Abstract
Background:
Vietnam has one of the highest hepatitis B virus (HBV) infection rates, with approximately 8 million people affected. Although antiviral drug resistance mutations have been reported in treatment-naïve patients with chronic hepatitis B, there is limited data on primary drug resistance mutations in circulating genotypes within this population.
Objectives:
This study aimed to investigate primary antiviral drug resistance mutations and common HBV genotypes in treatment-naïve patients with chronic hepatitis B, particularly in cases without well-characterized resistance profiles.
Design:
A cross-sectional study.
Methods:
We analyzed HBV genotypes and antiviral drug resistance mutations in 113 treatment-naïve patients with chronic hepatitis B in the Yenphong Medical Center, Bacninh Vietnam. The reverse transcriptase (RT) region of the HBV polymerase genes was sequenced to detect mutations.
Results:
Genotypes B, C, and G were identified in 85.0% (96/133), 14.1% (16/133), and 0.9% (1/133) of treatment-naïve patients with chronic hepatitis B, respectively. Mutations in the RT region associated with antiviral drug resistance were detected in 32.7% (37/113) of patients. In addition, the most frequent resistance mutations were rtV207M (89.2%, 33/37), followed by A194T, L180M + M204V, V173L + M204I + L80I, and A181T + V207M + A181T, each observed in 2.7% (1/37). Notably, no significant associations were found between resistance mutations and HBV genotype, gender, age, hepatitis B e-antigen status, baseline HBV DNA levels, or level of alanine aminotransferase, aspartate aminotransferase, gamma-glutamyl transferase.
Conclusion:
This study highlights the presence of primary resistance mutations in treatment-naïve patients and underscores the importance of genotypic screening prior to initiating therapy. These findings may inform treatment strategies and help reduce the risk of treatment failure, liver cirrhosis, and hepatocellular carcinoma.
Keywords
Introduction
Chronic hepatitis B virus (HBV) infection is a leading cause of liver decompensation, cirrhosis, hepatocellular carcinoma, and mortality worldwide, affecting an estimated 360 million individuals and accounting for approximately 1 million deaths annually.1–3 According to the World Health Organization, the global prevalence of HBV infection varies substantially, with the highest rates reported in Africa and the Western Pacific. 3 Vietnam, as a developing country in the Western Pacific region, has among the highest national HBV prevalence rates, with approximately 8 million individuals infected. 4 The distribution of HBV in Vietnam is highly heterogeneous due to variations in ethnicity, socioeconomic status, and regional healthcare access.5,6
Early detection, monitoring, and timely treatment of chronic hepatitis B infection are critical to reducing disease progression and mortality. 7 Significant advances have been made in the treatment of chronic hepatitis B, particularly through the use of interferon-alpha and nucleotide analogs (NAs). Currently, approved NAs include six antiviral drugs: Lamivudine (LAM), Adefovir dipivoxil (ADV), Tenofovir disoproxil fumarate (TDF), Entecavir (ETV), Tenofovir alafenamide (TAF), and Telbivudine (LDT).8–10 Despite their efficacy, NAs can cause adverse effects that complicate clinical management. Moreover, long-term use may exert selective pressure, allowing drug-resistant HBV variants to emerge and persist. 11 Notably, treatment with NAs has been shown to induce specific resistance mutations, including in treatment-naïve patients with chronic hepatitis B.12,13 These resistance mutations typically occur in the reverse transcriptase (RT) region of the HBV polymerase gene. Initial investigations of such mutations focused on patients receiving LAM, ADV, ETV, and LDT.14–17 Among them, LAM-resistant mutations are the most frequently reported, with the rtM204I/V mutation being the most prevalent. 18
The HBV genome is genetically diverse and has been classified into ten genotypes (A–J) based on a nucleotide divergence of ⩾8% across the entire genome. In addition, sub-genotypes differ from one another by at least 4%. The distribution of HBV genotypes varies significantly across racial groups and geographical regions. Genotype A is primarily found in Europe, where subgenotype A2 is predominant, and in Africa, where subgenotype A1 is more common. Other genotypes are distributed as follows: genotypes B and C in Asia; genotype D in Africa, Europe, the Mediterranean region, and India; genotype E in West and Southern Africa; genotype F in South and Central America; genotype G in France and the United States; genotype H in Mexico and Central America; genotype I in Vietnam and Laos; and the recently identified genotype J in the Ryukyu Islands of Japan.19,20 In Vietnam, genotypes B and C are most prevalent, with genotype B being more common than genotype C. 21 Conversely, genotypes A, I, and G are rarely observed, while sub-genotypes B2–B5 and C5–C16 are predominant. 21 Given this distribution, the present study aimed to investigate primary antiviral drug resistance mutations in circulating HBV genotypes among treatment-naïve patients with chronic hepatitis B in Bacninh, Vietnam. The study also sought to evaluate the clinical relevance of these genotypes in guiding treatment strategies.
