Abstract
Gonorrhea, a sexually transmitted disease caused by the bacteria Neisseria gonorrhoeae, is characterized by a large number of neutrophils recruited to the site of infection. Therefore, proper modeling of the N. gonorrhoeae interaction with neutrophils is very important for investigating and understanding the mechanisms that gonococci use to evade the immune response. We have used a combination of a unique human 3D tissue model together with a dynamic culture system to study neutrophil transmigration to the site of N. gonorrhoeae infection. The triple co-culture model consisted of epithelial cells (T84 human colorectal carcinoma cells), human primary dermal fibroblasts, and human umbilical vein endothelial cells on a biological scaffold (SIS). After the infection of the tissue model with N. gonorrhoeae, we introduced primary human neutrophils to the endothelial side of the model using a perfusion-based bioreactor system. By this approach, we were able to demonstrate the activation and transmigration of neutrophils across the 3D tissue model and their recruitment to the site of infection. In summary, the triple co-culture model supplemented by neutrophils represents a promising tool for investigating N. gonorrhoeae and other bacterial infections and interactions with the innate immunity cells under conditions closely resembling the native tissue environment.
Keywords
Introduction
The sexually transmitted disease (STD) gonorrhea caused by Neisseria gonorrhoeae is the second most common STD in the world. 1 More than 100 million new infections per year as well as the rapid increase of antibiotic resistance make gonorrhea a serious public health threat. 2 The mucosal layer of the urogenital epithelium is the first surface that encounters N. gonorrhoeae upon it entering the human body. The influx of a large number of neutrophils from the blood to the mucosal surface of the urogenital epithelium is one of the characteristics of gonorrhea.
N. gonorrhoeae contains various virulence factors, which stimulate the mucosal epithelium to release the pro-inflammatory cytokines such as interleukin (IL)-6, IL-8 and tumor necrosis factor alpha (TNFα), causing the recruitment of neutrophils, the frontline of innate immune defense.3,4 The induction of the expression of the cluster of differentiation molecule 11B (CD11b) in neutrophils occurs upon their activation and in response to inflammation. The trafficking process of neutrophils starts with adhesion of CD11b to intracellular adhesion molecule 1 (ICAM-1) present on the endothelial cell surface.5–9 After passing the endothelial layer, neutrophils continue migrating through the collagen fibers of the extracellular matrix (ECM) to reach the infected epithelium with assistance of collagenase granules.4,10 Neutrophils eliminate pathogens via various defense mechanisms such as phagocytosis, release of antimicrobial molecules, and formation of the neutrophil extracellular traps (NETs).11–13 However, N. gonorrhoeae modulate antibacterial activities of neutrophils by different strategies, which include delaying apoptosis, evading killing by NETs, or delaying the fusion of phagosomes with primary granules.14–16 Therefore, neutrophils are not able to completely clear gonococcal infection, resulting in survival of a population of N. gonorrhoeae that can replicate inside the phagosomes, allowing the bacteria to disseminate into deeper tissue layers or even the bloodstream.17,18
The early stages of N. gonorrhoeae infection with respect to the interaction with the immune system have been studied using transgenic animal models.19–21 In spite of the advantages of animal models for studying host-pathogen interaction, they cannot fully recreate all the features of the N. gonorrhoeae infection in humans.22–24 Recently, various epithelial cell monolayer models using Transwell® technology have been developed in order to study the key mechanisms that underline neutrophil transepithelial migration, including pathogen-epithelial interaction, the role of cell adhesion molecules in neutrophil recruitment, and infectious dose.25–27 Furthermore, the neutrophil-endothelial communication has been investigated using a monoculture of endothelial cells in Transwell® models, indicating neutrophil lateral crawling over the endothelial cell surface followed by transendothelial migration.28,29 In addition, sequential trafficking of the neutrophils across the bilayer of lung epithelial and endothelial cells presented the importance of more physiological and predictive models. 30 However, to the best of our knowledge, there has been no report so far of using epithelial and endothelial in vitro co-culture system for studying neutrophil transmigration upon N. gonorrhoeae infection.
Previously, we have reported an establishment of 3D tissue models based on a co-culture of various cells of mucosal surfaces with primary human dermal fibroblasts (HDFib) on the small intestinal submucosa (SIS) scaffold, which enabled us to study N. gonorrhoeae infection using a biomimetic platform. 31 Here, we introduce a novel 3D model of an increased level of complexity. This tissue model is based on the triple co-culture of epithelial cells (T84 human colorectal adenocarcinoma cells), connective tissue (HDFib), and endothelial cells representing the inner lining of a blood vessel (human umbilical vein endothelial cells HUVEC) on the SIS scaffold, recreating the site of infection during the neutrophil recruitment. We used a perfusion-based bioreactor system to mimic the dynamic neutrophil trafficking from the blood to the site of infection, which included the interaction with and transmigration through the endothelial, ECM, and epithelial layer. The in vitro model presented here provides the opportunity to investigate the interaction between N. gonorrhoeae and neutrophils under conditions closely mimicking the situation found in the native tissues at the site of infection.
