Abstract
Pulmonary arterial hypertension is a devastating pulmonary vascular disease, in which the pathogenesis is complicated and unclear. Pulmonary arterial smooth muscle cells (PASMCs) proliferation is a key pathological feature of pulmonary arterial hypertension. It has been shown that ubiquitin-specific protease 7 (USP7) is involved in cancer cell proliferation via deubiquitinating and stabilizing E3 ubiquitin ligase mouse double minute 2 (MDM2). However, the effect of USP7 and MDM2 on platelet-derived growth factor (PDGF)-induced PASMCs proliferation is uncertain. This study aims to explore this issue. Our results indicated that PDGF up-regulated USP7 protein expression and stimulated PASMCs proliferation; this was accompanied with the increase of MDM2, forkhead box O4 (FoxO4) reduction and elevation of CyclinD1. While prior transfection of USP7 siRNA blocked PDGF-induced MDM2 up-regulation, FoxO4 down-regulation, increase of CyclinD1 and cell proliferation. Pre-depletion of MDM2 by siRNA transfection reversed PDGF-induced reduction of FoxO4, up-regulation of CyclinD1 and PASMCs proliferation. Furthermore, pre-treatment of cells with proteasome inhibitor MG-132 also abolished PDGF-induced FoxO4 reduction, CyclinD1 elevation and cell proliferation. Our study suggests that USP7 up-regulates MDM2, which facilitates FoxO4 ubiquitinated degradation, and subsequently increases the expression of CyclinD1 to mediate PDGF-induced PASMCs proliferation.
Keywords
Introduction
Pulmonary arterial hypertension (PAH) is a devastating disease characterized by a progressive increase in pulmonary vascular resistance. 1 Persistent pulmonary vasoconstriction, pulmonary vascular remodeling and thrombosis in situ have been considered to be the pathological mechanisms of PAH, 2 in which pulmonary vascular remodeling caused by pulmonary arterial smooth muscle cells (PASMCs) proliferation and extracellular matrix deposition resulting in intima thickening and media hyperplasia, is a key pathological feature of PAH. 3 Therefore, it is meaningful to explore the molecular mechanisms underlying PASMCs proliferation and investigate new strategy for prevention of pulmonary vascular remodeling and management of PAH.
Ubiquitin proteasome system, a major protein quality and quantity control system, has been previously shown to be associated with the pathogenesis of PAH. 4 –6 Ubiquitin-specific proteases (USP) constitute the largest subfamily of deubiquitinating enzymes, which deconjugate ubiquitin chains from specific protein substrates, and thereby inhibit the degradation of the targeting substrates or the regulation of their subcellular localization. 7 , 8 USP7, the first identified deubiquitinating enzymes, has been implicated in cell cycle, apoptosis and tumourigenesis through stabilization of the associated proteins. 9 –13 Over-expression of USP7 has been found to be associated with lymph node metastasis and poor prognosis in epithelial ovarian cancers. 14 , 15 Study has shown that USP7 deubiquitinates and stabilizes E3 ubiquitin ligase mouse double minute 2 (MDM2), leading to ubiquitinated degradation of p53 and thereby promoting cell growth in ovarian cancer cells. 16
MDM2, an important E3 ubiquitin ligase, functions as an oncoprotein to regulate cell cycle and proliferation, control DNA replication and checkpoint response. 17 –20 It has been found that MDM2 is overexpressed in various human tumors. 21 –23 Previous studies have shown that the gene amplification/gain of copy number of MDM2 or up-regulation of its expression is associated with the pathogenesis of cancers. 21 , 24 , 25 Furthermore, MDM2 has been recently shown to be involved in the development of several types of non-oncologic disease, such as PAH, cardiovascular disorder and diabetic kidney disease. 26 –28 MDM2 is known to interact with WT p53 and inhibit p53-mediated transcriptional activation through degradation of WT p53 due to its ubiquitin ligase function. 17 , 29 , 30 Recently, a study has also found that MDM2 interacts with forkhead box O4 (FoxO4) and mediates its ubiquitinated degradation, and thereby promotes tumorigenesis. 31
FoxO, subfamily of Fox proteins, consists of four members (FoxO1, FoxO3, FoxO4 and FoxO6) and plays an important role in cell cycle progression, cell survival, tumor suppression and DNA repair pathways through interaction with other transcription factors to regulate target gene expressions. 32 , 33 Post-translational modifications (including various levels of phosphorylation and ubiquitination) regulate the function of FoxO. 34 , 35 FoxO4 has been reported to play a role in carcinogenesis. 36 The expression of FoxO4 has been described to be decreased in gastric cancer tissues. 37 Moreover, down-regulation of FoxO4 is highly correlated with decreased expression of E-cadherin in non-small cell lung cancer. 38 While the activation of FoxO4 has been shown to reduce the expression of CyclinD1, it also suppresses cervical cancer cell growth and survival. 39 It has been elaborated that MDM2 mediates FoxO4-ubiquitinated degradation and subsequently promotes cancer cell proliferation. 31 However, it remains uncertain whether up-regulation of MDM2 by USP7 facilitates FoxO4-ubiquitinated degradation and subsequently promotes PDGF-induced PASMCs proliferation.
Previous studies have reported that USP7 deubiquitinates and stabilizes E3 ubiquitin ligase MDM2, thereby promoting cell growth in ovarian cancer cells. 16 However, it is unclear whether up-regulation of USP7 elevates MDM2 and subsequently promotes cell proliferation in PDGF-stimulated PASMCs. Furthermore, MDM2 has been found to decrease FoxO4 by increasing ubiquitinated degradation. 31 Moreover, activation of FoxO4 has been shown to reduce the expression of CyclinD1 and subsequently inhibit cervical cancer cell growth and survival. 39 However, it is unclear whether up-regulation of MDM2 by USP7 facilitates FoxO4 ubiquitinated degradation and subsequently up-regulates CyclinD1 to promote PDGF-induced PASMCs proliferation. To address these issues, primary cultured PASMCs were prepared from male SD rats; the effect of USP7/MDM2/FoxO4/CyclinD1 axis on PDGF-induced PASMCs proliferation was examined.
