Abstract
Background:
To evaluate the potential clinical value of cerebrospinal fluid (CSF) circulating tumor DNA (ctDNA) in the diagnosis and monitors the central nervous system (CNS) lymphomas.
Methods:
This was a prospective study of 17 consecutive patients with B-cell lymphoma: 10 patients with CNS lymphomas and 7 patients with B-cell lymphomas at high clinical risk of CNS relapse. Genomic profiles were performed on the CSF and plasma samples of patients by next-generation sequencing.
Results:
In patients with CNS lymphomas, ctDNA was detected in 70.0% of CSF and 60.0% of plasma. The detection rate and gene mutation abundance of CSF were higher than plasma (p = 0.016). CSF had a unique genetic profile. Furthermore, we newly found that gene mutations consistent with plasma or lymphoma-related were also detected in the CSF of the high-risk group without CNS involvement. Analysis of paired plasma and CSF samples from three patients at different time points, changes of CSF ctDNA abundance occurred at the same time or earlier than clinical disease changes, which could timely monitor the therapeutic response and relapse trend.
Conclusion:
The detection rate of ctDNA in CSF is higher than that in plasma. The dynamic monitoring of ctDNA in CSF has hint significance for therapeutic response of CNS lymphoma patients.
Keywords
Introduction
Central nervous system (CNS) involvement remains to be the difficulty in the treatment of B-cell lymphomas. It occurs in about 2%–6% of patients with diffuse large B-cell lymphoma (DLBCL) and mantle cell lymphoma (MCL) or high-grade B-cell lymphoma. 1 CNS lymphoma includes primary and secondary subtypes. In contrast to primary CNS lymphoma (PCNSL), secondary CNS lymphoma (SCNSL) is caused by a lymphoma that originated elsewhere and metastasized to the CNS. Lymphoma cells can enter CNS through hematogenous, neuroinvasion, or direct invasion of adjacent structures. 2 PCNSL usually arises in the brain parenchyma, while approximately two-thirds of the SCNSL patients present with leptomeningeal spread and one-third, with parenchymal disease. CNS lymphoma is a fatal disease with a poor prognosis, although it is a curable brain tumor. 3 Therefore, early recognition of CNS involvement and high-risk groups, followed by timely intervention is key to improving patient outcomes. Unfortunately, the predictive capabilities of current clinical prognostic models are still limited, such as the CNS-International Prognostic Index (IPI), as almost half of CNS involvement occurs in medium- or low-risk groups. 4 The diagnosis of CNS lymphomas is typically based on MRI, brain tissue biopsies/resections, and cytopathological examination of the cerebrospinal fluid (CSF). 5 However, MRI and CSF cytopathological examination have poor specificity and sensitivity, respectively, and brain biopsies are invasive. Thus, there is a pressing need to find less-invasive and more reliable methods for diagnosing and monitoring patients with CNS lymphomas.
A promising innovative approach for analyzing circulating tumor cells or circulating nucleic acids in biofluids is known as liquid biopsy. In B-cell lymphomas, circulating tumor DNA (ctDNA) in the blood plasma or CSF is one of the most extensively studied analytes. 6 ctDNA consists of short DNA fragments released by tumor cells and is theorized to reflect the complete spectrum of genomic alterations present in the tumor. 7 Improvements in polymerase chain reaction (PCR) methods and next-generation sequencing (NGS) technologies based on the detection of tumor genomic aberrations in ctDNA provide new methods for early detection of minimal residual disease during or after therapy, allowing for early prediction of disease relapse and timely therapeutic intervention. 8
In CNS lymphomas, CSF is in close contact with the tumor cells. Only a limited number of current studies are available of the CSF ctDNA. In this study, we sought to determine whether analysis of CSF ctDNA can be used to detect individual mutations in CNS lymphoma patients and evaluate therapeutic response.
Materials and methods
Patients
A total of 17 patients with B-cell lymphomas who were diagnosed and treated in the Hematology Department at Fujian Medical University Union Hospital were included in the study. The histologic subtype of each lymphomas sample was determined according to World Health Organization criteria. All clinical data were obtained by consulting medical records and contacting patients by telephone.