Material and methods
Cell lines
Human dermal fibroblasts (HDFib) were isolated from human foreskin biopsy as described previously 32 and cultured in Dulbecco’s Modified Eagle Medium (DMEM) (Gibco/Thermo Fisher scientific, Massachusetts, USA). T84, the human colorectal carcinoma cells (ATCC® CCL-248™) were cultured in DMEM/F12 (Gibco/Thermo Fisher scientific, Massachusetts, USA). All media were supplemented with 10% heat-inactivated Fetal Calf Serum (FCS) (Sigma/Merck, Darmstadt, Germany) and 1% Penicillin/Streptomycin (Gibco/Thermo Fisher scientific, Massachusetts, USA). Human Umbilical Vein Endothelial Cells (HUVEC) were cultured in M200 (Gibco/Thermo Fisher scientific, Massachusetts, USA) supplemented with 10% Low Serum Growth Supplement (LSGS) and 1% Penicillin/Streptomycin (Gibco/Thermo Fisher scientific, Massachusetts, USA).
Establishment of human 3D tissue model
In order to study neutrophil transmigration to the site of infection, we modified the previously established co-culture 3D tissue model by triple co-culturing of human primary fibroblast cells, human colorectal carcinoma cells and human umbilical vein endothelial cells. For this purpose, models were generated as previously described. 31 The basal side of the SIS scaffold was coated with 5 μg/cm2 fibronectin (Sigma Aldrich, Missouri, USA). On day 7, the co-culture models were inverted and HUVECs (6 × 105 cell/cm2) were seeded on the basolateral side of the SIS scaffold (Figure 1) using co-culture medium. The model was then transferred to an orbital shaker (50 rpm/min) in order to assist the maturation of endothelial cells. 33

The workflow of the 3D tissue model establishment, infection and neutrophil circulation: (a) the preparation of the triple co-culture tissue model on the SIS scaffold. After preparation of the SIS scaffold, the scaffold was fixed between two cylinders in the cell crown. Primary human dermal fibroblasts (HDFib) cells were then seeded on the SIS scaffold and cells were incubated (37°C/5% CO2) (Day-2). After 2 days of incubation, the epithelial cells were seeded on the apical side of the SIS scaffold (Day 0). Endothelial cells (HUVECs) were seeded on the basolateral side 1 week later (Day 7) and cultivated for 5 days. The co-culture medium was exchanged every 2 days until the mature tissue model was obtained. After establishing the 3D tissue model, the triple co-culture model (T84/HDFib/HUVECs) was infected with N927-GFP N. gonorrhoeae under static conditions. After 24 h of infection, the infected tissue was transferred to the bioreactor system. The freshly isolated human neutrophils were administered to the endothelial side (basal compartment) via perfusion in the bioreactor system, (b) schematic, and (c) setup of the perfusion-based bioreactor system, comprising a main chamber, an external reservoir for culture medium, and neutrophil reservoir. The tissue model is placed in the main chamber and the culture medium is circulated through the system using peristaltic pumps.
Barrier integrity
The barrier integrity of the triple co-culture model was assessed using two different methods. Transepithelial electrical resistance (TEER) was measured using Millicell® ERS-2 Volt-Ohm Meter (Millicell ERS-2, Millipore, Massachusetts, USA) to evaluate the barrier integrity of 3D tissue model during the cultivation time. Afterwards, the model was evaluated for barrier integrity using 4 kDa fluorescein isothiocyanate (FITC)-dextran permeability assay (Sigma Aldrich, Missouri, USA). 31
N. gonorrhoeae culture
N. gonorrhoeae N927 (PorBIA, Pili-, Opa-) expressing GFP was grown on the GC agar (Gibco/Thermo Fisher scientific, Massachusetts, USA) plates supplemented with 1% vitamin mix for 14–17 h at 37°C in 5% CO2. The mature triple co-culture model was then infected with N. gonorrhoeae (MOI 20) under static condition for 24 h. We performed the infection in phosphate and serum-free medium (HEPES) in order to permit the phosphate-sensitive interaction with PorBIA.
Neutrophil isolation
Primary human neutrophil isolation from the blood was performed based on Ficoll® (Sigma Aldrich, Missouri, USA) separation method. Briefly, the whole blood was slowly layered on Ficoll®, which was followed by the centrifugation. The neutrophil layer was then separated, and residual erythrocytes were lysed.
Bioreactor setup
A perfusion-based bioreactor system was designed for studying the immune response (neutrophils) to the pathogens under dynamic culture condition. The details of the bioreactor have previously been described by Schweinlin et al. 34 Briefly, the bioreactor consists of a main chamber including two parts, apical and basal, an extra reservoir and a neutrophil reservoir (Figure 1). After maturation on the cell crowns and infection with N. gonorrhoeae, tissue models were carefully transferred into the bioreactor and two sides of the bioreactor were assembled, generating the apical and basal compartment of the main chamber, which are separated by the tissue model (Supplemental Figure S1). The bioreactor was then transferred to a custom-designed incubator (37°C and 5% CO2). The culture medium was circulated (perfusion rate of 0.5 mL/min) through the apical part (epithelial cells) and the isolated human neutrophils were delivered to the endothelial cells in the basal chamber via a perfusion pump (perfusion rate of 2.5 mL/min). Freshly isolated human neutrophils (200,000 cells/mL) were administered to the endothelial side by adding the neutrophils to the reservoir of the basal chamber compartment. The bioreactor system was allowed to run for 2 h and the culture media were then taken for downstream analysis. The setup enables simultaneous run of up to four bioreactors.
