Abstract
Background:
Hepatitis B virus (HBV) remains highly endemic in many low- and middle-income countries (LMICs), where challenges such as antiviral resistance and immune/vaccine-escape mutations complicate disease management.
Objective:
This review synthesizes current data on HBV drug resistance, including the prevalence and distribution of resistance-associated mutations (RAMs), vaccine-escape mutations (VEMs), and immune-escape mutations (IEMs) in LMICs.
Design:
Systematic review and meta-analysis.
Data sources:
PubMed, Scopus, and Web of Science.
Methods:
A systematic search was conducted from 2014 to 2025 (last search date: August 04, 2025), and data were extracted from eligible studies. A random-effects meta-analysis was conducted to estimate pooled resistance rates and prevalences.
Results:
A total of 43,834 participants from 47 studies conducted in 28 LMICs were included. The overall antiviral resistance rate was 7.87% (95% CI (5.13–11.11)), with very high heterogeneity (I2 = 99.7%). Lamivudine (3TC) and telbivudine (LdT) had the highest resistance rates. Asia consistently reported the greatest resistance, with rates of 32.45% for 3TC and 51.92% for LdT. In a subgroup analysis on economic regions, Lower-middle-income countries showed the highest pooled resistance rate at 10.61% (95% CI (5.21–17.51); I2 = 96.9%). The most frequently detected RAMs were rtM204V/I, rtV173L, and rtL180M. The pooled prevalence of vaccine-escape mutations was 8.92% (95% CI (5.02–13.64); I2 = 78.4%). Immune-escape mutations were much more common, with a pooled prevalence of 55.66% (95% CI (28.10–81.53); I2 = 98.1%). The most prevalent mutations were sG145R/A/K, sQ129H/R, sD144E/A, sS143L/T, and sP120S/T. The pooled estimates in this study are reported and interpreted descriptively, given the substantial heterogeneity observed.
Conclusion:
The remarkably high drug resistance rates and prevalence of immune and vaccine-escape mutations suggest notable burdens of resistance, especially for drugs with low genetic barriers such as lamivudine and telbivudine, underscoring the growing challenges in achieving the WHO 2030 reduction goals, and highlighting the need for enhanced surveillance, adoption of potent antiviral regimens such as tenofovir disoproxil fumarate, and continuous vaccine efficacy monitoring. However, the results should be interpreted with caution due to the very high heterogeneity observed in the study.
Trial registration:
International prospective register of systematic reviews (PROSPERO) under the ID: CRD420251030302.
Plain language summary
Hepatitis B virus (HBV) infection is considered a major public health problem, especially in low- and middle-income countries (LMICs). Managing HBV becomes challenging when the virus acquires mutations that confer resistance to antiviral therapy. Similarly, certain mutations classified as vaccine or immune-escape variants enable the virus to evade vaccine-induced or naturally acquired immunity. This study reviewed and combined existing research to understand how common these mutations are and the extent of drug resistance, vaccine and immune-escape in LMICs.
The research team searched scientific databases for studies that reported HBV drug resistance and escape mutations in LMICs. We analyzed data from 47 studies involving 43,834 individuals and employed statistical methods to estimate rates and regional differences in drug resistance and mutation patterns.
The overall rate of drug resistance across all antiviral drugs was 7.87%, but it was much higher for older drugs in use, such as lamivudine (3TC) and telbivudine (LdT). Asia had the highest levels of resistance, with 32.45% for 3TC and 51.92% for LdT. Among economic regions, lower-middle-income countries showed the highest resistance rates (10.61%). The most common drug resistance mutations were rtM204V/I, rtV173L, and rtL180M. Vaccine-escape mutations (VEMs) were found in 8.92% of cases, while immune-escape mutations (IEMs) were far more common (55.66%). Frequent escape mutations included sG145R/A/K, sQ129H/R, sD144E/A, sS143L/T, and sP120S/T.
The high rates of drug resistance and immune or vaccine-escape mutations indicate that resistance remains a major challenge, particularly for lamivudine and telbivudine, which have a lower barrier to mutation. These findings highlight the growing difficulty in meeting the WHO 2030 targets and emphasize the need for stronger surveillance systems, wider use of more effective antiviral drugs such as Tenofovir Disoproxil Fumarate (TDF), and ongoing monitoring to ensure vaccines continue to provide protection. However, the results should be interpreted with caution due to the wide variability between studies.
Keywords
Introduction
Hepatitis B virus (HBV) is a DNA virus that targets the liver, causing both acute and chronic hepatitis. 1 Transmission occurs through exposure to infected blood and bodily fluids, as well as perinatally from mother to child at birth.2,3 Although initial infection may be asymptomatic or mild, chronic HBV infection can progress over time, often without warning, to severe liver complications such as cirrhosis, hepatic failure, and hepatocellular carcinoma, ultimately leading to death.2,4 In fact, one-third of all liver cancer fatalities worldwide are due to HBV infections. 5 The effects of HBV are compounded by barriers such as inadequate screening programs, late diagnosis, and limited treatment access, which together increase the risk of adverse outcomes and further perpetuate high prevalence.
According to the Global Hepatitis Report by the World Health Organization (WHO) in 2022, an estimated 296 million people are living with chronic HBV, and approximately 1.1 million die each year as a result of HBV-related liver disease. 6 Additionally, reports estimate that 83% of all viral hepatitis deaths are due to HBV. Most of these deaths occur in low- and middle-income countries (LMICs), where healthcare resources are limited, and HBV prevalence remains highest. 7 For instance, in the WHO African region, where most countries are low to middle-income, the prevalence of hepatitis B infections among the general population in 2019 was 7.5% compared to 0.5% in the Americas region. 8 Hence, the WHO’s global hepatitis strategy is to reduce new hepatitis infections by 90% and deaths by 65% by 2030. 9
Although there is no specific treatment for acute hepatitis B, chronic hepatitis B can be treated with a range of antiviral medications. The introduction of nucleos(t)ide analogues (NAs) such as tenofovir disoproxil fumarate and entecavir has allowed for effective suppression of viral replication, which in turn lowers the risk of developing serious complications. The antiviral drugs lamivudine (3TC), entecavir, telbivudine (LdT), adefovir, tenofovir disoproxil fumarate (TDF), and, more recently, tenofovir alafenamide (TAF) are the approved and commonly used drugs for antiviral therapy of HBV. 5 In HBV and HIV co-infected individuals, 3TC and TDF are administered as antiviral therapy for both conditions. Despite the availability of these drugs, reports indicate that only 3% of people diagnosed with chronic hepatitis B had received antiviral therapy at the end of 2022. 6
Nonetheless, incomplete viral suppression and the emergence of drug resistance remain a major concern in the management of chronic hepatitis B. Alarmingly, all approved anti-HBV agents have been associated with the development of antiviral resistance, often resulting from specific mutations in the reverse transcriptase domain of the viral polymerase. 10 These mutations can significantly reduce the effectiveness of antiviral therapy, allowing the virus to replicate despite ongoing treatment.