Materials and methods
Cell isolation and culture
Healthy male Sprague-Dawley (SD) rats weighing 100–150 g were used in this study. Primary PASMCs was isolated from main pulmonary arteries of rats with our laboratory’s previously described method. 40 Rats were anesthetized with intraperitoneal injections of 30 mg/kg of body weight chloral hydrate. No signs of peritonitis were exhibited after the administration of 10% chloral hydrate. The hearts and lungs were excised and placed in cold phosphate-buffered saline (PBS), and then transferred to an aseptic room. Rats were sacrificed through harvesting of vital organs under anesthesia and leading to exsanguination. Starting from right ventricle, segments of the main pulmonary arteries were separated from the lung, removed from connective and fat tissues surrounding the outer membrane, and then washed twice with cold PBS. The smooth muscle of the pulmonary arteries was dissociated away from the adventitia and intima by scraping off with a bended microscopic tweezers, then sliced into fragments (about 0.5–1 mm2) and placed into a culture flask.
PASMCs were cultured in complete medium consisting of high glucose Dulbecco’s Modified Eagle Medium (DMEM) (Gibco, Grand Isle, NY, USA) supplemented with 10% fetal bovine serum (FBS, Sijiqing, HangZhou, China), 100 U/ml penicillin and 100 μg/ml streptomycin at 37°C with 5% CO2. Cells were passaged by trypsinization using 0.25% trypsin (Genview, Carlsbad, CA, USA) and identified by immunofluorescence using an antibody against smooth muscle α-actin (α-SMA, Sigma-Aldrich, St. Louis, MO, USA). PASMCs were grown to passage 4–8 and then starved overnight by seeding with medium containing 1% FBS prior to use for all studies. Cells were induced to proliferate by platelet-derived growth factor (PDGF) (Peprotech, Rocky Hill, NJ, USA), which was dissolved in distillated water at 1000 μg/ml as the stock solution.
BrdU incorporation assay
PASMCs proliferation was evaluated by BrdU incorporation assay using BrdU Cell Proliferation ELISA Kit (Maibio, Shanghai, China) according to the standard protocol of the manufacturer. Cells were seeded into 96-well plates with the same density of 5 × 103 cells/well, cultured at 37°C for 24 h, and then serum starved (1% FBS in DMEM) overnight. BrdU solution was added into each well at the end of different treatments, followed by incubation at 37°C for 2 h. Thereafter, FixDenat solution was added for 30 min at room temperature after the medium was aspirated. After incubation with anti-BrdU monoclonal antibody conjugated to peroxidase for 90 min at room temperature, peroxidase substrate solution was added and kept at room temperature for 10 min in the dark. The absorbance at a wavelength of 370 nm was measured by a microplate reader (Bio-Rad, Richmond, CA, USA). The blank corresponded to 100 μl of culture medium with or without BrdU.
Small interfering RNA transfection
Small interfering RNA (siRNA) specific to USP7 and MDM2 as well as non-targeting control siRNA, designed and synthesized by GenePharm (Shanghai, China), were transfected to PASMCs using Lipofectamine™ 2000 transfection reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. In brief, cells were seeded into six-well plates for transfection. Thereafter, siRNA and lipofectamine were diluted in serum-free DMEM separately, incubated at room temperature for 5 min, subsequently mixed with each other and incubated at room temperature for 20 min. Finally, cells at 30–50% confluence were transfected with the mixture of siRNA and lipofectamine for 6 h in serum-free DMEM and cultured in complete DMEM for 48 h at 37°C in 5% CO2. The silencing efficiency of siRNA transfection was verified by Western blotting.
Western blotting
Western blotting was performed as previously described. 40 PASMCs were washed three times with ice-cold PBS, lysed with ice-cold RIPA lysate buffer containing 1% henylmethanesulfonyl fluoride (PMSF), protease inhibitors and phosphatase inhibitors for 30 min on ice and centrifuged at 13,000 r/min for 20 min at 4°C. The supernatants were collected as total protein, and protein concentrations were measured by BCA protein assay kit (Pierce, Rockford, IL, USA). Equal amounts of collected protein were separated by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and subsequently transferred onto a nitrocellulose (NC) membrane using Bio-Rad Trans-Blot system. The membranes were blocked with 5% powered non-fat dry milk in Tris buffer saline containing 0.1% Tween-20 (TBST) for 1 h on a rocking platform at room temperature and subsequently incubated with polyclonal or monoclonal primary antibodies against USP7(Abcam, Cambridge, UK; 1:1000 dilution), FOXO4 (Abcam, Cambridge, UK; 1:1000 dilution), MDM2 (Proteintech, Chicago, IL, USA; 1:500 dilution), CyclinD1(Cell Signaling Technology, Danvers, MA, USA; 1:1000 dilution) and β-actin (Santa Cruz Biotechnology, Santa Cruz, CA,USA; 1:1000 dilution) at 4°C overnight. After being washed with TBST, membranes were incubated with horseradish peroxidase-conjugated secondary antibody (Sigma-Aldrich; 1:10,000 dilution) at room temperature for 1 h. Visualization was carried out using the SuperSignal Chemiluminescent Substrate (Pierce Biotechnology, Rockford, IL, USA). The optical density of the bands was measured using Image J software.
Statistical analysis
SPSS software, version 18.0 (SPSS, Inc., Chicago, IL, USA) was used for statistical analysis. Values were presented as mean ± S.D. Data were analyzed using one-way ANOVA followed by a Tukey post hoc test. P < 0.05 was considered to represent significant differences between groups.
Results
PDGF stimulates PASMCs proliferation
To explore whether PDGF stimulates PASMCs proliferation, cells were treated with different concentrations of PDGF (0, 1, 3, 10, 30 ng/ml) for different times (12, 24, 48, 72 h), proliferation was assessed using BrdU incorporation assay. As shown in Fig. 1a, PDGF stimulated PASMCs proliferation in a dose-dependent manner; PDGF at a concentration of 10 ng/ml significantly induced cell proliferation, which increased to 1.97-fold compared with control (P < 0.05). Fig. 1b indicates that PDGF time-dependently induced cell proliferation; 10 ng/ml PDGF for 72 h triggered a 2.66-fold increase in BrdU incorporation compared with control (P < 0.05). These results indicate that PDGF dose- and time-dependently stimulates PASMCs proliferation.