Flow cytometry analysis
Following the 2 h of neutrophil circulation, the supernatant culture medium was collected in 50 mL falcon tube for further flow cytometry analysis to characterize the activation state of neutrophils using CD11b marker (MACS). GFP-positive neutrophils were also sorted using flow cytometry. Neutrophils were transferred into a falcon tube and centrifuged for 5 min, 250 × g to pellet the cells. The pellet was resuspended in 100 µL of the corresponding buffer and the antibody was added according to the manufacturer’s instruction. Measurements were performed with FACS Aria III and the data were analyzed by Flowing software (Turku Centre for Biotechnology).
ELISA
Measurements of the quantity of IL-8 in the cell culture medium was performed using Human IL-8/CXCL8 DuoSet ELISA kit from R&D systems according to manufacturer’s instructions.
Histological and immunohistochemical analysis
The 3D tissue model was fixed using 4% PFA and stained with different markers. After paraffin embedding, samples were sectioned to 6 μm thickness. Hematoxylin and eosin staining and immunohistochemistry staining were performed in order to stain different parts of tissue samples specifically. 31
Immunofluorescence analysis
Tissue models were fixed with 4% paraformaldehyde for 2 h at room temperature. The tissue models were then washed with phosphate-buffered saline (PBS), permeated using 1% saponin, blocked with 1% BSA in PBS and stained with primary antibodies overnight. The cells were stained with anti E-Cadherin (rabbit, 1:100, Proteintech, Illinois, USA), anti ZO-1 (rabbit, 1:100, Proteintech, Illinois, USA), anti-fibroblast (mouse, 1:100, Antikörper, Aachen, Germany), anti-neutrophil cytosolic factor 2 (rabbit, 1:100, Antikörper, Aachen, Germany), anti-VE cadherin (mouse, 1:100, Gibco/Thermo Fisher scientific, Massachusetts, USA), anti-CD31 (mouse, 1:100, Abcam, Cambridge, United Kingdom), and anti-MUC1 (rabbit, 1:100, Abcam, Cambridge, United Kingdom) antibodies. This was followed by decoration with fluorophore-coupled secondary antibodies (Cy5 and Cy3) (Dianova, Hamburg, Germany), Phalloidin (MoBiTec, Göttingen, Germany) and DAPI (Sigma Aldrich, Missouri, USA). Z-stacks of images were obtained through 25 μm from the top of the monolayer using Leica SP5 confocal system and processed by FIJI and FIJI Plugin 3D Viewer.35,36 The confocal image of the whole 3D tissue model was surface rendered by IMARIS Version 8.4.1.
Statistical analysis
All data were analyzed using GraphPad Prism 8.4.2 (GraphPad Software, LA Jolla, CA, USA). The details of each statistical analysis were reported in the caption of the figures.
Results
Establishment of a triple co-culture model
To establish a triple co-culture of T84, HDFib, and HUVEC cells (Figure 1), decellularized porcine SIS scaffold was populated with HDFibs, followed by the seeding of epithelial T84 cells 2 days later, as reported before. 31 After 7 days of epithelial cell culture, we seeded HUVECs on the basal side of the scaffold and allowed the cells to mature for another 6 days under orbital shear stress using an orbital shaker (50 rpm). The triple co-culture tissue model was then infected apically with N. gonorrhoeae for 24 h. The infected tissue model was then transferred from a Transwell-like setting (i.e. so called cell crowns) to a perfusion bioreactor system, which allowed the separate circulation of medium on the apical and basal side of the model (Supplemental Figure S1).
Characterization of the established triple co-culture model
Histological characterization of the established tissue model after fixation and paraffin embedding of the tissue model showed the presence of polarized epithelial monolayer on the apical side, fibroblasts in the middle and flat endothelial monolayer on the basal side of the tissue model (Figure 2(a)). The tissues were also stained with anti-E-cadherin, anti-fibroblast and anti-mucin 1 (Muc1) antibodies in order to visualize the epithelial cell-cell contacts, connective tissue and mucus production on the apical side (Figure 2(b)–(d)). In addition, the samples were stained for CD31 and VE-cadherin as specific markers of the endothelial cells present on the basal side (Figure 2(e) and (f)). Immunofluorescence analysis of the whole tissue models using confocal microscopy revealed that the polarized epithelial monolayer is located on the apical side and a confluent flat monolayer of endothelial cells is present on the basal side of the scaffold (Figure 3(a) and (b)), as shown on the scheme (Figure 3(c)).

Histological and immunofluorescence analysis of the triple co-culture model: (a) H&E staining of the triple co-culture 3D tissue model, showing three layers of cells including epithelial (T84, top layer), fibroblasts (HDFib, middle layer), and endothelial (HUVEC, basal layer) cells and (b–f) the tissue model was stained with anti-E-cadherin (red), anti-fibroblast (cyan), anti-MUC1 (magenta), anti-CD31 (green), anti-VE-cadherin (yellow) and DAPI (blue). The images show the formation of an epithelial (stained by E-cadherin) and endothelial (stained by VE-cadherin and CD31) monolayer and the presence of a connective tissue (stained by anti-fibroblast) and mucin layer (stained by anti MUC1). The scale bars are 50 µm.
