Abstract
Background:
Reporting serious adverse events (SAEs) is crucial to reduce or avoid toxicities that can lead to major consequences for patient’s health due to treatments tested in clinical trials. Its exhaustiveness is often inadequate, and we observe discrepancies between data published by pharmacovigilance organizations and clinical databases.
Objectives:
While the process of reconciliation aims at reducing these differences, it remains a very time-consuming and imprecise task. We propose a tool to automate this process.
Design:
We have developed and tested Reconciliaid, an application that compares the SAEs of the databases of clinical trials collected according to a standard inspired by the Clinical Data Interchange Standards Consortium, and of pharmacovigilance collected according to the international standards ICH-E2B (R3). It generates a reconciliation file that indicates precisely what information does not coincide in the two databases to facilitate the identification of inconsistencies.
Methods:
Reconciliaid was tested to create 13 reconciliation files, containing 290 SAEs. We inspected these files to determine their ability in identifying the inconsistencies and compared the manual and semi-automated reconciliation time. Four users answered the System Usability Scale (SUS) to measure its usability.
Results:
The application identified all variables of interest in all reconciliations. Different formats and libraries were automatically harmonized, allowing a perfect identification of inconsistencies for all variables. The matching of the same SAE in the two databases was correct in 97.2% of the reconciliations. Reconciliaid is six times faster than the manual approach for senior data managers (range = 3–24 times). A novice data manager performed three reconciliations 4.8 faster with the help of Reconciliaid than manually (29 min vs 134 min) and with fewer mistakes. Mean SUS score was 92.5.
Conclusion:
Reconciliaid has a high level of usability, can increase the quality of reconciliation, and reduces considerably the reconciliation time, allowing to increase the frequency of reconciliation processes and to focus resources on patient safety and medical assessment.
Plain language summary
Why was the study done? A serious adverse event (SAE) is a harmful reaction to treatment that can lead to hospitalization, disability, or death. Accurate reporting is essential for pharmacovigilance—the science of monitoring medication safety—to minimize risks in clinical trials. However, SAEs may be recorded inaccurately and a periodic manual reconciliation of clinical trial and pharmacovigilance databases is necessary to capture all adverse events correctly. Automating this process can save time and improve accuracy.
What did the researchers do? The researchers developed RECONCILIAID, a software that detects differences between data in the clinical and pharmacovigilance databases. RECONCILIAID identifies the common elements in the files exported from the databases, standardizes their formats, and automatically compares them. The comparison results are presented in a reconciliation file to easily identify differences. The software can accurately detect errors in just 0.1 to 4 seconds for studies with 1 to 189 SAEs.
What do the findings mean? The application offers several benefits: it automates, enhances quality, and reduces the cost of reconciliation, particularly for studies with a large number of SAEs. In a context of limited resources, the time saved by using RECONCILIAID allows healthcare staff to focus more effectively on patient safety and medical evaluation.
Introduction
Serious adverse events (SAE) during clinical trials can lead to serious consequences for the health of patients and their reporting is crucial to reduce or avoid the toxicity of the treatments tested. Pharmacovigilance aims at detecting, assessing, and preventing these adverse effects or any other possible drug-related problems to minimize their impact on patients. 1 Reporting of SAE contributes enormously to effective pharmacovigilance. In the United States, sponsors are currently required to report SAEs to the US Food and Drug Administration, 2 and in the European Union, reporting to the National Competent Authorities is mandatory, and online platforms for spontaneous reporting have been created.3,4 As a result, the reporting of SAEs increased over time in the last decades 5 and guidelines have been provided to improve consistency in reporting adverse events (AEs) and SAEs during clinical trials. 6
Effective pharmacovigilance requires ease of access to all sources of pertinent safety data. For drugs under development or recently launched, the clinical trials database represents a vital source of safety information from where pharmacovigilance organizations extract information to fill a database designed to comply with a different data standard. The independent and redundant methods for reporting SAEs require duplicate work on data processing, medical coding, and dictionary management, which could lead to the presence of inconsistencies that lower the exhaustiveness of regulatory reporting. Burnstead and Furlan discuss in detail the development of common data collection procedures that from Electronic Data Capture systems merge safety and clinical data into a single repository. 7 A single system for capturing all data would offer a convenient single source for the pharmacovigilance but, up to now, clinical operations and drug safety departments independently collect different sets of safety data from clinical studies and their reconciliation remains necessary.