Simultaneously, the growing emergence of vaccine-escape mutations (VEMs) has become increasingly prominent in the global effort to control hepatitis B. Since the introduction of the hepatitis B vaccine over 40 years ago, immunization campaigns have resulted in dramatic declines in HBV incidence, especially among children, with the WHO estimating that vaccine coverage prevented over 37 million infections between 2000 and 2019. 6 However, genetic mutations in the “a” determinant region of the hepatitis B surface antigen (HBsAg) have been shown to enable the virus to evade vaccine-induced immunity. 11
While high-income countries have systematically monitored and extensively reported the emergence of HBV drug resistance and vaccine-escape mutations, similar comprehensive data from LMICs remain fragmented or insufficiently explored as a result of inadequate resources, especially in resource-limited countries, for monitoring treatment and immunoprophylaxis.12,13 With anticipated expansions in antiviral treatment programs and increased vaccine rollout in these regions, bridging this critical knowledge gap is imperative. Therefore, this systematic review and meta-analysis aims to consolidate existing evidence on the prevalence, distribution, and underlying genetic factors influencing HBV drug resistance and vaccine-escape mutations in LMICs.
Methods
This systematic review was registered in the International Prospective Register of Systematic Reviews (PROSPERO) under the ID: CRD420251030302. The methodology also followed the guidelines set by the Preferred Reporting Items for Systematic Review and Meta-analysis (PRISMA; Supplemental Table 1). 14
Search strategy and screening
Literature searches were conducted across electronic databases, including PubMed, Scopus, and Web of Science, to identify studies reporting on HBV drug resistance-associated mutations and vaccine-escape mutations in LMICs over the past decade. We combined Medical Subject Headings (MeSH) and relevant keywords such as “Hepatitis B virus,” “HBV,” “drug resistance,” “antiviral resistance,” “vaccine-escape,” “immune-escape,” “surface antigen mutations” and “mutation,” along with the individual country names of the countries classified as LMICs by the World Bank. 15 Boolean operators were used to refine search results, and the strategy was tailored to suit the indexing systems of each database. Filters were applied to restrict results to studies published between January 1, 2014, and April 2, 2025, which was the last search date. There was, however, no restriction on language. The full detailed search strategy for each database is available in Supplemental Table 2.
The search records were retrieved from the various databases and exported to Rayyan, 16 where duplicates were manually reviewed and removed before screening. Titles and abstracts of all identified studies were first independently assessed by two reviewers to exclude those that did not meet the predefined inclusion criteria. Subsequently, full texts of potentially eligible studies were retrieved and independently reviewed in detail on Zotero by the same reviewers. Disagreements during the screening process were resolved through discussion, and when necessary, a third reviewer was consulted to reach consensus.
Key definitions of HBV mutations
For this systematic review and meta-analysis, HBV mutations were classified into resistance-associated mutations, vaccine-escape mutations, and immune-escape mutations based on established literature and the definitions applied in the included studies.
Where mutation classifications differed across studies, the definitions applied by the original authors were retained if supported by existing evidence. Classifications were reviewed for consistency during data extraction.
Eligibility criteria
We included research articles that presented primary data on HBV drug resistance-associated mutations, vaccine-escape mutations, and immune-escape mutations in human populations from LMICs. Eligible studies reported on the prevalence, frequency, or molecular characterization of these mutations in clinical samples, regardless of participants’ treatment or vaccination history. Regarding participants’ treatment status, studies that included both treatment-naïve (NA-naïve) and treatment-experienced (NA-experienced) individuals, and that reported data stratified by these two groups, were considered eligible. We considered observational study designs but excluded reviews, commentaries, letters, conference abstracts, animal studies, and laboratory experiments unrelated to clinical populations. Studies without a clear country context or focused exclusively on high-income settings and had a sample size below 15 were also excluded. In cases of overlapping datasets, only the most recent or comprehensive publication was included.
Quality assessment
The quality of the included studies was appraised using the STROBE-ME (Strengthening the Reporting of Observational Studies in Epidemiology—Molecular Epidemiology) checklist, which is specifically designed to address the unique aspects of molecular epidemiology research, such as laboratory protocols, specimen handling, and the reporting of genetic or mutation data. For each study, we reviewed every item on the checklist and assigned a score of “reported” (1) if the information was explicitly stated in the publication, “not reported” (0) if absent or insufficiently described, or marked as “not applicable” when the item was irrelevant to the study design. Studies were then assigned to quality categories in accordance with the grading thresholds outlined by Limaye et al. 17 : Excellent (⩾85%), Good (70%–84%), Fair (50–69%), or Poor (<50%; Supplemental Table 3).
Data extraction and synthesis
Data extraction was performed using a standardized form and compiled in Microsoft Excel (Version 2410). Extracted data included study characteristics (author, year of publication, country, study design, setting, and sample size), population details (clinical status of study participants, HBV treatment exposure, and HBV genotypes), and diagnostic methods used to detect RAMs and VEMs. Specific mutations reported, such as rtM204V/I, rtL180M, sG145R, were also recorded.
The meta-analysis was conducted using RStudio (version 4.5.0) with the meta package. Variances were standardized through the Freeman-Tukey double arcsine transformation, and pooled resistance rates and prevalences were estimated using the restricted maximum-likelihood method. A random-effects model was applied to generate forest plots, which illustrated study-specific effect sizes along with 95% confidence intervals (CIs) for pooled resistance rates and prevalences. Heterogeneity among studies was evaluated using the I2 statistic, where values of 0%, 25%, 50%, and ⩾75% represented no, low, moderate, and high heterogeneity, respectively. Publication bias was assessed visually using funnel plots and statistically through Egger’s regression test. Sensitivity analysis was also performed to identify potential sources of heterogeneity in resistance rates by excluding studies with a high risk of bias and recalculating resistance rates for hepatitis B antiviral drugs, which were then compared to the rates from all studies combined.
Results
Search results
A total of 1854 records were identified through database searches, including 693 from PubMed, 345 from Scopus, and 816 from Web of Science. After removing 361 duplicates, 1493 records remained for screening. Of these, 1394 were excluded based on titles and abstract screening, and 99 studies proceeded to full-text screening. Following full-text review, 52 studies were excluded for the following reasons: Not LMICs (n = 10), outside the decade (n = 9), sample size below 15 (n = 4), review articles (n = 9), duplicate data from previously included studies (n = 1), inaccessible full texts (n = 10), no prevalence reported (n = 7), and no mutations reported (n = 2). Ultimately, 47 studies met the inclusion criteria and were included in the final review18–64 (Figure 1).

PRISMA flow diagram of study selection: the initial search yielded 1854 studies. Of these, 361 duplicates were removed. The remaining 1493 studies were screened for title and abstract alignment, with 1394 studies excluded after screening. Full-text article screening was conducted on the 99 studies, and 47 studies were finally selected that met the eligibility criteria.