Patients were recruited into two groups: The first group included patients (n = 10) with CNS lymphoma of the CNS by an aggressive lymphoma, from whom CSF was collected prospectively at the time of diagnostic lumbar puncture; The second group included patients (n = 7) without CNS disease but risk factors for CNS relapse. Risk factors for CNS relapse defined according to the National Comprehensive Cancer Network (NCCN) guidelines. Prophylactic intrathecal methotrexate was administered every 3 weeks together with systemic chemotherapy in patients with risk factors for CNS relapse. All patients in this group underwent a lumbar puncture before administration of prophylactic intrathecal methotrexate. Evaluation of response was assessed using imaging examination at the end of treatment and/or at suspicion of disease progression.
DNA extraction, library preparation, and target sequencing
All CSF samples were obtained through lumbar puncture with approximately 10 mL CSF collected for NGS testing, and 8 mL plasma was collected for NGS simultaneously. As previously described, 9 a standardized approach was used to extract cell-free DNA (cfDNA) from CSF and plasma. Within 4 h of collection, the cellular fraction was removed by two-step centrifugation at 4°C including 1900g for 10 min, then 16,000 g for 10 min. Supernatants were removed, and 3.5 mL aliquots were stored in cryotubes at −80°C until analysis. Plasma samples were first separated with blood cell sediment after centrifuging whole blood samples at 4°C for 10 min 1600g, then recentrifuged at 4°C 16,000g for 10 min. Plasma was then aliquoted and stored at −80°C until use.
Both CSF and plasma-derived cfDNA and genomic DNA from peripheral blood lymphocytes was extracted with the Magnetic Serum/Plasma DNA Maxi Kit (DP710, TIANGEN Biotechnology, Beijing, China). DNA was quantified using the Qubit dsDNA HS Assay Kit (Invitrogen, THERMO Fisher Scientific, USA) and sheared into 150–200 bp fragments was subjected to library construction in prior to fragment length analysis using Qsep100 (BiOptic, New Taipei City, China). Fragmented DNA libraries were constructed using nano DNA library prep kit (for Illumina®) (Nanodigmbio, Nanjing, China) for end-repairing, adding A-tailing, and adapter ligation. DNA libraries were then subjected to PCR amplification and purification. The fragment quantification was performed using Qsep100 automatic nucleic acid analysis system (BiOptic, New Taipei City, China). Subsequent, the libraries were captured hybrid with a targeted sequencing gene panel, including 127 genes that are recurrently mutated in lymphoma (Lymcare®, Oriomics Biotech Inc., Hangzhou, China) using xGen® Hybridization and Wash Kit (IDT). The barcoded duplex adapters were eventually blocked by NadPrep NanoBlockers (for illumina; Nanodigmbio, Nanjing, China). The libraries were sequenced on a NovaSeq 6000 platform (Illumina) with a mean coverage depth of about 9000X for cfDNA samples.
Bioinformatics analysis
Sequencing data were processed as previously described. 9 In brief, the data were first demultiplexed and subjected to FASTQ file quality control to remove low-quality data or N bases. All clean paired-end reads were aligned to the human reference genome (19) using Burrows–Wheeler Aligner. Variant calling was performed using Mutect2 from Genome Analysis Toolkit (GATK 3.4.0, https://software.broadinstitute.org/gatk/; Broad Institute, Cambridge, MA, USA). Then, the detected mutations were further selected following established criteria: somatic mutations with allelic fractions of 1% and/or supported by two reads were disregarded. EXAC_EAS or genomAD_EAS should be less than 0.05. DNAs from leukocytes of the same patients were sequenced as germline controls.
CNVkit was selected to identify copy number variations (CNVs). The cutoff of log2 ratio was set at ±0.6 for copy number changes (corresponding to 1.5-fold copy number gain and 0.65-fold copy number loss).
Statistical analysis
Statistical analyses were conducted using R package (version 3.61) (The R Foundation for Statistical Computing, Vienna, Austria). Two-sided p value <0.05 was considered to represent a statistically significant difference.
Results
Baseline characteristics of patients
Table 1 displays the baseline clinical characteristics of the two groups of patients. The median age of all patients was 46 years (range, 22–69 years), and the male to female ratio was 12:5. Histologically, 15 (88.2%) patients were diagnosed with DLBCL, one (5.9%) patients with MCL, and one (5.9%) with High-grade B-cell lymphoma (HGBL). Twelve (70.6%) of 17 patients had stage IV lymphoma, and 4 patients (23.5%) had IPI scores of 3–5. The majority of patients received the R-COEP regimen as first-line therapy. Of the 17 patients, 11 had complete/partial response (CR/PR), 1 had stable disease (SD), and 5 had progressive disease.
Characteristics of patients at the time of enrolment and outcome.

