Abstract
Corona Virus Disease 2019 (COVID-19) is a pandemic caused by Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2). Exploiting the potentials of phytocompounds is an integral component of the international response to this pandemic. In this study, a virtual screening through molecular docking analysis was used to screen a total of 226 bioactive compounds from African herbs and medicinal plants for direct interactions with SARS-CoV-2 RNA-dependent RNA polymerase (RdRp). From these, 36 phytocompounds with binding affinities higher than the approved reference drugs (remdesivir and sobosivir), were further docked targeting the active sites of SARS-CoV-2, as well as SARS-CoV and HCV RdRp. A hit list of 7 compounds alongside two positive controls (remdesivir and sofosbuvir) and two negative controls (cinnamaldehyde and Thymoquinone) were further docked into the active site of 8 different conformations of SARS-CoV-2 RdRp gotten from molecular dynamics simulation (MDS) system equilibration. The top docked compounds were further subjected to predictive druglikeness and ADME/tox filtering analyses. Drugable alkaloids (10’–hydroxyusambarensine, cryptospirolepine, strychnopentamine) and flavonoids (usararotenoid A, and 12α-epi-millettosin), were reported to exhibit strong affinity binding and interactions with key amino acid residues in the catalytic site, the divalent-cation–binding site, and the NTP entry channel in the active region of the RdRp enzyme as the positive controls. These phytochemicals, in addition to other promising antivirals such as remdesivir and sofosbuvir, may be exploited towards the development of a cocktail of anti-coronavirus treatments in COVID-19. Experimental studies are recommended to validate these study.
Introduction
The Coronavirus Disease 2019 (COVID-19) is a pandemic caused by the outbreak of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) and has resulted in increasing mortality and socio-economic burden. It is associated with high fever, cough, severe shortness of breathing, nausea, vomiting, and diarrhea in human populations; and has caused + 10 M infections with more than 0.5 M deaths worldwide.1,2 The pathogen, which is a new member of the betacoronavirus genus, is a positive-sense and single-stranded RNA virus similar to SARS-CoV and MERS-CoV (the Middle East Respiratory Syndrome coronavirus). 3 Along with structural proteins (like spike glycoprotein and accessory proteins), these viral genomes encode non-structural proteins, including 3-chymotrypsin-like protease, papain-like protease, helicase and RNA-dependent RNA polymerase (RdRp). 4
At present, no preventive vaccines or established antiviral therapies are available against COVID-19. 5 However, several repurposed drugs such as remdesivir, hydroxychloroquine, and chloroquine phosphate have shown promising results.6,7 Virtual screening, which is associated with search space minimization, economic feasibility, and high versatility, may be particularly valuable for rapidly finding a potent inhibitor of the COVID-19 virus. In the search for anti-coronavirus agents, important druggable targets include 3-chymotrypsin-like protease (3CLpro), papain-like protease (PLpro), RNA-dependent RNA polymerase (RdRp), and spike (S) proteins. 8 The RdRp, a multi-domain protein, is a central component of coronaviral replication/transcription machinery that is able to catalyze RNA-template dependent formation of phosphodiester bonds between ribonucleotides in the presence of divalent metal ion.9–11 The polymerase domain is highly conserved and shares common structural features among coronaviruses, Hepatitis C Virus and other positive sense RNA viruses.12–14 Thus, the viral RdRp or its catalyzed polymerization process has been explored as targets for developing several anti-viral drugs for treating Hepatitis C, Zika and coronaviruses.13,15–17
Exploiting the potentials of phytomedicine, which is perhaps the oldest, and most assorted of all therapeutic systems, is an integral component of the international response to the COVID-19 pandemic, as highlighted by the WHO-China Joint Mission on COVID-19. Phytomedicine has efficacy against COVID-19 as many antiviral plants native to China have shown promising therapeutic potential against SARS-CoV and SARS-CoV-2 16,18,19 African herbs and medicinal plants provide a vibrant resource for chemopreventive phytochemicals as they could accumulate secondary metabolites more than plants from the northern hemisphere.20,21 African plants and phytochemicals with documented antiviral, antimicrobial, antimalarial, antifungal exploited towards the development of prophylactic and therapeutic agents against the COVID-19..22–25 This study focused on virtual screening of phytochemicals derived from African herbs and medicinal plants against the COVID-19 and related virus RNA-dependent RNA polymerase.
Materials and methods
Retrieval and preparation of protein structure
The three-dimensional (3 D) structures of SARS-CoV-2 RdRp [PDB ID: 7BTF], 1 SARS-CoV RdRp [PDB ID: 6NUR] 26 and HCV RdRp [PDB ID: 4WTG] 27 were retrieved from the Protein Data Bank (http://www.rcsb.org). All the crystal structures were prepared by removing existing ligands and water molecules, while missing hydrogen atoms were added using Autodock version 4.2 program (Scripps Research Institute, La Jolla, CA). Thereafter, non-polar hydrogens were merged, while polar hydrogen was added to each protein. The process was repeated for each protein and subsequently saved into a dockable PDBQT format for molecular docking. The two active site aspartates (D760 and D761) were treated as flexible during the docking
Ligands preparation
Structure Data Format (SDF) structures of the reference inhibitors in its active triphosphate form (S1: Remdesivir, and S2: Sofosbuvir) and 226 bioactive compounds derived from African plants were retrieved from the PubChem database (www.pubchem.ncbi.nlm.nih.gov). They were converted to mol2 chemical format using Open babel, 28 while compounds that were not available on the database were drawn with ChemDraw version 19, and converted to mol2 chemical format. Polar hydrogen charges of the Gasteiger-type were assigned to atoms, while the non-polar hydrogen molecules were merged with the carbons, and the internal degrees of freedom and torsions were set to zero. Ligand molecules were further converted to the dockable PDBQT format using AutoDock Tools.
Molecular docking study
Virtual screening
An initial virtual Screening of 226 bioactive compounds against SARS-CoV-2 RdRp [PDB ID: 7BTF] was performed by AutoDock Vina. 29 A larger grid box size of (60 Å × 60 Å × 60 Å) was used to locate different binding conformation around the active site of grid box size (30 × 39 × 52 Å). From the initial docking analysis, compounds with higher binding affinities and conformational poses that were docked into the active side region were selected for further analysis. The PDBQT form of individual protein and bioactive compounds were uploaded into their respective columns of AutoDock Vina. The software performed an exhaustive series of docking calculations across the protein surface marked by the grid box size (60 Å × 60 Å × 60 Å) in order to find the spots within the active site with the best binding affinities.
Active site targeted molecular docking
Base on the docking scores, binding poses and interaction to the catalytic residue, a hit-list of top 36 ranked compounds with binding affinities that are higher than the reference inhibitors was defined, from this list, the top ten compounds were selected. The compounds were further docked into the active sites of RdRp of SARS-CoV-2, SARS-CoV, and Hepatitis C Virus (HCV) using Autodock vina in PyRx0.8 30. The protein structures in PDB format were uploaded into PyRx 0.8. Ligands were imported, and energy minimization was performed via software OpenBabel. The active region of the SARS-CoV-2 RdRp defined by a grid box size of 30 × 39 × 52 Å centered at (x, y, z) of (119.0, 116.2, 119.4) Å was used for docking. All the other parameters were kept as default. The molecular interactions between proteins and selected compounds with higher binding affinity to the proteins were viewed with Discovery Studio Visualizer version 16.