Study characteristics
A total of 47 studies from 28 different LMICs were included in this review. Most studies were conducted in Africa (57.45%) and Asia (39.91%). Additionally, 87.23% (n = 41) of the studies were conducted in hospital-based settings, 5 were recruited from community settings, and 1 was from a research institution. The sample sizes ranged from 15 to 19,440 participants, resulting in an aggregate of 43,834 participants across all studies. Study populations primarily comprised HBV-infected persons who were receiving a single or combined nucleoside analogue (NA) therapy. In total, 30,612 (69.84%) were HBV+ while 2792 (6.37%) were co-infected with HBV and HIV. Additionally, 27,403 (62.52%) were nucleotide analogue (NA)-experienced, while only 5093 (11.62%) were NA-naïve. More than two-thirds of the studies (71.43%) adopted the Sanger method for sequencing, while others employed direct, next-generation, or hybridization-based sequencing approaches. Based on the quality assessment, 2.1% of the included studies were rated excellent, 8.5% were good, 74.5% were fair, and 14.9% were poor (Table 1).
Study characteristics.

‘—’ Data not available.
HBV, hepatitis B virus; NA, nucleotide analog.
HBV markers
In a random-effects meta-analysis, the pooled prevalence of HBsAg, which is a general marker of HBV infection, was 72.65% [k = 44, where k is the number of studies] (95% CI (56.44–86.30)). In contrast, the pooled prevalence of hepatitis B e Antigen (HBeAg) was notably lower at 27.83% [k = 21] (95% CI (14.65–43.28)). Moreover, HBV DNA, which is a marker of active viral replication and occult hepatitis B, had a prevalence of 41.65% [k = 34] (95% CI (26.56–57.57); Figure 2). “k”.
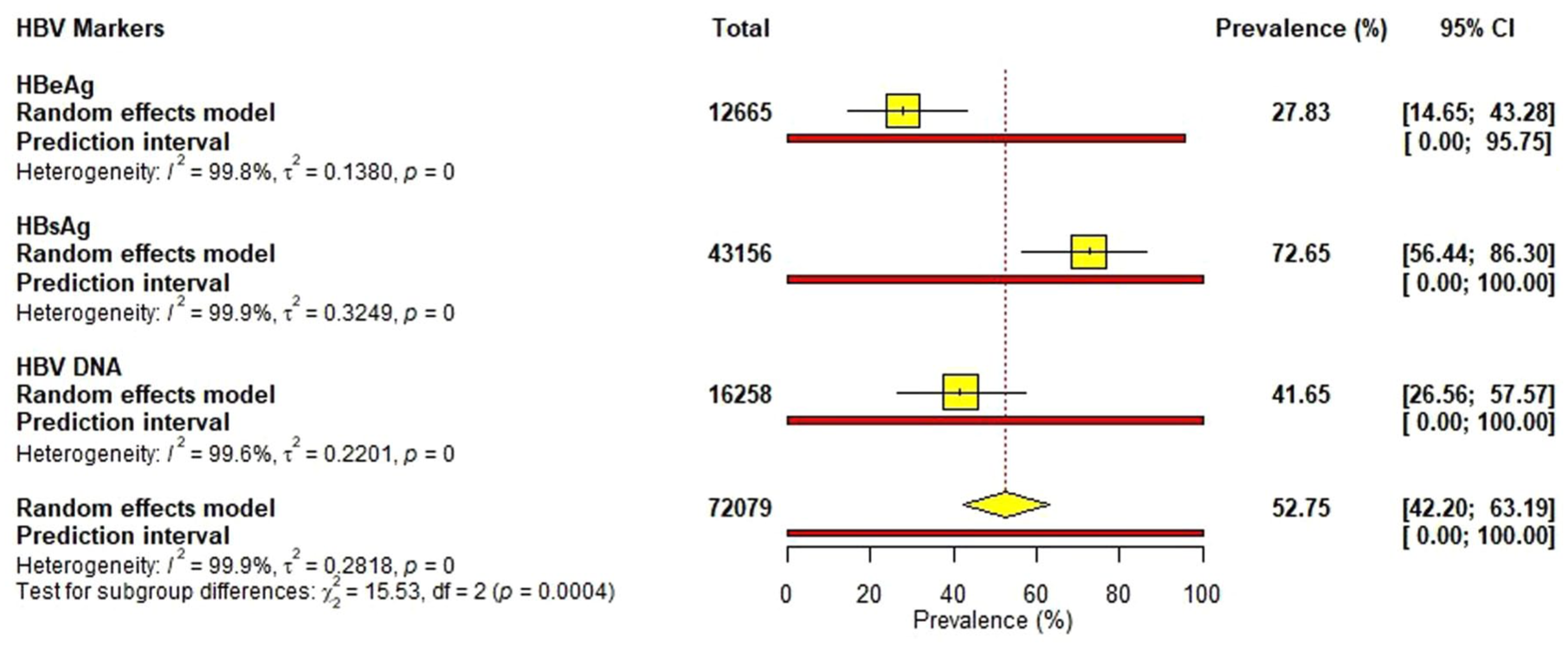
Pooled prevalence of HBV markers: The forest plot summarizes the prevalence (%) observed for HBsAg, HBeAg, and HBV DNA in a random-effects meta-analysis. The 95% CI is shown in brackets in the column on the right side of the plot. The prevalence is shown in the next column from the right. The Total column indicates the number (n) of participants contributing to the pooled estimate. Heterogeneity is represented I2 and t2 values. From the plot, very high heterogeneity is observed (99.6%–99.9%). Therefore, the results from this analysis are only reported descriptively rather than as population-level estimates.
Antiviral resistance
The overall resistance rate across all reported antiviral drugs for both NA-naïve and NA-experienced cohorts was 7.87% (95% CI (5.13–11.11), with very high heterogeneity (I2 = 99.7%, t2 = 0.0637). Among individual drugs, the highest rate was observed for telbivudine at 14.97% [k = 9](95% CI (1.20–39.09); I2 = 99.9%, t2 = 0.1748) and the lowest for tenofovir disoproxil fumarate at 2.01% [k = 12] (95% CI (0.55–4.20); I2 = 95.9%, t2 = 0.0100). Lamivudine resistance was 11.54% [k = 34] (95% CI (6.12–18.32); I2 = 99.7%, t2 = 0.0767), while adefovir and entecavir showed resistance rates of 5.42% [k = 14] (95% CI (2.13–9.96); I2 = 99.1%, t2 = 0.0234) and 5.44% [k = 17] (95% CI (1.78–10.75); I2 = 97.8%, t2 = 0.0389), respectively (Figure 3). Given the consistently high heterogeneity across analyses, pooled resistance estimates are presented to summarize this study’s findings and should not be interpreted as precise or generalizable population-level prevalence estimates.

Prevalence of resistance rates associated with hepatitis B antiviral drugs: The forest plot summarizes the resistance (%) prevalence for adefovir, entecavir, lamivudine, telbivudine, and tenofovir disoproxil fumarate across included studies. Pooled estimates were calculated using a random-effects meta-analysis, and 95% CIs are shown for each drug. The total number of participants (n) contributing to each estimate is indicated in the “Total” column. Prediction intervals, reflecting the expected range of resistance in a new study, are shown by horizontal red lines. Between-study heterogeneity is reported as I2 and t2. The overall pooled resistance across all antivirals is presented at the bottom of the figure. High heterogeneity (I2 > 95% for most drugs) indicates substantial variability among studies; therefore, all pooled estimates should be interpreted descriptively rather than as generalized population values.