CNS, central nervous system; COO, cell of origin; CR, complete response; CSF, cerebrospinal fluid; DLBCL, diffuse large B-cell lymphoma; F, female; GCB, germinal center B-cell-like; HGBCL, high-grade B-cell lymphoma; Histol, histology; ID, patient identification number; inv, involvement; IPI, International Prognosis Index; M, male; MCL, mantle cell lymphoma; N, no; NA, not available; PD, progressive disease; PR, partial response; Y, yes.
When comparing the two groups, CNS lymphoma group had seven males and three females with a mean age of 50.4 years, and high-risk CNS involvement lymphoma group had five males and two females with a mean age of 47.4 years. The IPI score in the CNS lymphoma group was significantly higher than in the high-risk CNS involvement lymphoma group. In CNS lymphoma group presented 7 out of 10 patients involved only brain parenchyma. Median cfDNA copy count per milliliter of acellular CSF was 0.143 (range, 0.067–0.234) for parenchymal-only disease and 0.613 (range, 0.346–16.1) for leptomeningeal disease. Table 2 shows high-risk characteristics of high-risk CNS involvement lymphoma patients.
High-risk characteristics of high-risk CNS involvement lymphoma patients.
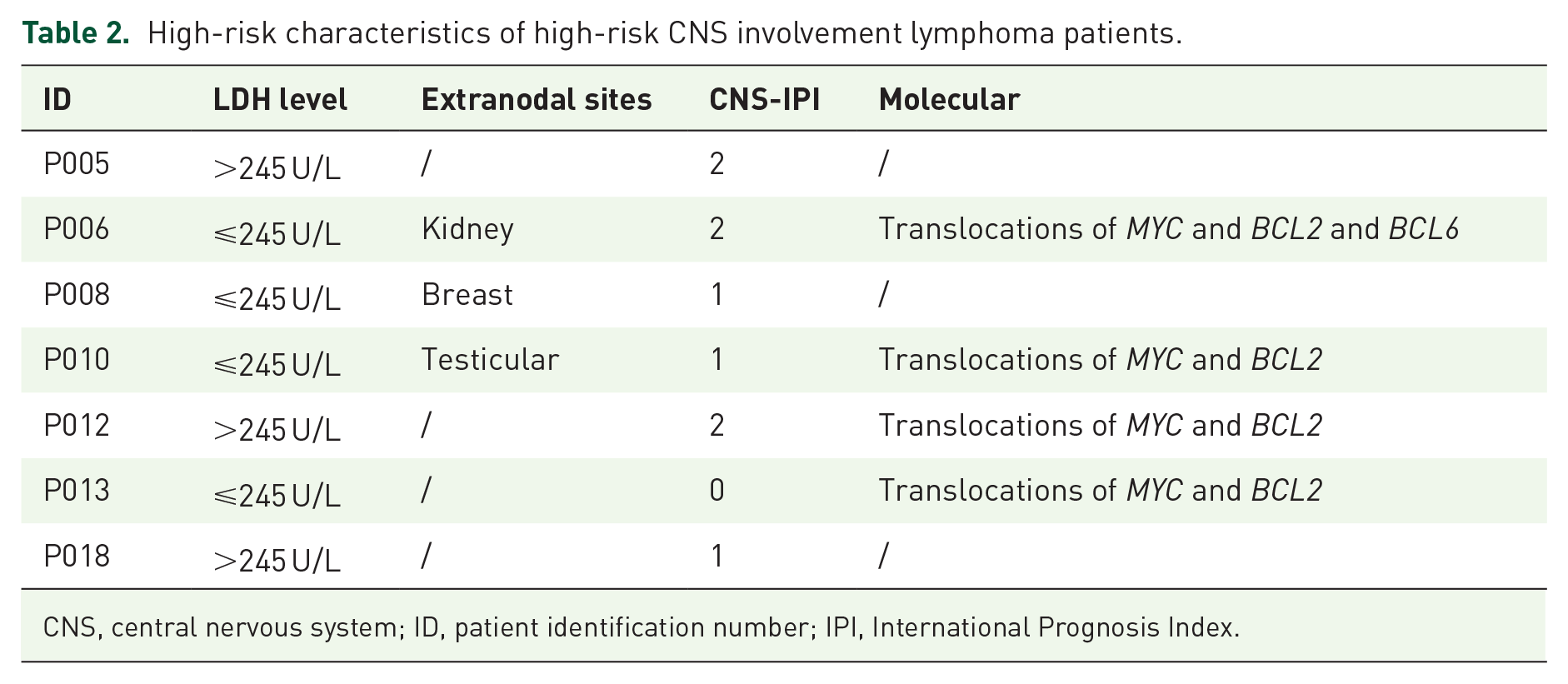
CNS, central nervous system; ID, patient identification number; IPI, International Prognosis Index.
CSF is superior in identifying gene mutations of CNS lymphoma than plasma
Here, we first sought to analyze the presence of ctDNA in the CSF and plasma from 10 patients with CNS disease at the time of enrollment. The results showed that ctDNA was detected in 70.0% (7/10) of CSF and 60.0% (6/10) of plasma. The detection rate of ctDNA in the CSF samples were higher than the paired plasma samples. In CSF, PIM1 (30%), MYD88 (30%), KMT2D (20%), and CD79B (20%) were the most common mutated genes, whereas in plasma, PIM1 (30%), MYD88 (30%), and CD79B (20%) were the most common mutated genes (Figure 1). A total of 46 genetic mutations were detected in CSF, while only 37 genetic mutations were detected in plasma (Figure 2). The gene mutation abundance of CSF were significantly higher than plasma (p = 0.016; Figures 3 and 4). Moreover, we identified CSF ctDNA had 35 unique gene mutations, which mainly included BIRC3, KMT2D, PIM1, and TSC2, among others. A full list of unique mutations and gene functions was shown in Supplemental Table 1. The most frequent mutation in 11 shared mutations is the MYD88 X160R mutation.

