at the end of learning, the adaptive version performs significantly better on unseen data, as shown in Table 5. This table presents the averaged values of the obtained results on the test set from our proposed models, in comparison to the models by Rahimzadeh et al. 49 and Nguyen et al., 54 which trained their models on this dataset.

The training accuracy (left) and training loss (right) comparison of AFEX-Net and AFEX-Net* on the COVID-CTset dataset. The horizontal axis shows the number of epochs, while the vertical axis shows the model accuracy and loss in the left and right plots, respectively. (a) Training accuracy; and (b) training loss. AFEX-Net: AFEX-Net: adaptive feature representation enhancement network.
Average of the obtained results from the six models on test data of COVID-CTset dataset (best obtained results are in bold).

AFEX-Net: Adaptive Feature EXtraction network; CC: COVID-Cancer-set.
* AFEX-Net without adaptive layers.
The visual comparison of our proposed models along with the other four models is presented in Figure 8. AFEX-Net achieved the best sensitivity, precision, and accuracy results and the second-best result for specificity.
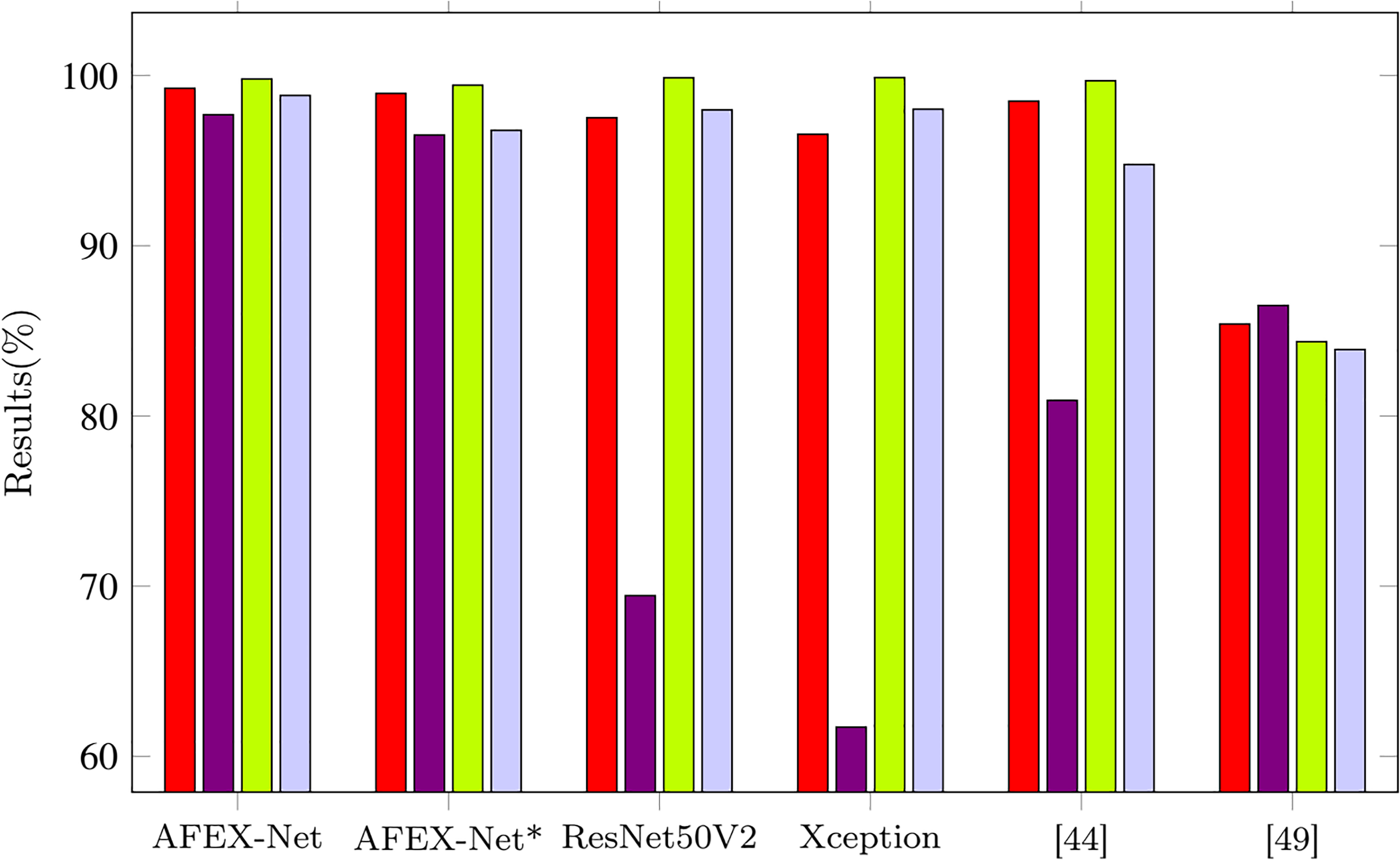
Visual comparison among the average performance of six models on the test set of COVID-CTset dataset.