Subgroup analyses by geographical region showed substantial variability in resistance patterns. Studies from Asia showed the highest resistance across all antiviral drugs, particularly for telbivudine (51.92 [k = 3], 95% CI (3.53–97.88); I2 = 98.7%, t2 = 0.2703) with notable resistance to lamivudine (32.45% [k = 10], 95% CI (15.08–52.70); I2 = 99.5%, t2 = 0.1053) and entecavir (15.37% [k = 5], 95% CI (0.37–43.42); I2 = 97.2%, t2 = 0.1216). In contrast, studies from Africa showed generally lower resistance, with lamivudine showing the highest resistance at 4.27% [k = 19] (95% CI (1.83–7.58); I2 = 94.2%, t2 = 0.0218), followed by adefovir (3.97% [k = 7], 95% CI (0.00–13.26); I2 = 96.8%, t2 = 0.0212) and telbivudine (3.24% [k = 5], 95% CI (0.00–12.52); I2 = 97.1%, t2 = 0.0404). Studies from South America showed comparatively low resistance across antivirals, with the highest prevalence observed for telbivudine (8.36% [k = 1], 95% CI (5.35–11.95); heterogeneity = not applicable) and lamivudine (6.02% [k = 3], 95% CI (1.85–12.26); I2 = 89.1%, t2 = 0.0083), and notably low rates for adefovir (0.42% [k = 2], 95% CI (0.00–1.33); I2 = 0.0%, t2 = 0.0000) and tenofovir (0.42% [k = 2], 95% CI (0.00–1.33); I2 = 0.0%, t2 = 0.0000; Supplemental Figures 1–5). Figure 4 shows a graphical presentation of the pooled antiviral resistance rates across the observed regions.
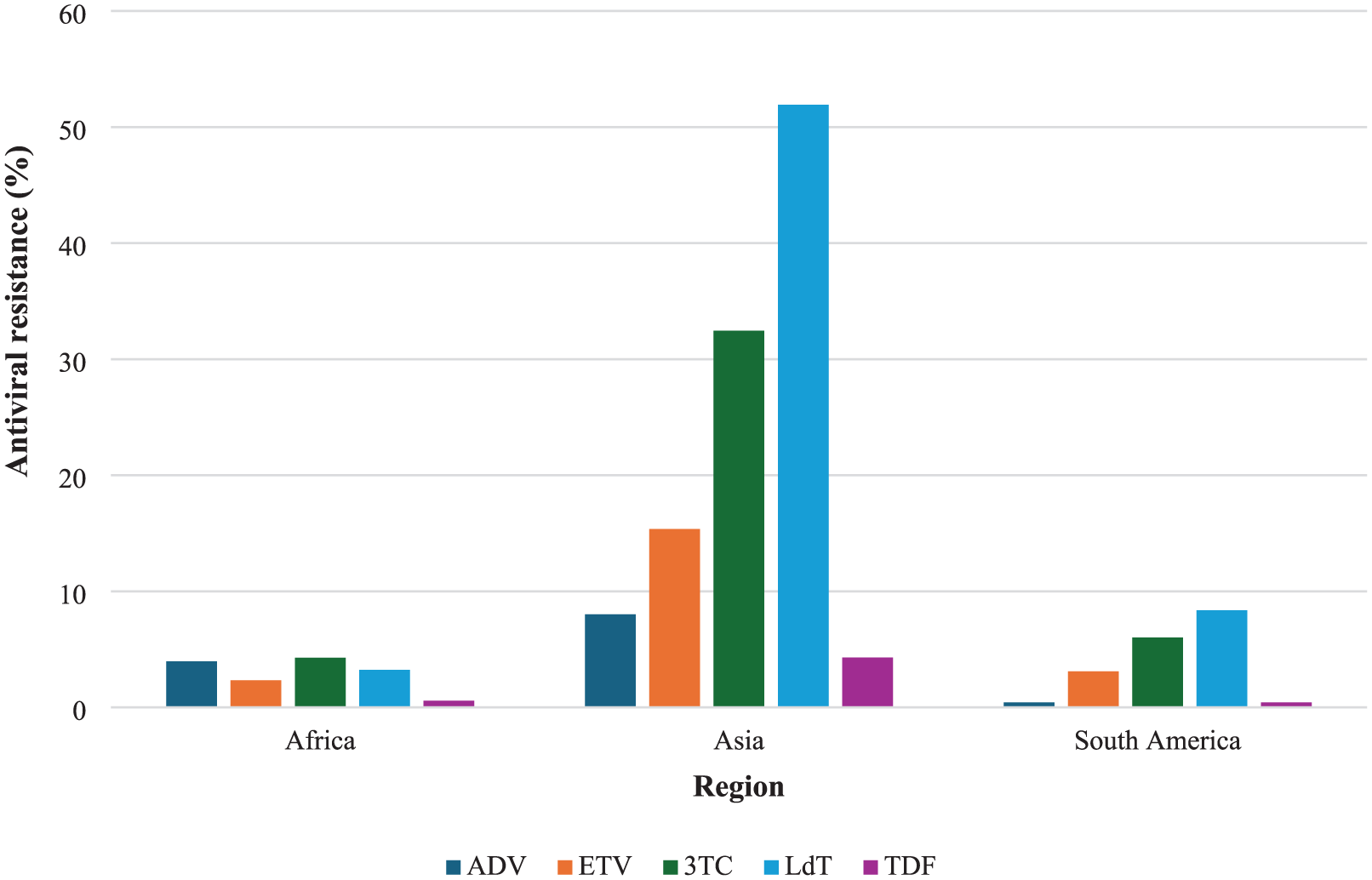
Pooled resistance rates of hepatitis B antiviral drugs stratified by region: The graph shows a summary of the antiviral resistance rate in included studies from Africa, Asia, and South America. The colored bars represent the percentage prevalence for the drugs adefovir, entecavir, lamivudine, telbivudine, and tenofovir disoproxil fumarate, each. The highest prevalence was observed in studies from Asia, particularly for telbivudine (51.92%).
Based on antiviral therapy history, it was observed that individuals with prior treatment experience had consistently higher resistance mutation rates to hepatitis B antiviral drugs, significantly in telbivudine, compared to those who were treatment naïve (Figure 5). Among NA-naïve patients [k = 25], resistance was lowest for tenofovir disoproxil fumarate (0.86%; 95% CI (0.24–1.76)) and highest for lamivudine (9.61%; 95% CI (4.00–17.21)). In contrast, NA-experienced individuals [k = 31] exhibited notably elevated resistance, with telbivudine reaching 25.35% (95% CI (1.85–62.19)) and lamivudine at 12.69% (95% CI (6.04–21.25); Supplemental Table 4).