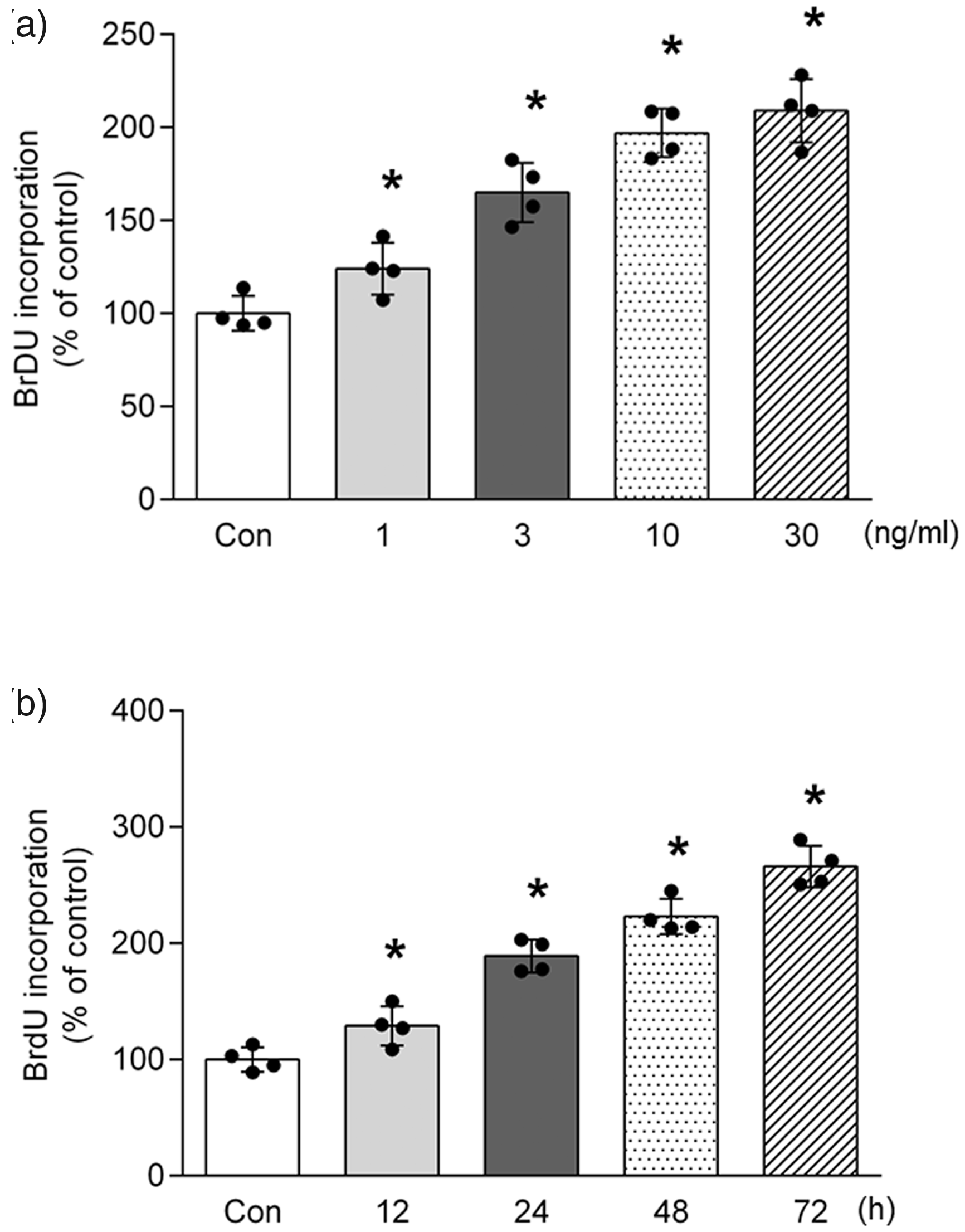
PDGF stimulates PASMCs proliferation. (a) Primary cultured PASMCs were stimulated with various concentrations (0, 1, 3, 10, 30 ng/ml) of PDGF for 24 h; cell proliferation was determined by BrdU incorporation assay (n = 4 per group). (b) Cells were treated with 10 ng/ml PDGF for different times (12, 24, 48, 72 h); cell proliferation was measured by BrdU incorporation assay (n = 4 per group). *P < 0.05 versus control.
PDGF up-regulates USP7 and MDM2, down-regulates FoxO4 and elevates CyclinD1 expression in PASMCs
To investigate whether PDGF regulates the protein expression of USP7, MDM2, FoxO4 and CyclinD1 in PASMCs, primary cultured PASMCs were incubated with 10 ng/ml PDGF for 24 h, and protein expression was measured using Western blotting. Fig. 2a indicates that PDGF increased the protein level of USP7 by 1.50-fold compared with control (P < 0.05). Fig. 2b shows that MDM2 protein was increased to 2.33-fold compared with control upon PDGF stimulation (P < 0.05). As depicted in Fig. 2c, PDGF dramatically decreased the protein level of FoxO4 to 0.56-fold compared with the control group (P < 0.05). Fig. 2d indicates that treatment of cells with PDGF increased CyclinD1 protein level to 2.60-fold over control (P < 0.05). This result suggests that PDGF up-regulates USP7, MDM2 and CyclinD1 protein expression while down-regulates FoxO4 expression in PASMCs.
