Abstract
Endothelial injury is a key factor initiating in-stent restenosis (ISR) following peripheral artery stent implantation. Genetically modified endothelial progenitor cells (EPCs) can promote reendothelialization of injured arteries and inhibit neointimal hyperplasia. However, the role of engineered EPCs overexpressing lncRNA H19 in these processes remains unclear. We constructed EPCs overexpressing lncRNA H19 and investigated their effects and mechanisms in promoting reendothelialization and inhibiting neointimal hyperplasia both in vitro and in vivo. Compared to the normal control group, ISR patients exhibited a significant reduction in circulating EPCs. Engineered EPCs overexpressing lncRNA H19 promoted reendothelialization and inhibited neointimal hyperplasia in injured arteries. Exogenous overexpression of lncRNA H19 significantly upregulated the endothelial repair-related gene S1PR3 in EPCs, while the opposite was also observed. Additionally, engineered EPCs overexpressing S1PR3 promoted reendothelialization and inhibited neointimal hyperplasia in injured arteries. S1PR3 overexpression enhanced EPCs proliferation, migration, and tube formation in vitro; these effects were lost with S1PR3 inhibition. Binding sites for H3K27 acetylation were identified on the S1PR3 promoter. Mechanistically, we found that lncRNA H19 directly interacted with HDAC2, a known H3K27ac deacetylase, disrupting its binding to H3K27 acetylation. Our findings suggest that lncRNA H19 positively regulates S1PR3 expression by disrupting HDAC2 / H3K27ac binding, thereby promoting reendothelialization of injured arteries and inhibiting neointimal hyperplasia.
Graphical Abstract
In the nucleus, H19 recruits HDAC2 and modulates H3K27ac levels at the S1PR3 promoter, thereby enhancing gene transcription. In the cytoplasm, S1PR3 activates PI3K/Akt signaling pathways, thereby promoting EPCs reendothelialization.
Highlights
H19-Overexpressing EPCs enhance reendothelialization and inhibit neointima in injured carotid arteries of nude mice.
S1PR3 activation of PI3K/AKT pathway modulates biological behaviors of EPCs.
lncRNA H19 inhibits HDAC2 to enhance H3K27ac and promote S1PR3 expression.
Background
Peripheral Artery Disease (PAD) refers to ischemic conditions stemming from structural and functional changes in the major arteries outside of the coronary and central vessels. 1 Over the past two decades, the prevalence of PAD has increased globally, posing a growing public health challenge. 2 The treatment of PAD has evolved from open surgeries to minimally invasive endovascular interventions, predominantly using stent angioplasty, due to its less invasive nature. 3 However, these procedures can damage endothelial cells (ECs), leading to vascular inflammation, and stimulate abnormal proliferation and migration of smooth muscle cells (SMCs) to the intimal layer, which results in neointimal hyperplasia (IH) and in-stent restenosis (ISR). 4 Despite the development of drug-eluting stents (DES), which deliver drugs like paclitaxel to locally inhibit the proliferation of vascular SMCs, the lack of cellular specificity also impedes the proliferation and reendothelialization of ECs, significantly affecting the endothelial repair process. 5 The clinical effectiveness of these interventions remains compromised with ISR rates exceeding 10% within one year. 6
Endothelial injury is the initial trigger for ISR. Promoting early reendothelialization post-stent implantation is key to preventing restenosis. 7 Endothelial progenitor cells (EPCs) have the potential to differentiate into ECs and play a crucial role in repairing the vascular endothelium. 8 Previous studies have explored approaches such as employing drugs that recruit EPCs, modifying stents with specific antibodies to capture EPCs, and directly transplanting EPCs to enhance rapid endothelial repair post-stent implantation and inhibit ISR.9 –11 However, these strategies have not achieved significant outcomes mainly due to insufficient EPC numbers and impaired function in PAD patients, poor biomolecular capturing abilities, and low survival rates of transplanted EPCs, hindering their clinical application. 12
Epigenetic modifications, regulated by mechanisms including non-coding RNAs and post-transcriptional histone modifications, have emerged as a new direction in disease research.13,14 Modifying EPCs to regulate their biological functions could enhance their reparative abilities, accelerate the reendothelialization process, and suppress ISR formation.15,16 LncRNA H19, a highly conserved long non-coding RNA, is significantly enriched in cardiovascular tissues and plays a critical role in maintaining cardiovascular stability. 17 In atherosclerosis, lncRNA H19 engages in multiple biological processes, including cell proliferation, migration, and angiogenesis, thereby affecting disease progression. 18 Additionally, lncRNA H19 is vital in stem cell differentiation, particularly in enhancing osteogenic differentiation. 19 Our previous studies have demonstrated that lncRNA H19 can promote the proliferation, migration, and vasculogenesis of EPCs. 20 The research progress on lncRNA H19 in cardiovascular diseases and stem cell differentiation provides important insights into its potential roles in stem cell transplantation therapies.
This study focuses on engineering EPCs to overexpress lncRNA H19. Their impact is assessed using a murine model of carotid artery guidewire injury, where their ability to promote endothelial repair and inhibit IH is evaluated. The research aims to elucidate the mechanisms by which lncRNA H19 enhances endothelial repair in these cells and explores their potential use in treating ISR.
Methods
Ethical approval of the study protocol
The research adhered to the Helsinki Declaration and received prior approval from the Institutional Ethics Committee (IEC) for Clinical Research and Animal Trials at the First Hospital of Sun Yat-sen University. Written informed consent was secured from all human subjects or their legal representatives before sample collection. Additionally, all procedures involving animals were conducted in accordance with the guidelines established by the National Institutes of Health (NIH) for the Care and Use of Laboratory Animals.
EPCs isolation and characterization
EPCs were derived from human peripheral blood as outlined in previous studies. 20 Blood from healthy volunteers, collected into sodium heparin tubes, was used to isolate peripheral blood mononuclear cells (PBMCs) via density gradient centrifugation with human Lymphocyte Separation Medium (TBD, LTS1077-1). These cells were then plated on six-well plates coated with Fibronectin (BD Biocoat, 356008) and cultured in EGM-2 BulletKit medium (Lonza, CC—3156 & CC—4176), which includes endothelial basal medium and SingleQuots Kit, supplemented with 20% FBS. The cultures were maintained at 37°C in an environment of 5% CO2 and 85% humidity. Media changes occurred first after 4 days and then every 2–3 days thereafter. EPCs from passages 3 to 6 were utilized for experiments. Phenotypic analysis of EPCs included flow cytometry to assess the expression of CD31, CD144, and KDR, using fluorescence-labeled monoclonal antibodies.