Molecular dynamics
The crystal structure of SARS-CoV-2 RNA-dependent RNA Polymerase (RdRp) was downloaded from the Protein DataBank (PDB ID: 7BTF). Hydrogen atoms were added, while non-relevant chains were removed using PyMOL software. 30 Molecular Dynamics Simulation (MDS) was performed on SARS-CoV-2 RdRp using NAMD software.31,32 This step was divided into two parts; the first was the minimization of the protein for 10,000 steps followed by 100 ns production run. Afterward, the implemented clustering method in Chimera software 33 was utilized to produce 8 clusters, and for each cluster, a representative frame was selected.
Targeted docking to different conformations from molecular dynamics
The trajectories files from molecular dynamics of the apo form of SARS-CoV-2 RdRps were clustered into 8 different conformational clusters from which 8 different cluster representatives were selected. AutoDock Vina was utilized to dock the test compounds (4 reference inhibitor and the top 7 phytocompounds) to the active sites of the 8 different representative structures of SARS-CoV-2 RdRp. The mean of binding affinities of each compound to the 8 different representative conformations was calculated alongside the standard error.29,34 Two positive controls (remdesivir and sofosbuvir), two negative controls (cinnamaldehyde and thymoquinone), and seven top docked compounds with binding affinities higher than the reference positive controls (10′-hydroxyusambarensine, cryptospirolepine, isocryptolepine, millettosin, oryzanol C, strychnopentamine, and usararotenoid A). The grid box size for the docking experiments is 30 × 30 × 30 Å3 centered at the active site residues (D760 and D761) for the different cluster representatives. Afterward, analysis of the interactions established after docking was performed utilizing the Protein-Ligand Interaction Profiler (PLIP). 35 The best conformations were selected for further analysis based on its binding affinity against SARS-CoV-2 RdRp.
Admet study
The drug-likeness prediction of the selected top 5 phyto-compounds that demonstrated highest binding affinity for active regions of the RdRp of SARS-CoV-2, were subjected to Lipinski filter in which an orally bio-active drug should comply to a minimum of four of the five laid down criteria for drug-likeness namely: cLogP, hydrogen donor and acceptor molecular mass, and molar refractive index.. 36 The predicted Absorption Distribution Metabolism, Excretion, and toxicity (ADME/tox) study were analyzed using the SuperPred webserver. 37 This tool was utilized for predicting important descriptors of drug-likeness. The SDF file and SMILES of the compounds were downloaded from the PubChem database to calculate ADMET properties using the default parameters.
Results and discussion
Molecular docking of compounds with the target protein
The results of docking analysis of 226 bioactive compounds alongside the reference drug against SARS-CoV-2 RdRp are presented in Table S1 (supplementary material). Remdesivir and sofosbuvir, the reference inhibitors, had binding affinities of –7.9 and –7.2 Kcal/mol for the RdRp of SARS-CoV-2, respectively. Ranking based on the negative and low value of ΔG, and comparing with the reference inhibitors, a hit list of 36 bioactive compounds were defined (Table S1). These topmost compounds, which comprise alkaloids, flavonoids, and terpenoids classes of compounds, display binding energy ranging from –10.4 to –8.1 Kcal/mol. Docking of these selected compounds with the active region of the SARS-CoV-2, as well as that of SARS-CoV and HCV, the best ten docked compounds were selected viz: 10-hydroxyusambarensine, cryptospirolepine, strychnopentamine, usararotenoid A, 12α-epimillettosin, 24-methylene cycloartenol, ekeberin, Isocryptolepine, dioncophylline B, and usararotenoid C (Figure 1). As reflected from the docking scores, these compounds, which belong to alkaloids, flavonoids, and terpenoids classes of phytochemicals (Table 1), can bind to the three viral RdRp active site regions with good binding energy (−6.3 up to −9.9 kcal/mol) (Figure 2).
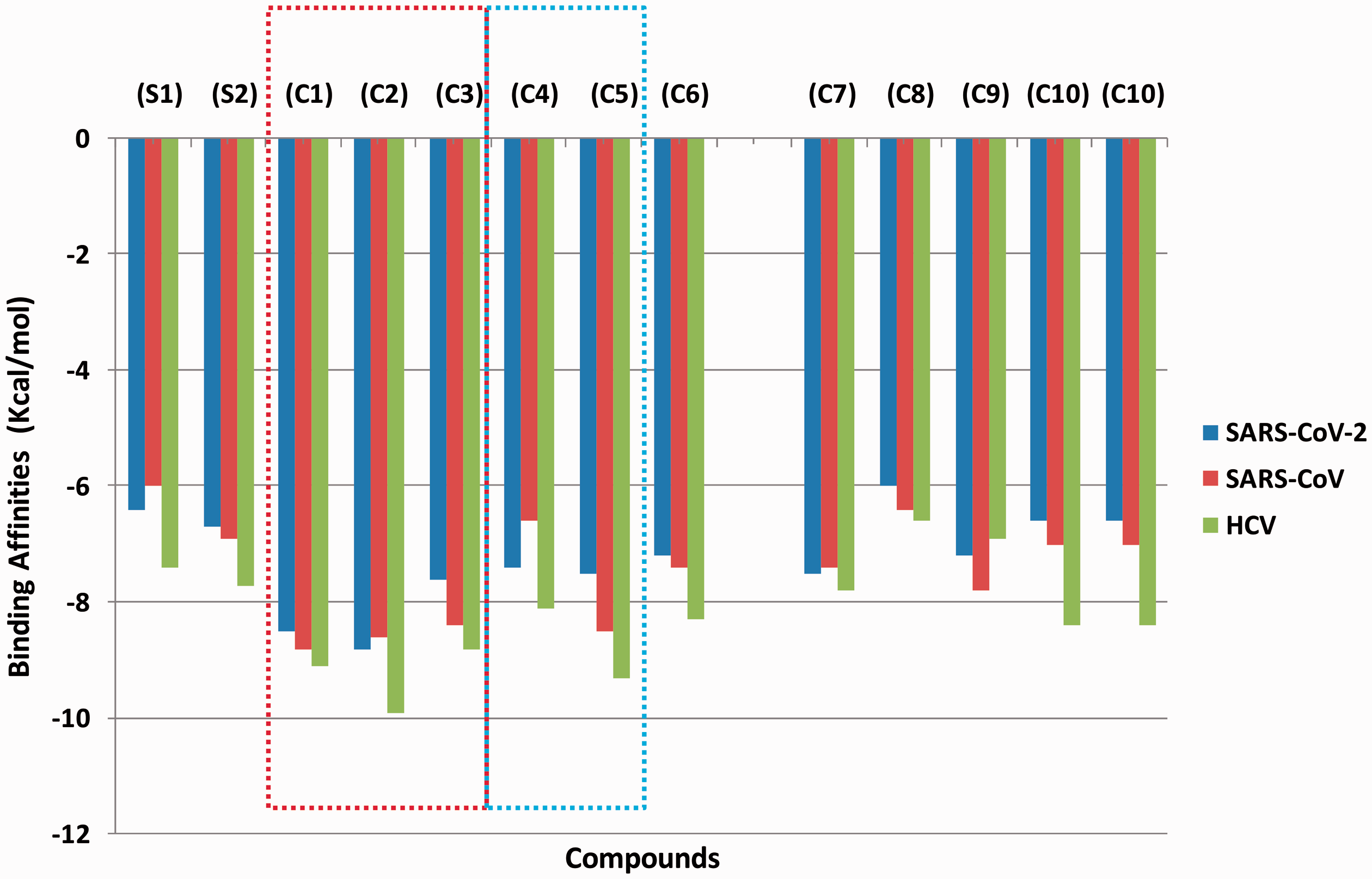
Binding affinities of top phyto-compounds and reference compounds to the active site residues of viral RNA-dependent RNA polymerase. The red dotted line shows the top 3 docked alkaloids while blue dotted lines show the top 2 docked flavonoids.