Genetic profiles in the spectra of plasma and CSF in 10 CNS lymphoma patients.

Concordance of mutations in CSF and plasma.

Distribution of allele fraction values for 11 shared mutations.

Comparison of allele fraction between CSF and plasma.
CSF ctDNA detection might indicate the CNS involvement with high-risk lymphomas
The majority of CNS relapses occurred within 1 year of initial remission in patients with a higher risk of CNS relapse. We then analyzed seven patients with systemic lymphoma without CNS involvement but with high risk for CNS relapse to see if CSF ctDNA could detect the signal of CNS disease. Much to our surprise, mutations in CSF ctDNA were detected in six of seven cases. KMT2D (28.6%) and TET2 (28.6%) were the most frequently mutated genes (Figure 5). Notably, in one case (P012), we detected MYD88 V172F mutation in both CSF and plasma. All the mutations detected in the CSF ctDNA were showed in the Table 3. These results indicate that ctDNA analysis of the CSF can improve the detection CNS involvement by conventional techniques.

Genetic profiles in the spectra of plasma and CSF in seven high-risk CNS involvement lymphoma patients.
Gene mutations identified in CSF of patients with high-risk CNS involvement lymphoma.

AF, allele fractions; CNS, central nervous system; CSF, cerebrospinal fluid; ID, patient identification number.
CSF ctDNA can be used to assess therapeutic response
