Abstract
Introduction
Maraviroc inhibits CCR5-tropic HIV-1 across different subtypes in vitro and has demonstrated efficacy in clinical trials. V3-loop amino acid variants observed in individual maraviroc-resistant viruses have not been found to be predictive of reduced susceptibility. Sequence-database searches have demonstrated that approximately 7.3% of viruses naturally encode these variants, raising concerns regarding potential pre-existing resistance. A study from Russia reported that combinations of these same amino acids are present in the V3 loops of the Russian variant subtype A (IDU-A, now A6) with a much greater prevalence (range: 74.4%–92.3%) depending on the combination. However, these studies and database searches did not include phenotypic evaluation.
Methods
Sixteen Russian HIV-1 isolates (including sub-subtype A6 viruses) were assessed for V3 loop sequence and phenotypic susceptibility to maraviroc.
Results
All 12 of the A6 viruses and 2/4 subtype B isolates encoded V3-loop variants that have previously been identified in individual virus isolates with reduced susceptibility to maraviroc. However, despite the prevalence of these V3-loop amino acid variants among the tested viruses, phenotypic sensitivity to maraviroc was observed in all instances. Similarly, reduced susceptibility to maraviroc was not found in virus from participants who experienced virologic failure in a clinical study of maraviroc in Russia (A4001101, [NCT01275625]).
Discussion
Altogether, these data confirm that the presence of individual or combinations of V3-loop amino acid residues in sub-subtype A6 viruses alone does not predict natural resistance to maraviroc and that V3-loop genotype analysis of R5 virus prior to treatment is not helpful in predicting clinical outcome.
Introduction
HIV binds to target cells initially through cellular CD4 and then to a cell-membrane coreceptor, CCR5 or CXCR4. Maraviroc, a CCR5 antagonist, inhibits CCR5-tropic HIV-1 variants, by binding to a specific intra-cellular membrane site on CCR5 and preventing the HIV-1 envelope (Env) glycoprotein (gp120) interaction. 1
Resistance to CCR5 antagonists is characterized by viral interaction with CCR5 and entry in the presence of the antagonist (compound-occupied receptor). Maraviroc binds deep within a pocket formed by CCR5 trans-membrane helices. The site is distinct from chemokine and HIV-1 envelope (Env) glycoprotein (gp120) binding sites 1 and is thought to stabilize a receptor conformation that cannot normally be recognized by maraviroc-sensitive HIV-1 isolates. Resistant viruses adapt to bind to the altered conformation of the maraviroc-occupied receptor, although with reduced affinity. As a result, EC50 does not change, because there is no change to the maraviroc occupancy of CCR5 at any given concentration, but resistant virus cannot be fully inhibited, even at high drug concentrations. This is characterized phenotypically by dose-response inhibition curves with plateaus showing the maximal percent inhibition (MPI) at less than 95%. 2
Resistance to CCR5 antagonists has been described in viruses selected in vitro and during treatment of HIV-infected persons.2,3 Although sequence changes spanning the entire env gene have been identified in resistant viruses, attention has focused on changes in the V3-loop region. V3-loop amino acid variants alone have been reported to confer a partial or complete resistance phenotype; however, they are context dependent and phenotypic characteristics are unique to the particular viral envelope. 4 Similarly, in the maraviroc registration trials, V3-loop changes were identified in resistant viruses, although the precise changes varied significantly between participants (Lewis et al, submitted paper). 5,6
Sequence database searches have identified viruses that naturally encode the V3-loop variants present in individual maraviroc-resistant viruses, raising concerns regarding potential preexisting resistance. However, published studies have not included parallel phenotypic analyses. 7 --9 A V3-loop sequence analysis of 323 samples from CCR5 antagonist–naive HIV-infected persons reported by Soulie et al.8 found that 18/235 CCR5-tropic viruses (7.3%) had V3-loop variants previously described as emerging in viruses from individuals treated with maraviroc. Vasilev et al. 9 also reported that combinations of these same amino acids are present in the V3 loops of the characteristic Russian form of subtype A1 (IDU-A or A6) with a much greater prevalence (range: 74.4%–92.3%). Since sub-subtype A6 is prevalent in HIV-infected persons throughout Russia, 10 it was important to determine whether these V3-loop sequences confer maraviroc resistance for the benefit-risk assessment for maraviroc use in Russia. In preparation for a Phase IV, multicenter, open-label clinical study of maraviroc in combination with zidovudine and lamivudine in participants infected with CCR5 tropic HIV-1 in Russia (A4001101 - A Multicenter, Open-label Study of Maraviroc, Zidovudine and Lamivudine Twice Daily for the Treatment of Antiretroviral Naive HIV-Infected Patients with R5 HIV-1 in Russia [NCT01275625]), HIV-1 isolates (including sub-subtype A6 viruses) from HIV-infected people from different geographic locations were assessed for susceptibility to maraviroc. Here we describe the V3 loop sequence and phenotypic assessment of maraviroc susceptibility in 16 HIV-1 isolates from maraviroc-naive participants. We also describe the virologic data for the 8 participants with virologic failure and sufficient HIV-1 RNA for analysis (500 c/mL) after 48 weeks in study A4001101.
