Abstract
Background:
In previous years, several cutaneous disorders have been associated with human papillomavirus (HPV); however, the exact role of HPV remains largely unknown. The lack of optimization and standardization of the pre-analytical phase forms a major obstacle. The aim of this study was to develop an accurate/patient-friendly sampling method for skin disorders, with cutaneous warts as a case study.
Methods:
Various sample processing techniques, pre-treatment protocols and DNA extraction methods were evaluated. Several sampling methods were examined, that is, skin scrapings, swabs and a tape-based method. Quantification of DNA yield was achieved by beta-globin real-time polymerase chain reaction (qPCR), and a wart-associated HPV genotyping qPCR was used to determine the HPV prevalence.
Results:
All samples tested positive for beta-globin. Skin scrapings had significantly higher yield than both swab and tape-based methods (
Conclusion:
Although skin scrapings produced the highest DNA yield, patient discomfort was an important limitation of this method. Considering that in combination with our optimized DNA extraction procedure, all samples gave valid results with the less invasive swab methods preferred. Standardization of the pre-analytical phase is the first step in establishing a link between HPV and specific skin disorders and may have significant downstream diagnostic as well as therapeutic implications.
Keywords
Background
The family of human papillomaviruses (HPVs) comprises over 200 different genotypes, classified into different genera according to their DNA sequence. Members of the genus alpha-HPV have a specific tropism for mucosal epithelium, while beta and gamma HPVs most frequently cause cutaneous lesions and are commonly referred to as cutaneous HPV types. 1 HPV is capable of causing a wide range of diseases from benign lesions to invasive tumours and although it is most commonly known for its involvement in the development of cervical cancer, there are also other mucosal as well as cutaneous disorders where HPV plays a role (see Table 1). Regarding cutaneous disorders, it still remains extremely challenging to determine the exact role of HPV in their development. Is HPV responsible for the onset of the disease? Is it a co-factor that acts with other carcinogens to amplify the risk of disease? Or is it merely an innocent bystander without any role in disease pathogenesis? 2
To demonstrate that a pathogen causes a disorder, we do not only need a plausible biological mechanism for pathogenesis but also convincing associative epidemiological evidence. 6
As for the epidemiological evidence of HPV involvement in skin disorders, to date, numerous studies are published with conflicting results. For example, more than 100 studies have investigated the relationship between HPV and cutaneous squamous cell carcinoma (SCC). While some studies have failed to find HPV in SCC, most studies do report HPV infection in some SCCs, although with variable percentages (3.2–85.7%). 2 These contradictory results can be attributed to several issues (see Table 2), one of the most prominent being a lack of standardization of the pre-analytical phase, that is, different studies use different sampling methods, sample processing protocols, pre-treatments and DNA extraction systems. The use of inconsistent methods results in differences in not only sensitivity but also specificity. 7 This high degree of between-study heterogeneity presents challenges to grouped study analysis and makes it problematic to establish distinct claims about the causal role of HPV in these disorders. 2
Issues causing conflicting results in studies regarding HPV prevalence in skin disorders.

ELISA, enzyme-linked immunosorbent assay; HPV, human papillomavirus; PCR, polymerase chain reaction.
Following the above-described reasoning, the first step in determining the exact role of HPV in skin disorders is to standardize and optimize the pre-analytical phase. To achieve this objective, there is need for a case study employing a straightforward, easily accessible cutaneous disorder where the causal role of HPV has already been ascertained, that is, cutaneous warts. 12
In this study, we performed a head-to-head comparison of different pre-analytic steps, including sampling methods, sample processing techniques, pre-treatment protocols and DNA extraction methods to develop an accurate, patient-friendly sampling method for skin disorders.
Materials and methods
The optimization of the pre-analytical phase included two separate stages: (1) optimization of the DNA extraction procedure and (2) development of a patient-friendly sampling method. The optimization was performed in preparation for a large clinical trial regarding cutaneous warts, that is, OVW-SA001 trial. 13 Samples were provided by patients visiting the Algemeen Medisch Laboratorium (AML) medical laboratory in December 2017 (Ethical approval number B300201734040).