To determine whether the CSF ctDNA could be a useful tool for evaluating the therapy response, we simultaneously analyzed sequential samples of CSF and plasma. Both blood and CSF samples were collected before starting treatment, or when progression or relapse of the disease was noticed. Tumor burden was evaluated by imaging examination.
Two individuals in our cohort (P004 and P012), we observed the CSF ctDNA levels decreased with response to treatment and undetectable after complete remission. In P004, following a disease-free interval of 11 months, the patient relapsed and ctDNA reappeared in CSF but not in plasma (Figures 6 and 7). The P012 showed recurrence of the lesion after a 4-month interval, ctDNA was detected in both plasma and CSF (Figures 8 and 9). These results are in line with what the imaging findings.

Dynamic monitoring results of patient P004. The patient’s condition was effectively controlled after taking zanubrutinib plus R-CHOP, and gene mutation were undetectable in the CSF and plasma at the second review; while at the third review, the patient’s clinical symptoms worsened and the ctDNA abundance increased, whereas it remained undetectable in plasma.

MRI image of patient P004. The arrow indicates the lesions observed before the treatment on February 3, 2021. A significant reduction in lesion size was observed on April 18, 2021. The arrow indicates that there is no abnormal enhancement in the area on June 3, 2021. Nodular callosal enhancement appears on May 27, 2022. On May 4, 2023, the range of the nodular callosal enhancement was larger than before.
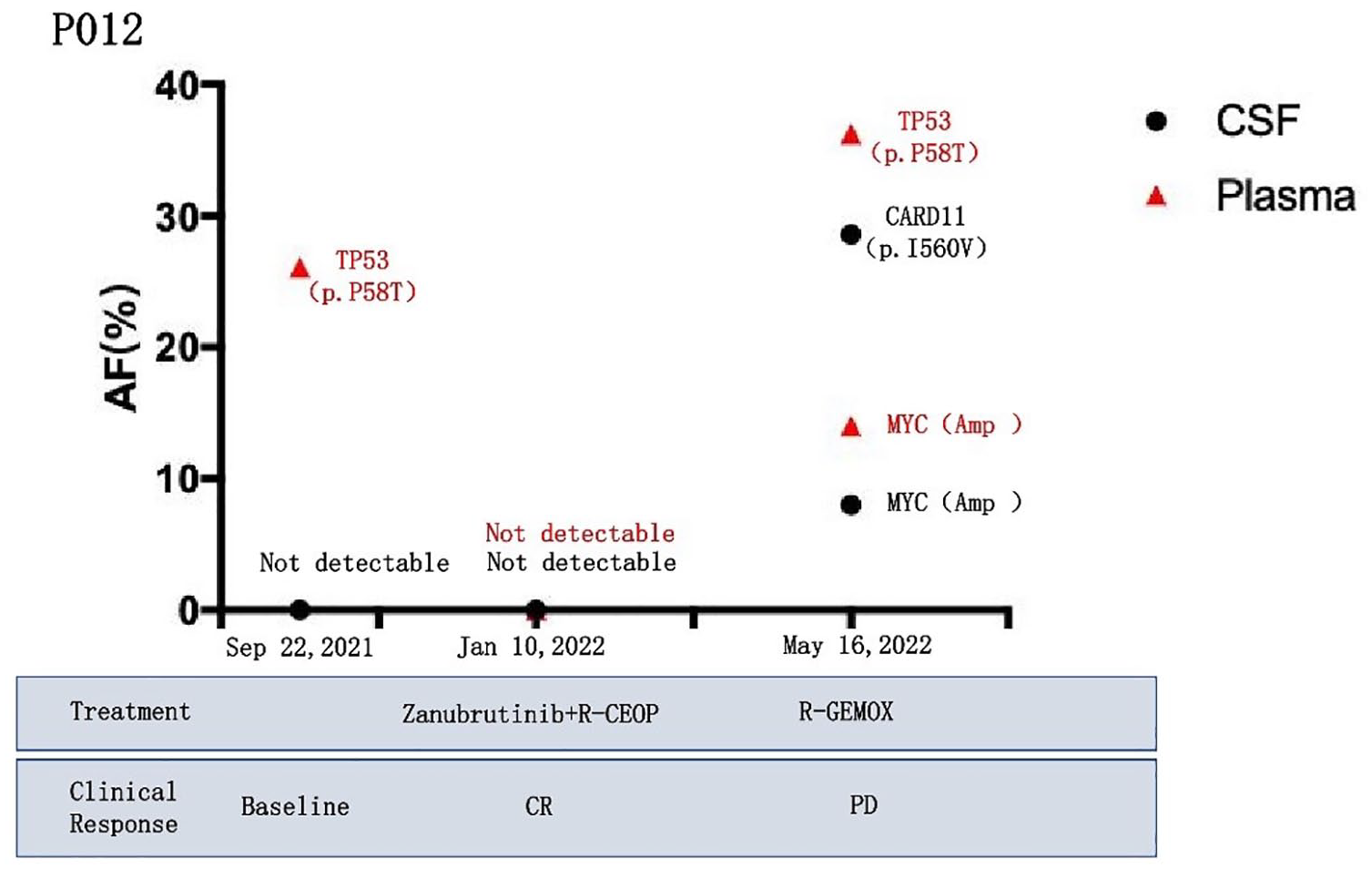