Top bioactive compounds with the active site residues of SAR CoV-2 RNA-dependent RNA polymerase.

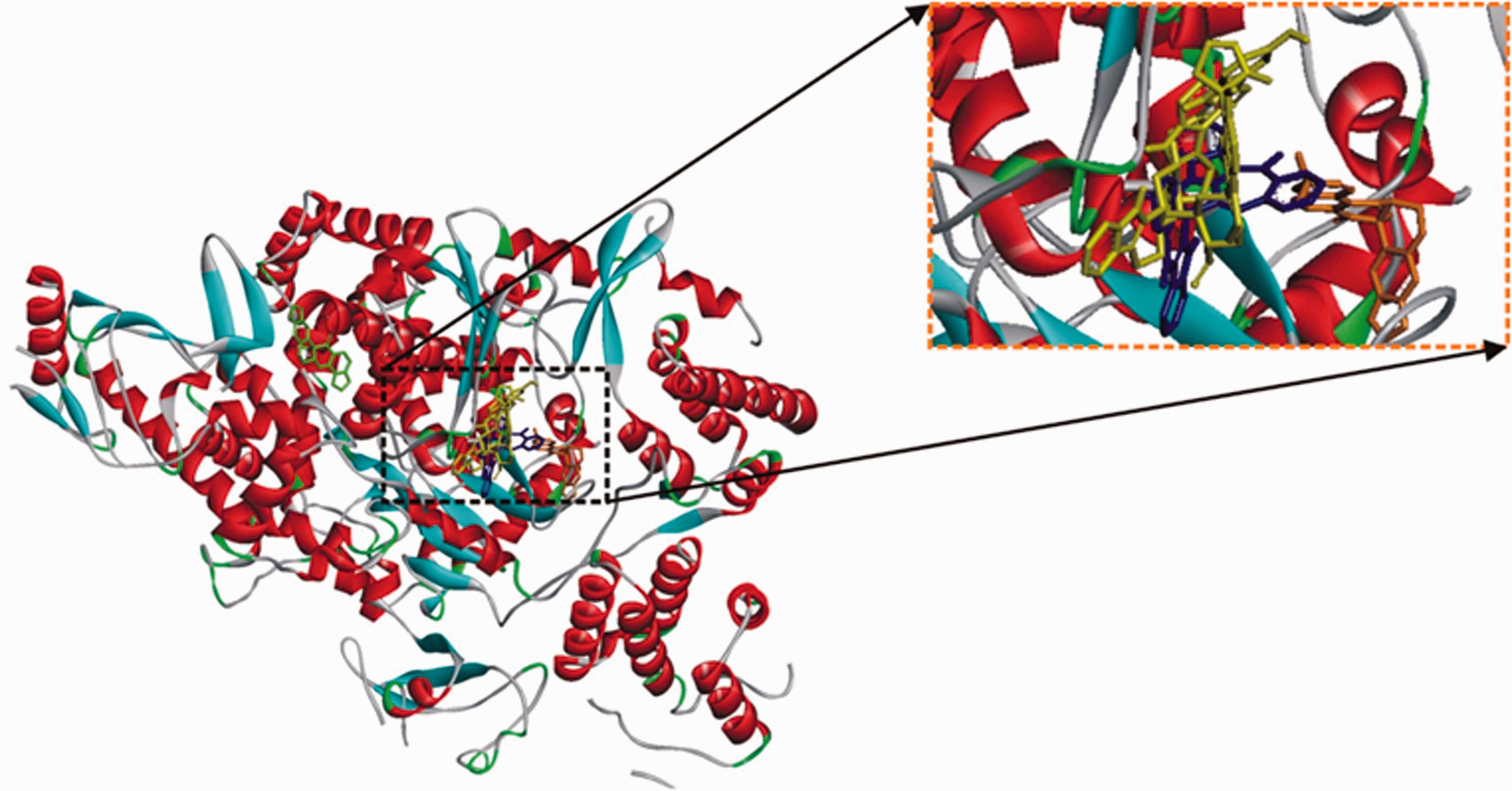
The top 5 phyto-compounds (10'-hydroxyusambarensine (gray), cryptospirolepine (blue), and strychnopentamine (yellow), usararotenoid A (green) and 12α-epi-millettosin (gold), docked into the active site of SARS-CoV-2 RdRp. The enlarged panel is depicted to show how the compounds are lying in the active site cavity of the protein.
The 3 top phyto-compounds docked into the active site of SARS-CoV-2 RdRp are alkaloids, namely: 10′-hydroxyusambarensine ( 9.6 kcal/mol), cryptospirolepine (-8.9 kcal/mol, and strychnopentamine (–8.7 Kcal/mol). It was observed that while 10-hydroxyusambarensine was the topmost docked compound to the RdRp of both SARS-CoV-2 (–9.6 Kcal/mol) and SARS-CoV (–8.8 Kcal/mol), cryptospirolepine had the highest binding affinity to HCV RdRp (-9.9 Kcal/mol) (Figure 1
The rotenoids usararotenoid A (−8.3 Kcal/mol) and 12α epi-millettosin (−8.0 Kcal/mol), are the top docked flavonoids with SARS-CoV-2 RdRp (Figure 1). 12α-epi-millettosin (−9.3 Kcal/mol) interacted more strongly with HCV than with coronaviruses.
Molecular interactions between the selected phyto-compounds and viral RdRp
Investigating the amino acid interactions of the selected phyto-compounds with viral RdRp revealed that, the ligands majorly interacted with the residues through hydrogen bonding and electrostatic interactions, with few hydrophobic interactions. The active site residues ofSARS-CoV-2 and HCV RdRps interacting with the topmost binding compounds are shown in Table 2 and Figure 3.
Interactions of top docked alkaloids and flavonoids with active site residues of viral RNA-dependent RNA polymerase.