Methods
HIV-1 env genes were amplified from stored plasma samples obtained from 16 maraviroc-naive persons attending 11 different regional AIDS centers across Russia. Population sequencing of the V3 loops was performed by Monogram Biosciences Inc. (South San Francisco, CA) and used to identify isolates with variant sequences of interest. All possible amino acid translations were imputed when there was ambiguity in the nucleotide sequence. The FASTA DNA sequences were aligned with reference sequences representative of subtypes A1, Russian A1 (now known as A6), D, and B, and a tree created using CLUSTAL. 11 A diagrammatic representation of the neighbor-joining tree was generated using MEGA (http://www.megasoftware.net/home). The location of the sequences on the branches determined the subtype of the sequence and this was confirmed using the BLAST search tool in the HIV Sequence database. In each case the best matches confirmed the subtype that had been allocated by the tree method. Sequences have been submitted to GenBank (National Institutes of Health, Bethesda, MD, USA) under accession numbers MW728314 to MW728329.
Pseudotyped viruses containing Env sequences derived from these clinical isolates were tested for maraviroc susceptibility in recombinant CD4 and CCR5-expressing cells (PhenoSense Entry, Monogram Biosciences Inc.). Maraviroc resistance was defined as MPI <95%.
For the virology analysis in clinical trial A4001101, all participants who discontinued early or who reached Week 48 with sufficient plasma HIV-1 RNA for analysis (≥500 copies/mL) were included. The Siemens TRUGENE HIV-1 genotyping assay (nucleoside reverse transcriptase inhibitor resistance assessment and subtype determination; Erlangen, Germany), a genotypic tropism assay and the PhenoSense Entry assay (for maraviroc susceptibility) was performed at screening and the last on-treatment time point available.
Study A4001101 was approved by independent ethics committees at study centres, and written informed consent was obtained from all participants. The study was performed in accordance with International Conference on Harmonization Good Clinical Practice guidelines and applicable local regulatory requirements and laws.
Results
Of the 16 viruses included in the survey, 12 were identified as A6 variants, all of which contained either 3 or 4 V3-loop amino acid variants previously proposed by others as being associated with maraviroc resistance (11S, 20F, 25D, and 26V; Table 1). 8 The remaining 4 isolates were classified as subtype B; 2 of these isolates encoded the V3-loop 26V variant, both also encoded 20F, and one of these also included 11S. Despite the presence of these V3-loop amino acid variants in the sub-subtype A6 viruses and 2/4 subtype B viruses, the HIV-1 isolates from all 16 participants were phenotypically sensitive to maraviroc (MPI >95%; Table 1).
Frequency of genotypic variants and their maraviroc susceptibility by individual mutation and viral subtype among the 16 samples of Russian origin.