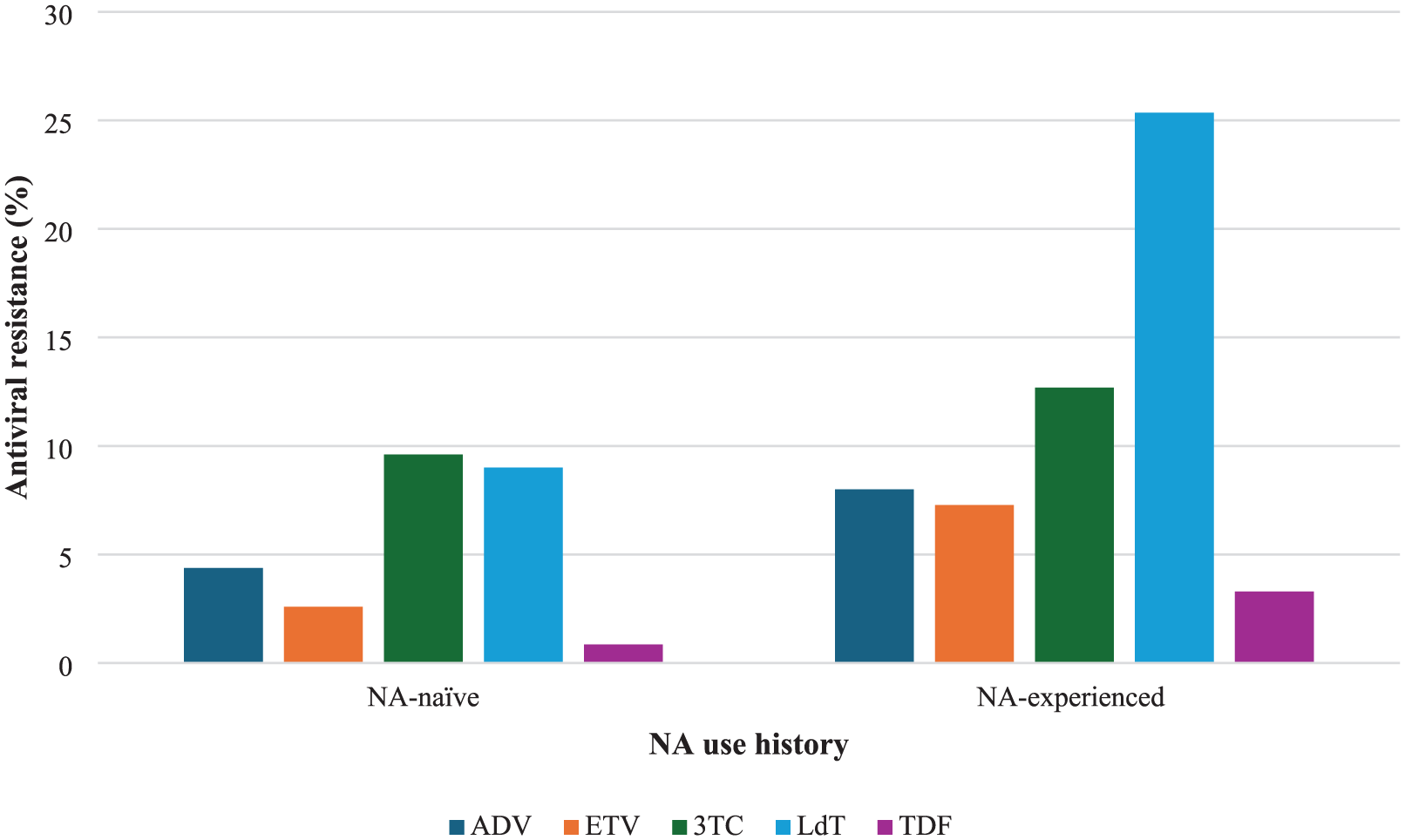
Pooled resistance rates of hepatitis B drugs based on antiviral use history: The graph shows a summary of the antiviral resistance rate in NA-naïve participants compared to NA-experienced participants from the included studies. The colored bars represent the percentage prevalence for the drugs adefovir, entecavir, lamivudine, telbivudine, and tenofovir disoproxil fumarate. The highest prevalence was observed in NA-experienced participants, particularly telbivudine (25.35%).
In the subgroup analysis by country income level, the highest resistance was observed in studies from LMICs, with a pooled prevalence of 10.61% [k = 14] (95% CI (5.21–17.51); I2 = 96.9%, t2 = 0.0889). Studies from upper-middle-income countries (UMICs) followed, with resistance estimated at 8.14% [k = 7] (95% CI (3.91–13.66); I2 = 99.8%, t2 = 0.0590), while studies from low-income countries (LICs) reported the lowest prevalence at 3.66% [k = 7] (95% CI (1.36–6.91); I2 = 96.2 %, t2 = 0.0246; Figure 6).

Subgroup analysis of HBV antiviral resistance by country income level: The figure presents pooled estimates of HBV drug resistance in low-income, lower-middle-income, and upper-middle-income countries. The pooled estimate of resistance (%) for each group is shown in yellow squares. Horizontal lines through the squares indicate the 95% CI. Heterogeneity within each subgroup was high, as indicated by I2 values: low-income countries (I2 = 96.2%), lower-middle-income countries (I2 = 96.9%), and upper-middle-income countries (I2 = 99.8%). Thus, all pooled estimates are descriptively reported and not interpreted as generalized values.
HBV genotypes
Genotypic analysis was reported in 45 of the 47 studies, revealing a predominance of genotype A (26%) followed by genotypes D (25%), E (22%), and C (12%). Two studies reported the recombinant genotypes E/A and A/E60,45 (Supplemental Figure 6). A map distribution of the prevalent genotypes in the countries included showed that genotype E was prevalent in the West African region (Ghana, Nigeria, the Republic of Guinea, and Côte d’Ivoire), the Central African region (Central African Republic, Cameroon, and DR Congo), and Southern Africa (Namibia). Genotype A was most prevalent in Brazil, EL Salvador, Gabon, Kenya, Ethiopia, Uganda, Botswana, and Malawi. Genotype C was reported in China. This was observed as the only country with this genotype being the most prevalent (Figure 7).

Distribution of predominant HBV genotypes across LMICs: The map illustrates the most frequently reported HBV genotype for each LMIC included in the analysis. Countries are shaded according to the predominant genotype identified (A–F), as indicated in the legend. For each country, the displayed genotype represents the genotype with the highest reported prevalence based on available data. Countries shown in grey indicate locations with no eligible genotype data or not included in the analysis. The map was created using mapchart.net. 65
Antiviral resistance was reported across all genotypes except telbivudine in genotype D. Tenofovir disoproxil fumarate recorded the lowest resistance rates (4.35%–6.54%), while lamivudine and telbivudine recorded the highest resistance rates (14.21%–72.69%; 2.38%–100%; Figure 8).

Pooled antiviral resistance by HBV genotype: The graph represents the antiviral resistance rates (5%) for genotype populations reported in the included studies. The genotypes reported, from the collection of included studies, were genotypes (A)–(F). The coloured bars represent the percentage prevalence for the drugs adefovir, entecavir, lamivudine, telbivudine, and tenofovir disoproxil fumarate. The highest prevalences were observed in genotypes (B), (C), and (F) for telbivudine (72.94%, 73.21%, and 100%, respectively). However, these results cannot be generalized due to the non-homogeneity of studies reporting these genotypes.