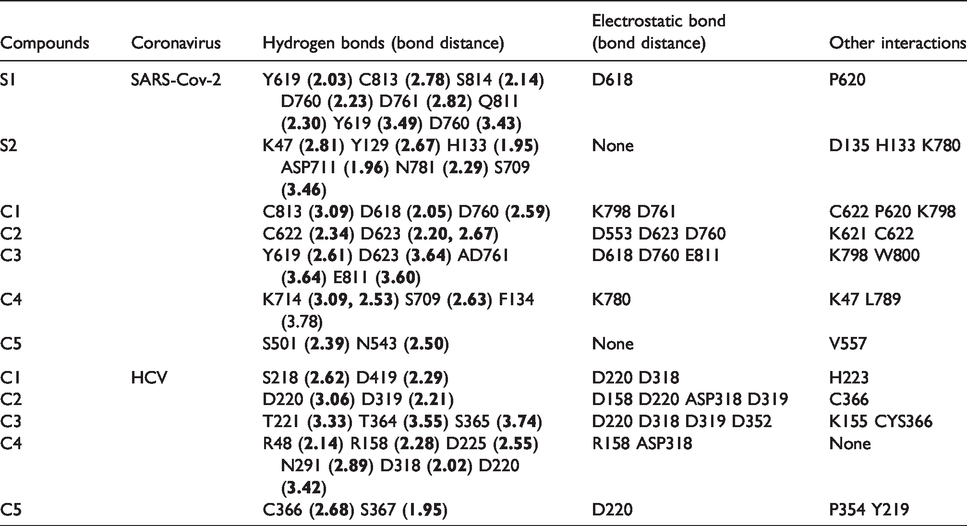
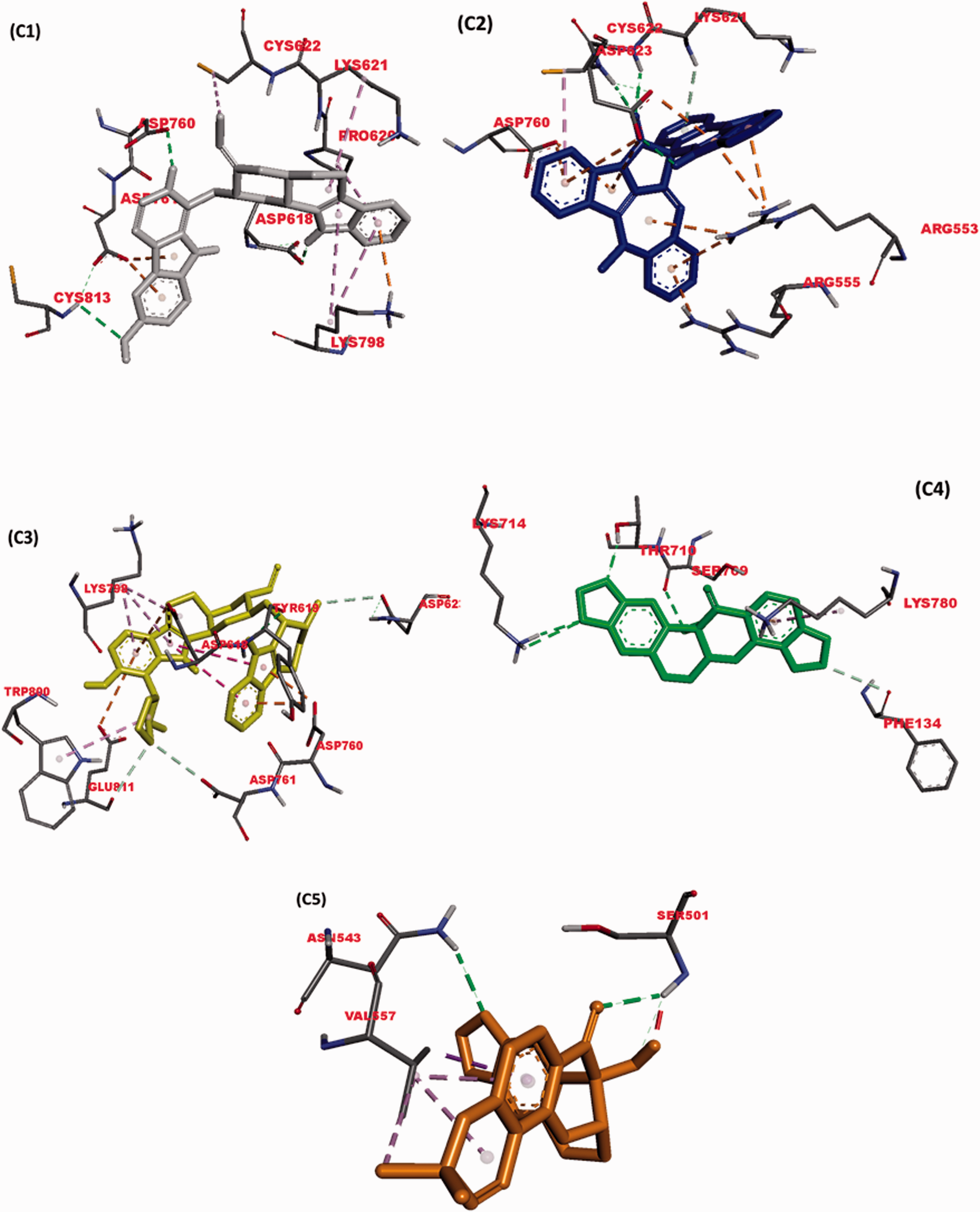
The interaction views of the best five compounds 10'-hydroxyusambarensine (gray), cryptospirolepine (blue), and strychnopentamine (yellow), usararotenoid A (green) and 12α-epi-millettosin (gold), against the active site of SARS-CoV-2 RdRp. H-bonds are depicted by the green dashed lines, while hydrophobic contacts in dashed-purple lines. Interacting residues are labelled with their three-letter codes.
The result obtained from the ligand-protein binding interaction showed that Remdesivir, the reference compound, was docked into the RdRp active site of SARS-CoV-2 by interacting with the binding and catalytic site residues through various types of bonds (Figure 3). It interacted via conventional hydrogen bonds to the catalytic site residues D760 and D761. It further interacted through electrostatic interaction with D618. Sofosbuvir, a second reference compound with a considerably higher binding energy (–7.3 kcal/mol) than Remdesivir, did not show significant binding to the active site residues of SARS-CoV-2 but utilized Hydrogen bond and other interactions to bind to several different amino acid residues.
10-Hydroxyusambarensine, an indole alkaloid with the highest binding affinity to SARS-CoV-2, was docked to the active site of the enzyme in a similar manner to the reference drug. It was observed that this compound interacted via a conventional hydrogen bond to the D618 and D760; Pi-Anion electrostatic bond with D761; Alkyl hydrophobic interaction with P620, K621, and C622 (Table 2, Figure 3(b)). 10-Hydroxyusambarensine exhibit highly conserved binding pattern to both SARS-CoV-2 and HCV RdRps as it also docked to the active site of HCV RdRp through several non-covalent interactions. Cryptospirolepine, a cryptolepines, also exhibited such a conserved binding pattern to both SARS-CoV-2 and HCV RdRps similar to that of 10-Hydroxyusambarensine. At the active site of SARS-CoV-2, it interacted with C622 and D623 through conventional hydrogen bonds; D760 and D623 through Pi-Anion electrostatic bonds; R553 and R555 through Pi-Cation electrostatic bonds; K 621 through Pi-Donor Hydrogen bond; K621 through Alkyl hydrophobic interaction; and C622 through Pi-Alkyl Hydrophobic interaction. Strychnopentamine, another indole alkaloid, was found to establish the highest number of hydrogen bonds with the SARS-CoV-2 RdRp active site (Table 2). Although rotenoids; usararotenoid A and millettosin had high binding affinities to the viral RdRp, they tend to interact selectively with HCV RdRp active site residues.
Molecular docking of top docked phyto-compounds with different conformation from molecular dynamics analyses
Figure 4 shows the average binding energies of each of the reference inhibitors and phytocompounds (in kcal/mol) against the 8 different representative structures of of 8 clusters of SARS-CoV-2 RdRp gotten from a 100 ns run MDS trajectories files. Error bars represent the standard deviation from the mean. Remdesivir and Sofosbuvir (red columns) are used as positive controls to judge the affinity of the natural compounds to SARS-CoV-2 RdRp. Additionally, Cinnamaldehyde and Thymoquinone are used as negative controls. As reflected from Figure 4, the two compounds 10'-hydroxyusambarensine, and cryptospirolepine (dark green columns) show the best average binding affinity to SARS-CoV-2 RdRp (−10.1 ± 0.38 kcal/mol and −10.5 ± 0.57 kcal/mol, respectively). Additionally, the compounds millettosin, oryzanol C, strychnopentamine, and usararotenoid A (light green columns) show better average binding affinities to SARS-CoV-2 compared to the positive controls.