Materials and methods
Patients
This study included 113 treatment-naïve patients diagnosed with chronic hepatitis B between 2022 and 2023 at the Yenphong Medical Center in Bacninh, Vietnam. Inclusion criteria were as follows: (1) patients with an HBV viral load ⩾103 copies/mL at the time of initial sampling during the study period; (2) no prior treatment with HBV antiviral drugs; (3) complete laboratory results for alanine aminotransferase (ALT), aspartate aminotransferase (AST), gamma-glutamyl transferase (GGT) and hepatitis B e-antigen (HBeAg) rapid test; (4) age 18 years or older. Exclusion criteria included: (1) co-infection with hepatitis C virus (HCV) and/or human immunodeficiency virus (HIV); and (2) diagnosis of hepatocellular carcinoma. All enrolled patients had tested positive for hepatitis B surface antigen (HBsAg) for at least 6 months and were confirmed negative for both HIV and HCV. In addition, each patient had an HBV viral load ⩾103 copies/mL and had not received any prior antiviral treatment. The study was approved by the Ethics Committee of the Hanoi University of Public Health (Approval No. 224/2023/YTCC-HD3), dated May 10, 2023, granted by the director of the Yenphong Medical Center, dated March 28, 2023. This study was conducted in accordance with the principles of the Declaration of Helsinki. Informed consent was obtained from all participants. In cases where patients were unable to provide consent due to medical conditions, cognitive impairments, or severe illness, consent was obtained from their next of kin. Following ethics approval on May 10, 2023, patients recruited prior to this date, specifically those who had visited the Yenphong Medical Center in 2022, were included using stored serum samples. These patients were subsequently contacted by phone or during in-person follow-ups at the medical center to obtain informed consent and additional clinical information. Especially, data including HBV viral load, treatment status, liver enzyme levels (ALT, AST, and GGT), HBeAg status, and age were retrieved from medical records.
Quantification of HBV
HBV DNA was detected and quantified using quantitative polymerase chain reaction (qPCR), in accordance with the manufacturer’s instructions for the TOPSENSI® HBV QPCR Kit (ABT Equipment Co., Ltd., Hanoi, Vietnam). All tests were performed automatically at the Yenphong Medical Center, Bacninh, Vietnam.
Polymerase chain reaction
HBV DNA was extracted from 200 µL of serum using the DNA/RNAprep MAGBEAD kit (Nam Khoa Trading and Services Co., Ltd., Ho Chi Minh city, Vietnam), following the manufacturer’s protocol. The RT region of the HBV gene (540 bp) was amplified via nested PCR using two sets of specific primers: CLC 882 for the first-round PCR, and CLC 188 m for the second-round PCR. Amplifications were performed on a SimpliAmp Thermo Cycler (Applied Biosystems, Thermo Fisher Scientific, Inc., Waltham, MA, USA). The thermocycling protocol included an initial denaturation at 95°C for 15 min, followed by 40 cycles of 95°C for 15 s, 57°C for 30 s, and 72°C for 30 s, with a final extension at 72°C for 10 min.