Guidewire-induced neointimal formation in arterial models
Eight-week-old male nude mice were obtained from the Animal Center of the First Affiliated Hospital of Sun Yat-sen University. These mice were anesthetized with 2% isoflurane and underwent a procedure where the superior thyroid and occipital arteries, branches of the left external carotid, were ligated with 8-0 prolene sutures. Blood flow in the internal and common carotid arteries was temporarily stopped with vascular clamps. A small incision was made in the external carotid artery to insert a 0.38 mm diameter guidewire, which was rotated three times to induce artery damage. After guidewire removal, the artery was ligated, blood flow restored, and the site sutured. Post-surgery, the mice recovered on a warming blanket before returning to their cages for further care.
Cell therapy
For cell therapy, 5 × 106 EPCs, either transduced with adenovirus or lentivirus, were resuspended in 200 μl PBS. Following labeling with DiR (Yeasen, 40757ES25, USA), these cells were administered intravenously through the tail vein using a 31-gauge needle into models. Controls included PBS and non-transduced EPCs. Cell therapy was performed 3 h after surgery.
Tracking endothelial progenitor cells in nude mice models
Three days after tail vein injection of DiR-labeled EPCs, the mice were anesthetized with 2% isoflurane. The small animal live imaging system was set up and adjusted to the DiR dye channel (Ex/Em 748 nm/780 nm). Anesthetized mice were placed in a prone position on the imaging platform inside a dark box. Images were automatically captured using the control software. After imaging, the mice were returned to their cages for recovery.
Carotid artery Evans blue staining in nude mouse models
Three days post-modeling, nude mice were anesthetized with a standard dose of 2% isoflurane. Once anesthetized, 200 μl of 5% Evans blue dye was injected via cardiac injection to stain the carotid arteries. After allowing 10 min for circulation, the mice were euthanized. The mice were dissected, and saline was injected at the apex of the heart to flush out the dye from the carotid arteries, which were then excised and further rinsed in saline. The carotid arteries were longitudinally opened, photographed, and the reendothelialization was quantitatively analyzed using ImageJ software.
HE staining of carotid artery in nude mouse models
On day 28 post-surgery, carotid artery tissues from nude mice were excised and fixed in 4% paraformaldehyde, followed by paraffin embedding. Tissues were sectioned into 5-micron slices and deparaffinized in xylene for 20 min. The sections were then rehydrated in a graded alcohol series (100%, 90%, 80%, and 75% ethanol, each for 5 min) and rinsed under running water. After soaking in hematoxylin for 3–5 min, the slices were washed, differentiated, and blued under running water. They were then stained with eosin for 15 s, dehydrated in 95% alcohol for 1 min, cleared in xylene for 2 min, and mounted with neutral resin. Tissue sections were examined under a microscope, images were captured, and analyses were conducted.
Immunofluorescence staining of carotid artery in nude mouse model
Sections were deparaffinized and rehydrated through xylene and graded alcohols, followed by TBS washes. Antigen retrieval was performed using EDTA buffer (pH 9.0) in a pressure cooker, followed by a rinse in distilled water. Sections were blocked with 10% donkey serum and incubated overnight at 4°C with primary antibodies against αSMA and CD31. The next day, sections were washed and incubated with fluorescent secondary antibodies at 37°C for 45 min, followed by DAPI staining for nuclear visualization. After final washes, slides were mounted with a fluorescent mounting medium and examined under a microscope.
CCK8 assay
Cell viability was assessed using the Cell Counting Kit-8 (CCK8, HangZhou FUDE Biological Technology Co. Ltd, FD3788). A 100 μl cell suspension containing 5000 cells per well was plated in a 96-well plate and incubated at different time intervals (0, 24, 48, and 72 h). At each time point, 10 µl of CCK8 solution was added to the wells and incubated for 2 additional hours. The absorbance was measured at 450 nm using an enzyme-linked immunosorbent assay reader.
EdU proliferation assay
EPCs proliferation was assessed using the EdU Proliferation Kit (Abbkine Scientific Co., Ltd, KTA2031). Approximately 1 × 105 treated EPCs per well were seeded in a culture plate and incubated for 18 h. Subsequently, the cells were exposed to 10 µM EdU for 2 h at 37°C. Following the incubation, cells underwent washing, fixing, permeabilization, and staining processes as per the kit instructions. Cell nuclei were counterstained with DAPI. Fluorescence microscopy (DMI8, Leica) was used to visualize and capture the images of the cells.
Transwell migration assay
The migration ability of EPCs was evaluated using a Transwell setup. Treated EPCs (3 × 105 cells in 200 μl of serum-free medium) were placed in the upper chamber of a Transwell insert (8 μm pore size, 24-well plate). The lower chamber was filled with 600 μl of medium enriched with 10% FBS. After a 12 h incubation, cells on the upper side of the insert were washed with PBS, fixed in 4% paraformaldehyde, and stained with crystal violet (HangZhou FUDE biological technology Co. Ltd, FD7608). Migrated cells were quantified by counting under a microscope.
Wound healing migration assay
EPCs migration was assessed using a wound healing assay. Confluent EPCs in a six-well plate were scratched with a 200-μl pipette tip to create a straight line wound. The cells were then rinsed twice with PBS and cultured in serum-free medium for 12 h. Microscopic images captured at 0 and 12 h post-wounding were analyzed with ImageJ to measure the wound closure area.
In Vitro tube formation assay
The vasculogenic potential of EPCs was assessed using a tube formation assay. Treated cells (5000 cells in 100 μl) were plated on Matrigel-coated 96-well plates (Corning, 356237) and incubated at 37°C. After 6 h, the formed tubular structures were imaged using a microscope. The lengths and areas of these structures were quantified using ImageJ software.
