Abstract
Background:
Empirical antibiotic therapy (EAT) in febrile neutropenia (FN) remains challenging due to multidrug-resistant (MDR) Gram-negative bacteria, often leading to inappropriate empirical antibiotic therapy (IEAT).
Objective:
To demonstrate that risk stratification based on machine learning (ML) and prior colonisation with MDR bacteria may support the tailoring of EAT in patients with haematological malignancies.
Design:
Retrospective proof-of-concept cohort study.
Methods:
All consecutive FN episodes in patients with haematological malignancies were retrospectively included from January 2020 to March 2023 at a tertiary-level university hospital. We compared real-world, clinician-driven empirical antibiotic use with a simulated approach guided by an ML-based risk stratification model combined with prior colonisation data. The main outcomes were antibiotic selection and rates of IEAT.
Results:
A total of 553 FN episodes in 398 haematological patients were analysed. Bloodstream infection (BSI) occurred in 141/553 episodes (25.5%). Anti-pseudomonal (PsA) beta-lactams were prescribed in 515/553 episodes (93.1%), with carbapenems in 406/553 (73.4%). The clinician-driven approach resulted in 16/70 (22.9%) GNB-BSI episodes receiving IEAT. The ML plus colonisation-guided approach would have reduced the use of meropenem by 29.7% (−2.08 days; 95% CI, −2.42 to −1.73;
Conclusion:
ML-based risk stratification combined with colonisation status would allow for personalised antibiotic therapy in FN, potentially reducing IEAT and improving antimicrobial use. These results support integrating these tools into clinical practice.
Keywords
Background
The empirical treatment of febrile neutropenia (FN) has become increasingly complex. The emergence of multidrug-resistant (MDR) Gram-negative bacteria poses significant challenges to antibiotic decision-making across numerous regions worldwide. Several studies report that, even when adhering to guideline-recommended regimens, the rate of inappropriate empirical antibiotic therapy (IEAT) in neutropenic patients with Gram-negative bloodstream infection (BSI) can reach up to 30%.1–3
Addressing this issue is challenging, as the empirical use of broad-spectrum antibiotic regimens, intended to cover even the most resistant pathogens, is often unnecessary for many patients. This approach may be costly, toxic, and can promote the development of antimicrobial resistance. Therefore, designing effective strategies to tailor antibiotic therapy in patients with FN is of paramount importance. Our research group has long focused on the development of artificial intelligence (AI) models aimed at improving personalised empirical treatment and clinical decision-making processes.4–8 In this context, we have recently developed a machine learning (ML) model designed to stratify the risk of BSI at the onset of fever in neutropenic patients. 8 This algorithm categorises patients into three distinct risk levels for BSI, with the following observed rates of BSI per FN episode within each group: low risk (4% overall BSI, 0% Gram-negative BSI), intermediate risk (27% overall BSI, 12% Gram-negative BSI), and high risk (38% overall BSI, 19% Gram-negative BSI). Notably, 75% of MDR-GNB BSI occurred in the high-risk group. A definition of the risk categories is provided in Table S1.
We hypothesise that this risk stratification can guide personalised antibiotic therapy strategies, optimising treatment in two key directions: narrowing the spectrum for patients at low risk of BSI, and conversely, identifying those who may benefit from broader-spectrum antibiotics to reduce the incidence of IEAT. This approach can be further enhanced by incorporating prior colonisation data with MDR organisms, as emphasised in the recent recommendations of the European Conference on Infections in Leukaemia (ECIL-10). 9 To test this hypothesis, we conducted a theoretical study comparing real-world physician-prescribed empirical regimens and associated IEAT rates in FN patients at our centre with a simulated scenario in which clinical decisions were guided by an ML-based BSI risk stratification model, supplemented with rectal colonisation screening data in high-risk patients to assign tailored antibiotic regimens by risk category.