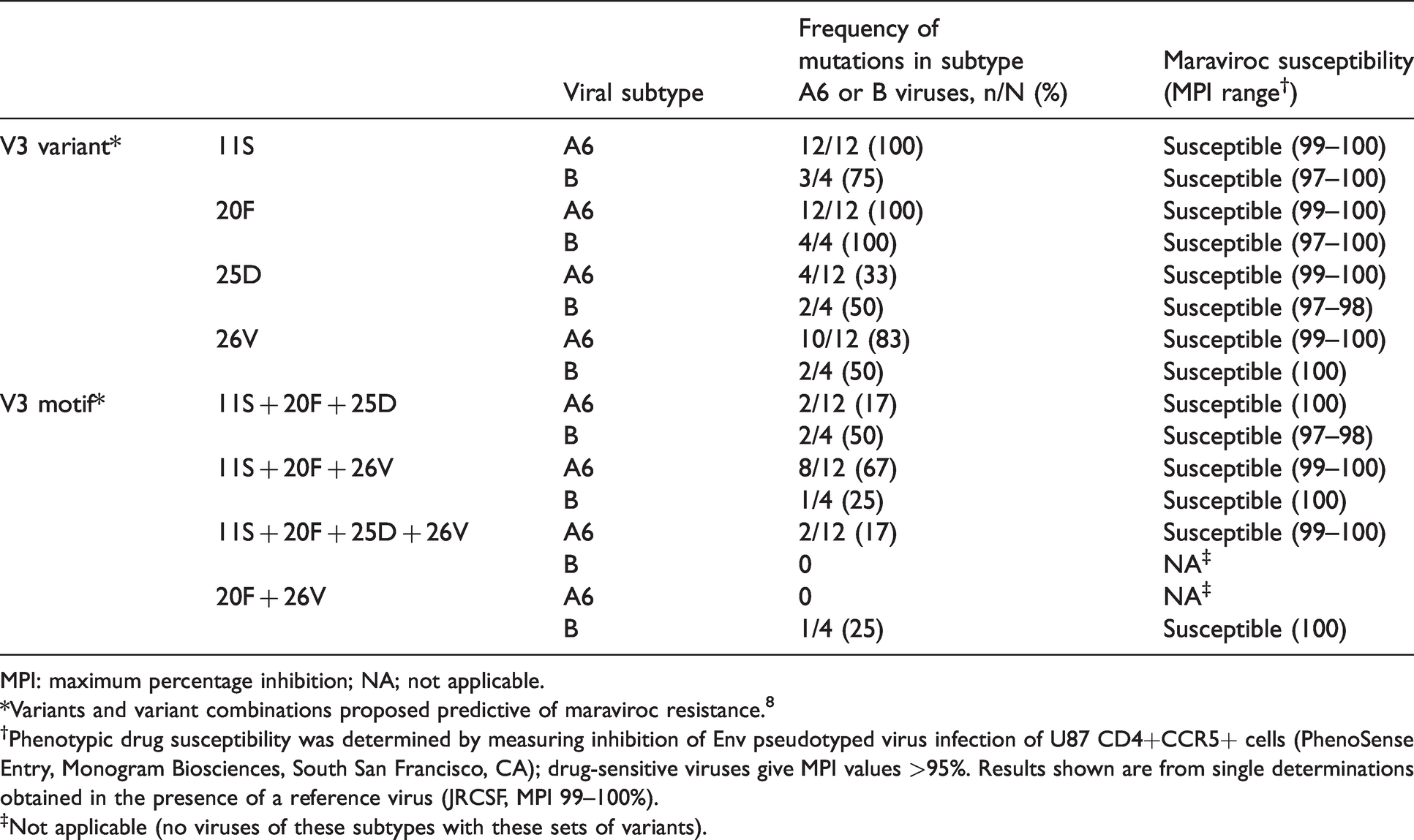
MPI: maximum percentage inhibition; NA; not applicable.
*Variants and variant combinations proposed predictive of maraviroc resistance. 8
†Phenotypic drug susceptibility was determined by measuring inhibition of Env pseudotyped virus infection of U87 CD4+CCR5+ cells (PhenoSense Entry, Monogram Biosciences, South San Francisco, CA); drug-sensitive viruses give MPI values >95%. Results shown are from single determinations obtained in the presence of a reference virus (JRCSF, MPI 99–100%).
‡Not applicable (no viruses of these subtypes with these sets of variants).
In study A4001101, 60/77 participants had HIV-1 RNA <50 copies/mL at Week 48. Eight participants had a valid on treatment virologic analysis (HIV-1 RNA ≥500 copies/mL). Of these, 5 were infected with HIV-1, originally identified as subtype A1, 2 viruses were identified as subtype B virus, and 1 was identified as circulating recombinant form, CRF_01 (AE). No changes in tropism were observed in virus from any participant. Maraviroc susceptibility testing of 5 screening samples (3 subtype A1 and 2 subtype B) and 3 on-treatment samples (1 subtype A1 and 2 subtype B); none showed reduced maraviroc susceptibility.