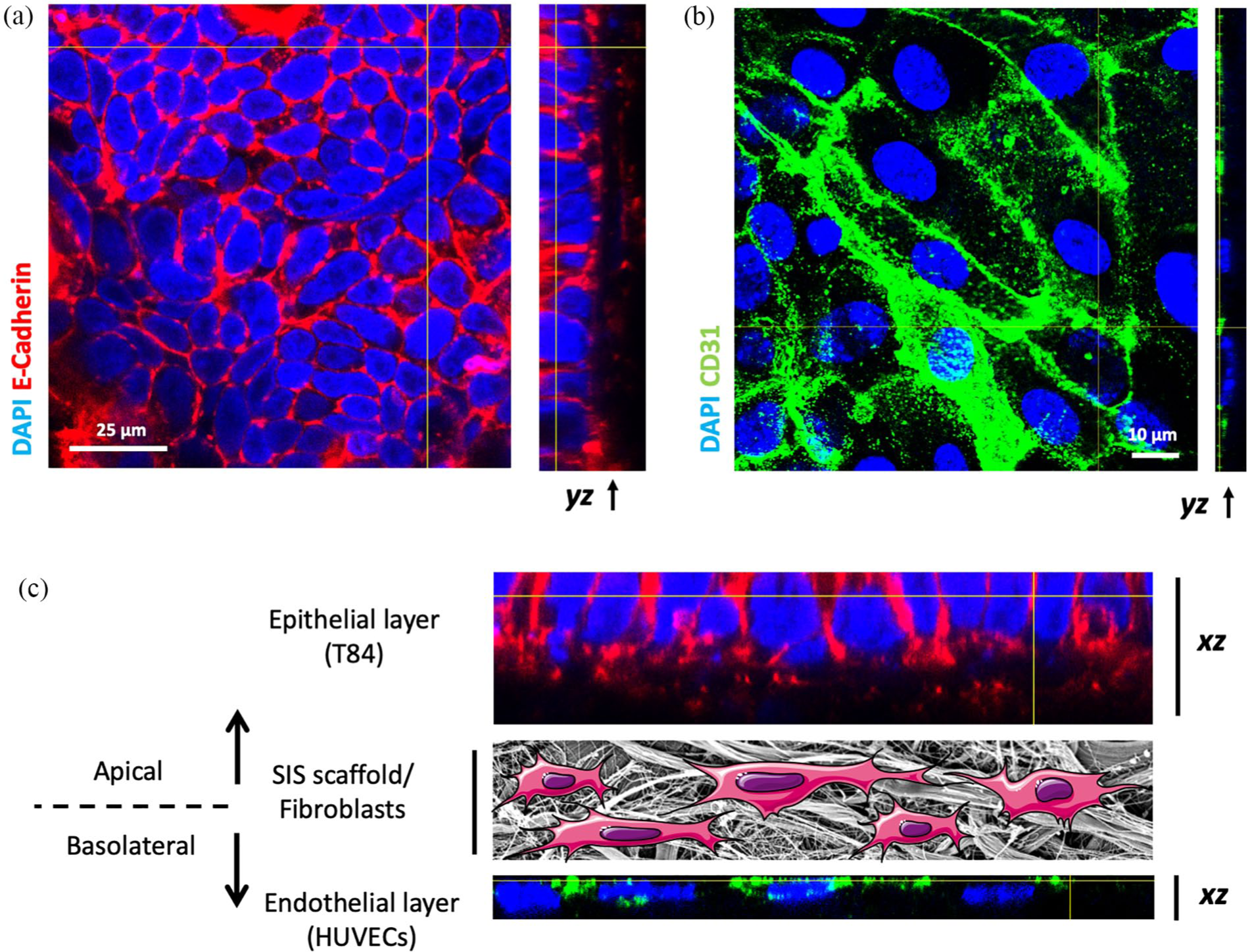
Immunofluorescence analysis using confocal laser microscopy of triple co-culture model. Immunofluorescence analysis of the triple co-culture model (T84/HDFib/ HUVECs) after maturation: (a) XY and YZ view of the apical side of the tissue model (epithelial, T84), showing E-cadherin junctions. Z-stacks were acquired using confocal fluorescence microscopy from the top of the epithelial layer and were reconstructed using FIJI and (b) XY and YZ view of the basal side of the tissue model (endothelial, HUVECs), depicting endothelial cell adhesion, which was stained by CD31. Z- stacks were made from the top of the endothelial layer to the beginning of the SIS scaffold. (C) XZ view of the triple co-culture model, showing the apical and basolateral side together. The middle layer is SEM image of the SIS scaffold, showing the fibrous structure, with fibroblasts represented by cartoons. E-Cadherin (red), CD31 (green), and DAPI (blue).