Methods
Study overview, population, and data collection
This retrospective analysis was based on a single-centre observational cohort study conducted at the Hospital Clinic of Barcelona, a 700-bed tertiary care academic institution. The study included all consecutive adult patients (⩾18 years) with hematological malignancies who developed FN and were hospitalised between January 2020 and March 2023. Episodes were excluded if clinical or microbiological data were incomplete or if patients received outpatient management. The primary aim was to simulate and compare two approaches to antibiotic management at FN onset: (1) the Clinician-Driven approach based on real-world clinical practice and (2) a hypothetical approach guided by an ML-based risk stratification model, supplemented with rectal colonisation surveillance data in the high-risk cluster when available. The analysis compared antibiotic prescription patterns and the incidence of IEAT between the two approaches.
Data for the clinician-driven approach were extracted from the electronic health records of the included patients. At our institution, EAT at FN onset is prescribed by haematology specialists and reviewed by the Infectious Diseases team as part of the antimicrobial stewardship programme. Different antibiotic recommendation strategies were tested according to patient risk stratification. The hypothetical antibiotic strategies were designed based on the following premises: (a) low-risk patients would potentially receive ceftriaxone (CTR), as no Gram-negative bacterial infections were documented in this subgroup in the reference studies 8 ; (b) intermediate-risk patients would receive antipseudomonal (PsA) beta-lactams as recommended by FN guidelines. Accordingly, the results were described based on the use of each of the following antibiotics: ceftazidime (CTZ), cefepime (CEP), piperacillin-tazobactam (PIT), or meropenem (MER); (c) high-risk patients would potentially receive one of the following options: (i) any of the aforementioned PsA beta-lactams; (ii) the same PsA beta-lactams, adjusted for documented prior colonisation with MDR bacteria when applicable; (iii) an initial course of ceftazidime-avibactam (CTV) for the first 48 h, followed by de-escalation to CTZ if blood cultures and other diagnostic results were negative; or (iv) CTV, tailored according to MDR colonisation status.
The study adheres to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines to ensure methodological rigor and transparency, with the checklist available in the Supplemental Material. 10
Definitions
FN was defined as a body temperature of ⩾38.0°C and an absolute neutrophil count (ANC) of < 0.5 × 109/L. BSI was defined by the growth of non-skin commensal bacteria in at least one blood culture. For common skin colonisers such as coagulase-negative staphylococci or
EAT was defined as the antibiotics initiated following the collection of blood cultures on Day 1 of FN. EAT was considered inappropriate if the bacteria isolated in blood cultures showed no
Antibiotic use was assessed during the first 7 days of each FN episode. In the Clinician-Driven approach, antibiotic prescriptions administered during routine clinical care were retrospectively recorded. In the ML-guided strategy, predefined regimens based on risk stratification were applied, assuming a fixed 7-day duration per episode. Exposure was defined as the number of days each antibiotic was used, and differences in antibiotic-days per episode were compared between strategies.
Microbiological methods
Blood cultures were processed using the BACTEC FX system (BD, Franklin Lakes, NJ, USA). Identification of isolates was performed using conventional microbiological methods and MALDI-TOF MS (Bruker, Bremen, Germany). Antimicrobial susceptibility testing was carried out by disc diffusion and broth microdilution (Phoenix system, BD). Rectal swabs were cultured on chromID ESBL and CARBAsmart media (bioMérieux, Marcy l’Etoile, France), with identification by MALDI-TOF MS. ESBL production was suspected based on MIC values and confirmed by the double-disc synergy test using cefotaxime, CTZ, and CEP discs placed near clavulanic acid. AmpC production was suspected in cefoxitin-resistant isolates and confirmed with boronic acid disc testing. Carbapenemase production was confirmed with the NG-test CARBA5 (NG Biotech, France). In case of a negative result, CRE were further evaluated using the modified carbapenem inactivation method (mCIM). All results were interpreted according to EUCAST guidelines (version 12.0; effective January 1, 2022).