Resistance-associated mutations
Of 47 studies, 35 reported resistance-associated mutation in at least one of the antiviral drugs (lamivudine, entecavir, adefovir, tenofovir disoproxil fumarate, and telbivudine). Of these studies, five25,33,44,46,61 reported VEMs, four31,37,39,48 reported immune-escape mutations (IEMs), and one study 25 reported both VEMs and IEMs. The rtM204V/I, rtV173L, and rtL180M mutations, which confer resistance to lamivudine, telbivudine, and in some cases entecavir, in the presence of the compensatory mutation rtL180M, were the most prevalent resistance-associated mutation overall, identified in both treatment-experienced and treatment-naïve individuals. It was observed either in isolation or in conjunction with other polymorphisms, including rtL80I/V, rtV173L, rtL180M, rtA181S, rtT184S, rtA200V, and/or rtS202S. The details of RAMs identified in the studies are listed in Supplemental Table 5.
Vaccine-escape mutations and immune-escape mutations
Vaccine-escape mutations were reported in 11 studies, whereas immune-escape mutations were identified in 12 studies. Two studies24,64 reported both VEMs and IEMs. Vaccine-escape mutations had a pooled prevalence of 8.92% [k = 11] (95% Cl (5.02–13.64); I2 = 78.4%, t2 = 0.0119). Immune-escape mutations had a pooled prevalence of 55.66% [k = 12] (95% Cl (28.10–81.53); I2 = 98.1%, t2 = 0.1918; Figure 9). Due to the absence of data on vaccination history in the included studies, interpretation of the pooled estimates cannot be generalized.

Pooled prevalence of VEMs and IEMs: The forest plot summarizes the prevalence (%) observed for IEMs and VEMs in a random-effects meta-analysis. The 95% CI is shown in brackets in the column on the right side of the plot. The prevalence is shown in the next column from the right. The Total column indicates the number (n) of participants contributing to the pooled estimate. Heterogeneity is represented I2 and t2 values. From the plot, very high heterogeneity is observed.
Most of the mutations were similarly identified either as a VEM or an IEM, including the common escape mutations sG145R/A/K, sQ129H/R, sD144E/A, sS143L/T, and sP120S/T (Supplemental Table 6). These mutations were found in the S gene.
Publication bias and heterogeneity
There was no publication bias due to a wider variation in sample sizes of the included studies. This was confirmed by Egger’s regression test (p-value = 0.0552; Supplemental Figure 7). Estimated resistance rates and prevalences were significantly heterogeneous (H > 1 and I2 ⩾ 75%).
Sensitivity analysis
The sensitivity analysis of resistance rates for Hepatitis B antiviral drugs: adefovir, entecavir, lamivudine, telbivudine, and tenofovir showed that excluding low-quality studies had little impact on the results. For most drugs, resistance rates remained consistent (e.g., adefovir: 6.58% vs 5.42%; tenofovir: 2.51% vs 2.01%), with overlapping 95% confidence intervals indicating robustness. Telbivudine showed no difference (14.97% vs 14.97%). The overall pooled resistance rate was nearly identical (8.56% vs 7.87%) across scenarios. These findings are reliable and not heavily influenced by study quality (Supplemental Table 7).
Discussion
This systematic review and meta-analysis reports the distribution and prevalence of HBV drug resistance, RAMs, VEMs/IEMs, and HBV genotypes from 47 studies in LMICs. Significantly high heterogeneity was observed across all analyses conducted. This indicates substantial variability in effect estimates between studies rather than chance alone. A key contributing factor is likely the wide variation in sample sizes among the included studies. The regional representation was skewed towards Africa (57.45%), followed by Asia (39.91%), reflecting the high burden and endemicity of HBV in these regions. 19 This mirrors WHO estimates, which report HBV to be most prevalent in Africa. 19 A significant proportion of participants (69.83%) were HBV mono-infected, while approximately 6.37% were co-infected with HIV, a known modifier of HBV disease progression and treatment response. 27 A substantial proportion of participants (62.78%) were experienced with NA, compared with only 11.73% who were naïve to NA. This indicates that the study population predominantly reflected individuals with prior antiviral exposure, which is an important context when interpreting RAM prevalence.
Over two-thirds of the studies (71.43%) employed the Sanger method, a gold-standard approach for detecting dominant viral populations. Sanger sequencing is widely used in resource-limited settings due to its relatively low cost, established protocols, and interpretability. It provides high-quality, long-read sequences, making it suitable for identifying well-established RAMs and genotypes. However, the method demonstrates limited sensitivity for minority variants, typically detecting only those comprising over 20% of the quasispecies pool. 31 This limitation is particularly important in the context of treatment-experienced patients, where resistant mutants may persist as low-frequency variants and later re-emerge under drug pressure. However, the use of direct sequencing, next-generation sequencing (NGS), hybridization, and bidirectional techniques, as employed in some of the studies, allows for the identification of minor variants and coexisting mutations, enhancing sensitivity and mutation profiling depth. Notably, the emergence of nanopore sequencing makes it increasingly viable for surveillance of HBV genotypes, RAMs, and escape variants. Its portability, speed, and scalability make it particularly attractive for decentralized testing in LMICs, especially in outbreak or high-burden settings. 13
Although most individuals (72.65%) tested positive for HBsAg, indicating current or prior exposure, only about one quarter (27.83%) exhibited HBeAg, which typically reflects high-level viral replication and infectivity. This serological profile normally characterizes later stages of infection, following the host’s immune-mediated clearance of HBeAg. The transition from HBeAg-positive to HBeAg-negative chronic hepatitis B is often associated with the emergence of precore (G1896A) or basal core promoter mutations (A1762T/G1764A) that suppress HBeAg expression without halting viral replication. 66 As such, HBeAg negativity does not always equate to viral quiescence; rather, it can coexist with fluctuating or elevated HBV DNA levels and ongoing hepatic necroinflammation, especially in genotype D and E infections that are common in Africa and the Mediterranean region. 66 Geographically, the predominance of HBeAg-negative profiles reported in this review aligns with prior studies from Africa and Asia, where patients frequently acquire HBV perinatally or in early childhood and progress to the immune control phase by adulthood. Studies in African cohorts have shown that 70%–90% of HBsAg-positive individuals can be HBeAg-negative by their third or fourth decade of life. 67 Similarly, in Asia, where genotype B and C dominate, delayed seroconversion and fluctuating HBV DNA levels in HBeAg-negative phases are commonly reported. 62 Moreover, the pooled prevalence of detectable HBV DNA, which is a direct measure of viral replication, was 41.65%, indicating that a significant proportion of individuals, despite low HBeAg prevalence, still harbour replicating viruses. This finding reinforces the importance of HBV DNA testing, particularly in detecting occult hepatitis B infection (OBI), a condition characterized by the presence of HBV DNA in the absence of detectable HBsAg.68,12
Across all antiviral drugs, the pooled resistance prevalence was 7.87% (I2 = 99.7%, t2 = 0.0637), with substantial heterogeneity across studies. Resistance was highest for lamivudine and telbivudine, consistent with their low genetic barrier and rapid selection of RAMs, particularly rtM204V/I in the tyrosine-methionine-aspartate-aspartate (YMDD) motif of the HBV polymerase.69,70 By contrast, resistance to tenofovir disoproxil fumarate (TDF) remained low at 2.01% (I2 = 95.9%, t2 = 0.0100), supporting evidence that TDF has a high barrier to resistance. 71 However, a few cases of resistance have been reported in both TDF-naïve and treatment-experienced individuals, with non-adherence identified as a potential contributing factor.72,73 Entecavir and adefovir demonstrated intermediate resistance levels (5.44% (I2 = 97.8%, t2 = 0.0389) and 5.42% (I2 = 99.1%, t2 = 0.0234), respectively), which may partly reflect their use as second-line therapies or following lamivudine failure, situations known to predispose to cross-resistance. 74 The higher resistance observed with older antivirals underscores the rationale for prioritizing high-barrier-to-resistance drugs, such as tenofovir-based regimens, in national treatment guidelines. In many LMICs, TDF is often administered as part of HIV antiretroviral therapy, increasing exposure among HIV-HBV coinfected individuals. While TDF is generally effective, potential nephrotoxicity remains a concern, highlighting the need for monitoring during long-term therapy. TAF, by contrast, demonstrates a higher genetic barrier and potency, with no resistance or nephrotoxicity reported to date.73,75 Emerging therapies such as ATI-2173, a liver-targeted molecule for chronic HBV, have shown promise in early-phase trials, with most participants achieving sustained viral suppression and reductions in covalently closed circular DNA biomarkers. Adverse effects were generally mild, primarily headache, but the evidence is currently limited to phase Ib studies, and the drug has not yet received FDA or WHO approval.76,77 These findings suggest potential future therapeutic options but require further validation in larger and more diverse populations provide more homogenous estimates to enhance policymaking.