Western blot analysis
EPCs were lysed using RIPA buffer (Beyotime Biotechnology, P0013B) and the proteins were separated by 10% SDS-PAGE. The proteins were then transferred to PVDF membranes (Millipore Sigma, ISEQ00010) and blocked with 5% skimmed milk in TBST for 1 h. Membranes were incubated overnight at 4°C with primary antibodies against PI3K, PPI3K, AKT, PAKT, S1PR3, and GAPDH, diluted in HangZhou FUDE’s antibody diluent (FD0040). Following three washes in TBST, the membranes were incubated with horseradish peroxidase-conjugated secondary antibodies (Proteintech, SA00001—1 and SA00001—2) for 1 h at room temperature. Protein bands were visualized using an ECL detection reagent (Millipore Sigma, WBKLS0500) and analyzed with Image-Pro Plus Version 6.0 software relative to GAPDH.
Transcriptomic mRNA sequencing and analysis
EPCs were infected with an adenovirus overexpressing lncRNA H19 and harvested after 72 h. Total RNA was extracted using Trizol reagent according to the manufacturer’s instructions. High-quality RNA samples were then used to construct the mRNA sequencing library with a kit, and sequencing was performed on the Illumina NovaSeq 6000 system by MegaGen Biotech (Shanghai, China). Differential mRNA expression was analyzed using the “DESeq2” R package, identifying significant differences with |LogFC| > 1 and p < 0.05. Differential genes were annotated using Diamond software for GO terms and ID mapping tools for KEGG pathways. Enrichment analyses for GO functions and KEGG pathways were conducted using GOATOOLS and KOBAS software, respectively, with significance set at p_fdr ⩽ 0.05 after multiple testing corrections.
Identification and validation of target genes for endothelial repair
Initially, the top 1000 genes associated with endothelial repair were extracted from the GeneCards database (https://www.genecards.org/). Additionally, 406 genes were identified as endothelial repair-related through the STRING database (https://cn.string-db.org/). Subsequently, these genes were cross-referenced with the top 3000 upregulated and downregulated differentially expressed genes from transcriptomic mRNA sequencing to create a Venn diagram, isolating 38 candidate genes related to endothelial repair. A protein-protein interaction (PPI) network of these 38 genes was then constructed using STRING. The candidate genes were scored, and the top 10 genes according to network centrality metrics—Betweenness Centrality, Degree, and Closeness Centrality—were further analyzed through Venn diagram, yielding four key genes for endothelial repair validation.
RNA pulldown assay
Following the acquisition of the H19 overexpression plasmid (Figure S2(e)) and its subsequent linearization to create a DNA template (Attachments 1 and 2), a large quantity of Biotin-labeled RNA molecules was synthesized using the RiboTM RNAmax-T7 Biotin-labeled Transcription Kit (R11074.3, RiboBio, China), according to the manufacturer’s instructions. For the subsequent RNA pulldown experiment, the protocol provided by the Pierce™ Magnetic RNA-Protein Pull-Down Kit (catalog no. 20164, Thermo Scientific, USA) was followed. Briefly, the Biotin-labeled RNA was initially bound to streptavidin magnetic beads. The RNA-bound beads were equilibrated in protein-RNA binding buffer before adding cell lysates, which were prepared using RIP lysis buffer. After incubation, the beads were washed thoroughly by adding the appropriate buffers, followed by vortexing and magnetic separation. Finally, the isolated protein samples were analyzed by western blot to verify the results.
RIP assay
RNA-protein immunoprecipitations (RIP) were performed using the Magna RIP kit (17-700, Millipore, USA). Briefly, cells were cultured until approximately 90% confluence. The cells were then scraped and collected using cold PBS, followed by resuspension in RIP lysis buffer containing protease and RNase inhibitors. Two 10 μl aliquots from each sample were reserved as input controls. Magnetic protein A/G beads were conjugated with 5 μg of rabbit anti-HDAC2 antibody or normal rabbit IgG as a negative control for each sample. The RIP immunoprecipitation buffer was prepared according to the manufacturer’s instructions. The bead-antibody complexes were then mixed with the RIP lysate supernatant, ensuring thorough mixing, and incubated overnight at 4°C on a rotator. The following day, RNA was purified from the immunoprecipitates for subsequent analysis of the precipitated RNA.
CHIP assay
Chromatin immunoprecipitation (ChIP) assays were performed using the ChIP-IT High Sensitivity Kit (53040, Active Motif, China). According to the experimental protocol, treated cells were cross-linked using a specially formulated formaldehyde buffer, which preserves protein/DNA interactions. The cells were then sonicated to shear DNA into small fragments and incubated with antibodies (HDAC2 & H3K27ac) targeting DNA-binding proteins. The antibody-bound protein/DNA complexes were immunoprecipitated using protein G agarose beads, followed by washing through gravity filtration. After immunoprecipitation, the cross-links were reversed, and proteins were removed with proteinase K. The DNA was then recovered and purified. ChIP-enriched DNA was subsequently used for gene-specific analysis.
Statistical analysis
Experiments were conducted with a minimum of three independent replicates. Unless otherwise noted, results are presented as mean ± SD. Statistical comparisons were made using an unpaired Student’s t-test or one-way ANOVA followed by Tukey’s post hoc test, performed with GraphPad Prism 9.0 software. A p-value of less than 0.05 was considered statistically significant.
Results
Transplantation of H19-overexpressing EPCs enhances reendothelialization and inhibits neointima formation in wire-injured carotid arteries of nude mice
Compared to healthy individuals, the number of circulating EPCs in the peripheral blood of ISR patients was significantly reduced (Figure S1(a) and (b)). Based on previous research, we isolated mononuclear cells from the peripheral blood of healthy donors and induced the formation of primary EPC lines (Figure S1(c) and (d)). Subsequently, we examined the expression profile of lncRNA H19 in these EPCs and found that it was predominantly localized in the nucleus (Figure S1(e), (f), (g)). To explore the potential impact of lncRNA H19 on endothelial repair in wire-injured carotid arteries, we designed an animal study (Figure 1(a)). Three days post-transplantation of EPCs overexpressing lncRNA H19 via tail vein into the injured carotid artery model in nude mice, in vivo imaging demonstrated a significant increase in the homing of cells at the injury site (Figure 1(b) and (c), Figure S3(a) and (b)). Evans Blue staining indicated that lncRNA H19-overexpressing EPCs significantly enhanced reendothelialization of the injured carotid artery (Figure 1(d) and (e)). After 28 days, further evaluation through HE and immunofluorescence staining assessed the long-term effects of these treated EPCs on neointima formation post-injury. Immunofluorescence staining confirmed that lncRNA H19-overexpressing EPCs more effectively promoted reendothelialization of the injured carotid artery (Figure 1(f) and (g)). Furthermore, the results showed that lncRNA H19-overexpressing EPCs significantly suppressed neointima formation compared to control EPCs (Figure 1(h) and (i)).