We also monitored the barrier integrity of the triple co-culture tissue model using electrical TEER measurement and FITC-dextran permeability assay. The TEER values for the triple co-culture model of T84/HDFib/HUVECs showed a slight increase (≈400 Ω × cm2) in comparison to the co-culture model of only T84/HDFib (≈320 Ω × cm2) after 12 days (T84/HDFib/HUVECs vs T84/HDFib; p-value: 0.0018; two-way ANOVA followed by Tukey’s multiple comparison test), which might be due to the presence of the endothelial monolayer on the basal side (Figure 4(a)). The permeability of the model was also tested on day 12 using the FITC-dextran permeability assay. The results indicated that the triple co-culture model is less permeable (permeability ≈ 0.5%) than the co-culture model (permeability ≈ 1%) (Figure 4(b)). These values did not significantly change after 14 days of culture indicating the maximal epithelial tightness after day 12 of culture.

Functional analysis of the triple co-culture model (T84/HDFib/HUVECs): (a) (top panel) TEER measurement of HDFib, co-culture (T84/HDFib) and triple co-culture (T84/HDFib/HUVECs) 3D tissue models on the SIS scaffold on days 0, 7, 12, and 14 of cell culturing. * shows the statistical significance in TEER between co-culture (T84/HDFib) and triple co-culture (T84/HDFib/HUVECs) at days 12 and 14. **<0.002, ***<0.001. (bottom panel) Statistical analysis of the TEER values (n = 5) were performed by two-way ANOVA followed by Tukey’s multiple comparison test and (b) barrier permeability assessment using FITC-dextran (4 kDa) of HDFib, co-culture (T84/HDFib) and triple co-culture (T84/HDFib/HUVECs) model on the SIS scaffold on day 12. Statistical analysis of the permeability values (n = 3) were performed by one-way ANOVA followed by Tukey’s multiple comparison test. The graphs represent the mean values ± SD. ****≤0.0001.
Pathogen-induced neutrophil transmigration
After establishing a stable triple co-culture model, we transferred the tissue to a perfusion-based bioreactor system (Figure 1) to introduce immune cells to the tissue model. Freshly isolated human neutrophils were added to the culture medium in the external chamber of the bioreactor system (Figure 1(b)). Before addition, we controlled the purity of the isolated neutrophils using confocal microscopy (Supplemental Figure S2). On average, we added 200,000 neutrophils per ml of medium. The neutrophils were isolated from the blood of different donors, male and female, and we applied cells from one donor per bioreactor. We ran two to four bioreactors simultaneously, depending on the setup, in several experimental rounds (n > 3). The bioreactor system enabled us to mimic the blood flow by circulating medium over the basal, endothelium-covered side of the model. At the same time, medium was circulated slowly over the infected epithelial side of the model (apical side) to imitate the conditions on the mucosal surfaces. Our previous experiments in the static system identified 2 h as the earliest time point after addition of neutrophils to tissue models at which we could reproducibly isolate bacteria from the fraction of transmigrated neutrophils. We therefore circulated neutrophils for 2 h and then stopped the experiment by taking the tissues out of the bioreactor, fixing and staining them. The immunofluorescence analysis (Figure 5) and FITC-dextran measurements (not shown) of the infected triple co-culture 3D tissue model with N. gonorrhoeae showed that the epithelial and endothelial cell monolayers are still intact. As it is technically challenging to do complete microscopy in the z-direction due to the thickness of the tissue model (≈150 µm), we analyzed the apical and basal sides of the stained tissue using separate models, which were generated under identical conditions. Confocal images demonstrated zonula occludens (ZO)-1 staining of the tight junction (TJ) on the epithelial side (top layer, Figure 5(a)) and CD31 on the endothelial side (basal layer, Figure 5(b)). The cell layers appeared intact, showing that the perfusion did not disturb the integrity of the cells. At the same time, the measurement of IL-8, a cytokine associated with neutrophil signalling, 37 showed that IL-8 quantities increased upon infection, as well as after 2 h of neutrophil circulation (Supplemental Figure S3). Interestingly, despite the relatively short incubation time of 2 h we were already able to successfully demonstrate the transmigration of the neutrophils through the endothelial up to the epithelial layer. Moreover, transmigrated neutrophils associated with gonococci on the epithelial side of the model within this limited time window as shown in confocal imaging (Figure 5(c) and (d); Supplemental Movie 1). The deformation of neutrophils translocating across the endothelial-fibroblast-epithelial layer (Figure 5(c)) corresponds to the description of transmigrating neutrophils in other studies.38,39

Confocal laser scanning microscopy of triple co-culture 3D tissue model infected by N. gonorrhoeae under dynamic conditions: (a) surface rendering of the CLSM image of the triple co-culture tissue model from the top side of the tissue (perspective view). The tissue model was stained with ZO-1 (magenta), anti-N. gonorrhoeae antibody (green), NCF2 (red), and DAPI (blue). The confocal image of the whole 3D tissue model was surface rendered by IMARIS. The scale bar is 5 µm, (b) orthogonal view of the basal side of the tissue model, displaying XY, YZ, and XZ view of the tissue model. The tissue model was stained with CD31 (green), NCF2 (red) and DAPI (blue). The images were analyzed and reconstructed using FIJI. The scale bar is 20 µm. (c–d) higher magnification of the surface rendered image of the tissue model from the top side, showing the deformation of transmigrating neutrophils and co-localization of the red and green signals, indicating the interaction of N. gonorrhoeae with neutrophils. ZO-1 (magenta), NCF2 (red), anti-N. gonorrhoeae antibody (green), and DAPI (blue). The scale bar is 10 µm and 5 µm, respectively. See also Supplemental Movie 1.