Unfortunately, there is no uniform approach to reconciliation and, despite its importance, it remains an inefficient process that relies on emails and spreadsheets. It is an expensive and very time-consuming task that requires the manual comparison of the records. As a result, considerable discrepancy is often observed between the data published by pharmacovigilance organizations and the clinical bases. Better strategies for the reconciliation of these records are necessary to ensure data consistency. Thanks to data standards designed for organizing and exchanging clinical data, automatic methods could exploit matching rules to reconcile the common data in the two databases more efficiently and effectively but, to our knowledge, they are not exploited in routine medical settings.
In our center, the reconciliation process consisted of manually rearranging the Excel file containing the safety database extraction to match the same formatting of our standardized eCRF extraction tables (same number of columns and a single line per SAE) and display the records in a side-by-side view to identify the same event in the clinical database. Once detected, the event is inserted in the line below the matching safety information to more easily compare the two records and detect after a visual inspection the incoherencies. The whole process took up to 6 h for a single study with 187 reported SAEs and had to be performed every 3 months for the whole duration of the study (142 SAEs were already reported at the end of 2017). This repetitive, laborious, and uninteresting task is prone to fatigue-related errors and reduces the time for effective data management of current clinical trials.
We have developed and tested Reconciliaid, an automatic system to reconcile SAEs to reduce: (1) the risk of human error in the manual method due to the difference in formats in the two databases and (2) the considerable time needed to compare all data elements. It is based on a deterministic record linkage strategy that uses the SAE identifier to couple the data elements in the two bases, harmonize their format and automatically detect disagreements thanks to a reconciliation matching rules dictionary. Reconciliaid creates the reconciliation file to clearly show the discrepancies between the clinical and safety database that will be discussed in a meeting for deciding the corrective actions.
Study context
The goal of Reconciliaid is to compare the SAEs reported in the electronic Case Report Forms (eCRFs) and in the pharmacovigilance databases. It was designed to reduce the manual work and to provide a single reconciliation file indicating precisely the differences between the two databases. The application was developed and tested at the Department of Epidemiology, Biostatistics, and Health Data (DEBDS) of the Centre Antoine Lacassagne (CAL), a cancer treatment center in Nice, France. This tertiary care facility treats over 6200 patients annually and the DEBDS ensures the data quality for more than 40 clinical trials conducted at this center.
Reconciliaid is a home-grown system developed specifically for the needs of clinical trials and pharmacovigilance teams. It manages records of SAEs reported during clinical trials as well as the information related to the clinical trial and to patients. It was developed to reproduce the manual reconciliations carried out on 10 current clinical studies at the CAL and to minimize random operator errors for increasing the quality of reconciliations. It also supports pharmacovigilance monitoring as can detect SAEs that have not been declared and can be used as an audit trail to track data changes across the study.
Based on previous manual reconciliations, the list of common data elements in the clinical and pharmacovigilance databases are known for each clinical study and routinely exported in two separate files by data managers and pharmacovigilance officers. After review, we fixed the structure of the safety database export so that a standardized output, as shown in Figure 1, was available for each reconciliation regardless of the particular study.