The differences in resistance rates across regions highlight the heterogeneous landscape of HBV management. Asia showing the highest drug resistance rates overall, particularly for telbivudine—51.92% (heterogeneity; I2 = 98.7%, t2 = 0.2703) and lamivudine—32.45% (heterogeneity; I2 = 99.5%, t2 = 0.1053), is likely attributable to the region’s historically high HBV burden, early adoption of nucleos(t)ide analogues, and prolonged monotherapy regimens. 78 Africa and South America, contrastingly, reported relatively lower resistance rates, possibly reflecting the delayed introduction of antiviral therapy and limited access to HBV monotherapy outside of HIV programs. 12 In Europe, lamivudine resistance from one study was notably high—36.79% (heterogeneity; not applicable), which may be associated with historical prescription before guideline updates recommending high-barrier antivirals, such as TDF, as first-line therapy. Sparse data from North America, showing low resistance rates, likely reflect the dominance of tenofovir-based regimens and robust monitoring systems. This may also reflect insufficient data from this region to give a fuller picture of the current resistance rates.
Treatment-experienced patients exhibited significantly higher resistance than treatment-naïve individuals, with telbivudine resistance peaking at 25.35%. The persistence of lamivudine resistance in NA-naïve patients suggests primary resistance transmission, a concern that has been increasingly recognized in high-prevalence regions. 79 Exposure to any NA selects for polymerase mutations, and sequential therapy increases the risk for multidrug resistance. The European Association for the Study of the Liver reports that entecavir resistance in nucleoside-naïve patients is ~1.2% at 5 years but exceeds 50% in patients with prior lamivudine exposure. Moreover, guidelines recommend against re-using lamivudine or entecavir in lamivudine-refractory patients, and favor switching to or adding tenofovir in the case of breakthrough. 80 Treatment experience is a key driver of resistance, reinforcing that first-line choices matter for long-term viral control.
Notably, this survey found low-income countries showed the least resistance—3.66% (heterogeneity; I2 = 96.2%, t2 = 0.0246) compared to lower-middle-income countries—10.61% (heterogeneity; I2 = 96.9%, t2 = 0.0889) and upper-middle-income countries—8.14% (heterogeneity; I2 = 99.8%, t2 = 0.0590). The high drug resistance in lower-middle-income countries may reflect greater antiviral drug availability in low-income settings, yet less robust treatment monitoring and resistance testing infrastructure than in upper-middle-income countries. 6 Furthermore, this may be because low-income settings have only recently scaled up tenofovir-based HBV therapy. In contrast, middle-income programs may still be “catching up” after earlier use of lamivudine. However, the high heterogeneity within each subgroup indicates that the income classification alone does not explain these differences, even as the data used in the analysis is from a subset of LMICs, and not from all LMICs.
The distribution of HBV genotypes A through G showed that genotypes A, D, and E, were the most predominant, accounting for 26%, 25%, and 22%, respectively. The most prevalent genotypes observed across LMICs included depicts a pattern which is consistent with previous global and regional epidemiological studies indicating that genotype A is prevalent in Sub-Saharan Africa (particularly Eastern, Southern, and Central Africa), Europe, and parts of South Asia, while genotypes B and C dominate in East and Southeast Asia, genotype D in the Mediterranean and parts of India, genotype E in West Africa.31,81 The widespread presence of resistance-associated mutations across all genotypes, as revealed in this review, raises considerable concerns regarding the efficacy and long-term sustainability of current antiviral regimens, particularly in resource-limited settings. Lamivudine resistance was strikingly high, ranging from 14.21% to 72.69% across genotypes A through to F. This finding is not surprising given lamivudine’s low genetic barrier to resistance, wherein a single point mutation—most commonly rtM204V/I in the YMDD motif of the polymerase gene—can confer high-level resistance. 82 While the overall resistance rates to newer antivirals such as tenofovir disoproxil fumarate and entecavir were comparatively lower (4.35%–6.54% and 4.7%–12.61%, respectively), the cross-genotype emergence of RAMs in response to these agents is particularly troubling. Genotype-specific polymorphisms may influence the rate and pattern of RAM development. Studies have suggested that genotype C is more prone to developing A181T/V and N236T mutations, associated with adefovir resistance, as was seen in two of the studies reporting these mutations in individuals with HBV genotype C.41,50 Genotype A may more rapidly accumulate L180M + M204V mutations under lamivudine pressure.23,27,30,34,56 These genotype-resistance relationships have important clinical implications, as they may dictate differential responses to therapy and the likelihood of virological breakthrough. Furthermore, the detection of RAMs in both treatment-naïve and experienced individuals in this review suggests potential transmitted drug resistance mutations or undocumented prior exposure to nucleoside analogues, especially in regions with poorly regulated drug distribution.