N. gonorrhoeae-neutrophil interaction
We previously observed that among different strains and derivatives of N. gonorrhoeae, N927 strongly damaged the epithelial barrier integrity upon prolonged infection of the tissue models. This derivative of the MS11 strain contains porin PorBIA, and the colonies were selected to be both devoid of pilus (Pili-) and to not express opacity associated proteins (Opa-), to limit the initial interaction of gonococci with the tissue only to PorBIA-mediated adhesion. 40 N927 was shown to induce cytotoxicity and inflammatory cytokine release in a co-culture model of T84 and HDFibs. The initially Opa- N927 gonococci were also shown to revert to Opa+ phenotype during tissue model infection, which would enable these bacteria to interact with neutrophils. 31 A variant of N927 expressing green fluorescent protein (N927-GFP) was used for the experiments to simplify the detection of the infected neutrophils (the GFP signal) using flow cytometry.
The flow cytometry analysis was applied to both assess the percentage of infected neutrophils and analyze their surface receptors, as well as to sort the population associating with GFP-expressing N927 N. gonorrhoeae. The latter was shown to increase after 2 h of circulation, indicating that the neutrophils successfully associated with the bacteria present on the other side of the tissue (Figure 6(a)). We also measured the neutrophil activation (expression of CD11b) of only neutrophils (control), neutrophils in the non-infected, and infected tissue. The results showed that up to 90% of neutrophils were activated in the presence of N. gonorrhoeae (CD11b positive), whereas the circulation alone did not affect neutrophil activation (≈20% for both control and non-infected) (Figure 6(b)).
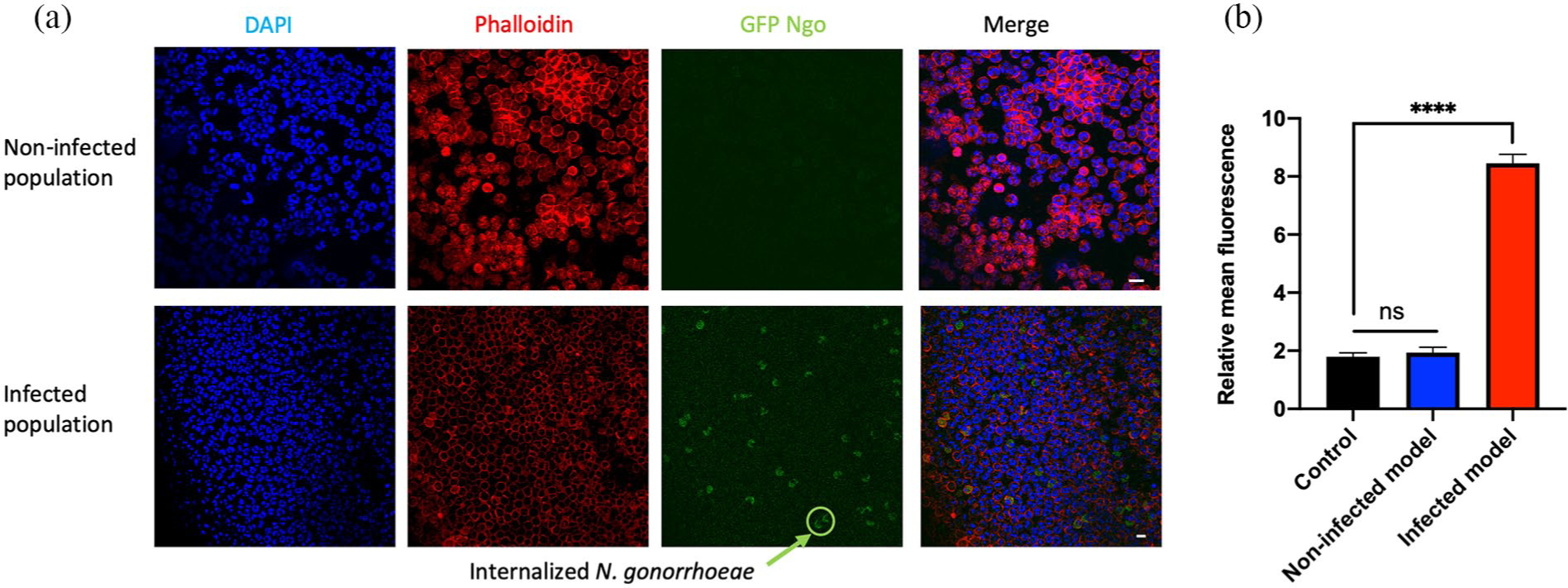
Neutrophil characterization upon infection study: (a) immunofluorescence analysis of FACS-sorted neutrophils. FACS-sorted neutrophils were stained with Phalloidin (red) and DAPI (blue) and were then imaged using confocal fluorescence microscopy. The green signal is GFP expressed by N. gonorrhoeae. No green signal was detected in the non-infected population (top row). Considerable numbers of GFP-positive neutrophils were observed in the infected neutrophils (bottom row, arrow). The scale bar is 25 µm and (b) neutrophil activation was evaluated by the expression of CD11b (in relative mean fluorescence of the signal) for control (neutrophils without circulation), neutrophils circulated through the non-infected tissue model, and neutrophils circulated through an infected tissue model in the perfusion-based bioreactor system after 2 h. The data represent the mean values ± SD from at least three independent replicates.