Transplantation of H19-overexpressing EPCs enhances reendothelialization and inhibits neointima formation in wire-injured carotid arteries of nude mice: (a) Schematic illustration of EPCs implantation into wire-injured carotid arteries of nude mice, and workflow for animal experiments, (b, c) Representative images and quantification of in vivo fluorescence imaging of the neck region in mice from each group 3 days post-cell injection therapy (n = 3), (d, e) Representative images and quantification of Evans Blue-stained carotid arteries from each group of mice at 3 days post-surgery (n = 3), (f, g) Representative images and quantification of immunofluorescence staining to evaluate reendothelialization in carotid arteries of each group of mice 28 days post-cell injection therapy (n = 4), (h, i) Representative images of HE staining evaluating neointima formation in carotid arteries of each group of mice 28 days post-cell injection therapy, along with quantification of the neointima area (I) to media area (M) ratio (n = 4).
S1PR3 identified as a target of lncRNA H19
To further explore the mechanisms by which lncRNA H19 regulates endothelial progenitor cells (EPCs) to enhance their endothelial repair functions, we overexpressed lncRNA H19 in EPCs in vitro and performed transcriptomic mRNA sequencing. The results indicated significant changes in mRNA expression profiles in EPCs post-overexpression of lncRNA H19 (Figure 2(a) and (b)). GO and KEGG pathway analyses revealed that pathways related to endothelial repair, such as PI3K-Akt signaling pathway, Wnt signaling pathway, and so on, were significantly enriched following the overexpression of lncRNA H19 (Figure 2(c)–(f)). By analyzing endothelial repair-related genes from the GeneCards and STRING databases and DEGs identified through sequencing, we generated a Venn diagram that revealed 38 candidate genes involved in endothelial repair (Figure 2(g)). A protein-protein interaction network for these genes was visualized using STRING (Figure 2(h)). After scoring these genes based on network centrality metrics, four key genes were identified (Figure 2(i)). Transcriptomic data showed significant upregulation of S1PR3 and AKT1 and downregulation of VCAM1 and CXCL12 in lncRNA H19-overexpressing EPCs (Figure 2(j)). Considering the trends of these four key genes and findings from the literature review, we hypothesize that S1PR3 may be a stable downstream target gene in EPCs following overexpression of lncRNA H19. Validation at the protein level confirmed that S1PR3 expression was significantly affected by the modulation of lncRNA H19 in EPCs, showing increased levels with overexpression and decreased levels with suppression (Figure 2(k)–(n)).

S1PR3 identified as a target of lncRNA H19: (a) Volcano plot illustrating mRNA expression changes in EPCs following lncRNA H19 overexpression. Each dot represents an mRNA, with red indicating upregulation, blue indicating downregulation, and gray indicating non-significant changes, (b) Heatmap showing the top 100 upregulated and downregulated mRNAs between the oe-H19 and oe-NC groups. The color intensity reflects the differences in expression levels, (c) bubble plot of BP enrichment factors for differentially expressed genes. Each dot represents a BP term, with size indicating the number of genes associated with that term. (d) Bubble plot of CC enrichment factors for differentially expressed genes. Each dot represents a CC term, with size indicating the number of genes associated with that term. (e) Bubble plot of MF enrichment factors for differentially expressed genes. Each dot represents an MF term, with size indicating the number of genes associated with that term. (f) Bubble plot of KEGG pathway enrichment factors for differentially expressed genes. Each dot represents a KEGG term, with size indicating the number of genes associated with that term, (g) Venn diagram showing 38 candidate genes related to endothelial repair, identified using transcriptome mRNA sequencing data, Genecards, and STRING databases, (h) Protein-protein interaction network of the 38 candidate genes constructed using the STRING analysis tool, (i) Venn diagram of four key genes identified through a combined screening using Betweenness Centrality, Degree, and Closeness Centrality scoring systems, (j) expression levels of the four key genes in transcriptome mRNA sequencing under lncRNA H19 overexpression conditions (n = 4), (k, l) protein expression of S1PR3 in EPCs after siRNA-mediated knockdown of lncRNA H19 (n = 5), (m, n) protein expression of S1PR3 in EPCs after adenovirus-mediated overexpression of lncRNA H19 (n = 3).
Enhanced reendothelialization and reduced neointima formation in wire-injured carotid arteries of nude mice following transplantation of S1PR3-overexpressing EPCs
To assess the impact of S1PR3 on endothelial repair, we conducted a study with wire-injured carotid arteries in nude mice (Figure 3(a)). EPCs overexpressing S1PR3 were transplanted via the tail vein 3 days post-injury. In vivo imaging showed increased cell homing to the injury site (Figure 3(b) and (c), Figure S4(a) and (b)), and Evans Blue staining revealed that these EPCs substantially improved reendothelialization (Figure 3(d) and (e)). Evaluations 28 days later using immunofluorescence staining and HE showed that S1PR3-overexpressing EPCs effectively enhanced endothelial recovery (Figure 3(f) and (g)) and reduced neointima formation compared to controls (Figure 3(h) and (i)).

Enhanced reendothelialization and reduced neointima formation in wire-injured carotid arteries of nude mice following transplantation of S1PR3-overexpressing EPCs: (a) Diagram of EPCs implantation into wire-injured carotid arteries of nude mice, and the workflow for animal experiments, (b, c) images and quantification of in vivo fluorescence imaging of the neck region in mice from each group 3 days post-cell injection therapy (n = 3), (d, e) images and quantification of Evans Blue-stained carotid arteries from each group at 3 days post-surgery (n = 3), (f, g) images and quantification of immunofluorescence staining to assess reendothelialization in carotid arteries of each group 28 days post-cell injection therapy (n = 4), (h, i) HE staining images evaluating neointima formation in carotid arteries of each group 28 days post-cell injection therapy, with quantification of the neointima area (I) to media area (M) ratio (n = 4).