Sequencing the HBV-RT region
The size of the PCR products was assessed by electrophoresis on a 2% agarose gel using a PowerPac Universal Bio-Rad electrophoresis machine (Bio-Rad Laboratories, Inc., Hercules, CA, USA). PCR products were purified using the Illustra ExoStar Enzymatic cleanup system (Cytiva, Marlborough, MA, USA). Sequencing reactions were then performed on the target gene segment using a single primer (CLC 887 and CLC 188 m) and dideoxynucleotide (ddNTP) labeled with fluorescent dyes: green for adenine, red for thymine, black for guanine, and blue for cytosine. Capillary electrophoresis was conducted on the ABI 3130xl Genetic Analyzer (Applied Biosystems, Thermo Fisher Scientific, Inc., Waltham, MA, USA), following the manufacturer’s protocols. The resulting sequences were analyzed using Sequencing Analyzer software, version 5.2 (Applied Biosystems, Thermo Fisher Scientific, Inc., Waltham, MA, USA).
Sequence analysis
To confirm the specificity of each amplicon, the obtained sequences were analyzed using the GenBank Basic Local Alignment Search Tool (BLAST). Sequences from the same patient were aligned to a reference sequence, and any discrepancies were manually corrected through visual inspection using Sequencing Analyzer software, version 5.2 (Applied Biosystems, Thermo Fisher Scientific, Inc., Waltham, MA, USA). The consensus sequence of each HBV isolate was then submitted to https://blast.ncbi.nlm.nih.gov/Blast.cgi to determine the HBV subgenotype and to predict phenotypic resistance based on genotype-specific mutations in the polymerase gene (RT mutations).
Statistical analysis
All statistical tests were 2-tailed, and a p-value of <0.05 was considered the threshold for statistical significance. Data management and statistical analyses were performed using SPSS software, version 26.0 (IBM Corp, Armonk, NY, USA).
Results
Clinical characteristics of treatment-naïve patients with chronic hepatitis B
A total of 113 treatment-naïve patients with chronic hepatitis B were enrolled in this cross-sectional study at the Yenphong Medical Center, Bacninh, Vietnam. The clinical characteristics of the study population are presented in Table 1. Among these patients, 53.1% were male and 46.9% were female, with a mean age of 36 years (range: 18–71 years). Elevated levels of serum liver enzymes were observed in the following proportions of patients: ALT ⩾ 41 U/L in 49.6%, AST ⩾ 37 U/L in 41.6%, and GGT ⩾ 50 U/L in 26.5%, with corresponding mean values of 59.4, 51.9, and 56.3 U/L, respectively. The mean serum HBV DNA level was 5.82 log10 copies/mL (range: 2.8–8.98), and 39.8% of these patients were HBeAg seronegative. The distribution of HBV genotypes was as follows: genotype B in 96 patients (85%), genotype C in 16 patients (14.1%), and genotype G in one patient (0.9%). Mutations associated with antiviral drug resistance were identified in 37 patients (32.7%).
Clinical characteristics of treatment-naïve patients with chronic hepatitis B at the Yen Phong Medical Center, Bacninh, Vietnam from 2022 to 2023 (n = 113).

ALT, Alanine aminotransferase; AST, Aspartate aminotransferase; GGT, Gamma-glutamyl transferase; HBeAg, Hepatitis B e antigen; HBV, Hepatitis B virus.
Mutation analysis in the HBV-RT region of treatment-naïve patients with chronic hepatitis B
Among the 117 treatment-naïve patients with chronic hepatitis B at the Yenphong Medical Center, Bacninh, Vietnam, a total of 37 (32.7%) exhibited mutations in the RT region of the HBV polymerase gene that may be associated with resistance to antiviral drugs such as LAM, TDF, and/or ADV (Table 2). Mutations associated with LAM resistance were identified in 36 of the 37 samples. In addition, mutations linked to resistance to ADV and TDF were detected in two and one samples, respectively. The identified resistance-associated mutations included rtV207M, rtV173L + rtM204I + rtL80I, rtA181T + rtV207M + rtA181T, rtA194T and rtL180M + rtM204V (Table 2). Among these, the most frequent mutation was rtV207M, observed in 33 of the 37 samples, all of which were associated with LAM resistance. The combined rtL180M + rtM204V mutations were found in one of the 37 samples. Notably, the rtA194T mutation, detected in one sample, was associated with resistance to TDF but remained sensitive to LAM. Furthermore, two combined mutation patterns: rtV173L + rtM204I + rtL80I and rtA181T + rtV207M + rtA181T conferred resistance to both LAM and ADV, each identified in one of the 37 samples (Table 2).