DNA extraction
Formalin-fixed, paraffin-embedded (FFPE) wart biopsies, skin scrapings, tape and swabs from normal and callous skin were used to optimize the DNA extraction procedure. Multiple target genes were analysed to determine the optimal DNA quantification method for skin samples. Several manuals as well as automatic DNA extraction methods were examined, together with different preservatives, pre-treatment and lysis buffers, and protocol amendments (Table 3). For these latter experiments, multiple skin scrapings from four different patients were used. Skin scrapings were first weighted and divided into equal aliquots before further processing.
Summary of various parameters tested during optimization of the DNA extraction procedure.

Sampling method
Several sampling methods were examined employing cutaneous warts from different individuals, that is, skin scrapings (
Likewise, the performance of two different swabs, that is, cotton (Abbott MC Specimen Collection) and flocked (FLOQSwab Copan Diagnostics, Murietta, CA, USA), was analysed. FLOQSwabs contain short hydrophilic nylon fibres attached to plastic. Due to their design without an internal absorbent core, they do not disperse and entrap the specimen and should therefore provide better DNA yield. 25 In total, 45 warts were sampled by both types of swabs in an alternating order to account for interpatient variability. All samples were stored at 4°C and subsequently extracted according to the optimized DNA extraction protocol (see further). Quantification of the DNA yield was achieved by beta-globin real-time polymerase chain reaction (qPCR) (cell control) 26 and a newly developed HPV qPCR genotyping assay capable of detecting the most prevalent wart-associated HPV types (i.e. HPV1, 2, 3, 4, 7, 10, 27, 41, 57, 60, 63, and 65) was used for HPV detection. 24
Statistical analysis
Data analysis was conducted using MedCalc version 20.111 (MedCalc Software Ltd, Ostend, Belgium). The paired student’s
Results
DNA extraction
The first step to optimize the DNA extraction procedure was to determine the analysis method, that is, DNA quantification method. For this purpose, a total of 21 samples (seven FFPE wart biopsies, seven skin scrapings and swabs from normal skin) were analysed with three different housekeeping genes: two cellular genes, that is, HMBS (hydroxymethylbilane synthase) and beta-globin, and one mitochondrial gene, that is, MTCOI (mitochondrial cytochrome oxidase subunit 1). All samples as well as the negative control tested positive for MTCOI. In concordance with previous research (unpublished data), MTCOI seems to be ubiquitous and requires the use of cut-off values for data analysis. This makes unambiguous analysis challenging and MTCOI is therefore not recommended for further cutaneous sample evaluation.
The results of DNA quantification by HMBS and beta-globin are depicted in Figure 1. Although beta-globin consistently exhibits higher DNA yield than HMBS, the DNA yields were only significantly different in biopsy and swab samples (paired student’s

Quantification of the DNA yield of several sample types with two different target genes, that is, beta-globin and HMBS. The beta-globin DNA quantification was significantly higher in biopsy and swab samples in comparison with HMBS quantification (
The next step was the comparison of several DNA extraction systems. For this purpose, skin-scraping aliquots were used. The results of this experiment are depicted in Table 4. The lowest average DNA yield was obtained
Summary of the analysis of two DNA extraction parameters (DNA extraction system and sample preservative).

In total, 70 skin scrapings from four different patients were used for these experiments. The scrapings were weighted and aliquoted in equal proportions prior to testing. beta-globin is used as cellular control of a successful DNA extraction and all sample should test positive. The percentage of beta-globin positive samples is depicted in the third column together with the total amount of samples tested between brackets. The average DNA yield (ng/µl) per protocol variable is depicted in the fourth column.