Dynamic monitoring results of patient P012. The patient’s condition was effectively controlled after taking zanubrutinib plus R-CHOP, and gene mutation were undetectable in the CSF and plasma at the second review; while at the fourth review, the patient’s clinical symptoms worsened and the CSF and plasma ctDNA abundance increased.

PET/CT image of patient P012. The arrow indicates hypermetabolic lymphadenopathy on September 22, 2021. The lesions disappeared after treatment with zanubrutinib plus R-CHOP on June 3, 2021. The arrow indicates that the bilateral cervical had new multiple hypermetabolism foci on May 27, 2022.
Moreover, we also observed that the increase in the CSF ctDNA level can precede the detection of relapses by imaging evaluation and clinical manifestations. Another case of high-risk CNS involvement lymphoma (P013) treatment with orelabrutinib + R-CEOP/R-GEMOX achieved a CR, although the CSF ctDNA levels remained elevated. The P013 patient’s CSF GNA13 gene mutation abundance increased to 6.76% at the third review, and the imaging examination and clinical manifestations results suggested disease recurrence (Figures 10 and 11). This finding may indicate that rigorous clinical follow-up should be done to monitor disease progression and recurrence in such patients.

Dynamic monitoring results of patient P013. The patient’s condition was effectively controlled after taking orelabrutinib plus R-CHOP, and gene mutation were undetectable in the CSF at the second review; while at the third review, the patient’s clinical symptoms worsened and the ctDNA abundance increased, whereas it remained undetectable in plasma.