In Table 5, we can see that AFEX-Net performs noticeably on the COVID-CTset dataset in terms of both accuracy and sensitivity of the COVID-19 category, where it obtains 0.76% and 0.71% improvement compared to the second-best model proposed by Rahimzadeh et al. 49
The confusion matrices for maximum accuracy obtained from the four models on the CCs dataset and the two versions of AFEX-Net on the COVID-CTset dataset are presented in Figures 9 and 10, respectively, to have more scrutiny look at classification results by these trained networks.

Confusion matrices for the four trained networks on the CCs dataset: (a) confusion matrix for the proposed AFEX-Net; (b) confusion matrix for the proposed AFEX-Net*; (c) confusion matrix for the ResNet50; and (d) confusion matrix for the proposed VGG16. AFEX-Net: AFEX-Net: adaptive feature representation enhancement network; CC: COVID-Cancer-set.

Confusion matrices for the two trained networks on the COVID-CTset dataset: (a) AFEX-Net; and (b) AFEX-Net*. AFEX-Net: AFEX-Net: adaptive feature representation enhancement network.
Based on Figure 9(a), one can infer that there were rare COVID-19 samples (just four among 1954 images) that were misclassified as cancer and normal in the proposed AFEX-Net model. Just two cancer images were classified as COVID-19, and just one sample of normal images was not correctly predicted. Therefore, AFEX-Net can be considered a perfect network for this classification. To see the impact of adaptive layers used in AFEX-Net on network performance, we have to compare the AFEX-Net confusion matrix with the one resulting from AFEX-Net* and shown in Figure 9(b). Here, 13 COVID-19 samples got the wrong cancer label, while seven cancer samples were wrongly considered as COVID-19. Although these differences are numerically subtle, they are important due to the consequences of misdiagnoses in medicine. Undoubtedly, these differences mirror the role of adaptive pooling layers and adaptive non-linear activation functions used in AFEX-Net.
Figure 9(c) demonstrates the ResNet50 model confusion matrix, where 57 COVID-19 samples (
Finally, the confusion matrix of the VGG16 model is shown in Figure 9(d). As was mentioned before (due to Table 4), this network achieved the second-best result in COVID-19, cancer, and normal image classification. Also, VGG16 achieved the best classification in the cancer category with no misclassification. Although VGG16 performance for the aim of lung images classification is reasonable, it is worth mentioning again that it is a massive network with many parameters, and accordingly, it requires a long training time and equipped hardware. Regarding the confusion matrix obtained from the COVID-CTset (see Figure 10), it is evident that the AFEX-Net classified both classes robustly, even in the presence of imbalance in data.
Discussion
Results of the evaluation on two datasets demonstrate that the proposed AFEX-Net can be considered a small, lightweight CNN with adaptive feature extraction layers for classifying lung CT images into COVID-19, cancer, and normal categories. Furthermore, the parameters of AFEX-Net are reasonably efficient (almost 4.5 thousand) for implementation in medical clinics. It is hopeful that this developed network can be re-tuned for successful application in similar contexts.