S1PR3 activation of PI3K/AKT pathway modulates biological behaviors of EPCs
To investigate the regulatory effects of S1PR3 on EPCs, we modulated S1PR3 expression through lentiviral overexpression and siRNA-mediated knockdown. Knockdown of S1PR3 significantly reduced EPCs proliferation (Figure 4(a), (c), (d)), migration (Figure 4(g), (i), (m), (n)), and tube formation (Figures 4(k), 3(o)), affirming its crucial influence on these biological processes. Conversely, overexpression of S1PR3 substantially enhanced these activities (Figure 4(b), (e), (f) for proliferation; Figure 4(h), (j), (p), (q) for migration; Figure 4(l), (r) for tube formation), confirming S1PR3’s role in promoting EPCs functions. Given previous findings linking H19 to the PI3K/AKT pathway through GO enrichment of downstream differential genes and the known ability of S1PR3 to modulate this pathway, 21 we further examined the expression levels of related proteins. Knockdown of S1PR3 significantly inhibited the activation of the PI3K/AKT pathway in EPCs, while overexpression of S1PR3 markedly enhanced its activation (Figure 4S and T). These results substantiate the roles of H19 and S1PR3 in regulating EPCs behavior and highlight their impact on the PI3K/AKT signaling pathway essential for vasculogenic functions.

S1PR3 activation of PI3K/AKT pathway modulates biological behaviors of EPCs: (a) CCK8 assay evaluating the proliferation capacity of EPCs after S1PR3 knockdown, (b) CCK8 assay evaluating the proliferation capacity of EPCs after S1PR3 overexpression, (c, d) representative images and quantification bar graph of EdU assay assessing the proliferation capacity of EPCs after S1PR3 knockdown, (e, f) representative images and quantification bar graph of EdU assay assessing the proliferation capacity of EPCs after S1PR3 overexpression, (g, m) representative images and quantification bar graph of scratch assay evaluating the migration ability of EPCs after S1PR3 knockdown, (h, p) representative images and quantification bar graph of scratch assay evaluating the migration ability of EPCs after S1PR3 overexpression, (i, n) representative images and quantification bar graph of Transwell assay assessing the migration ability of EPCs after S1PR3 knockdown, (j, q) representative images and quantification bar graph of Transwell assay assessing the migration ability of EPCs after S1PR3 overexpression, (k, o) representative images and quantification bar graph of in vitro tube formation assay evaluating the angiogenic function of EPCs after S1PR3 knockdown, (l, r) representative images and quantification bar graph of in vitro tube formation assay evaluating the angiogenic function of EPCs after S1PR3 overexpression, (s, t) representative images and quantification bar graph of PI3K/AKT signaling pathway activation after S1PR3 knockdown/overexpression.
S1PR3 knockdown mitigates the enhancing effects of H19 overexpression on the biological functions of EPCs
To evaluate the impact of abnormal S1PR3 expression on EPCs, siRNA was used to target S1PR3. Significantly, reducing S1PR3 levels counteracted the increases in proliferation (Figure 5(a), (e), (i)), migration (Figure 5(b), (c), (f), (g)), and tube formation (Figure 5(d), (h)) caused by overexpressing H19. Corresponding changes were also noted in the PI3K/AKT pathway (Figure 5(j) and (k)). These findings strongly support the role of S1PR3 as a key mediator of H19’s effects in EPCs.
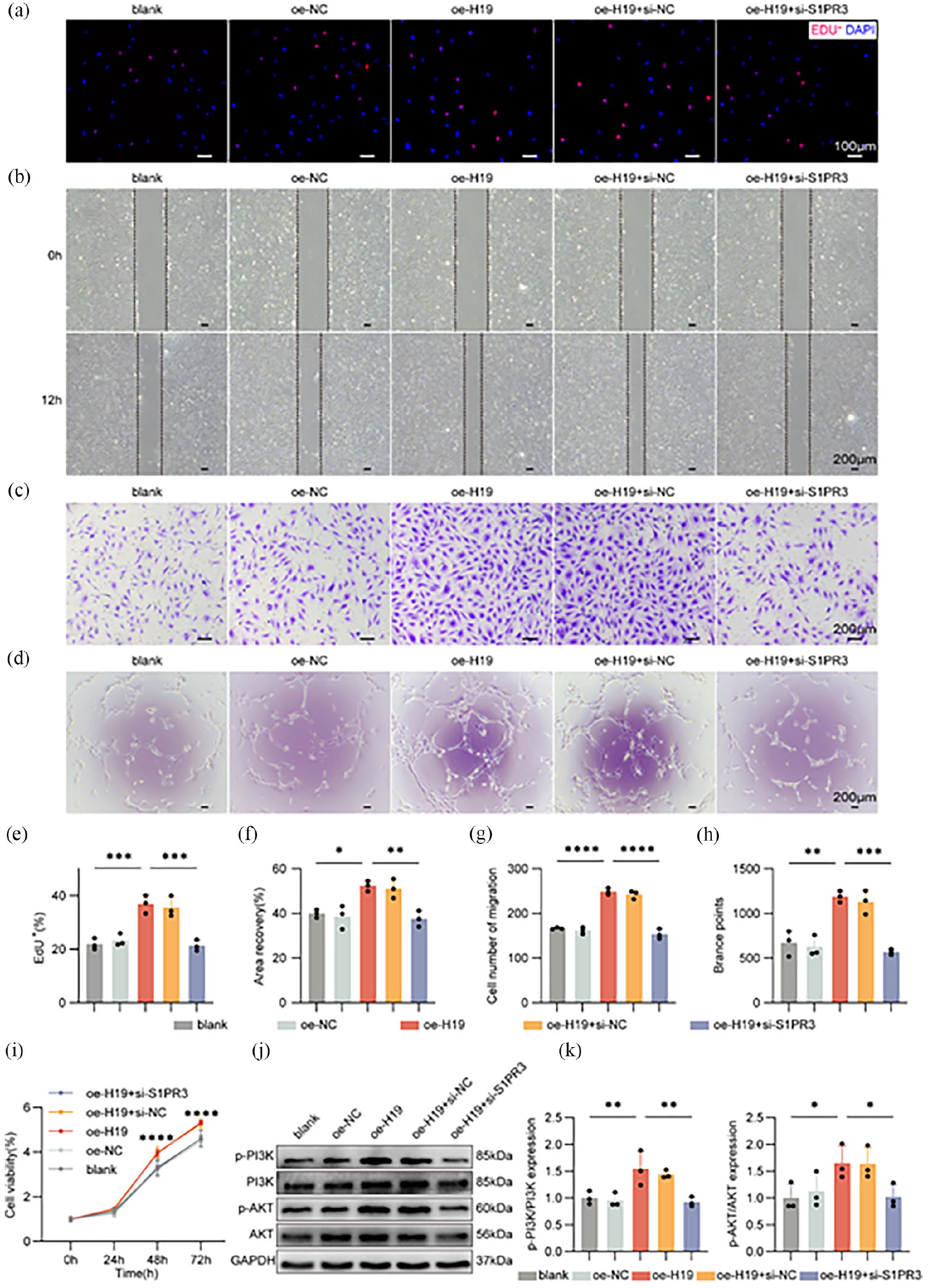
S1PR3 knockdown mitigates the enhancing effects of H19 overexpression on the biological functions of EPCs: (a, e) Representative images of EdU assay evaluating the proliferation capacity of EPCs, (b, f) representative images of scratch assay assessing the migration potential of EPCs, (c, g) representative images of Transwell assay determining the migration potential of EPCs, (d, h) representative images of tube formation assay evaluating the angiogenic capacity of EPCs, (i) CCK-8 assay assessing the proliferation capacity of EPCs, (j, k) representative images and quantification bar graphs of western blot analysis for PI3K/AKT signaling pathway activation.
LncRNA H19 enhances acetylation of histone 3 at lysine 27 (H3K27ac) by inhibiting HDAC2 to promote S1PR3 expression