Discussion
Results from this survey of representative clinical Env sequences found that, despite several viruses harboring combinations of the V3-loop variants 11S, 20F, 25D, and 26V, the plasma viruses from both major viral strains in Russia (sub-subtype A6 and subtype B) retained full maraviroc susceptibility. This finding is consistent with V3-loop variants’ dependence on the broader context of the specific Env amino acid structure for any effect on maraviroc susceptibility. Thus, as described for the maraviroc-resistant viruses from the MOTIVATE trials, 4 variants in the V3 loop that result in maraviroc resistance are likely selected based on the need to preserve several other essential functional elements in Env. This leads to great diversity in combinations of mutations conferring resistance between isolates. Furthermore, the findings are consistent with the clinical response observed in study A4001101, which assessed the efficacy and safety of maraviroc and lamivudine/zidovudine in a treatment-naive Russian population infected with HIV-1. In that study, 60/77 (78%) treatment-naive participants had <50 copies/mL plasma HIV-1 RNA at Week 48. None of the 8 failures with virologic analysis showed a tropism change, and, of 3 with successful MPI determination, none showed reduced susceptibility to maraviroc.
The findings further support the observation that the specific variants (11S, 20F, 25D, and 26V) in the V3 loop of gp120, although observed in individual viral isolates with maraviroc resistance, are not inherently predictive of susceptibility reduction. 4 This was confirmed for 4 viruses with maraviroc resistance when site-directed mutagenesis was used to reverse the V3-loop substitutions in the resistant clones to match the sensitive baseline clones and vice versa; the sensitive and resistant phenotypes were typically not restored.5,12 Studies with other CCR5 antagonists also found variable routes to resistance.3,13 A study of virus resistant to vicriviroc from 5 treatment-experienced participants in the VICTOR_E1 study demonstrated different substitutions in the V3 loop for each virus studied. However, when site-directed mutagenesis was used to reverse the V3-loop substitutions, the sensitive and resistant phenotypes were not restored. 3 This observation suggests that some changes outside the V3-loop region might be required to support a resistant phenotype. Changes were seen across gp160 as well as in the V3 loop in clones, demonstrating reduced susceptibility to aplaviroc. 13 However, as with maraviroc and vicriviroc, a consistent mutational pattern was not revealed, either in the V3 loop substitutions or in the gp160 changes, that could predict reduced susceptibility.
These data confirm that the presence of individual or combinations of V3-loop amino acid residues alone does not predict natural resistance to maraviroc and that V3-loop genotype analysis of R5 virus prior to treatment is not helpful in predicting clinical outcome.
Data accessibility
The data generated and analyzed in this study are available from the corresponding author upon reasonable request.
Footnotes
Acknowledgements
All listed authors meet the criteria for authorship set forth by the International Committee of Medical Journal Editors. We are grateful to Dr Dimitry Kuevda for helping with the provision of samples.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: MEL, CC, and EvdR were employees of Pfizer at the time of the submitted work but are now contracted by The Research Network Ltd. to perform consulting work for Pfizer. CC is also an employee of Research Virology Consulting Ltd and has received contractor fees from ViiV Healthcare. BJ was an employee of Pfizer at the time of the submitted work but is now an employee of Covance Inc. PS was an employee of Pfizer at the time of the submitted work but is now an employee of AstraZeneca. MW was an employee of Pfizer at the time of the submitted work but is now an employee of Centauri Therapeutics Ltd. SLB was an employee of Pfizer at the time of the submitted work but is now an employee of Takeda Pharmaceuticals International. AL, DK, and MB have nothing to disclose.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by ViiV Healthcare. The authors acknowledge the contribution of the Ministry of Science and Higher Education of the Russian Federation (unique project identifier RFMEFI61019X0020). Editorial assistance was provided under the direction of the authors by Sherri Damlo, ELS, MedThink SciCom, and was funded by ViiV Healthcare. The authors have authorized the submission of this manuscript by Rupali Ullal, MedThink SciCom, whose submission assistance was funded by ViiV Healthcare.