The limitation of this study is that the results obtained by the proposed network are specific to the dataset used. No other dataset was found to include both COVID-19 and lung cancer (or other lung diseases) images. Therefore, the performance of the proposed method may vary slightly when applied to other datasets containing images from different lung diseases.
The future directions for research could include the following:
Validation and testing: Further validation and testing of AFEX-Net on a larger and more diverse dataset to assess its performance across a wider range of lung diseases and conditions. This could involve collaborating with multiple medical institutions to gather a more comprehensive dataset that includes a variety of lung diseases, including rare and less common conditions. Clinical trials: Conducting clinical trials to evaluate the real-world application of AFEX-Net in medical settings. This would involve assessing its effectiveness in aiding radiologists and clinicians in making accurate diagnoses and treatment decisions based on chest CT images. Interpretability and explainability: Exploring methods to enhance the interpretability and explainability of the classifications made by AFEX-Net. Understanding the features and patterns that the network uses for classification could provide valuable insights for medical professionals and improve trust in the system. Generalization to other modalities: Investigating the potential for generalizing the approach to other medical imaging modalities, such as X-rays or MRI, to create a more comprehensive diagnostic tool for respiratory diseases. Integration with clinical workflow: Exploring the integration of AFEX-Net into the clinical workflow, including considerations for user interface design, data security, and regulatory compliance to ensure seamless and safe adoption in medical practice. These future research directions would contribute to the continued development and refinement of AFEX-Net as a tool for early detection and diagnosis of lung diseases, ultimately improving patient outcomes and advancing the field of medical imaging and artificial intelligence.
Conclusion
This work presented a rapid lightweight CNN with adaptive feature extraction layers, called AFEX-Net, to classify the lung CT images. To get sure about the fairness and unbiasedness of images, a robust preprocessing step was applied to the lung CT images to remove any noise and unrelated background. Then the proposed AFEX-Net was trained, which included adaptive pooling layers and adaptive nonlinear activation functions. The role of adaptive layers is to help the network adapt its con uration to the input data and learn the required features faster and more accurately.
Two datasets were utilized to investigate the performance of the proposed model. The CC dataset is a non-public one containing three classes of COVID-19, lung cancer, and normal images. Concerning the importance of distinguishing between COVID-19 and cancer from CT images, this study was the first attempt to collect the COVID-19 and lung cancer images from one origin to eliminate probable biases in images and the obtained results. The COVID-CTset dataset is a public dataset containing two CONID-19 and normal classes. According to the two utilized datasets, the proposed model was then benchmarked against three networks: the non-adaptive version of AFEX-Net and the two state-of-the-art ResNet50 and VGG16 networks for the CC dataset and five networks, the non-adaptive version of AFEX-Net and other four state-of-the-art models by Rahimzadeh et al. 49 and Nguyen et al. 54 for CODID-CTset. Considering several evaluation metrics, AFEX-Net achieved higher results than the its non-adaptive version and all the other models, regarding the influence of adaptive layers and feature extraction. Although the performance of the VGG16 network was very similar to the proposed adaptive AFEX-Net in terms of evaluation metrics, the privilege of our proposed model relies upon having much fewer parameters (almost 24 times fewer parameters) and low computational complexity (3 times faster training). Hence, being lightweight, accurate, and fast make AFEX-Net a unique method for diagnosing the abnormalities in chest CT images. Also, it can be successfully re-tuned to get applied in similar applications.
Footnotes
Contributorship
The authors confirm contribution to the paper as follows: Roxana Zahedi Nasab: study conception and design; Mahdieh Montazeri: data collection; Sobhan Amin: data labeling; Hadis Mohseni and Fahimeh Ghasemian: analysis and interpretation of results; Roxana Zahedi Nasab: draft manuscript prepration. All authors reviewed the results and approved the final version of the manuscript.
Data availability
The data and code that support the findings of this study are available upon request.
Declarations of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
This paper was approved by the Ethics Committee of Kerman University of Medical Sciences (IR.KMU.REC 99000303).
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Informed consent
In this study, we only extracted the necessary data items from the patients’ records, and the demographic information of the patients was not extracted.