Statistical analysis
Descriptive statistics were used to summarise demographic, clinical, microbiological, and antibiotic-related variables. Categorical variables were compared using the chi-square or Fisher’s exact test, as appropriate. Paired
Results
Patients and bloodstream infection epidemiology
A total of 553 consecutive episodes of FN were documented among 398 haematological patients. Demographic and baseline characteristics of these episodes are detailed in Table S2. Most patients had leukemia or lymphoma as an underlying disease and/or had undergone a haematopoietic stem cell transplant (HSCT). Clinically significant BSI was documented in 141/553 (25.5%) FN episodes, of which 70/141 (49.6%) were caused by GNB. Table S3 summarises the microorganisms involved in these BSI episodes in the overall cohort and stratified by risk cluster.
Recent colonisation data were obtained for 45/92 (48.9%) episodes of BSI within the high-risk cluster. Figure 1 displays the isolates categorised by resistance profile.
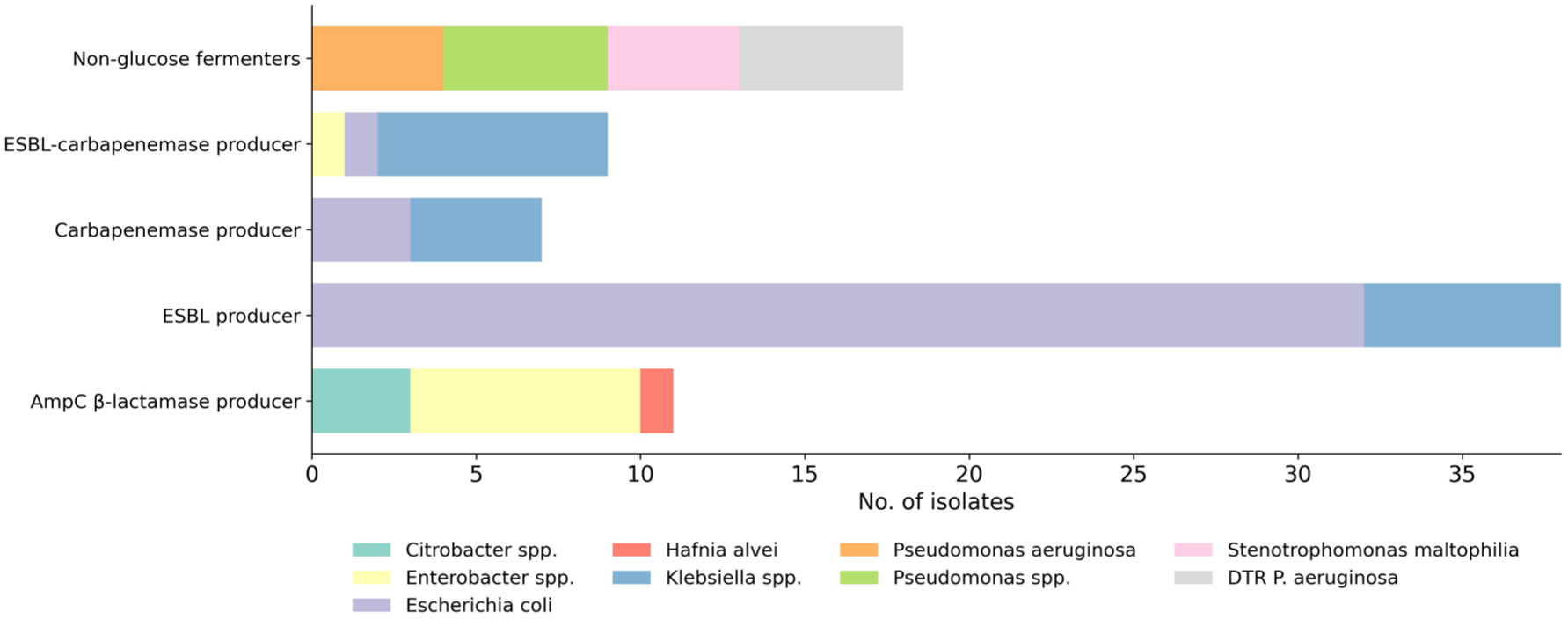
Distribution of multidrug-resistant Gram-negative bacilli isolates from rectal swabs in the high-risk cluster by resistance profiles.