To investigate the transcriptional regulation mechanisms of S1PR3 by lncRNA H19, we utilized the UCSC database to identify enriched H3K27 acetylation in the upstream region of S1PR3 (Figure 6(a)). Further analysis using the catRAPID database predicted 2063 potential interacting proteins for H19. After compiling all 18 enzymes associated with histone acetylation, a Venn diagram was generated, suggesting a potential interaction between H19 and HDAC2 (Figure 6(b)). This interaction suggests that H19 could influence S1PR3 expression by interacting with HDAC2, thereby affecting H3K27 acetylation at the S1PR3 promoter. Bioinformatics analysis confirmed a high binding affinity between H19 and HDAC2 (Figure 6(c) and (d)), further supporting this interaction. Next, we treated EPCs with Mocetinostat, a relatively selective HDAC2 inhibitor, and found that S1PR3 expression was significantly increased (Figure 6(e)–(h)). Additionally, treating EPCs with H19 overexpression adenovirus significantly increased S1PR3 expression and elevated H3K27ac levels (Figure 6(i)–(l)). ChIP assays revealed that the promoter region of S1PR3 was enriched with HDAC2 binding and H3K27ac signals (Figure 6(m)). Given that HDAC2 is a key enzyme in regulating chromatin acetylation, we investigated its role in S1PR3 regulation. As hypothesized, reducing HDAC2 expression using siRNAs significantly increased S1PR3 mRNA and protein levels (Figure 6(n)–(p)). These data suggest that HDAC2 can influence S1PR3 transcription in EPCs by binding to the promoter and modifying histone acetylation. To explore the role of H19 in HDAC2 regulation of S1PR3, RNA-protein pull-down assays indicated that H19, but not antisense H19, could distinctly retrieve HDAC2 from EPCs total protein extracts in vitro (Figure 6(q)). Furthermore, RIP assays confirmed substantial enrichment of H19 using an anti-HDAC2 antibody compared to the negative control (Figure 6(r)). These findings strongly suggest that H19 can significantly bind HDAC2. Additionally, ChIP assays demonstrated that H19 knockdown decreased HDAC2 enrichment and increased H3K27ac at the S1PR3 promoter region (Figure 6(s) and (t)). Therefore, we conclude that H19 binding to HDAC2 reduces HDAC2 enrichment, thereby increasing histone acetylation at the promoter and inducing S1PR3 transcription. In summary, H19 recruits HDAC2 in the nucleus and modulates H3K27ac levels at the S1PR3 promoter, thereby enhancing gene transcription. In the cytoplasm, S1PR3 activates the PI3K/Akt signaling pathways, promoting EPC-mediated reendothelialization (Figure 7).

lncRNA H19 enhances acetylation of histone 3 at lysine 27 (H3K27ac) by inhibiting HDAC2 to promote S1PR3 expression: (a) Layered H3K27 acetylation enrichment in the upstream region of S1PR39 as shown in the UCSC database, (b) a Venn diagram suggesting a potential interaction between H19 and HDAC2, with 2063 potential interacting proteins for H19 predicted by the catRAPID database and all 18 histone acetylation enzymes, (c) the interaction profile of H19 showing interaction scores across different nucleotide positions as analyzed by the catRAPID database. Higher scores indicate stronger interactions, (d) the interaction matrix depicts the interaction between RNA nucleotides and protein residues as analyzed by the catRAPID database. Each point represents an interaction, with the intensity of color indicating the interaction strength, (e, f, g, h) detection and quantification of S1PR3 and histone 3 acetylation following 24-h treatment with 500 nM Mocetinostat (n = 3), (i, j, k, l) detection and quantification of S1PR3 and histone 3 acetylation following 48-h treatment with 50 nM H19 overexpression adenovirus (n = 3), (m) ChIP-qPCR analysis showing HDAC2 and H3K27ac enrichment at the S1PR3 promoter (n = 3), (n) RT-qPCR analysis of S1PR3 expression in si-HDAC2 treated EPCs (n = 3), (o, p) Western blot analysis of S1PR3 expression in EPCs following si-HDAC2 treatment (n = 3), (q) RNA pull-down assay demonstrating that H19 retrieves HDAC2 in EPCs, as detected by western blot. Input represents the total cellular protein used, (r) RIP experiment demonstrating that the anti-HDAC2 antibody can precipitate H19 in EPCs (n = 3), (s, t) ChIP-qPCR analysis revealed that H19 knockdown decreased the enrichment of HDAC2 and increased the level of H3K27ac acetylation at the S1PR3 promoter region (n = 3).