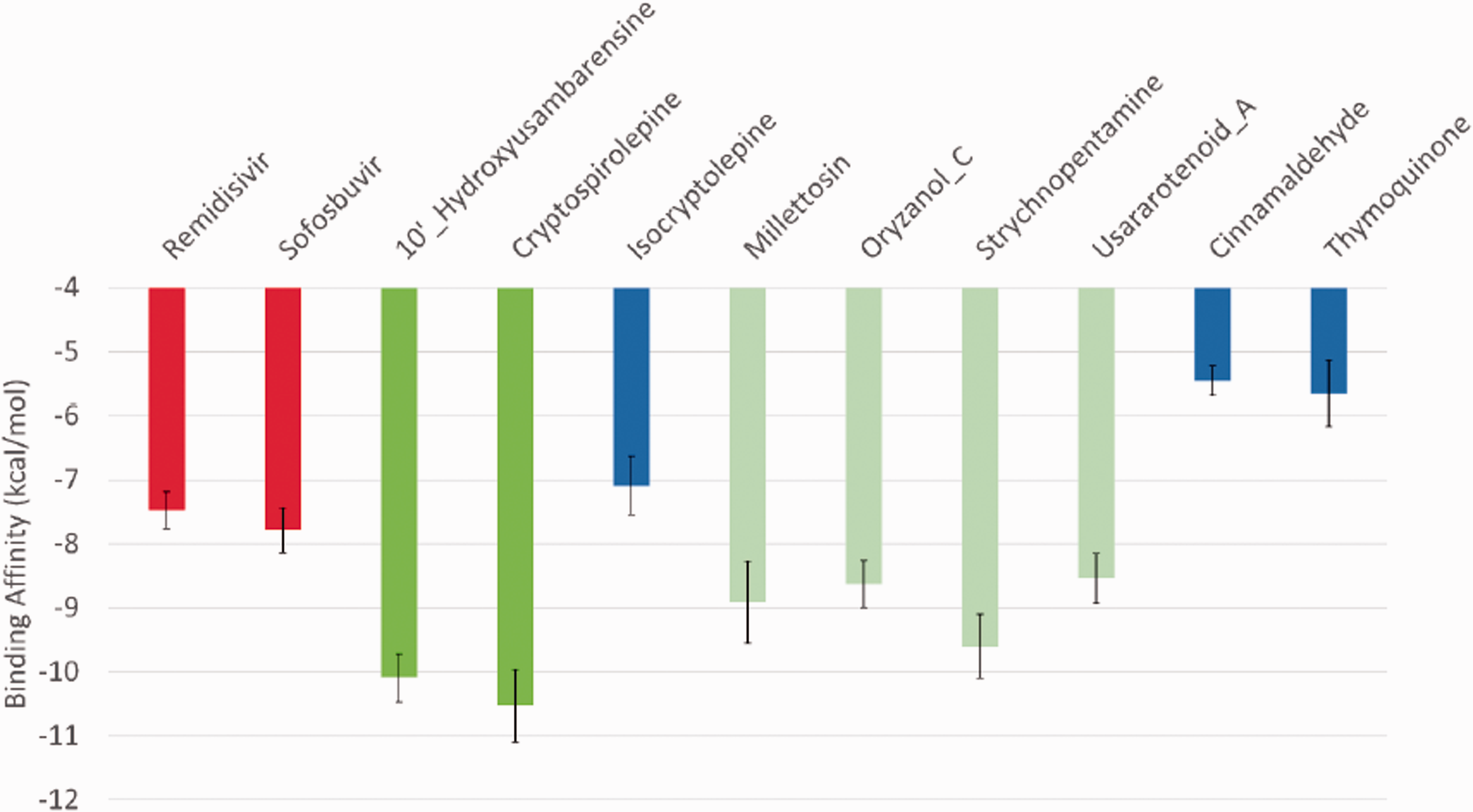
The average binding energies (in kcal/mol) of the 11 compounds against SARS-CoV-2 RdRp active site residue D760 and D761. Error bars represent the standard deviation (SD). The best two compounds are 10'-Hydroxyusambarensine and Cryptospirolepine (dark green columns) with average binding affinities of −10.08 Kcal/mol and −10.525 Kcal/mol, respectively.
The result of the docking analysis of the reference inhibitors and 7 selected compounds to each of the 8 different clustered conformations from the 100 ns run MDS trajectories files of SARS-CoV-2 RdRp is presented in Table 3. Each compound formed 8 complexes with the 8 different clustered conformation, from these, the best representative complex with the highest binding mode and affinity was selected for further analysis using PLIP webserver. Table 3 shows the established interactions upon docking the compounds and inhibitors into the active site of the selected representative complexes of SARS-CoV-2 RdRp. Hydrophobic contacts: H-bonds, salt-bridge, π-stacking (blue), and π-cation (green) interactions are reported and recorded in Table 3. Amino acids involved in these interactions are also listed in the table with the corresponding docking scores in kcal/mol. The most-reported amino acids interacting with the compounds are K551 and S814 (bold). As reflected from the table, at least three hydrophobic contacts are established for all the complexes (7 and 6 contacts for the 10'-hydroxyusambarensine, and cryptospirolepine, respectively). On the other hand, the main driving force for the binding for the positive controls is the H-bonding (5 and 9 H-bonds for remdesivir and sofosbuvir, respectively).
The interaction pattern of the selected complexes. The AutoDock Vina scores are listed among the number of H-bonds, Salt Bridges, and hydrophobic contacts and the residues that interact. Residues that interact with π-stacking are blue-colored, and residues that interact with π-cation are green-colored.
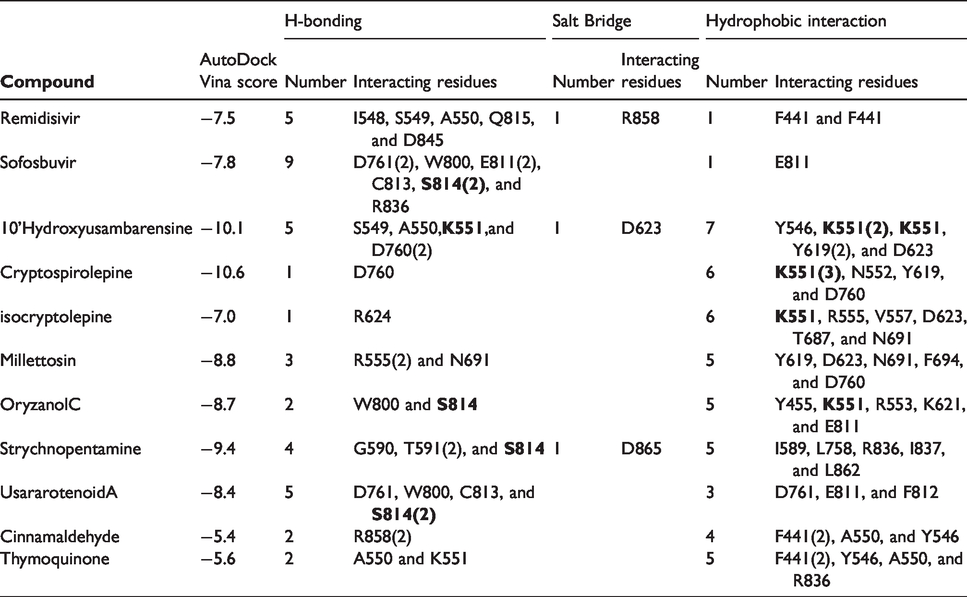
Figure 5 shows the interaction patterns for the best three compounds based on binding affinities (10'-hydroxyusambarensine, cryptospirolepine, and strychnopentamine). For each of the test compounds that was docked to the 8 different representative structure to the 8 clusters, the complex with the best binding conformation and binding affinities were selected for further interactive analysis. Hydrophobic interactions (dashed gray lines) and Hydrogen bonding (blue lines) are the most common types of communications. Salt-Bridges (dashed yellow line between two yellow balls) are found only in 10'-hydroxyusambarensine (D 623) and strychtopentamine (D865), while π-cation interaction is only found in 10'-hydroxyusambarensine (K551).