The following step was to determine the optimal sample preservative. All samples, that is, the previously described skin scraping aliquots, were stored in their respective preservatives for a period of 5 days at 4°C and extracted using the NucliSens easyMAG platform (bioMérieux, Boxtel, The Netherlands). The results depicted in Table 4 demonstrate that the DNA yield varied significantly according to the preservative used (Friedman test
The results of the analysis of several pre-treatment buffers, lysis buffers and protocol amendments are not further discussed in detail. In summary, the final optimized DNA extraction protocol involved sample storage in MC medium (Abbott Molecular Inc., Des Plaines, IL, USA) at 4°C, and overnight digestion in a buffer containing 1 mg/ml proteinase K and 0.38 M EDTA (pH 8) at 56°C and 1400 rpm (Thermo–Shaker TS-100C, Biosan, Riga, Latvia), followed by automated extraction on the NucliSENS® easyMAG® system (Generic 2.0.1 protocol, bioMérieux, Boxtel, The Netherlands) and DNA quantification with beta-globin.
Sampling method
After the DNA extraction protocol was optimized, we performed a comparison of the different sampling methods. All samples tested positive for beta-globin and were considered valid. Skin scrapings had significantly higher yield than both swab and tape-based methods (Kruskal–Wallis test;
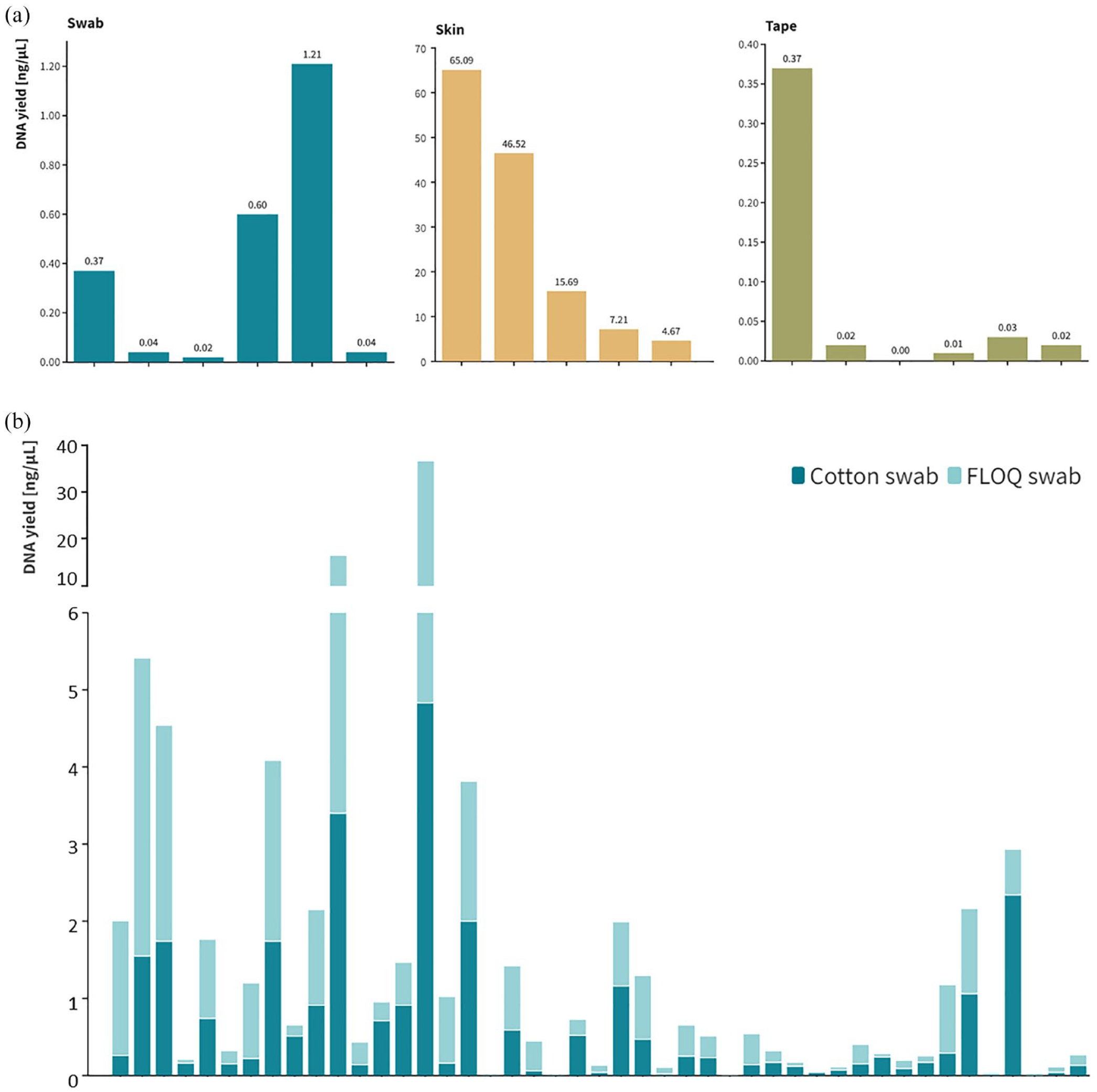
(a) Comparison of the DNA yield (ng/µl) of all the examined sampling methods, that is, swab (
Overview of qualitative comparison between HPV prevalence in cotton and flocked swabs.