Discussion
One of the hallmarks of gonorrhea is a robust neutrophil influx to the site of infection in order to promote pathogen clearance. 23 N. gonorrhoeae can survive in the presence of neutrophils and modify the anti-bacterial activities of these immune cells. 3 It has been shown that N. gonorrhoeae may replicate in the phagosome of a sub-population of neutrophils. 17 Such neutrophils harboring bacteria might return to the blood vessels and contribute to the inflammation distribution to some extent. 41
Several non-human species studies have been used to study the gonorrhea disease development such as rabbit, guinea pigs, hamsters, mice, and chimpanzee.42,43 Although animal models provide a general insight into the disease development they are not ideal for studying the invasion and dissemination of N. gonorrhoeae followed by the corresponding immune response.42–44
The classicaly used 2D cell culture relies on cell lines and monoculture systems. However, gonococcal infection does not limit itself to the surface of the tissues, but the bacteria can pass into the deeper tissue layers and even the bloodstream, causing systemic infection. Therefore, a variety of cells beneath the epithelium and in underlying tissue layers such as fibroblasts and endothelial cells are also involved in host-pathogen interaction.45,46 Fibroblasts are responsible for the structure and shape of the connective tissue by producing and secreting the ECM components such as collagen, glycosaminoglycans (GAGs), and proteoglycans and play a key role in immune responses by producing and responding to various cytokines.47,48
Recently, in vitro tissue models opened a new chapter for studying N. gonorrhoeae.24,31 The advancement of tissue engineering and bioengineering offers numerous novel in vitro models of human organs, which can be applied in the field of infectious disease research.49,50
It has been shown that the crosstalk between epithelial and endothelial monolayer has concomitant effects on neutrophil transmigration such as promotion of IL-6 and IL-8 release from epithelial and endothelial cells. 30 However, studies of N. gonorrhoeae interaction with human immune cells have been limited to Transwell inserts 26 that involved epithelial and endothelial cells without accounting for the connective tissue. We therefore aimed to improve the already established epithelial and fibroblast co-culture tissue model in our group 31 to mimic functional and morphological features of the site of gonococcal infection in the human body. To replicate the process of neutrophil extravasation and movement to the site of infection, we enhanced the complexity of the co-culture tissue model by introducing endothelial HUVEC cells on the basal side of the SIS scaffold. In the triple co-culture model of T84/HDFib/HUVECs on the SIS scaffold, a polarized monolayer of epithelial cells formed on the apical side, supported by fibroblast in the middle layer and in addition, a flat monolayer of endothelial cells was present on the basal side of the tissue model (Figures 2 and 3). To ensure the adherence and proliferation of endothelial cells, we coated the basal side of the SIS scaffold with fibronectin, an ECM protein that enhances binding sites of cells to substrate. 51 Many cell types respond to mechanical forces such as shear stress that is induced by blood circulation in the body. Shear stress is playing a crucial role in the cell proliferation and function of the endothelial cells. 52 We therefore grew the models upon addition of the endothelial cells under shear stress provided by an orbital shaker to support the endothelial cell proliferation. 33 The endothelial cells improved the barrier function (higher TEER value and lower permeability) compared to the co-culture model (Figure 4), which agrees with other established tissue models showing that endothelial cells increased TEER value in co-culture model with lung epithelial cell line NCI-H441 53 and human alveolar epithelial type II cells. 54
For circulation of neutrophils, a perfusion-based bioreactor system was utilized to circulate the culture medium through the tissue model, mimicking the blood flow. This dynamic cell culture model therefore resembles the situation that the neutrophils in the blood vessel encounter when recruited to the site of infection. Gradients of pro-inflammatory cytokines produced in response to N. gonorrhoeae infection cause recruitment and transmigration of neutrophils to the site of infection. 3 The infection of the tissue models with N927-GFP was first allowed to proceed for 24 h under static condition prior to the transfer of models to a bioreactor and neutrophil circulation, to ensure the increase in the production of IL-6, TNFα, and IL-8, as we have observed in our previous study 31 and were able to show for IL-8 in this work (Supplemental Figure S3).