Schematic diagram illustrating that lncRNA H19 positively regulates S1PR3 expression by disrupting HDAC2/H3K27ac binding in EPCs.
Discussion
Endovascular stent angioplasty has increasingly become the preferred treatment for patients with symptomatic PAD, largely due to its minimally invasive nature and effective vascular reconstruction. 22 However, a major limitation is ISR, which typically occurs months after the procedure. 23 ISR involves a narrowing of the vessel lumen by more than 50% near the stent, often leading to treatment failure and the need for additional interventions. 24 IH, the accumulation of SMCs and extracellular matrix in the intima, is driven by an imbalance between inhibitory and stimulatory factors, including inflammatory mediators and coagulation disruptions. 25 This process is a key biological mechanism behind ISR. Recent studies suggest that EPCs can promote reendothelialization and potentially prevent restenosis.26 –28 Nevertheless, current methods to enhance reendothelialization—through mobilization, recruitment, or transplantation of EPCs—have not yet produced significant results.29,30 Engineering EPCs in vitro presents a promising new direction for improving ISR treatments.
Our previous studies have demonstrated that H19 is significantly enriched in EPCs. 20 H19-overexpressing EPCs (oe-H19 EPCs) exhibit enhanced vasculogenesis capacity in PAD patients under ischemic, hypoxic, and inflammatory conditions, effectively restoring blood flow to ischemic limbs. We investigated whether these oe-H19 EPCs also possess superior homing and endothelial repair abilities compared to regular EPCs. If confirmed, this could significantly promote reendothelialization post-stent implantation, thereby inhibiting IH formation and reducing ISR incidence. Using a carotid artery wire injury model to simulate post-stent endothelial injury, 31 we administered early tail vein injections of oe-H19 EPCs. Fluorescent dye tracking revealed significantly increased homing of these EPCs to the injury site. Further analysis with Evans Blue and HE staining indicated that oe-H19 EPCs effectively promoted reendothelialization of the damaged vessel wall. They significantly inhibited IH formation and maintained lumen patency. These findings provide strong evidence for the early transplantation of oe-H19 EPCs to inhibit IH formation and reduce ISR risk after vascular stent implantation. Previous studies showed that CXCR7-overexpressing EPCs helped suppress IH formation in diabetic rat carotid artery injury models. 32 Our study used human peripheral blood-derived EPCs, offering a more practical approach for future clinical applications. This method avoids species-specific immune rejection and provides a convenient source of EPCs for therapeutic use. In the future, we can further explore the potential of transplanting humanized autologous peripheral blood-derived EPCs engineered to overexpress H19.
Transcriptomic sequencing on oe-H19 EPCs was further conducted, revealing significant enrichment in endothelial repair pathways, including the PI3K/AKT pathway, Wnt signaling pathway, and cell adhesion molecules. Literature review and experimental validation identified S1PR3 as a downstream gene regulated by H19. Overexpression of H19 significantly upregulated S1PR3 expression in EPCs. Sphingosine-1-phosphate (S1P), a signaling lipid synthesized by sphingosine kinases (SPHK1 and SPHK2), exerts its effects through receptors (S1PR1, S1PR2, S1PR3, S1PR4, S1PR5) expressed differentially across tissues, influencing cardiovascular functions in various ways.33,34 Among the five S1P receptors, S1PR1, S1PR2, and S1PR3 are predominantly expressed in cardiovascular tissues, while S1PR4 is enriched in the lymphatic system and S1PR5 in the immune and nervous systems.35,36 S1PR1 and S1PR3 are the main receptor types in ECs. Activation of the S1P/S1PR1/S1PR3 axis enhances endothelial barrier integrity and cell-cell junctions, promoting vasodilation via eNOS activation, thus exhibiting anti-atherosclerotic effects. 37 Injury-induced TGFβ1 upregulation promotes S1P secretion, which enhances VEGFA expression and angiogenesis in the mouse cornea through the S1PR3/PI3K/AKT pathway. 38 Additionally, S1P can significantly enhance bone marrow-derived EPCs proliferation and reduce apoptosis caused by oxidative stress via the S1PR3-mediated PI3K/AKT pathway. 21 In this study, we found that S1PR3 activation significantly increased EPCs proliferation, migration, and tube formation capabilities. Compared to normal EPCs, S1PR3-overexpressing EPCs showed enhanced homing to injured vessels, effectively promoted reendothelialization, inhibited IH formation, and maintained lumen patency. These findings suggest that targeting S1PR3 for EPCs modification could provide new strategies and options for enhancing EPC-based therapies.
Histone acetylation modifications impact various cellular physiological processes, including transcription, mitosis, and differentiation. 39 They play significant roles in diseases such as cancer, immune disorders, and vascular diseases.40,41 Increased levels of histone acetylation reduce the affinity between histones and DNA, leading to a more relaxed chromatin nucleosome structure. 42 This relaxation facilitates the binding of transcription factors to DNA, thereby promoting the transcription and expression of corresponding genes. 43 Conversely, decreased acetylation levels exert the opposite effect. H3K27ac, acetylation at the 27th lysine residue of histone H3, is associated with higher transcriptional activation and is found near both proximal and distal transcription start sites (TSS). 44 Mediating chromatin remodeling and histone modifications to influence downstream gene expression is one of the classical regulatory mechanisms of long non-coding RNAs. 45 Through the UCSC database, we identified significant enrichment of H3K27ac peaks in the promoter region of the H19-regulated downstream gene S1PR3. Building on this foundation, we further revealed that lncRNA H19 increases the enrichment of H3K27ac and histone acetylation at the S1PR3 promoter region by interacting with HDAC2, thereby inducing the transcription of S1PR3. Previous studies have suggested that HDAC2 can inhibit H19 expression by deacetylating H3K27, thereby suppressing cancer metastasis.46,47 However, in this study, we found that in EPCs, lncRNA H19 can interact with HDAC2 to increase histone acetylation at the promoter region of downstream genes, promoting their transcription. This discrepancy may be due to different mechanisms of action in various cell types.