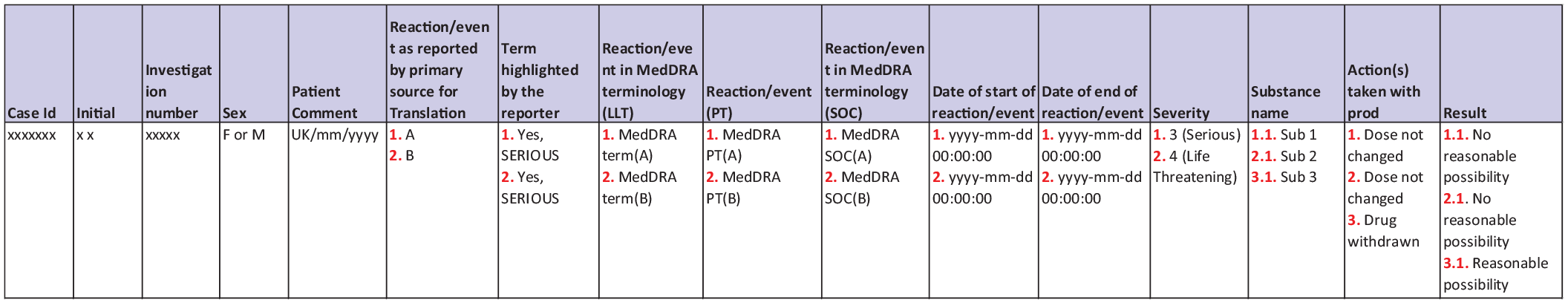
Standardized export from the pharmacovigilance database.
As in the manual process, the structure of the tables needs to be harmonized. Indeed, all SAEs reported in the same standardized form developed by the Council for International Organizations of Medical Sciences (CIOMS), the CIOMS form 8 , appear in the same cell as a numbered list in the safety file (as reported in red in Figure 1) while the clinical table contains a row for each SAE. Furthermore, to replicate the manual process, we need a dictionary of the standardized terms for comparing the sponsored-defined controlled terminology in the clinical database, as Clinical Data Interchange Standards Consortium (CDISC) Controlled Terminology has not been defined yet, and the International Council for Harmonisation of Technical Requirements for Pharmaceuticals for Human Use (ICH) ICH-E2B terminology. This dictionary should therefore include the relationship between the SAE and the study treatment and the action taken with the treatment in response to the SAE. As our center takes part in international as well as single-country clinical trials, the dictionary also needs to take into account the translation of terms in English to French, such as for the SEX (M/F to Homme/Femme) and AESER (Yes, serious to Oui) items as well as missing codes (Not Applicable to Non-Applicable).
We also need to correctly pair SAEs in the two tables according to different items, as the same CIOMS identifies a list of SAEs in the safety database. For this reason, the two files are merged and reorganized according to a predefined order as in the manual process: CIOMS number, reaction/event in MedDRA terminology (System Organ Class [SOC] and Preferred Term [PT] 9 ), and starting date of the event.
Methods
Methods for data acquisition and measurement
Safety data were collected in an eCRF and managed using Ennov Clinical® Software (Ennov, Paris, France), a secure, web-based application designed to support data capture for research studies. The eCRFs were designed according to the Clinical Data Acquisition Standards Harmonisation (CDASH) standard developed by the CDISC 10 and data were exported according to the CDISC Study Data Tabulation Model (SDTM) format. Data stored in the safety databases were collected according to the international standard ICH-E2B (R3). 11 The Common Terminology Criteria for Adverse Events (CTCAE) versions 5.0 and 4.03 were used as scales for the classification and severity scale of the AEs reported.
Common data in the two databases are exported in two separate files for comparison. From the clinical database, the exported file contains the SDTM AE domain for all AEs, including the not serious ones, and additional data from the Subject Characteristics (SDTM SC) and Demographics (SDTM DM) domains that are used for the full understanding and assessment of the events in drug safety departments of sponsors. Specific data elements of the ICH-E2B (R3) fields are added to the table. The full list is reported in Table 1.
Data elements exported from the clinical database for the reconciliation process.

Asterisk refers to fields specific to the ICH-E2B (R3).
CIDSC, Clinical Data Interchange Standards Consortium; SDTM, Study Data Tabulation Model.
To have a single line for each SAE in the reconciliation file, the elements Adverse Event Relationship to Study Treatment (AEREL) and Adverse Event Action Taken (AEACN) are exported in a separate column for each treatment used in the clinical trial. Safety data available in both the safety and clinical databases are extracted as reported in Table 2.
Data elements exported from the pharmacovigilance database for the reconciliation process.