CD11b is a receptor on the neutrophil surface crucial for neutrophil migration to the site of infection. The overexpression of CD11b occurs because of the inflammatory response (e.g. IL-8 production) to the infection.6,55 In particular the expression level of CD11b plays a major role in the transepithelial migration of neutrophils through the T84 monolayer5,8 and transendothelial crawling through vascular endothelial cells. 7 Therefore, after 2 h of circulation, we analyzed the expression of CD11b, as a marker for activation of neutrophils and transendothelial migration upon infection. 56 The results showed that the majority of neutrophils (⩾90%) were activated (CD11b+), while the activation level of neutrophils was less than 20% in the absence of infection (Figure 6). This is in agreement with another study that showed that the neutrophils that migrated across the alveolar epithelial cell layer (A549 cells) had a higher CD11b expression level compared to the neutrophils that did not. 57 Flow cytometry analysis showed an increase in the GFP signal in a small part of neutrophil population (ca. 5%) that circulated over the infected tissue model (Figure 6). The presence of a highly GFP-positive subpopulation of neutrophils among the majority of uninfected cells reflects the main characteristic of urethral exudate from gonorrhea patients. 58 The confocal microscopy analysis demonstrated the elongated morphology of neutrophils present throughout the tissue model (Figure 5(c)), which has an important role both in the transmigration across the endothelium and phagocytic activity.59,60 However, association of neutrophils with bacteria, demonstrated by the co-localization of the signals, is mainly observed on the surface of the tissue model (Figure 5(d)), indicating that N. gonorrhoeae can recruit the neutrophils from the basal compartment (endothelial layer) to the apical side (epithelial layer) of the tissue model. This is similar to neutrophil migration across the T84 single epithelial cell layer in response to a chemoattractant observed by Chu and colleagues. 25
Neutrophil-bacteria interaction in gonorrhea is not well understood due to the absence of suitable in vitro models for studying infection under in vivo-like conditions. 22 Culturing neutrophils on the collagen scaffold allowed them to adhere and release granule content, which therefore provided a suitable in vitro model for studying the role of virulence factors of N. gonorrhoeae in infection in the presence of neutrophils.17,58 Nevertheless, establishing more complex in vitro tissue models could help us to understand the interactions between host cells and bacteria. To our knowledge, this is the first report on establishing a 3D tissue model including triple co-culture of epithelial, fibroblast and endothelial cells to study neisserial infection and immune response under more physiological conditions. We used T84 cells as model epithelial cells that attain high polarisation and formation of tight junctions. Considering that gonococci infect urogenital tract mucosal surfaces, it would be possible to use other cell lines of endometrium, cervix or urothelium, or to resort to primary cells as a way of advancing our model.
The tissue model established in this study is nevertheless superior to the previous models, which mainly used Transwell® inserts and culture dishes. Our triple co-culture tissue model including neutrophils allows us to observe features of gonococcal infection reported by other groups, but additionally offers an increased level of complexity, mimicking closely human tissues at the site of infection. Furthermore, the established 3D tissue model in this study enables us to study the role of neutrophil reverse migration, which is not possible under static culture conditions. Since the contribution of neutrophil reverse migration to the dissemination of pathogen is not clear, the introduced tissue model offers a promising tool with a great potential for studying N. gonorrhoeae interaction with the cells of the innate immune system.
One of the obvious limitations of the model we introduce is the usage of a cancer cell line such as T84, which does not fully recapitulate the mucosal surface of cervix or endometrium. This can be improved either by using immortalized cell lines or, even better, isolated human primary cells. However, despite the fact that primary human cells enable better mimicking of the epithelial surface of the infection site, their application has its limitations as well due to their low proliferation rate, cross-contamination during cell isolation, and heterogenicity.22,61 Therefore, the establishment of primary cell-based models remains an effort for the future. In addition, one other future direction of this work could be a detailed study of the cellular and molecular basis for pathogen-epithelial interaction. This could be achieved by combining complex tissue models such as the one we present here with different technologies, including transcriptome and single and dual RNA-seq analysis. Moreover, using CRISPR/Cas9 technology we will be able to modify and study the function of specific genes in the infection process.
Conclusion
Overall the results of the present study show an establishment of a novel biomimetic triple co-culture tissue model that can be employed to further comprehend the cross-talk between epithelial cells and immune cells during the gonorrhea infection. Our proposed 3D tissue model could be an alternative to animal experiments, which would not only help to reduce the number of animals testing, but also represent a better alternative for human-specific pathogens.
Supplemental Material
sj-docx-1-tej-10.1177_2041731420988802 – Supplemental material for Triple co-culture and perfusion bioreactor for studying the interaction between Neisseria gonorrhoeae and neutrophils: A novel 3D tissue model for bacterial infection and immunity
Supplemental material, sj-docx-1-tej-10.1177_2041731420988802 for Triple co-culture and perfusion bioreactor for studying the interaction between Neisseria gonorrhoeae and neutrophils: A novel 3D tissue model for bacterial infection and immunity by Motaharehsadat Heydarian, Matthias Schweinlin, Thomas Schwarz, Ravisha Rawal, Heike Walles, Marco Metzger, Thomas Rudel and Vera Kozjak-Pavlovic in Journal of Tissue Engineering
Footnotes
Acknowledgements
The authors thank the blood donors for their significant contribution to the study by donating the blood for the neutrophil isolation. Artworks in Figures 1 and 3 were reproduced and modified from the open source public database Servier Medical Art (![]() ), under a Creative Commons Attribution 3.0 Unported License.
), under a Creative Commons Attribution 3.0 Unported License.
Author contributions
MH and RR performed the experiments, TS, HW, and MM provided essential material, MH, MS, HW, MM, TR, and VK-P designed experiments and analyzed and interpreted the data, MH and VK-P prepared the figures and wrote the manuscript, TR and VK-P provided necessary funding. All authors have read and approved the submitted manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Deutsche Forschungsgemeinschaft (DFG) in the Research and Training Group 2157 “3D Infect” to VK-P and TR. This publication was funded by the German Research Foundation (DFG) and the University of Wuerzburg in the funding program Open Access Publishing.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