Virological and resistance mutation profile of treatment-naïve patients with chronic hepatitis B to antiviral drug (n = 37).
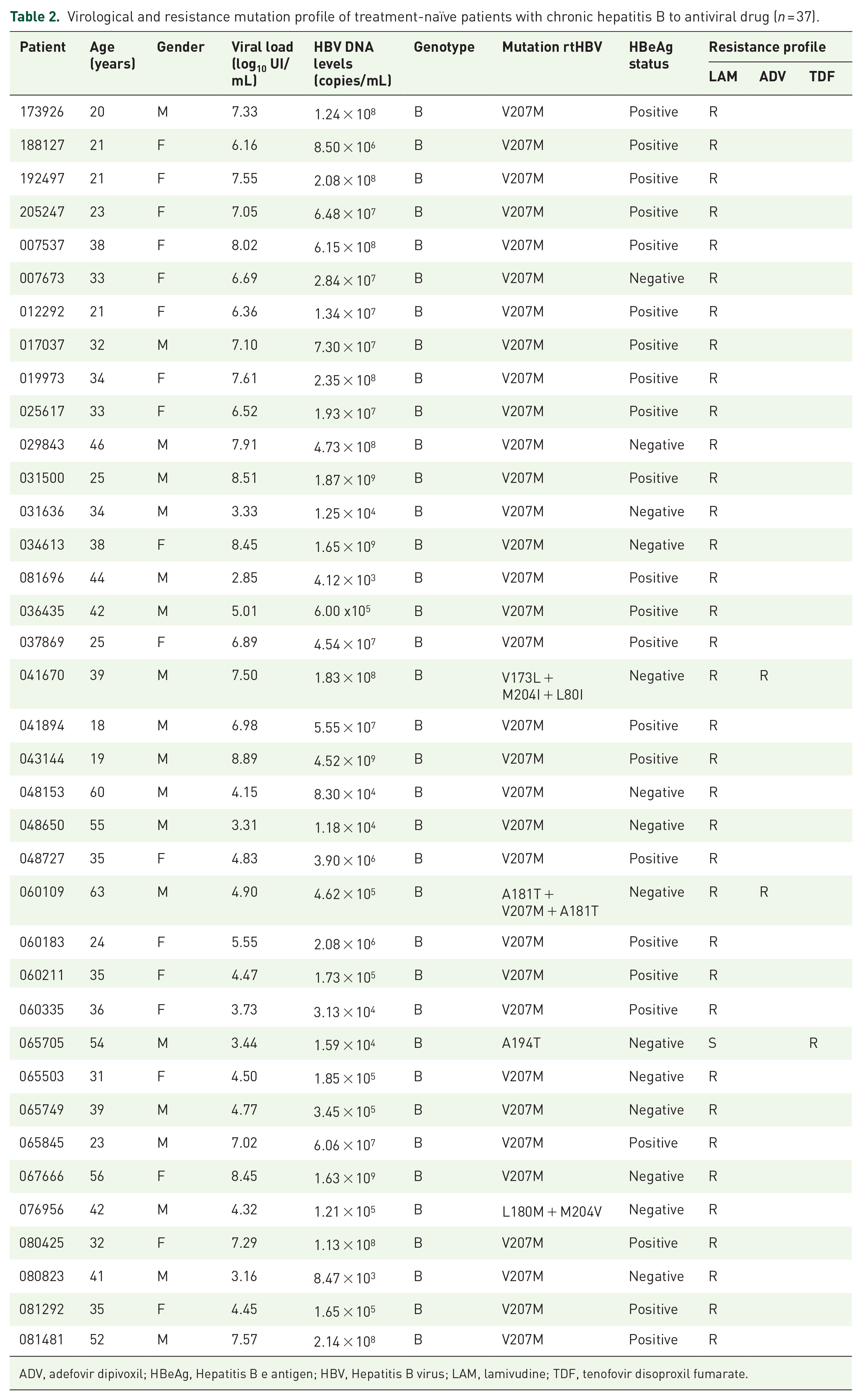
ADV, adefovir dipivoxil; HBeAg, Hepatitis B e antigen; HBV, Hepatitis B virus; LAM, lamivudine; TDF, tenofovir disoproxil fumarate.