PET/CT image of patient P013. The arrow indicates hypermetabolic lymphadenopathy on May 18, 2021. The lesions disappeared after treatment with orelabrutinib plus R-CHOP on April 19, 2022. The arrow indicates that the bilateral cervical had new multiple hypermetabolism foci on September 21, 2022.
Discussion
B-cell lymphoma can infiltrate the CNS with a poor prognosis, especially at relapse. 10 Diagnosing and monitoring can be challenging in CNS lymphoma. The presence of the MYD88 L265P mutation is a strong indicator of CNS lymphoma, a perspective supported by numerous prior studies.11–13 In CSF ctDNA from patients with CNS lymphoma, the MYD88 L265P mutation was found, demonstrating that the CSF ctDNA may be employed as a complementary diagnostic tool. ctDNA was detectable in almost all CSF specimens in this study and demonstrated that CSF ctDNA has the potential to improve both current diagnostics and therapeutic response monitoring in CNS lymphoma.
In terms of CNS lymphoma, tumor cells release ctDNA fragments shed into the CSF directly. The blood–brain barrier could prevent ctDNA from entering the circulation, resulting in the low detectability of ctDNA in plasma. 14 Moreover, a significant quantity of free DNA from normal tissue exists in peripheral blood, and ctDNA accounts for a relatively small proportion of the test, which can be easily interfered. 15 Previous research has indicated that CSF ctDNA can more accurately reflect the genomic alterations associated with intracranial lesions in patients with CNS tumors, thereby aiding in the guidance of clinical targeted therapies. 16 In our study, we observed that the detection rate and gene mutation abundance of CSF were higher than plasma. Although the sample size of this study was small, this trend was consistent with the results of previous studies.17,18 Therefore, CSF sampling is a more invasive method of fluid biopsy than a simple blood test but is still necessary to detect CNS lymphoma by CSF ctDNA.
In our research, lymphoma-related gene mutations could be detected by CSF in the high-risk group without CNS involvement in imaging. This finding suggests that CSF ctDNA testing may be able to identify molecular level changes earlier than imaging in the high-risk group. According to three separate investigations, serial ctDNA monitoring of DLBCL patients in CR helps the diagnosis of lymphoma relapse in the vast majority of cases, with a ~3–6 months lead time prior to radiographic imaging.19,20 Imaging studies are often noninformative or slow to reflect progression and provide no genetic information. 21 Beyond that, a previous study has confirmed that cancer containing about 50 million malignant releases sufficient DNA for detection in the circulation, whereas cancer of that size is far below that required for definitive imaging. 22 A better sensitivity of CSF ctDNA (100%) than imaging examination (62.86%) was also demonstrated by Zhao et al. 7 Thus, CSF ctDNA as liquid biopsy technology may serve as a supplement to diagnose CNS relapse with negative imaging.
Monitoring the response to therapeutic treatments remains a challenging question in CNS lymphoma. Tumor biomarkers and imaging are widely used to assess the effectiveness of chemotherapy treatment. However, these methods may not be very accurate in evaluating therapeutic responses and do not provide information about genetic alterations in tumors. 15 Six patients with brain tumors had their CSF-ctDNA examined in studies by De Mattos-Arruda et al., and they discovered that the mutant allelic frequencies of DNA decreases with surgical resection or response to systemic therapy and increased with the tumor progressed. 17 Similarly, our study found that CSF ctDNA is a promising biomarker. Dynamic detection of gene mutations in CSF may be used to assess the therapeutic response in real-time and guide the choice of the therapeutic regimen.
There are some limitations to our study that are worthy of mention. Because of the challenge of CNS lymphoma patients accessibility, we conducted the tests in a relatively small sample volume analyses only. It will be necessary to validate the current study’s findings in a larger cohort of patients. Moreover, patients participating in these studies had different types of lymphoma and received a variety of different protocols, which in some cases may have eradicated the microscopic source of clonotypic DNA in the CSF. A cohort with uniform histology and treatment regimen is required in the future analysis.
Conclusion
In conclusion, utilizing CSF ctDNA as liquid biopsy technology to detect cancer-associated gene mutations may be useful in monitoring response to treatment and identifying disease relapse. Additionally, our data supported that CSF ctDNA is a promising diagnostic tool for PCNSL and high-grade lymphoma.
Supplemental Material
sj-docx-1-tah-10.1177_20406207251321721 – Supplemental material for Next-generation sequencing of circulating tumor DNA in cerebrospinal fluid for detecting gene mutations in central nervous system lymphoma patients
Supplemental material, sj-docx-1-tah-10.1177_20406207251321721 for Next-generation sequencing of circulating tumor DNA in cerebrospinal fluid for detecting gene mutations in central nervous system lymphoma patients by Haiying Fu, Taiqin Wang, Yong Yang, Chenxi Qiu, He Wang, Yanyan Qiu, Jianzhi Liu and Tingbo Liu in Therapeutic Advances in Hematology
Supplemental Material
sj-docx-2-tah-10.1177_20406207251321721 – Supplemental material for Next-generation sequencing of circulating tumor DNA in cerebrospinal fluid for detecting gene mutations in central nervous system lymphoma patients
Supplemental material, sj-docx-2-tah-10.1177_20406207251321721 for Next-generation sequencing of circulating tumor DNA in cerebrospinal fluid for detecting gene mutations in central nervous system lymphoma patients by Haiying Fu, Taiqin Wang, Yong Yang, Chenxi Qiu, He Wang, Yanyan Qiu, Jianzhi Liu and Tingbo Liu in Therapeutic Advances in Hematology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