Asterisk refers to fields coded in MedDRA terminology: Lowest Level Term (LLT), Preferred Term (PT), and System Organ Class (SOC).
All substances in the safety database are reported as a numbered list in the same cell. Related information in the other data elements is numbered accordingly. A different order for other events can be found in the same database extraction so that the number cannot be associated with a particular substance.
The application, developed in Python, uses the tkinter package to interface to the Tcl/Tk Graphical User Interface (GUI) toolkit. It displays an easy-to-use GUI that allows importing the two Excel files extracted from the two databases. Once the pharmacovigilance file is selected, it is imported as a Pandas data frame from which all substances are identified (case-insensitive) and shown on the interface. Once the clinical database file is selected, it is shown on the right side of the GUI to allow the user to find the column numbers reporting the measures implemented after the appearance of the SAEs (Action) and their relationships (Relation) for each substance if these are available in both databases (Figure 2).

Interface of Reconciliaid. The user can upload the two files exported from the clinical and pharmacovigilance databases, choose the name of the reconciliation file, and select if compare the action and the result of the assessment (relation) related to each substance in the Pharmacovigilance database.
Once all fields are completed, by clicking on the Reconciliate button, the Reconciliation file is created on the user’s desktop.
The reconciliation file contains the matched data from the two bases separated by two lines: Validation and Comment. The first line shows whether the two lines contain consistent data (green cell OUI) or not (red cell NON). Consistency is based on an exact match or on a plausible match of the two lines in a column. An exact match is required for all variables except Sex, Term highlighted by the reporter, Actions, and Results. Dates reported in this file have been formatted to match a common standard: Unknown/Month/Year with leading zeros (UK/MM/YY) for the Date of Birth, and pandas.Timestamp (YY-MM-DD HH:mm:ss) for Date of start and Date of end for reaction/event. As missing days or months were sometimes reported in these two dates, we used a function that converts dates from the format DD/MM/YYYY to YYYY-MM-DD 00:00:00, while handling missing days or months represented by NK (Not Known). The function first splits each date string using the “/” delimiter to separate the day, month, and year. If the day or month is marked as NK, it is replaced with “00” to indicate the missing value. The function then reassembles the date components into the desired format YYYY-MM-DD 00:00:00.
The comparison is insensitive to case, blank spaces, and punctuation for Initials and uses the dictionary previously described for Sex, Actions, and Results. The user fills the second line, Comment, after the verification of the line Validation. Indeed, the Term highlighted by the reporter is not considered in the reconciliation as it usually contains terms in English and French that cannot be automatically compared and need manual verification. If the line Validation is completely green, the user can add the comment Reconciliate (RECONCILIE), otherwise does not Reconciliate (NON RECONCILIE). The comment is not compulsory but can be used to prove that the file has been validated by the data manager.
An example of output is reported in Figure 3.

Reconciliation file created by Reconciliaid and inspected by the data manager. The row Validation is filled automatically and differences in entries in the pharmacovigilance database (row “base PV”) and in the clinical database (row “eCRF”) are depicted in red. The row “Commentaire” is filled by the user after verifying the results.
Methods for data analysis
Two data managers first tested Reconciliaid between June and December 2021 at the CAL. At that time, both had more than 7 years of experience in manual reconciliations. They did not receive specific training but were provided with a written procedure for the correct use of the application and its features. As Reconciliaid was developed to reproduce the manual reconciliations that they performed on the clinical studies under their responsibility, they could easily evaluate its performance.
We expected a very significant difference between reconciliation time using our tool and manual reconciliation. A calculation based on a comparison of means of the duration of the reconciliation of an SAE, with alpha and beta risks of 1%, an expected difference where manual reconciliation takes five times longer than Reconciliaid (2 min per AE for manual reconciliation vs 0.4 min for Reconciliaid), and a standard deviation of 1, indicates that 22 AEs are sufficient to determine whether Reconciliaid is significantly faster than the manual process. However we included more SAEs to provide more experience to the data managers to better comment on their experience, so they finally performed 13 reconciliations containing a total of 290 SAEs with the help of Reconciliaid.