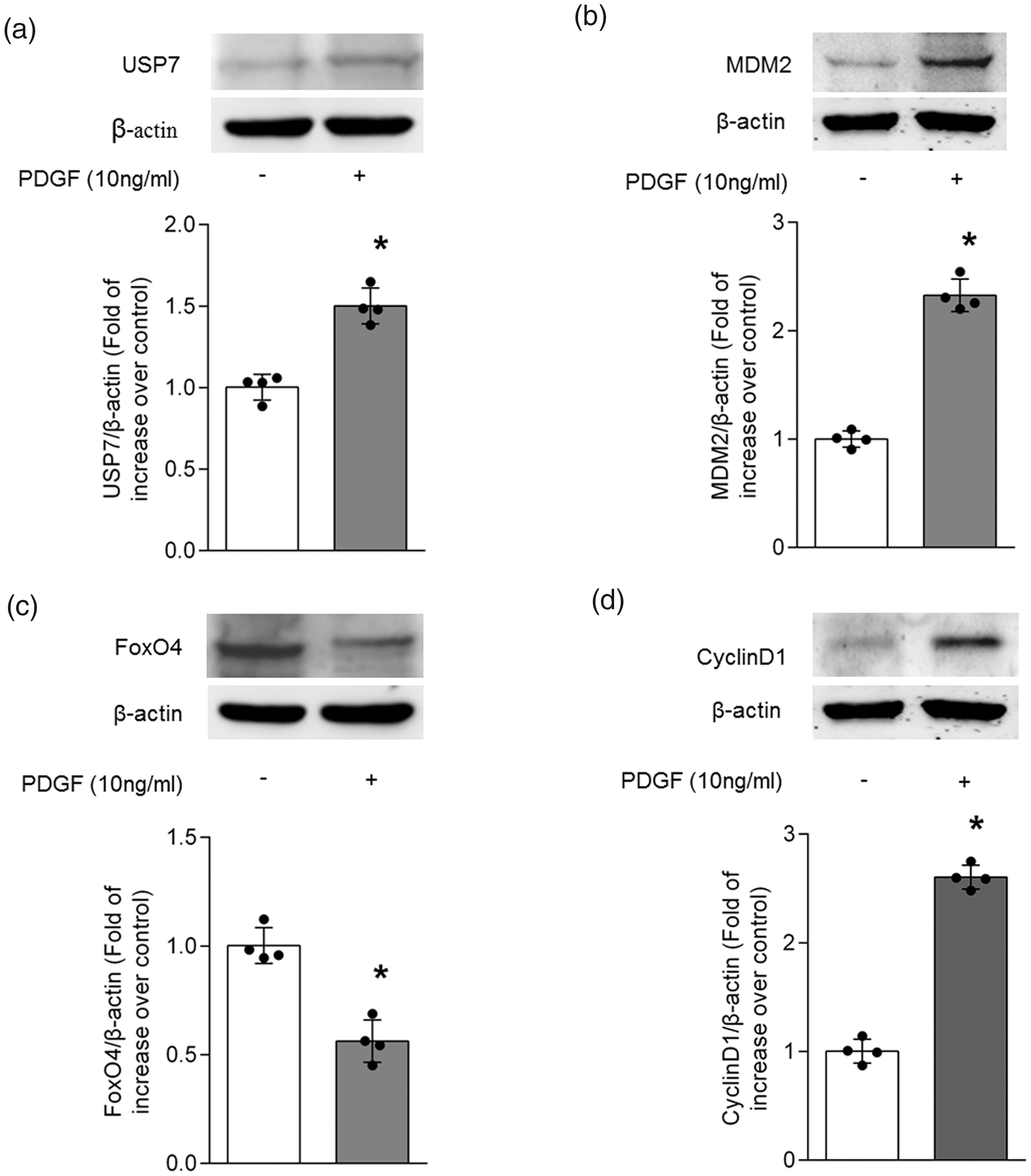
PDGF up-regulates USP7 and MDM2, down-regulates FoxO4 and elevates CyclinD1 expression in PASMCs. PASMCs were stimulated with 10 ng/ml PDGF for 24 h, protein levels of USP7 (a), MDM2 (b), FoxO4 (c) and CyclinD1 (d) were measured using Western blotting. β-actin served as a loading control. A representative blot and quantification of bands are shown (n = 4 per group). *P < 0.05 versus control.
USP7 mediates PDGF-induced up-regulation of MDM2, down-regulation of FoxO4 and elevation of CyclinD1 in PASMCs
To determine whether USP7 mediates PDGF regulation of MDM2, FoxO4 and CyclinD1 expression in PASMCs, cells were first transfected with USP7 sequence-specific siRNA for 24 h and then treated with 10 ng/ml PDGF for 24 h; the protein level of MDM2, FoxO4 and CyclinD1 was measured using Western blotting. As shown in Fig. 3a, transfection of USP7 siRNA for 48 h dramatically reduced USP7 protein level to 33.8% of control (P < 0.05), whereas non-targeting siRNA did not affect USP7 protein level.