The rtM204V/I, rtV173L, and rtL180M mutations emerged as the most prevalent RAMs across the included studies, indicating their central role in lamivudine resistance. Located in the highly conserved YMDD motif of the HBV reverse transcriptase domain, substitutions at rtM204 to either valine or isoleucine disrupt lamivudine binding, effectively abrogating its antiviral efficacy.69,70 These mutations also confer cross-resistance to other L-nucleoside analogues, such as telbivudine and emtricitabine, complicating subsequent therapeutic options. The mutation rtM204V/I often emerge in conjunction with compensatory mutations, most commonly rtL180M and rtV173L, as observed in this systematic review, which help restore viral replication fitness that is otherwise compromised by the primary resistance mutation.30,54,12 These secondary mutations enhance HBV polymerase stability and viral replication capacity, enabling resistant variants to outcompete wild-type strains even in the absence of continued drug pressure. This makes their co-selection particularly concerning, as it not only fosters sustained high-level resistance but also increases the potential for drug-resistant viruses. The persistence of rtM204V/I variants even in the absence of selective pressure has also been attributed to their incorporation into covalently closed circular DNA (cccDNA) reservoirs within hepatocytes. As a result, these resistant variants can be reactivated upon immunosuppression or treatment failure, contributing to virological rebound and complicating treatment switching strategies. 83 This highlights the importance of baseline resistance testing, particularly in populations with a high prevalence of RAMs, and supports the global transition toward high genetic barrier therapies such as tenofovir disoproxil fumarate and tenofovir alafenamide, which maintain efficacy against most lamivudine-resistant strains.
VEMs and IEMs present a significant challenge to global HBV control, particularly in regions of high endemicity. 84 In this review, the pooled prevalence of VEMs was 8.92%, while IEMs were considerably higher at 55.66%, also showing high heterogeneity across studies (VEMs: I2 = 78.4%, t2 = 0.0119; IEMs: I2 = 98.1%, t2 = 0.1918). These estimates should be interpreted descriptively, given the high heterogeneity, absence of data on vaccination coverage, differences in circulating genotypes, and prior antiviral exposure among study populations. The most frequently reported escape mutations (sG145R/A/K, sQ129H/R, and sD144E/A) occur within the “a” determinant of the HBsAg, a key region spanning amino acids 124–147 in the major hydrophilic domain. 61 Mutations in this domain can alter the conformation of the antigenic loop, reduce antibody binding, and enable immune evasion.74,84 The mutation sG145R has been documented to escape detection by diagnostic assays and to infect individuals despite prior vaccination.47,53 The high prevalence of IEMs suggests a substantial burden of naturally occurring or therapy-driven mutations that impair effective immune responses. While some IEMs overlap with VEMs, others occur in human leukocyte antigen-restricted T-cell epitopes within surface, polymerase, or core regions (e.g., L60V, P25S), potentially compromising antigen presentation and cytotoxic T-cell recognition.4,74 In long-standing HBV endemic areas, particularly in LMICs where vaccine coverage may be suboptimal, immune pressure may drive selection and persistence of such variants, contributing to reduced vaccine efficacy, diagnostic challenges, and increased risk among immunocompromised populations. These findings highlight the need for ongoing surveillance of HBV escape variants and periodic evaluation of vaccine formulations. Evidence supports the potential benefit of next-generation vaccines with broader epitope coverage or incorporation of preS1 and preS2 regions to improve immunogenicity.82,85 Furthermore, maintaining high vaccination coverage, ensuring robust birth-dose delivery, and strengthening laboratory capacity for molecular testing and viral genotyping are critical, particularly in low- and lower-middle-income countries, to enable earlier identification of emerging mutations and to guide public health interventions.
The meta-analysis in this review demonstrates wide variability in the prevalence of hepatitis B antiviral drug resistance and vaccine/immune-escape mutations across low- and middle-income countries. Although pooled estimates provide an overall indication of resistance patterns, interpreting these findings must be approached with considerable caution, given the consistently high heterogeneity observed across all analyses (I2 values > 95%). Such high heterogeneity indicates that the included studies differ substantially in population characteristics and treatment histories, which limits the reliability and comparability of the pooled percentages. This also highlights the importance of strengthening national and regional HBV surveillance systems, particularly in low- and middle-income countries where routine monitoring is limited. Standardized diagnostic protocols, improved sequencing capacity, and harmonized reporting frameworks are needed to generate more reliable data. The substantial proportion of studies rated as fair quality in the STROBE-ME assessment introduces a notable risk of bias, potentially leading to an underestimation of the pooled prevalence and reducing the overall certainty of the findings. Considering the wide prediction intervals and the predominance of fair-quality studies, these pooled estimates should be interpreted as indicative trends rather than precise national prevalence values. Policymakers may therefore consider using these results to inform general planning of treatment strategies and resource allocation, rather than as definitive measures.
We acknowledge some limitations. The high degree of heterogeneity across all analyses emphasizes the limitations of synthesizing highly diverse datasets. Although this review provides valuable insight into the burden of hepatitis B drug resistance and escape mutations in low- and middle-income countries, substantial inter-study heterogeneity limits the overall precision and interpretability of the pooled results. Moreover, the variation in quality reflects common methodological limitations in HBV research from LMICs, including sample sizes, cross-sectional designs, incomplete NA use history, and insufficient control for confounding factors such as vaccination status and genotype distribution. Finally, the evidence base was skewed toward Asian studies, with limited representation from South America, Europe, and other LMIC regions.
Conclusion
This review provides an overview of the prevalence of hepatitis B antiviral drug resistance and vaccine/immune-escape mutations across low- and middle-income countries, highlighting substantial variation across settings. Although pooled estimates suggest notable burdens of resistance, especially for drugs with low genetic barriers such as lamivudine and telbivudine, the interpretation of these findings is constrained by the extremely high heterogeneity observed across studies. The same pattern of variability was seen for both immune and vaccine-escape mutations. These results underscore that the current evidence base is highly fragmented, and that resistance and escape mutation prevalence cannot be reliably generalized across diverse contexts.
The overall data also point to the urgent need for high-quality, prospective, community-based studies, particularly in underrepresented regions such as South America, North America, the Middle East, and parts of North Africa. The expansion of NGS and more rigorous study design, enhanced reporting on antiviral therapy exposure, vaccine history, and immune status, will be crucial in monitoring resistance trends and guiding public health interventions. Additionally, this underscores the need for frequent detection of YMDD motif mutations and S-gene escape variants, adoption of potent antiviral regimens, and continuous vaccine efficacy monitoring. Addressing challenges in cost, accessibility, and data interpretation remains critical for the global control of HBV. The use of high genetic barrier antivirals such as TDF and TAF, personalized medicine, resistance testing, enhanced surveillance, and research on emerging cost-effective diagnostic methods will further strengthen the fight against HBV.
Supplemental Material
sj-docx-1-tai-10.1177_20499361261431606 – Supplemental material for A systematic review and meta-analysis of hepatitis B virus drug resistance and escape mutations in low- and middle-income countries
Supplemental material, sj-docx-1-tai-10.1177_20499361261431606 for A systematic review and meta-analysis of hepatitis B virus drug resistance and escape mutations in low- and middle-income countries by Esther M. E. Asamoah, Anastasia A. Asantewaa, Samuel N.-A. Yartey, Nicholas I. Nii-Trebi and Eric S. Donkor in Therapeutic Advances in Infectious Disease
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