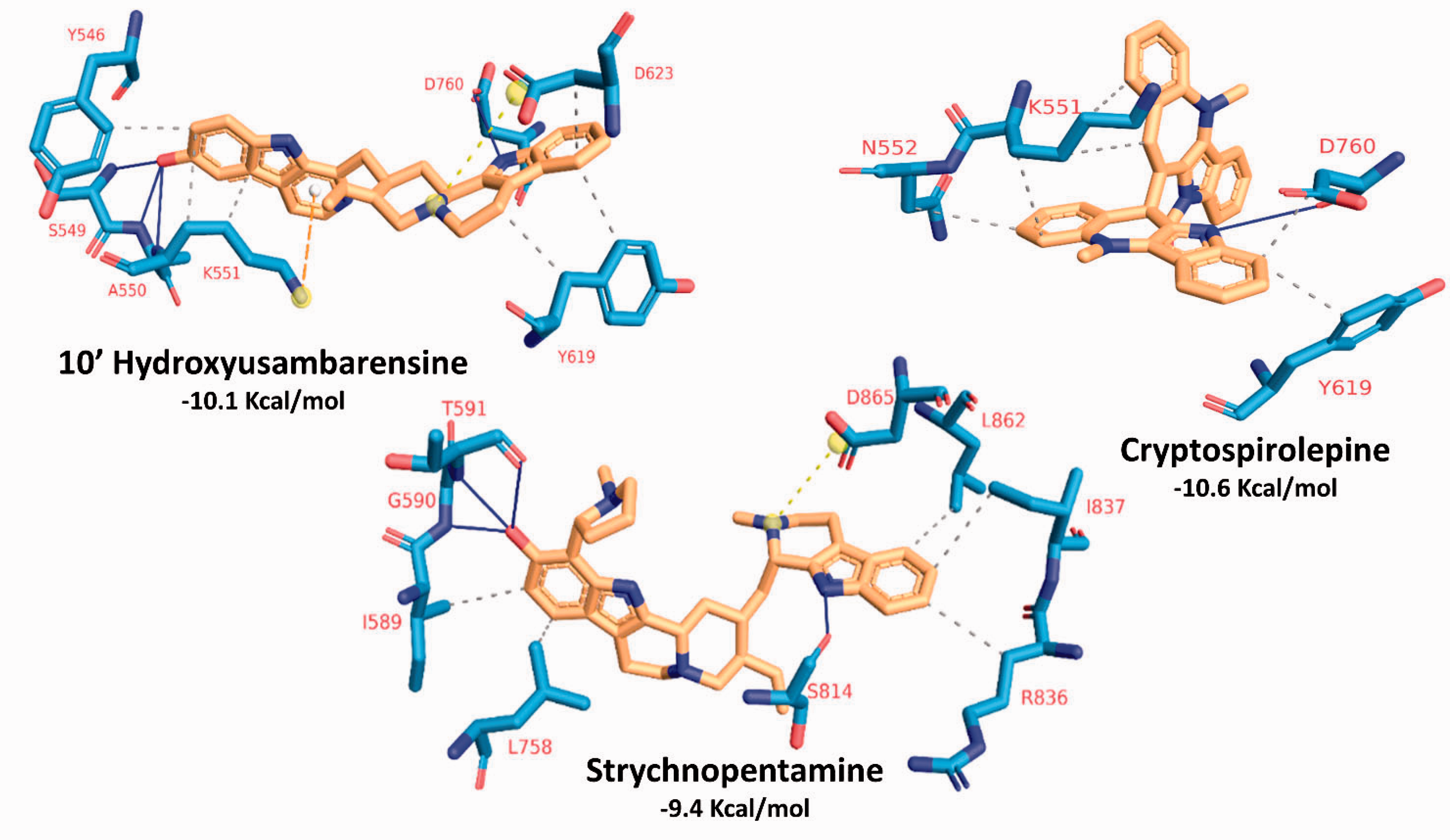
The interaction pattern for the docking of the best three compounds (10'_hydroxyusambarensine, cryptospirolepine, and strychnopentamine) against the active site of SARS-CoV-2 RdRp. H-bonds are depicted by blue lines, while hydrophobic contacts in dashed-gray lines. Salt-bridges are depicted in dashed-yellow lines between two yellow balls. π-cation interactions are in dashed-orange lines. Interacting residues are labelled with their one-letter codes and red-coloured. Coloured sticks represent both the compounds (yellow) and the interacting residues from SARS-CoV-2 RdRp (blue).
Figure 6 shows the superposition of the three top complexes, with 10'-Hydroxyusambarensine as green sticks, cryptospirolepine as cyan sticks, and strychtopentamine as magenta sticks. The active site aspartates (D760 and D761) are labeled and represented in colored sticks.
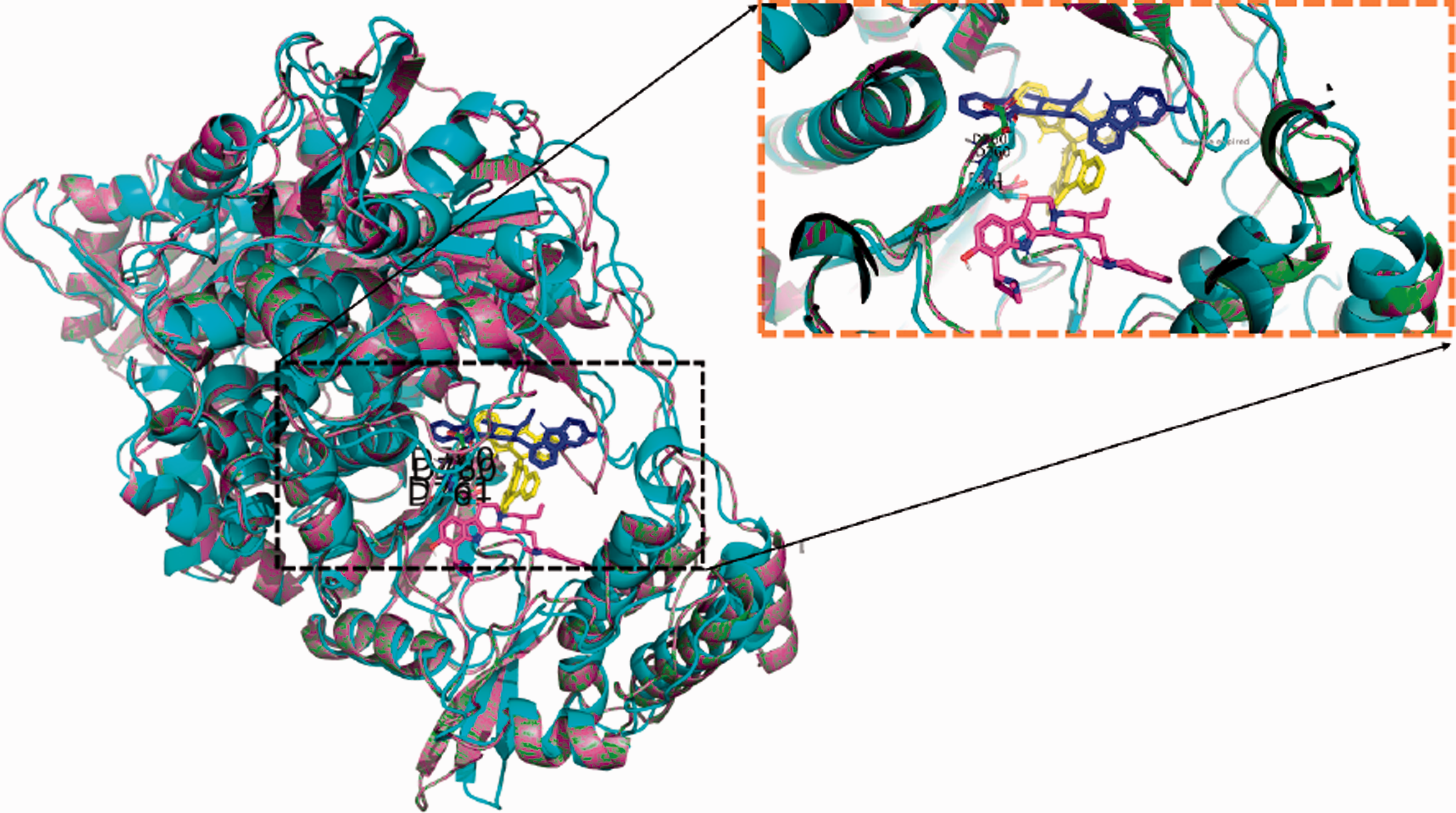
The superposition of the docking complexes of the 10'-hydroxyusambarensine (yellow), cryptospirolepine (blue), and strychnopentamine (magenta), docked into the active site of SARS-CoV-2RdRp. The enlarged panel is depicted to show how the compounds are lying in the active site cavity of the protein.