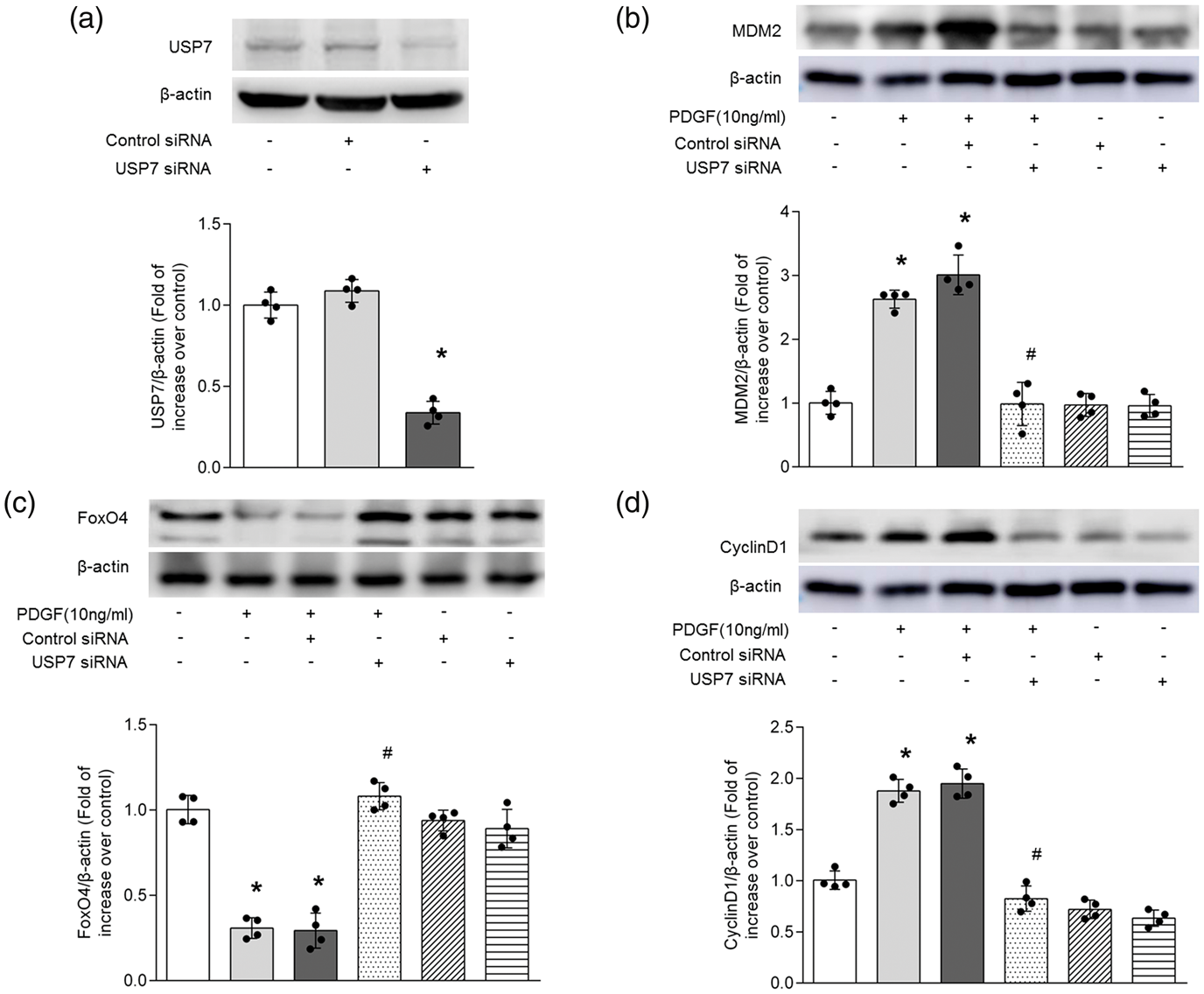
USP7 mediates PDGF-induced up-regulation of MDM2, down-regulation of FoxO4 and elevation of CyclinD1 in PASMCs. (a) PASMCs were transfected with USP7 siRNA and non-targeting siRNA for 48 h. The protein level of USP7 was determined by Western blotting (n = 4 per group). β-actin served as a loading control. A representative blot and quantification of bands are shown (n = 4 per group). *P<0.05 versus control siRNA transfection. (b–d) Cells were transfected with USP7 siRNA or non-targeting siRNA for 24 h before stimulation of PDGF (10 ng/ml) for 24 h, and the protein levels of MDM2 (b), FoxO4 (c) and CyclinD1 (d) were determined by Western blotting (n = 4 per group). β-actin served as loading control. A representative blot and quantification of bands are shown (n = 4 per group). *P<0.05 versus control. #P< 0.05 versus PDGF-treated cells.
Fig. 3b indicates that PDGF dramatically increased MDM2 protein level to 2.63-fold over control, while pre-silencing of USP7 reduced MDM2 protein level to 0.99-fold over control (P < 0.05 versus PDGF-treated cells). In addition, knockdown of USP7 did not change MDM2 protein level in cells without PDGF treatment.
We next examined the effect of loss of USP7 on PDGF-induced FoxO4 expression. Fig. 3c shows that treatment of cells with PDGF significantly decreased the protein level of FoxO4 (P < 0.05 versus control), whereas this effect was reversed by prior transfection of USP7 siRNA, FoxO4 protein level increased from 0.31-fold to 1.08-fold over control (P < 0.05 versus PDGF-treated cells). Moreover, silencing of cells with USP7 did not affect FoxO4 protein level in the absence of PDGF.
Finally, we determined CyclinD1 protein expression in cells lacking USP7. Fig. 3d shows that CyclinD1 protein level was increased to 1.88-fold over control in PASMCs stimulated with PDGF (P < 0.05), whereas loss of USP7 by siRNA transfection reduced PDGF-induced the increase of CyclinD1 protein, which declined to 0.82-fold over control (P < 0.05 versus PDGF-treated cells). In addition, silencing of cells with USP7 did not impact the protein level of CyclinD1 in the absence of PDGF. These results suggest that USP7 mediates PDGF regulation of the protein expression of MDM2, FoxO4 and CyclinD1 in PASMCs.
Up-regulation of MDM2 mediates reduction of FoxO4 via the ubiquitin proteasome pathway, and thereby up-regulates CyclinD1 in PASMCs treated with PDGF
To explore whether MDM2 mediates PDGF-induced down-regulation of FoxO4 via proteasome degradation in PASMCs, cells were pre-silenced with MDM2 sequence-specific siRNA for 24 h or treated with proteasome inhibitor MG-132 (10 μM) for 12 h before stimulation with 10 ng/ml PDGF for 24 h. As shown in Fig. 4a, MDM2 siRNA transfection for 48 h in PASMCs dramatically reduced MDM2 protein level to 26.4% of control (P < 0.05), whereas non-targeting siRNA did not affect MDM2 protein level. Fig. 4b indicates that PDGF notably reduced the protein level of FoxO4 (P < 0.05), while pre-knockdown of MDM2 reversed PDGF-induced FoxO4 reduction, which rose to 1.08-fold over control (P < 0.05 versus PDGF-treated cells). At the same time, prior incubation of cells with proteasome inhibitor MG-132 also restored FoxO4 protein level to 1.05-fold over control (Fig. 4d; P < 0.05 versus PDGF-treated cells), while knockdown of MDM2 or MG-132 did not change the protein level of FoxO4 in PASMCs without PDGF. These results suggest that MDM2 mediates FoxO4 down-regulation by ubiquitin-mediated degradation in PDGF-treated PASMCs.