Association between mutations and epidemiological characteristics of treatment-naïve patients with chronic hepatitis B
The association between resistance mutations and epidemiological characteristics, including HBV genotype, gender, age, HBeAg status, baseline HBV DNA levels, and serum levels of ALT, AST, and GGT, was not statistically significant among the 113 treatment-naïve patients with chronic hepatitis B (p > 0.05) (Table 3).
Association of epidemiological characteristics of patients with resistance mutations and without resistance mutations in the HBV-RT region.

ADV, adefovir dipivoxil; ALT, alanine aminotransferase; AST, aspartate aminotransferase; GGT, gamma-glutamyl transferase; HBeAg, hepatitis B e antigen; HBV, hepatitis B virus; LAM, lamivudine; TDF, tenofovir disoproxil fumarate.
Discussion
Antiviral drug-resistant mutations have been reported even in treated treatment-naïve patients with chronic hepatitis B, and they carry important clinical implications. Monitoring both primary and secondary resistance mutations in treatment-naïve individuals is essential for determining the most effective therapeutic strategies and for preventing progression to cirrhosis and hepatocellular carcinoma.13,22 In this study, we analyzed the distribution of HBV genotypes and the presence of primary resistance mutations in 113 treatment-naïve patients with chronic hepatitis B at the Yenphong Medical Center, Bacninh, Vietnam. Three HBV genotypes were identified: B, C, and G. Resistance to LAM was common among these patients, with genotype B showing the highest prevalence of the rtV207M mutation.
The predominant HBV genotypes observed in Vietnam are B and C, while genotypes A, I, and G are relatively rare. Sub-genotypes commonly found include B2–B5 and C5–C16. 21 Genotyping plays a crucial role in evaluating patient prognosis, diagnosis, and treatment, as the HBV transcription process has a high error rate, resulting in the emergence of numerous viral variants. Notably, Liu et al. demonstrated that the location of the mutation is genotype-dependent. 23 In this study, HBV genotypes were determined in all 113 treatment-naïve patients with chronic hepatitis B. Among these, genotype B accounted for 85% of cases, genotype C for 14.1%, and genotype G for 0.9%. Although genotype G was rarely detected, genotypes B and C remain the most epidemiologically and clinically significant in Vietnam. Previous studies have shown that genotypes B and C are commonly associated with vertical transmission. 24 In addition, Lin et al. reported that genotype C is associated with a higher risk of cirrhosis and hepatocellular carcinoma compared to other genotypes. 19 For patients infected with genotypes B and C who have not previously received antiviral treatment with ETV or TDF is recommended that treatment is indicated. 25 However, treatment decisions should be made with caution, as HBV genotype may influence the frequency and pattern of treatment-emergent resistance mutations in treatment-naïve patients.
Further investigations into the presence of antiviral drug resistance mutations in treatment-naïve patients with chronic hepatitis B are warranted due to its significant clinical implications. In this study, direct sequencing of HBV DNA targeting the RT region of the polymerase gene was conducted to monitor resistance-associated mutations. Resistance mutations were identified in 37 of the 113 treatment-naïve patients (32.7%). These findings are consistent with previous reports, which have documented RT mutations in treatment-naïve patients at the rate of 56%, 26 38%, 27 46%, 28 and 47.9%. 29 However, in contrast, a study conducted in the Northern and Northeastern regions of Brazil found that only 2.6% (5/189) of treatment-naïve patients harbored primary resistance-associated mutations. 30 The variation in mutation prevalence among studies may be attributed to factors such as sample size, geographic region, and patient lifestyle. In Asia, the reported prevalence of resistance mutations among treatment-naïve patients ranges from 1% to 20%. 30 In Vietnam, where chronic HBV infection remains highly endemic, approximately 6% of the adult population are carriers, a situation influenced by the country’s tropical climate and developing status. This presents ongoing challenges to public health and social infrastructure.
Based on the 42 known RT mutation sites and the classification system proposed by Liu et al., 23 the mutations identified in this study were primarily categorized as primary drug resistance mutations, those that directly confer resistance by reducing drug susceptibility. These included amino acid substitutions such as A194T, A181T, M204I, and M204V. The secondary or compensatory mutations, which compensate for the impaired replication fitness defects caused by primary mutations, included L80I, L180M, and V173L. In addition, the V207M mutation, while associated with drug resistance, is currently considered a putative resistance mutation, as its role has yet to be validated through in vitro studies.