Pharmacokinetic properties of selected alkaloids and flavonoids
The result generated from the Lipinski and ADME/tox filtering analyses are represented in Figure 7 and Table 4.
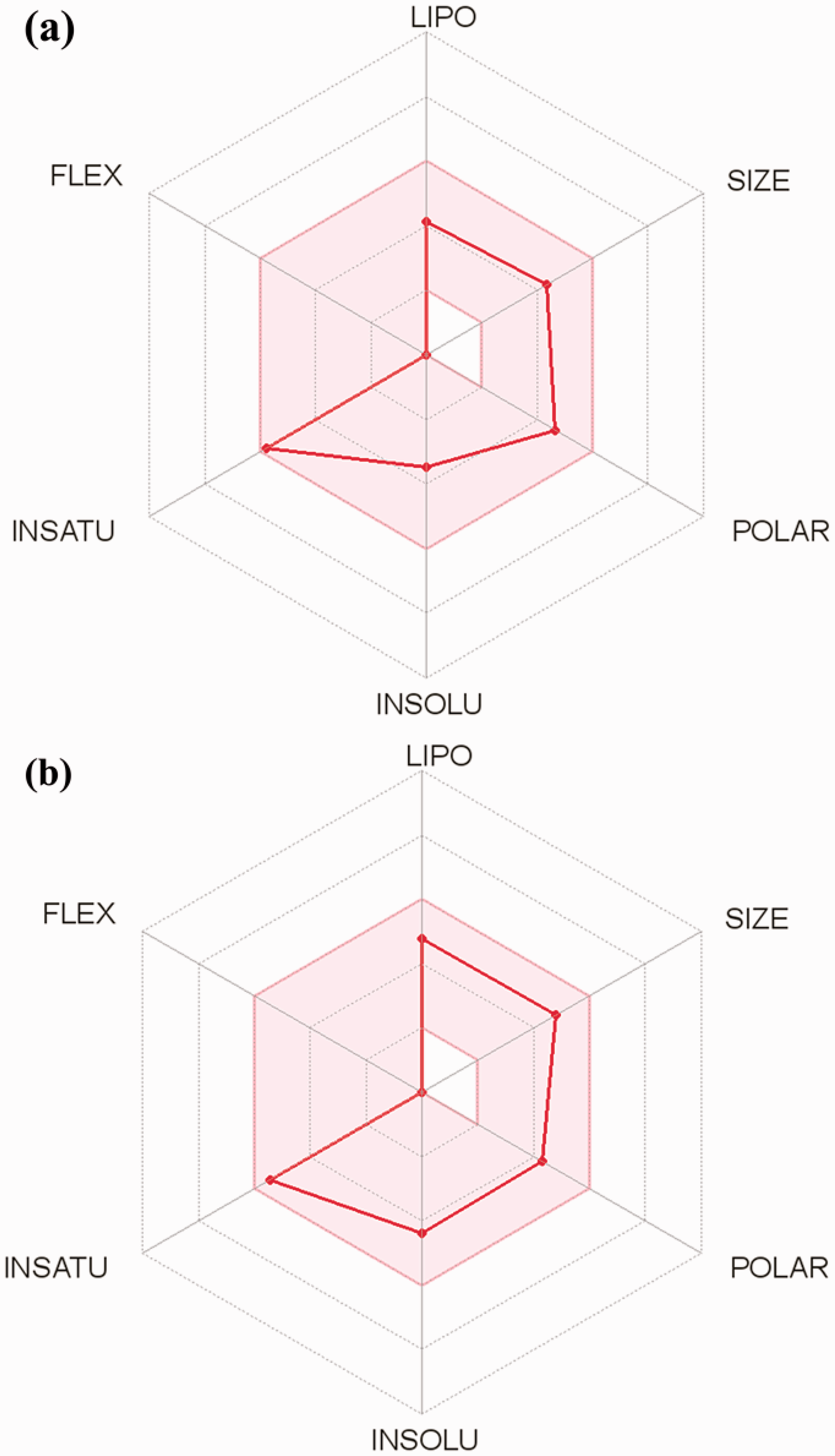
Summary of pharmacokinetic properties of the top, binding phytocompounds from African plants to the
Physicochemical properties of the top, binding phyto-compounds from African plants.
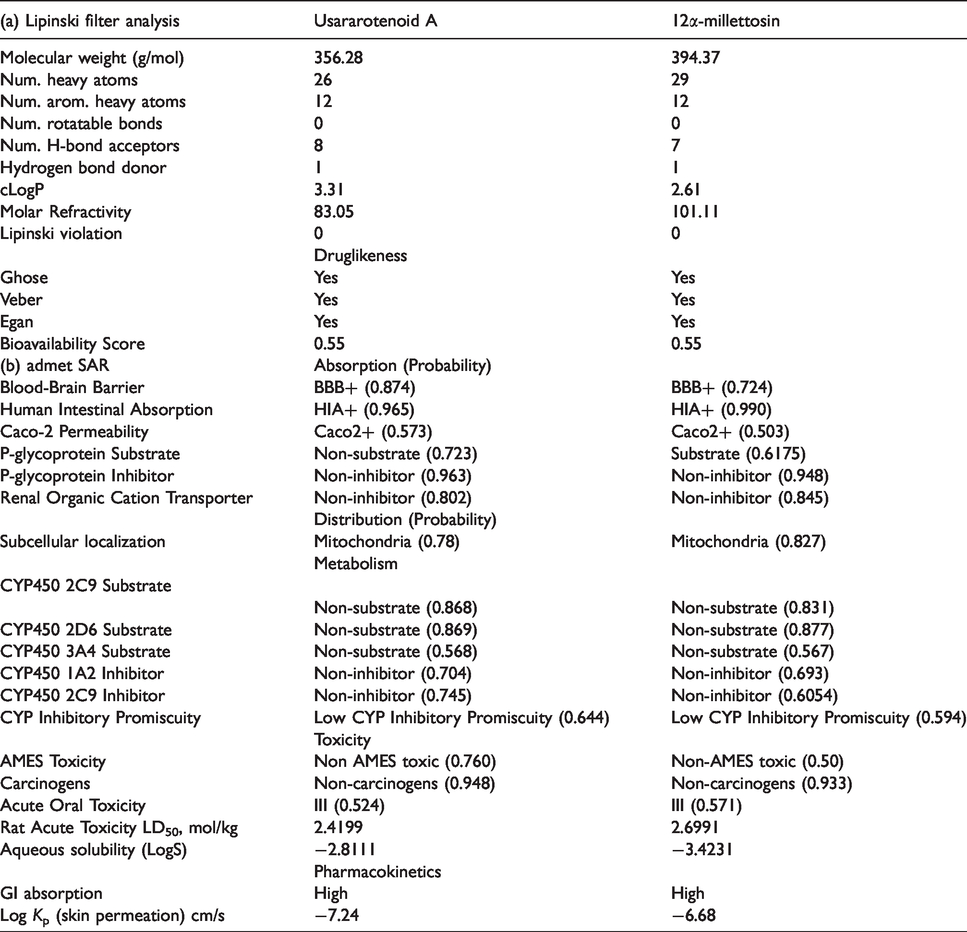
The five (5) top Phyto-compounds fulfilled the requirement for Lipinski analysis of the rule of-five and other drug-likeness parameters such as Ghose, Veber, and Egan with corresponding favorable predicted ADME/tox parameters. From the selected 5 top docked compounds, the three alkaloids (10-hydroxyusambarensine, cryptoquindoline, and strychnopentamine) have been reported in our previous studies 24,25 , while the two flavonoids (usararotenoid A and 12α-epi-millettosin) are represented in Figure 7 and Table 4. The predicted physicochemical properties for bioavailability of the lead compounds were further described in Figure 5.