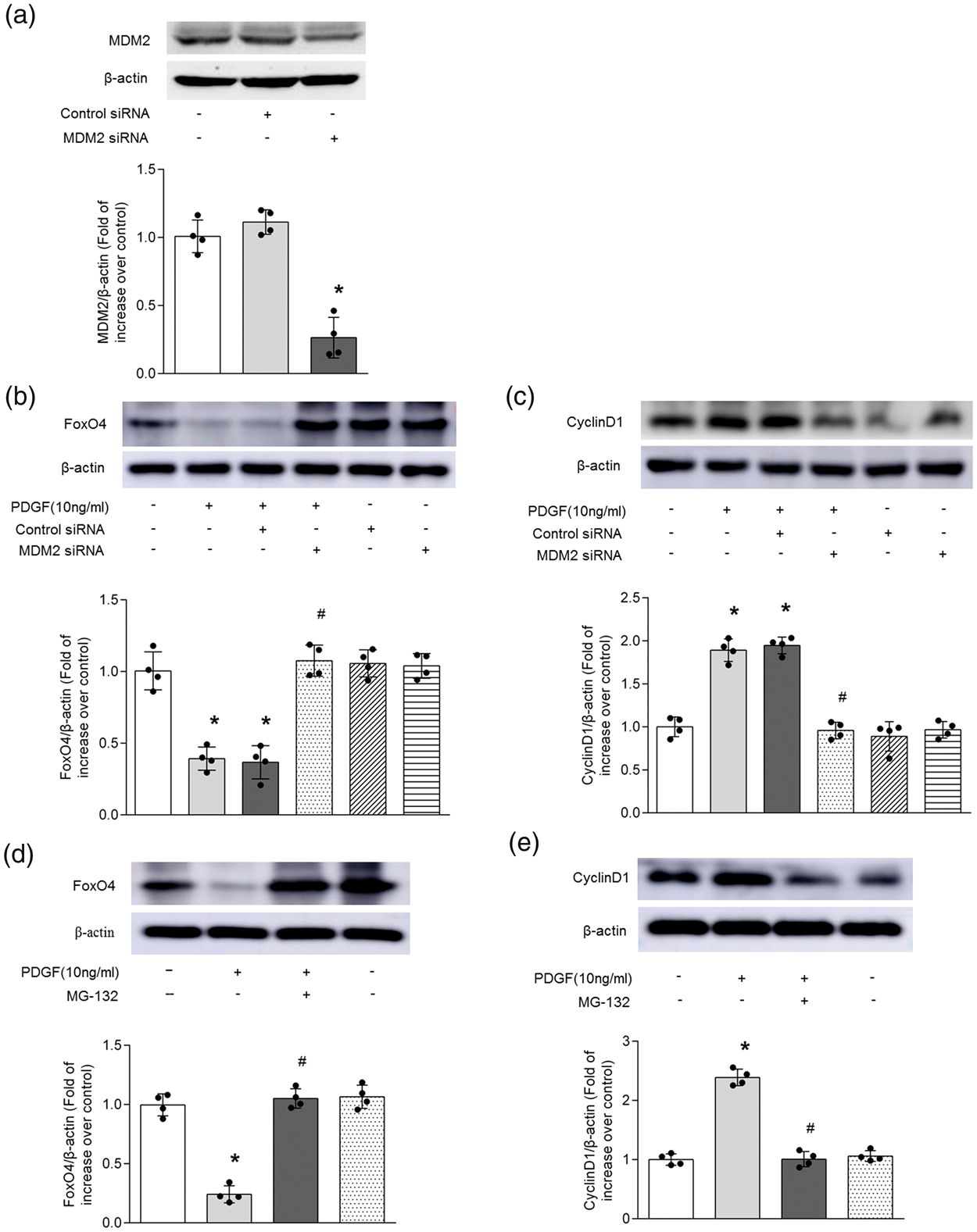
Up-regulation of MDM2 mediates reduction of FoxO4 via ubiquitin proteasome pathway and thereby up-regulates CyclinD1 in PASMCs treated with PDGF. (a) PASMCs were transfected with MDM2 siRNA and non-targeting siRNA for 48 h, and MDM2 protein level was measured by Western blotting (n = 4 per group). β-actin served as loading control. A representative blot and quantification of bands are shown (n = 4 per group). *P < 0.05 versus control siRNA transfection. (b–c) PASMCs were transfected with MDM2 siRNA or non-targeting siRNA for 24 h before PDGF stimulation (10 ng/ml) for 24 h, and the protein levels of FoxO4 (b) and CyclinD1 (c) were examined using Western blotting (n = 4 per group). (d–e) Cells were treated with MG-132 (10 μM) for 12 h before PDGF stimulation (10 ng/ml) for 24 h, and the protein levels of FoxO4 (d) and CyclinD1 (e) were examined using Western blotting (n = 4 per group). β-actin served as loading control. A representative blot and quantification of bands are shown (n = 4 per group). *P < 0.05 versus control. #P < 0.05 versus PDGF-treated cells.
To further examine the effect of MDM2 on PDGF-induced the increase of CyclinD1 expression, PASMCs were transfected with MDM2-specific siRNA for 24 h or pre-incubated with proteasome inhibitor MG-132 (10 μM) for 12 h, and then stimulated 10 ng/ml PDGF for 24 h, the protein level of CyclinD1 was determined by Western blotting. As shown in Fig. 4c and e, the loss of MDM2 or inhibition of proteasome function by MG-132 reduced PDGF-induced CyclinD1 protein elevation to 0.96-fold and 1.01-fold over control, respectively (both P < 0.05 versus PDGF-treated cells). Whereas transfection of MDM2 siRNA or utilization of MG-132 did not change the protein level of CyclinD1. These results indicate that MDM2 up-regulates the expression of CyclinD1 by promoting the ubiquitinated degradation of FoxO4 in PASMCs treated with PDGF.