Comparison of empiric antibiotic therapy guided by a clinician-driven approach versus a hypothetical ML-based risk stratification model and prior colonisation
The description of the Clinician-driven EAT approach is provided in Table 1. The most frequently administered antibiotics were anti-PsA beta-lactams, particularly carbapenems (406/553, 73.4%). Using this approach, 16/70 (22.9%) episodes of GNB BSI received IEAT. Of these, 14/49 episodes (28.6%) occurred in the high-risk cluster, while 2/21 episodes (9.5%) were in the intermediate-risk cluster. Table S4 details the causative agents and resistance phenotypes of BSI isolates across clusters in episodes receiving IEAT.
Clinician-driven empirical antibiotic therapy and associated rates of IEAT.

Anti-Pseudomonal (PsA) beta-lactams = cefiderocol, ceftazidime, cefepime, ceftazidime-avibactam, ceftolozane-tazobactam, piperacillin-tazobactam, aztreonam, or meropenem. Methicillin-resistant
Vancomycin-resistant
BSI, bloodstream infection; GNB, Gram-negative bacilli; IEAT, inappropriate empirical antibiotic therapy; MRSA, methicillin-resistant
Following a hypothetical empirical antibiotic prescribing approach guided by an ML–based risk stratification model, the 143/553 (25.9%) episodes classified as low risk would have been treated with CTR. None of these episodes would have received IEAT based on microbiological culture results. Tables 2 and 3 present the IEAT outcomes associated with the different antibiotics recommended by FN guidelines, which would have been used in the intermediate- and high-risk groups, respectively.
Hypothetical empirical antibiotic strategies and associated rates of IEAT, guided by an ML-based risk stratification model in intermediate-risk cluster patients.

One case was attributed to the absence of coverage against
BSI, bloodstream infection; 3GCephRE, 3rd generation cephalosporin-resistant Enterobacterales.
Hypothetical empirical antibiotic strategies and associated rates of IEAT, guided by an ML-based risk stratification model with and without prior colonisation data in high-risk cluster patients.

CTZ (
PIT (
MER (
CTV (
One case was attributed to the absence of coverage against
Both CRE cases exhibited co-production of VIM and ESBL, detected in
AZT, aztreonam; BSI, bloodstream infection; CDE, cefiderocol; CRE, carbapenem-resistant Enterobacterales; CTT, ceftolozane/tazobactam; CTV, ceftazidime-avibactam; CTZ, ceftazidime; MER, meropenem; PIT, piperacillin-tazobactam; 3GCephRE, 3rd generation cephalosporin-resistant Enterobacterales.
All regimens in the high-risk cluster combined with rectal colonisation guidance would have reduced IEAT rates compared to their use without guidance: CTZ (18.4% vs. 57.1%,
Figure 2 compares the 7-day antibiotic usage between the Clinician-Driven approach and the hypothetical approach guided by an ML-based risk stratification model—recommending CTR for the low-risk cluster, MER for the intermediate-risk cluster, and CTV with empiric therapy guided by rectal colonisation for the high-risk cluster. The ML-guided strategy would have significantly reduced the use of MER by 29.7% (−2.08 days; 95% CI, −2.42 to −1.73;

Antibiotic-days in the clinician-driven approach and the selected hypothetical approach guided by an ML-based risk stratification model.
Discussion
This study highlights the clinical feasibility and potential benefits of implementing a personalised antibiotic strategy for FN in hematological patients, based on risk stratification determined by a ML-derived model and complemented by prior colonisation data. Current clinical practice largely adheres to standardised protocols for empirical antibiotic prescribing, resulting in uniform treatment across patients, irrespective of individual risk profiles.12,13 This highlights a key limitation of existing FN guidelines, which lack sufficient criteria for risk stratification and do not support personalised therapeutic decision-making.
Our findings demonstrate that ML-driven stratification would allow for a more precise alignment of empirical antibiotic regimens with predicted epidemiological patterns. A major outcome is the significant reduction in the use of broad-spectrum antibiotics among low-risk patients, aligning with antimicrobial stewardship goals and minimises unnecessary exposure to potentially toxic agents.