A third data manager, who was naïve of the manual reconciliation, performed three reconciliations in 2024, both manually and with Reconciliaid, to provide preliminary results of the effect of experience on the time saved by using the proposed tool. We also compared the reconciliation files generated both manually and with the aid of the app to the reconciliation file generated by a senior data manager to evaluate the errors.
All users answered the French version of the System Usability Scale (F-SUS). 12 It is a simple, 10-item questionnaire used to evaluate the usability of a system. It provides a quick, reliable way to measure user satisfaction, efficiency, and effectiveness. Scores range from 0 to 100, with higher scores indicating better usability.
The reporting of this study conforms to the Statement on Reporting of Evaluation Studies in health informatics (STARE-HI) statement. 13
Results
Based on the inspection of the output files, the application precisely identified all variables of interest in all reconciliations and correctly placed them as separate columns. According to the variable, the different date formats and libraries were automatically harmonized, allowing a perfect identification of inconsistencies for all variables. The matching of the same SAE in the two databases (linkage in lines), necessary to perform the reconciliation, was correct in 97.2% of the reconciliations. Incorrect matching was mainly due to differences in CIOMS values in the two databases, which was the first variable used for the coupling. In other instances where the same CIOMS referred to several SAEs, we observed a mismatch in cases where the MedDRA code of the event or the starting date differed in the two databases. The reconciliation file needed therefore a manual correction to displace the line under the corresponding SAE and a manual reconciliation of the SAE. Reconciliaid produced the reconciliation file in a time comprised between 0.1 and 4.4 seconds (mean = 0.6 s) for studies reporting between 1 and 189 SAEs (mean = 22.3) as shown in Table 3.
List of clinical studies on which we tested Reconciliaid and comparison of the manual and automatic execution times.

Asterisks refer to studies performed by a less experienced user and are execution time is not considered for the evaluation of the average results.
SAE, serious adverse event.
For experienced data managers, this file needed a manual verification (a few seconds per SAE) and, when necessary, a correction which added approximately 1–2 min per incorrectly coupled SAE, while the manual reconciliation required between 3 and 360 min (average = 46.1 min). This difference was due mainly to the time needed to format the reconciliation file (addition of lines after each SAE), format the clinical database export (deleting unnecessary columns and reorder them to match the clinical database order), and search the same SAE in the clinical database export. An average of 24 s was needed to Reconciliate each SAE by Reconciliaid and the manual verification (range = 5 s to 1 min) while the manual process takes an average of 2.5 min (range = 1.7–3 min), meaning that Reconciliaid is six times faster than the manual approach (range = 3–24 times). When a novice user performed the reconciliations of 41 SAEs both manually and with the help of Reconciliaid, we observed again an important time reduction. The manual reconciliations took a total of 134 min, while using Reconciliaid reduced the total time to 29 min—a 4.6-fold decrease in time required. We observed two missing SAEs in the manual reconciliation file and errors in the manual comparison of five elements compared to the reconciliation file created by a senior data manager. When comparing the file generated with the help of Reconciliaid, all SAEs were present and we observed two errors in the reconciliation file.
The overall SUS score was 92.5, indicating a high level of usability. According to standard benchmarks, this score suggests that Reconciliaid is user-friendly and effectively meets the needs of its users, enhancing their experience during the reconciliation process.
Indeed, all users commented positively on the simplicity of the interface design and suggested minor improvements regarding the choice of colors and the organization of the fields, in particular the position of the entry field for the file name on the left side. They commented on the absence of feedback from the interface and, as suggested, a new version of the GUI will include an automatic message to help the user understand the issues in the input files that could present missing columns or misspelled words in the labels, preventing the app to create the Reconciliation file. Currently, a manual verification of the two exports from the two databases is required before using the app, which is described in its documentation. However, this guide is currently available in the app folder and, as suggested by the data managers, the next version of the app will include a menu with the help tab for more convenient access to the documentation.