USP7/MDM2/FoxO4/CyclinD1 cascade mediates PDGF-induced PASMCs proliferation
To explore the novel mechanisms underlying PDGF-induced PASMCs proliferation, BrdU incorporation assay was performed. Cells were pre-transfected with USP7 and MDM2-specific siRNA for 24 h separately, or pre-incubated with proteasome inhibitor MG-132 for 12 h before stimulation with PDGF for 24 h. As shown in Fig. 5, PDGF triggered a 1.98-fold increase in cell proliferation compared with control (P < 0.05), while knockdown of USP7 by siRNA transfection dramatically suppressed PDGF-induced PASMCs proliferation from 1.98-fold to 1.51-fold over control (P < 0.05 versus PDGF-treated cells), pre-silencing of MDM2 also reversed the effect of PDGF on cell proliferation from 1.98-fold to 1.51-fold over control (P < 0.05 versus PDGF-treated cells), whereas non-specific siRNA did not affect PDGF-induced cell proliferation. Furthermore, pre-treatment of cells with proteasome inhibitor MG-132 reduced PDGF-induced cell proliferation, which declined from 1.98-fold to 1.57-fold over control (P < 0.05 versus PDGF-treated cells). However, loss of USP7 or MDM2, or inhibition of proteasome function with MG-132 did not impact cell proliferation in the absence of PDGF. These results suggest that USP7 up-regulates MDM2, and subsequently reduces FoxO4 expression by MDM2-mediated ubiquitinated degradation, thereby up-regulates CyclinD1, finally promotes PDGF-induced PASMCs proliferation.
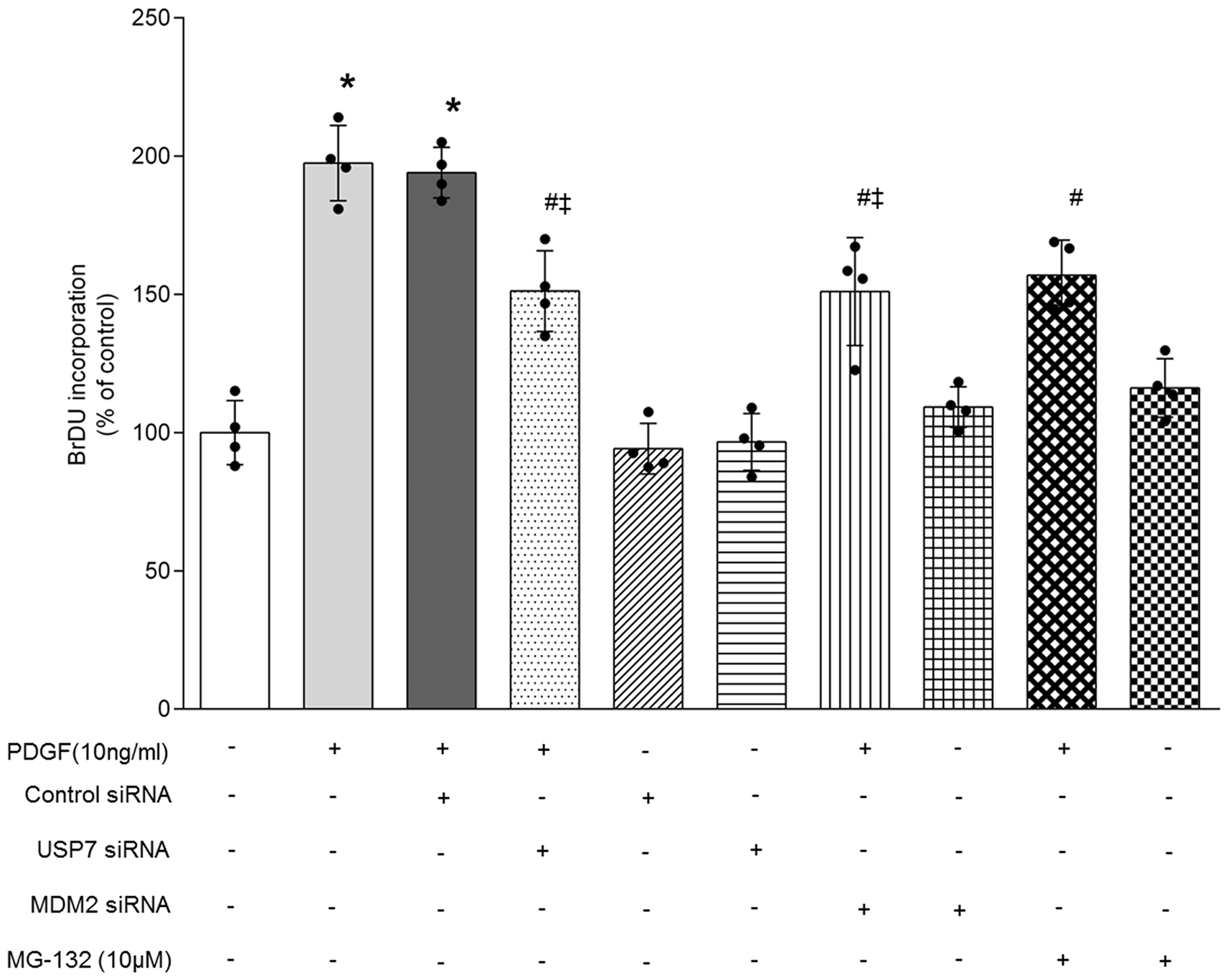
USP7/MDM2/FoxO4/CyclinD1 cascade mediates PDGF-induced PASMCs proliferation. PASMCs were transfected with USP7-specific siRNA or MDM2-specific siRNA for 24 h, or treated with proteasome inhibitor MG-132 (10 μM) for 12 h before stimulation of PDGF (10 ng/ml) for 24 h, and cell proliferation was measured by BrdU incorporation assay (n = 4 per group). *P < 0.05 versus control; #P < 0.05 versus PDGF-treated cells; ‡P < 0.05 versus control siRNA transfection in PDGF-treated cells.