The ADME/tox and pharmacokinetic properties from the filtering analyses suggested the top 5 phyto-compounds with a high probability of absorption, subcellular distribution, and low toxicity parameter. The top 5 phyto-compounds were indicated to be non-carcinogenic, with very low acute toxicity and aqueous solubility of < 0. The gastrointestinal absorption index was indicated to be high for usararotenoid A and 12α-epi-millettosin (Table 4).
Discussion
Exploiting the potentials of phytochemicals is an integral component of the international response to the COVID-19 pandemic. Herein, phytochemicals from African herbs and medicinal plants are reported as potential inhibitors of the SARS-CoV-2 RdRp. Since this enzyme plays a central role in coronaviral replication/transcription machinery, it is accepted as an excellent target for new therapeutics for which lead inhibitors such as remdesivir has been approved by the FDA..1,13 The topmost five bioactive compounds, which comprise three alkaloids and two flavonoids, were found to establish multiple non-covalent interactions with the active sites of RdRp of both SARS-CoV-2 and HCV. These compounds can bind more tightly than the two drugs (remdesivir and sofosbuvir) used as a reference in this study. Thus, these compounds may be able to bind to the new coronavirus strain RdRp tightly and hence compromise the polymerase function.
All the top docked compounds from this study exhibited a close conserved binding pattern. The compounds like the reference inhibitors (positive controls) were all directed into the conserved polymerase motifs A to G of RdRp of both SARS-CoV-2 and HCV. The strongest binding tendency and protein interaction exhibited by alkaloids: 10’-hydroxyusambarensine, cryptoquindoline, and cryptospirolepine (the top docked compounds) portrays alkaloids as potential inhibitors RdRp of SARS-CoV-2, SARS-CoV and, HCV.
The alkaloids especially the indole alkaloids (10-Hydroxyusambarensine and Strychnopentamine) from
The stability of the complexes formed stemmed from the vast number of interactions with the catalytic residues in the Motif C of the active site of the enzyme. Motif C, which comprises amino acid region 753 to 767, contains the catalytic residues S759, D760, and D761 in the β-turn structure. 1 These catalytic residues are highly conserved in most viral RdRps, e.g., G317, D318, and D319 in HCV ns5b and S327, D328, and D329 PV 3Dpol, with the first residue being either serine or glycine. These two highly conserved aspartate residues protruding from a beta-turn structure, making them surface accessible through the nucleotide channel (free nucleotides can pass through), are formed by the conserved polymerase motifs A to G in the palm domain and configured like other viral RNA polymerases. Besides, these potential alkaloid inhibitors interacted with the classic divalent-cation–binding residue D618 and other key amino acid residues in Motif A, which comprises amino acid residues 611 to 626. The divalent-cation–binding residue D618, is also conserved in most viral polymerases, including hepatitis C virus (HCV) ns5b (residue AD220) 27 and poliovirus (PV) 3Dpol (residue D233). 39
Rotenoids usararotenoid A and (+)12α epi-millettosin, the two top docking flavonoids have different binding modes from the alkaloids as they preferably interacted with amino acid residues around the NTP entry channel of the SARS-CoV-2 RdRp. This channel is formed by a set of hydrophilic residues, including K545, R553, and R555 in motif F. In addition to the interactions of the rotenoids with SARS-CoV-2, they tend to exhibit more selective binding to the HCV RdRp active site residues. This suggests a broad spectrum binding of the rotenoids to viral RdRp.
A comparative analysis of the interactions obtained from the docking of the top docked phyto-compounds to a single conformation of SARS-CoV-2 RdRp, and clusters from different conformations obtained during MDS analysis further established the stability of the docked complexes. The compounds were anchored in a similar pattern, interacting with similar amino acid residues with close binding affinities. Furthermore, a competitive molecular docking study that used both positive and negative controls as a reference in the analysis showed that the phyto-compounds portrayed inhibitory potential 40
The Lipinski, other drug-likeness predictive parameters and the ADME/tox filtering analyses, revealed that the top five docked compounds were non-toxic, druggable natural compounds that bind to the active region of SARS-CoV RdRp. They showed relatively good lipophilicity as the logP values were less than 541,42. The result from the predicted filtering analyses of the five compounds showed parameters that suggest favorable ADME/tox and pharmacokinetic properties. This further indicates the druggability potential of the best-docked alkaloids and flavonoids
Conclusion
The current study revealed that specific alkaloid and flavonoid compounds from African herbs and medicinal plants are potential inhibitors of SARS-CoV-2. They are capable of establishing strong and favorable binding interactions with the active site of SARS-CoV-2 RdRp as well as that of SARS-CoV and HCV, thus may compromise the catalytic functions of this enzyme. These compounds show favorable predicted ADME/tox parameters and may be exploited towards the development of a cocktail of anti-coronavirus treatments that potentially can be used in the prevention and management of COVID- 19 pandemic. Such naturally existing phyto-compounds may provide a wealth of chemical structure diversity, that can help in the development of nutraceuticals, pharmaceutical intermediates, and chemical entities for synthetic or semisynthetic as prophylactic and therapeutic agents against the SASR-CoV-2. However, experimental studies are suggested to validate the possible preclinical and clinical efficacy of these agents for the prevention and treatment of COVID-19.
Supplemental Material
sj-pdf-1-avc-10.1177_2040206620984076 - Supplemental material for Alkaloids and flavonoids from African phytochemicals as potential inhibitors of SARS-Cov-2 RNA-dependent RNA polymerase: an in silico perspective
Supplemental material, sj-pdf-1-avc-10.1177_2040206620984076 for Alkaloids and flavonoids from African phytochemicals as potential inhibitors of SARS-Cov-2 RNA-dependent RNA polymerase: an
Footnotes
Acknowledgement
The authors are grateful to the Department of Biosciences, Salem University, Lokoja Nigeria for providing an enabling environment to the BioNet-AP:Bioinformatics Networkon for African Phytomedicine for COVID-19 research.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