Discussion
Pharmacovigilance is fundamental to identify as quickly as possible new adverse reactions, gather more knowledge on suspected adverse reactions and the corresponding risk factors, with the aim to improve clinical practice. Quality of data collection, management, and coding needs to be controlled, and the reconciliation process between clinical trial databases and pharmacovigilance databases is therefore undertaken.
Reconciliaid is an automated tool designed to reconcile discrepancies between these databases. The results of this study demonstrate that Reconciliaid successfully identifies inconsistencies between the two databases. It harmonized different formats and libraries automatically, allowing it to identify all variables of interest and detect inconsistencies in 100% of the reconciliations. Moreover, the matching of SAEs between the databases was highly accurate, with a success rate of 97.2%. In terms of efficiency, Reconciliaid significantly reduced the time required for reconciliation for both senior and novice data managers and reduced the errors of the novice user. The reconciliation file indicates precisely what information does not coincide in the two databases, using a common format independent of the operator or the execution period, thus facilitating the identification of inconsistencies. Reconciliaid saves considerable time, potentially increasing the frequency of reconciliation processes and focusing resources on patient safety and medical assessment. The application does not need informatics knowledge to make it operational and can lead to an economic benefit on care management, especially when the clinical studies present a high number of SAEs. According to these results, we believe that Reconciliaid proves to be an effective tool for the reconciliation of SAEs, improving both the accuracy and speed of the process compared to manual methods.
To the best of our knowledge, there are no published studies addressing the automation of SAE reconciliation between clinical trial databases and pharmacovigilance databases. Prior research has highlighted the challenges and inefficiencies in manual reconciliation processes, particularly the time-consuming nature and risk of human error, but none has offered an automated solution. Our results are therefore novel in demonstrating the feasibility and effectiveness of such an approach. While we cannot directly compare our findings to similar tools, our results, in particular the reduction of the time required for the process and the mean SUS score of 92.5, support the adoption of automated tools for pharmacovigilance.
As a future development, we plan to convert it into an open-source application. It will be developed in Python and made available for download via GitHub. Given the sensitive nature of the data it handles, the application will not be web-based to ensure strict compliance with data protection and security standards. It will be designed specifically for desktop environments as handling sensitive patient data on mobile devices poses significant security risks. Users will be able to download and run the application locally on their computers.
Our study provides an initial approach to automating SAE reconciliation, with the potential to improve data accuracy and efficiency in clinical trials and pharmacovigilance efforts. While further validation is needed, we hope these findings will encourage additional research into automating reconciliation processes and enhancing data quality in this area.
Limitations
A limitation of this study is the lack of an exhaustive dataset for computing the frequency of manual reconciliation errors. It is indeed not feasible to estimate the error rate from manual reconciliations conducted before the development of Reconciliaid as the pharmacovigilance officer only focused on discrepancies, and any incorrect reconciliations would have gone unnoticed. Additionally, we are unable to apply Reconciliaid retrospectively to these older reconciliations, as the exported files were formatted differently from the structure required by Reconciliaid (different columns and formats). This limitation restricts our ability to comprehensively assess the effectiveness of Reconciliaid against the manual process but will be analyzed in future work.
From a technical perspective, a limitation of the system is its dependence on a fixed dictionary and future developments are required to adapt the application to the inevitable changes in libraries in the databases. In particular, in addition to the improvements suggested by users, the next version of Reconciliaid will use a separate file containing the list of synonyms that are currently hardcoded in the source code, which will allow changes specific to a particular study but also its use in different languages.
Conclusion
Reconciliaid demonstrates significant improvements in the reconciliation of SAEs between clinical trials and pharmacovigilance databases. By automating the comparison process, this tool has the potential to minimize random operator errors and enhance the overall quality of reconciliation. Its ability to harmonize different formats and libraries allows for precise identification of inconsistencies, achieving a remarkable accuracy rate of 97.2% in matching SAEs. Additionally, Reconciliaid drastically reduces the time required for reconciliation. With a high usability score of 92.5, Reconciliaid not only simplifies the reconciliation process but also enables more frequent assessments, ultimately allowing healthcare professionals to dedicate more resources to patient safety and medical evaluations.