The current limitations of autologous EPCs transplantation in ISR patients mainly include the insufficient number of circulating EPCs, functional impairments of EPCs, and challenges associated with their isolation, culture, and expansion.10,48 For experimental convenience, we utilized EPCs from healthy donors and conducted transplantation experiments in immunodeficient nude mice to evaluate the therapeutic effects of the modified EPCs, thus avoiding the influence of immune responses. In actual clinical scenarios, prior to stent implantation in patients with PAD, autologous EPCs could potentially be harvested from a certain volume of the patient’s peripheral venous blood. Using tissue engineering strategies, these autologous EPCs could be isolated, extensively expanded, and genetically modified to enhance their biological properties, such as suppressing neointimal hyperplasia.9,49,50 This approach could facilitate the clinical translation and application of EPC-based therapies for ISR prevention and treatment. Furthermore, by elucidating the mechanisms of action of these engineered EPCs, future studies could pave the way for acellular therapeutic strategies, such as treatments utilizing extracellular vesicles derived from modified EPCs.51,52 This transition from EPCs transplantation to acellular therapies may offer significant advantages for clinical applications by overcoming the challenges associated with cell-based therapies.
Despite revealing that EPCs with high H19 expression can inhibit IH by promoting early reendothelialization, this study has some limitations. First, we did not evaluate IH on days 7 and 14 after EPCs injection therapy, only assessing long-term IH at day 28. Additionally, since SMCs proliferation and migration play a crucial role in IH, further research is needed to determine whether the EPCs homing to the injury site and promoting early reendothelialization influence SMCs behavior. Finally, to achieve greater breakthroughs in the prevention and treatment of ISR using EPCs with high H19 expression, more extensive studies involving larger sample sizes and more large animal models are necessary.
Conclusion
In conclusion, this study demonstrates that lncRNA H19 promotes S1PR3 expression through histone acetylation modifications, activating the PI3K/AKT pathway and significantly enhancing EPCs proliferation, migration, and tube formation capabilities. Transplantation of engineered EPCs with high H19 expression promotes reendothelialization of injured vessels and inhibits IH. Our findings provide a solid scientific basis for the potential use of EPCs transplantation therapy to prevent ISR following vascular stent implantation.
Supplemental Material
sj-docx-1-tej-10.1177_20417314251315959 – Supplemental material for Transplantation of engineered endothelial progenitor cells with H19 overexpression promotes arterial reendothelialization and inhibits neointimal hyperplasia
Supplemental material, sj-docx-1-tej-10.1177_20417314251315959 for Transplantation of engineered endothelial progenitor cells with H19 overexpression promotes arterial reendothelialization and inhibits neointimal hyperplasia by Yanchen Ye, Lin Huang, Kangjie Wang, Yunhao Sun, Zhihao Zhou, Tang Deng, Yunyan Liu, Rui Wang, Ridong Wu and Chen Yao in Journal of Tissue Engineering
Supplemental Material
sj-docx-2-tej-10.1177_20417314251315959 – Supplemental material for Transplantation of engineered endothelial progenitor cells with H19 overexpression promotes arterial reendothelialization and inhibits neointimal hyperplasia
Supplemental material, sj-docx-2-tej-10.1177_20417314251315959 for Transplantation of engineered endothelial progenitor cells with H19 overexpression promotes arterial reendothelialization and inhibits neointimal hyperplasia by Yanchen Ye, Lin Huang, Kangjie Wang, Yunhao Sun, Zhihao Zhou, Tang Deng, Yunyan Liu, Rui Wang, Ridong Wu and Chen Yao in Journal of Tissue Engineering
Supplemental Material
sj-docx-3-tej-10.1177_20417314251315959 – Supplemental material for Transplantation of engineered endothelial progenitor cells with H19 overexpression promotes arterial reendothelialization and inhibits neointimal hyperplasia
Supplemental material, sj-docx-3-tej-10.1177_20417314251315959 for Transplantation of engineered endothelial progenitor cells with H19 overexpression promotes arterial reendothelialization and inhibits neointimal hyperplasia by Yanchen Ye, Lin Huang, Kangjie Wang, Yunhao Sun, Zhihao Zhou, Tang Deng, Yunyan Liu, Rui Wang, Ridong Wu and Chen Yao in Journal of Tissue Engineering
Supplemental Material
sj-docx-4-tej-10.1177_20417314251315959 – Supplemental material for Transplantation of engineered endothelial progenitor cells with H19 overexpression promotes arterial reendothelialization and inhibits neointimal hyperplasia
Supplemental material, sj-docx-4-tej-10.1177_20417314251315959 for Transplantation of engineered endothelial progenitor cells with H19 overexpression promotes arterial reendothelialization and inhibits neointimal hyperplasia by Yanchen Ye, Lin Huang, Kangjie Wang, Yunhao Sun, Zhihao Zhou, Tang Deng, Yunyan Liu, Rui Wang, Ridong Wu and Chen Yao in Journal of Tissue Engineering
Footnotes
Acknowledgements
None.
Author contributions
The study was conceptualized and designed by Chen Yao, Ridong Wu, and Rui Wang. Yanchen Ye, Lin Huang, and Kangjie Wang performed the in vitro experiments, whereas Tang Deng and Yunyan Liu conducted the in vivo experiments. Zhihao Zhou and Yunhao Sun were in charge of the sequencing data analysis. Yanchen Ye and Lin Huang led the manuscript preparation. All authors reviewed and approved the final version of the manuscript.
Availability of data and materials
The data underpinning the results of this study are available from the corresponding author upon reasonable request.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the National Natural Science Foundation of China (grant number: 82070495), the National Natural Science Foundation of Guangdong Province (grant number: 2023A1515011602), and the China Postdoctoral Science Foundation Fellowship (2022M723645), and the Guangdong Basic and Applied Basic Research Foundation of provincial and municipal joint fund (2023A1515111014).
Consent for publication
Not applicable.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