While anti-PsA antibiotics have historically been the cornerstone of FN management, as supported by seminal studies,14,15 the evolving clinical landscape necessitates a reassessment of this practice. FN is increasingly heterogeneous, encompassing non-infectious entities such as CAR-T therapy-associated cytokine release syndrome, which further challenges traditional empirical antibiotic approaches. 16 CAR-T-associated fever, often driven by inflammatory cytokines rather than infection, exemplifies a scenario in which broad-spectrum anti-PsA coverage may not always be warranted, particularly in low-risk patients. 17 However, future studies are needed to validate this hypothesis in real-world settings and to ensure that this strategy does not compromise patient outcomes.
A critical concern in FN management is the rate of IEAT, particularly in high-risk patients with MDR-GNB BSI.1–3 Our study highlights the ability of the ML model to stratify these patients effectively, would enable timely administration of broad-spectrum antibiotics when necessary and would reduce the incidence of IEAT. Importantly, additional consideration of colonisation status in high-risk patients further refines empirical therapy decisions. Colonisation with MDR-GNB frequently precedes BSI, and concordance between colonising strains and subsequent bloodstream pathogens is well-established in the literature.18–21 Leveraging colonisation data as a clinical adjunct in high-risk patients allows for empirical regimens to be more precisely tailored to local microbiological profiles, thereby potentially decreasing the risk of IEAT.
This study has several limitations. First, it is a single-centre analysis, and the findings may not be generalisable to institutions with different patient populations and resistance profiles. Because the ML-based model was applied to an expanded dataset from the same institution in which it was originally developed, external validation in independent cohorts is warranted to confirm its performance in other clinical settings. Second, the study is theoretical, based on retrospective data, necessitating prospective validation to confirm the safety and efficacy of ML-guided strategies. The low number of MDR-GNB BSI events also limits the generalisability of the findings, particularly in regions with higher antimicrobial resistance rates. Third, it is unknown whether rectal colonisation status influenced clinical decisions, limiting comparison with the proposed strategy. Another critical area requiring further investigation is the safety of treating low-risk patients with negative cultures using antibiotics lacking coverage against
Conclusion
This study demonstrates that ML–based risk stratification models could enable the tailoring of antibiotic regimens, reducing unnecessary broad-spectrum antibiotic use in low-risk patients and optimising coverage in high-risk groups, with a consequent reduction in IEAT. Further studies are needed to confirm in real-world settings that this personalised antibiotic approach does not compromise clinical outcomes and, on the contrary, may help reduce infections caused by multidrug-resistant organisms.
Supplemental Material
sj-docx-1-tai-10.1177_20499361251411061 – Supplemental material for Towards personalised empirical antibiotic therapy in febrile neutropenia: a theoretical model based on machine learning and prior colonisation with multidrug-resistant gram-negative bacilli – a retrospective proof-of-concept cohort study
Supplemental material, sj-docx-1-tai-10.1177_20499361251411061 for Towards personalised empirical antibiotic therapy in febrile neutropenia: a theoretical model based on machine learning and prior colonisation with multidrug-resistant gram-negative bacilli – a retrospective proof-of-concept cohort study by Antonio Gallardo-Pizarro, Christian Teijón-Lumbreras, Tommaso Francesco Aiello, Patricia Monzó-Gallo, Ana Martinez-Urrea, Cynthia Terrones-Campos, Guillermo Cuervo, Laura Morata, Pedro Castro, Climent Casals, Mateu Espasa, Jordi Esteve, Josep Maria Nicolas, Josep Mensa, Alex Soriano and Carolina Garcia-Vidal in Therapeutic Advances in Infectious Disease
Footnotes
Acknowledgements
We are grateful for the support provided to TFA, who received funding from Instituto de Salud Carlos III, under the file code CM23/00277, pursuant to the ‘Resolución de la Dirección del Instituto de Salud Carlos III, O.A., M.P.’ dated December 13, 2023, which awards the Rio Hortega Contracts, also co-funded by the European Union.
Declarations
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