In total, 45 warts were sampled with both swabs. These swabs were examined for the presence of 12 distinct cutaneous HPV types (i.e. a total of 540 single-plex reactions). The results are shown in the form of frequencies (the number of samples that simultaneously satisfy the specific criteria indicated in the column and row). Although there were some discrepancies in HPV prevalence between both swabs, an overall good strength of agreement was found (kappa,
HPV, human papillomavirus.
Discussion
In this study, we describe a comprehensive optimization of the pre-analytical phase of cutaneous samples determined for HPV evaluation. A robust, standardized protocol for sample processing and DNA extraction of several types of skin samples (i.e. skin scrapings, swabs, tape and FFPE biopsies) was devised. Various head-to-head comparisons between different non-invasive sampling techniques were performed to determine the optimal sampling method.
Currently, tissue biopsy is considered the gold standard in sampling skin disorders for HPV detection. Biopsies contain DNA derived from not only superficial but also deeper epithelial layers and can yield information about the infectious HPV reservoir in basal stem cells.
6
In addition, they also provide histological background to a skin disorder, that is, identification of affected skin layers and histological localization of viral particles by immunohistochemistry with HPV-specific antibodies. However, a skin biopsy is a rather invasive procedure, which requires skilled personnel and is accompanied by a moderate cost. These issues often discourage subject participation and make biopsies impractical for large studies.
6
A study by de Koning
While there have been some comparisons between certain skin sampling methods, 16 there have not been, to our knowledge, any direct comparisons between non-invasive sampling techniques. As regards the current study, as far as sampling is concerned, although a somewhat better DNA yield was found in skin scrapings, patient discomfort was an important limitation of this method. Seeing that in combination with the optimized DNA extraction procedure all samples gave valid result, with the less invasive methods preferred.Tape sampling is quick and straightforward; however, the subsequent DNA extraction is more challenging due to the adhesiveness and rigidity of the tape. 17 According to current literature, tape-based methods also exhibit low reproducibility caused by variable operator sampling techniques. 6 The last non-invasive method analysed was skin swabs. Swabs comprise a very straightforward sampling method, allowing for quick, painless sampling that can be repeated multiple times with little risk and patient inconvenience. 6 An additional advantage of swabs is the option for automated pre-analytical processing, which is not feasible with the alternative methods. Accordingly, the performance of both cotton and flocked swabs was also demonstrated to be equal.
As described in current literature, the main disadvantage of all non-invasive methods is that they only access the superficial epithelial layers, making it difficult to assess if a positive sample represents contamination, carriage, transient or persistent infection.
6
As previously mentioned, a study by de Koning
In conclusion, a robust pre-analytical phase is the first step necessary to establish an unambiguous link between HPV and certain skin disorders. Furthermore, this optimized sampling technique can also be employed for HPV detection in other mucosal HPV-associated disorders such as head-and-neck, penile as well as anal tumours. Clarification of the viral mechanism in HPV-related disorders may lead to more targeted treatment modalities, reduction in disease burden/healthcare costs, and overall better patient outcomes.2,6 Future studies should not only examine the effects of specific HPV genotypes and viral loads in patients but also explore the longitudinal, subsequent development of the underlying skin disorders.