The rtA194T mutation may be associated with resistance to TDF. In vitro studies have shown that this mutation reduces sensitivity to TDF, particularly when it co-occurs with LAM-resistant mutations such as rtM204V and rtL180M. However, the clinical significance of the rtA194T mutation remains unclear and warrants further investigation through long-term follow-up studies.16,31 HBeAg-negative patients may be at greater risk of developing the rtA194T mutation and, consequently, resistance to antiviral therapy. In such cases, ETV may represent a more suitable option. 32 Among the mutations associated with LAM resistance, rtL180M + rtM204V and rtV207M are particularly notable. The rtM204V mutation is considered a primary resistance mutation due to its direct effect on reducing HBV susceptibility to LAM. In contrast, rtL180M is classified as a compensatory or secondary mutation, as it may restore or enhance the replication capacity of the virus impaired by primary mutations.33,34 The rtM204V mutation is one of the most frequently observed in studies of treatment-naïve patients. 35 Along with the V207M mutation, it is strongly associated with LAM resistance. Nevertheless, according to current guidelines for the diagnosis and treatment of chronic hepatitis B, LAM is no longer recommended as a first-line antiviral agent due to a low genetic barrier to resistance. 25 The rate of antiviral drug resistance with LAM can reach approximately 80% after 5 years of therapy.36,37 In accordance with recommendations from the Vietnamese Ministry of Health, ETV and TDF are the preferred first-line agents for chronic hepatitis B who have not yet developed cirrhosis. While resistance mutations to ADV are more commonly observed in patients previously treated with LAM, they are relatively rare in treatment-naïve individuals. 38 Previous studies have identified seven key mutations associated with ADV resistance: rtL80V/I, 39 rtS85A,40,41 rtV84M, 41 rtA181V/T,41,42 rtV214A,40,41 rtQ215S/H,40,41 and rtN236T.41,42 In contrast, findings from the present study revealed the presence of the rtL80I and rtA181T mutations in treatment-naïve patients with chronic hepatitis B.
Patients with antiviral drug resistance mutations were compared with those without mutations, and no significant differences were observed. These findings are consistent with those of the previous study. 30 However, Zhang et al. conducted a meta-analysis to evaluate the prevalence of naturally occurring drug resistance mutations in treatment-naïve patients with chronic hepatitis B. 43 Their analysis demonstrated that male patients, individuals infected with HBV genotype C, and those who were HBeAg-negative exhibited a higher rate of natural mutations compared to female patients, those with HBV genotypes B or D, and HBeAg-positive individuals. 43 In addition, prior studies have shown that serum levels of ALT, AST, and GGT are not significantly associated with the presence of antiviral resistance mutations. 43 In Vietnam, only a limited number of studies have investigated drug resistance mutations in treatment-naïve patients.5,23 Therefore, further research involving larger cohorts of treatment-naïve patients with chronic hepatitis B is warranted to better understand the evolving patterns and clinical relevance of resistance mutations.
In conclusion, genotypic monitoring is essential for identifying antiviral drug resistance mutations in treatment-naïve patients with chronic hepatitis B, as preexisting mutations in the RT region of the polymerase gene are associated with reduced drug susceptibility. The therapeutic efficacy of antiviral agents may be compromised by various factors, including infection with drug-resistant HBV variants. Consequently, antiviral treatment may not always be effective, even in patients who have not previously received therapy. Implementing HBV genotyping prior to treatment initiation may aid clinical decision making, minimize treatment failure, and reduce the risk of disease progression to cirrhosis or hepatocellular carcinoma.
Footnotes
Acknowledgements
We are grateful to Dr. Bon Trinh, UVA School of Medicine (https://med.virginia.edu/pathology/contact/bon-trinh-ph-d/) and Dr. Thi Thanh Thoa Tran, IU School of Medicine (![]() ) for revising any grammar mistakes and oversights, and for enhancing readability, clarity, and tone of the manuscript.
) for revising any grammar mistakes and oversights, and for enhancing readability, clarity, and tone of the manuscript.