Discussion
In this study, we have shown that USP7 mediates PDGF-stimulated PASMCs proliferation by up-regulation of MDM2 to increase MDM2-mediated FoxO4 ubiquitinated degradation, and consequently elevation of CyclinD1. These results provide novel molecular mechanisms for PDGF-stimulated PASMCs proliferation and suggest that targeting USP7/MDM2/FoxO4/CyclinD1 axis might have a potential value in the management of PAH.
UPS, mediating protein destabilization by proteasome, plays a crucial role in cellular physiology and pathophysiology, such as cell cycle progression, proliferation, apoptosis and signaling transduction. Ubiquitination is a highly dynamic process in which proteins are targeted for degradation by covalent ligation to ubiquitin. The conjugation of ubiquitin to targeted proteins is reversible due to the presence of deubiquitinating enzymes, which remove ubiquitin chains from targeted proteins. 41 USPs, the largest subfamily of deubiquitinating enzymes, with more than 60 members in humans, are emerging as potential targets for pharmacologic interference based on their protease activity and involvement in the pathogenesis of many cancers, such as non-small cell lung cancer and liver cancer. 11 , 42 USP7, also known as herpesvirus-associated ubiquitin-specific protease, has been found to be over-expressed in epithelial ovarian cancers, which promotes cell cycle progression and cell proliferation. 14 , 15 In the present study, we found that PDGF increased the expression of USP7; silencing of USP7 attenuated PDGF-induced PASMCs proliferation, suggesting that USP7 mediates PDGF-stimulated PASMCs proliferation.
USP7 has been reported to deubiquitinate several targeted proteins such as MDM2, UHRF1, DNMT1, claspin and PTEN, 43 and thereby suppress their ubiquitinated degradation or the regulation of their subcellular localization. 44 –46 A study has shown that USP7 deubiquitinates and stabilizes E3 ubiquitin ligase MDM2, and thereby promotes MDM2-mediated ubiquitinated degradation of p53 and cell growth in ovarian cancer cells. 16 USP7 silencing has been shown to increase steady-state p53 level by promoting MDM2 degradation. MDM2, an E3 ubiquitin ligase, has been found to exert its oncogenic activity to increase cell growth and invasion, decrease cell apoptosis and chemosensitization. 47 A study has shown that MDM2 is overexpressed in various human tumors. Furthermore, up-regulation of MDM2 has been recently observed to promote PASMCs proliferation and migration via p53 signaling pathway. 26 Our study indicated that MDM2 was up-regulated; this was accompanied with down-regulation of FoxO4, elevation of CyclinD1 and cell proliferation in PASMCs stimulated with PDGF, while knockdown of USP7 attenuated the changes of MDM2, FoxO4 and CyclinD1 as well as PASMCs proliferation caused by PDGF. These results suggest that up-regulation of USP7 stabilizes MDM2 to mediate PDGF-induced PASMCs proliferation.
Fox proteins, a family of transcriptional factors, contain a DNA-binding domain (DBD), which are characterized by forkhead box or winged helix domain. 48 FoxO proteins, evolutionally conserved transcription factors, consisting of FoxO1, FoxO3a, FoxO4 and FoxO6 in humans, function as tumor suppressors by regulating expressions of targeted genes which involved in cell cycle arrest, apoptosis, DNA repair and oxidative stress resistance. 32 , 33 Post-translational modifications regulate FoxO functions by ubiquitination and phosphorylation. 34 , 35 It has been shown that PI3K/Akt, NF-κB and ERK/MAPK signaling cascades regulate FoxO via interacting with and phosphorylating FoxO. 49 –51 Furthermore, FoxO proteins have been shown to be recognized by E3 ligases SKP2 and MDM2, leading to their ubiquitinated degradation in human primary tumors and cancer cell lines. 31 , 52 FoxO4 has attracted increasing interest due to its role in cancer carcinogenesis. 36 The expression of FoxO4 has been described to be decreased in gastric cancer tissues. 37 Study has provided evidence that the E3 ubiquitin ligase MDM2 is involved in FoxO4 ubiquitinated degradation, thereby reducing the expression of FoxO4, 31 which down-regulates the expression of CyclinD1 to inhibit cancer cell growth. 39 The present study demonstrated that PDGF caused a significant decrease of FoxO4 protein level, an elevation of CyclinD1 and proliferation of PASMCs, while loss of MDM2 reversed PDGF-induced reduction of FoxO4 and subsequent elevation of CyclinD1, thereby suppressing cell proliferation. In addition, pre-treatment of cells with proteasome inhibitor MG-132 also changed the expression of FoxO4 and CyclinD1 as well as cell proliferation in the presence of PDGF. Our results confirm that up-regulation of MDM2 by USP7 decreases ubiquitinated degradation of FoxO4, thereby up-regulates CyclinD1 leading to PDGF-stimulated PASMCs proliferation.
Conclusion
In this study, we have shown that PDGF-stimulated PASMCs proliferation by USP7-mediated MDM2 up-regulation and further MDM2-mediated FoxO4 ubiquitinated degradation, and finally CyclinD1 elevation. These results provide novel molecular mechanisms for PDGF-stimulated PASMCs proliferation and suggest that interfering above targets might prevent and treat the development of PAH.
Footnotes
Conflict of interest
The author(s) declare that there is no conflict of interest.
Authors’ contribution
Yanting Zhu and Manxiang Li designed this study. Yanting Zhu, Qianqian Zhang, Xin Yan and Lu Liu performed the research. Yanting Zhu, Cui Zhai, Qingting Wang and Limin Chai performed statistical analysis using appropriate software. Yanting Zhu and Manxiang Li wrote this paper. All authors have read and approved the article.
Ethical approval
All animal care and procedures were performed in accordance with Xi'an Jiao Tong University Animal Care Policy following the Guide for the Care and Use of Laboratory Animals. Ethics approval for the experimental protocols was received from the Laboratory Animal Care and Use Committee of Xi’an Jiao Tong University.
Acknowledgements
None.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (Grant No. 81330002 and No. 81670051).
