Abstract
Background
With the advent of next generation integrase strand transfer inhibitors, the rates of virologic failure in treated subjects are expected to decrease. In this study, we analyzed the mutation patterns leading to virologic failure before and after starting integrase strand transfer inhibitor-based regimen as first-line or salvage therapy.
Methods
Between 2016 and 2019, blood samples were received from 258 patients with HIV-1 infection. Plasma HIV-1 RNA concentrations, and pol gene sequences were determined at baseline, and 16–48 weeks of treatment with integrase strand transfer inhibitor-based regimen. Only patients who did not achieve viral suppression at 48 weeks of integrase strand transfer inhibitor-based treatment were eligible for the current study.
Results
Virologic failure was observed in seven patients on raltegravir-based regimen. All patients with virologic failure but one were infected with CRF01_AE virus subtype. Raltegravir based-regimen was offered as first-line therapy for four patients, and as salvage therapy for three patients. M184V mutation associated with high level resistance to lamivudine and emtricitabine was detected in six out of seven patients. Primary mutations (Y143C, N155H, T66I, G118R, E138K) conferring high level resistance to raltegravir were detected in only three patients. Pre-existing polymorphic integrase mutation (T97A) was detected in two patients. Furthermore, two patients reported low adherence to treatment.
Conclusions
Emergence of primary mutations in the integrase gene can account for virologic failure in less than half of patients on raltegravir-based regimen. Low adherence to treatment, pre-existing accessory mutations, and resistance to reverse transcriptase inhibitors may have some role in virologic outcome.
Introduction
Integrase strand transfer inhibitor (InSTI)-based regimen is generally recommended as first-line antiretroviral therapy (ART). 1 There are currently five InSTIs, four approved by the US Food and Drug Administration, raltegravir (RAL), elvitegravir (EVG), dolutegravir (DTG), and bictegravir (BIC), and an investigational drug, cabotegravir (formerly S/GSK-1265744). 1 BIC and DTG have higher genetic barrier to resistance than the first-generation InSTIs (RAL and EVG), and do not require pharmacologic boosting. 1 The efficacy of DTG was clearly shown in HIV-1 patients with evidence of resistance to RAL or EVG.2,3 The InSTI-based regimen generally consists of one InSTI plus two nucleoside reverse transcriptase inhibitors (NRTIs). For instance, DTG in combination with tenofovir alafenamide (TAF) or tenofovir disoproxil fumarate (TDF) and emtricitabine (FTC), or BIC in combination with TAF and FTC, are recommended for rapid initiation. Consequently, more than 90% of patients receiving the DTG/TDF/FTC regimen are expected to achieve viral suppression at week 48, as reported in previous studies.4,5
Mutations in the HIV-1 pol gene conferring resistance to InSTIs have been reported following ART start.6–8 ART-resistance mutations are grouped into major and minor types. Those appearing first during treatment failure and generally conferring ART resistance are defined as major or primary mutations; whereas those occurring later, modulating ART susceptibility, compensating for fitness defects or presenting sometimes as polymorphisms, are defined as minor, accessory or secondary mutations. 8 Major mutations have been mainly reported in ART-experienced patients,6,7,9 whereas minor mutations have been described in both ART-naïve and -experienced patients.6,7,9–11
Although the first-generation InSTIs (RAL and EVG) are potent well tolerable drugs, 12 major mutations resulting in reduced susceptibility to InSTIs and virologic failure are detected in up to 60% of highly treatment-experienced patients. 13 Mutations at positions 92, 143, 148, and 155 of the integrase gene are the most common mutations to arise during failure of first-generation InSTI-based therapy.14–16
In our previous studies, we reported the detection of major mutations conferring resistance to nucleoside and non-nucleoside reverse transcriptase inhibitors (NNRTIs) in 12.5% of 64 ART-naive patients, and about 30% of 64 treatment-experienced patients. 17 Major non-polymorphic mutations that confer resistance to InSTIs were not detected in 53 InSTI-naïve patients. 18 Given the scarce information on HIV-1 drug resistance in real clinical settings, especially in the Arabian Gulf region, we aimed in this report to characterize the patterns of mutations detected among patients who did not achieve viral suppression following 48 weeks of treatment with InSTI-based regimen.
Materials and methods
Study population
Patients infected with HIV-1 and treated with InSTI-based regimen were followed up for 48 weeks. There were all recruited from Infectious Disease Hospital, Ministry of Health, Kuwait. The study period was from January 2016 to December 2019. An informed consent was obtained from each participant before blood sample collection. The research study was carried out in accordance with the recommendations of the Ethical Decision Committee of the Research Administration, Faculty of Medicine, Kuwait University, and the 2008 Declaration of Helsinki.
HIV-1 RNA concentrations
HIV-1 RNA concentrations in the plasma samples of patients were measured within 16–48 weeks of treatment, using the COBAS AmpliPrep/COBAS TaqMan HIV-1 test v2.0 (Roche Diagnostic Systems, Branchburg, NJ). Viral suppression is defined when the viral load is below the limit of detection (<50 copies/mL). Virologic failure is defined as viral load above 200 copies/mL on at least two consecutive measurements. 1
HIV-1 genotyping and drug resistance assessment
The MagNa Pure LC 2.0 system (Roche Diagnostic Systems) was used to isolate total RNA from plasma samples. Two nested reverse transcription polymerase chain reactions (RT-PCR) were performed to amplify the protease/reverse transcriptase region, and the integrase region in the HIV-1 pol gene, as described previously.19,20 The Wizard SV GEL and PCR Clean-Up System kit (Promega Corporation, Madison, WI) was used to purify the PCR products. The ABI 3500 Genetic Analyzer (Applied Biosystems, Foster City, CA) was used to determine the nucleotide sequences of 5' and 3' DNA strands as described previously. 18 The identification of HIV-1 subtype and mutations associated with resistance to protease inhibitors, reverse transcriptase inhibitors and InSTIs, was done using the Stanford University genotypic resistance interpretation algorithm. 21
Statistical analysis
The differences in the HIV-1 RNA concentrations at 24 and 48 weeks of treatment were assessed using the Wilcoxon signed-rank test. The statistical analysis was performed using the IBM SPSS Statistics for Windows, version 25 (IBM Corp., Armonk, NY).
Results
From January 2016 to December 2019, a total number of 258 blood samples were received for routine HIV-1 drug resistance testing. The samples were collected from 191 patients newly diagnosed with HIV-1 infection, and 67 ART-experienced patients. Among patients treated with InSTI-based regimen, virologic failure was observed in a total of seven patients on RAL-based regimen, while viral suppression was achieved in all patients on EVG-based regimen (Stribild or Genvoya), or DTG-based regimen (Tivicay + Descovy or Truvada or Epzicom). Table 1 shows the demographic and viral findings in patients who did not achieve viral suppression at 48 weeks of treatment. Their median age was 40 years (range, 4–45 years). All patients were Kuwaiti including two females and five males. One patient was infected with HIV-1 subtype C, and the rest with subtype CRF01_AE. Patients “1” through “3” were offered RAL-based regimen as second-line therapy, following a virologic failure with NNRTI-based regimen. Patients “4” through “7” were offered RAL-based regimen as first-line therapy. The median viral load after 24 weeks of treatment with RAL-based regimen (2.50 × 102 copies/mL), was significantly lower than after 48 weeks of treatment (9.57 × 104 copies/mL, p = 0.018).
Demographic and viral findings in InSTI-treated patients with virologic failure.
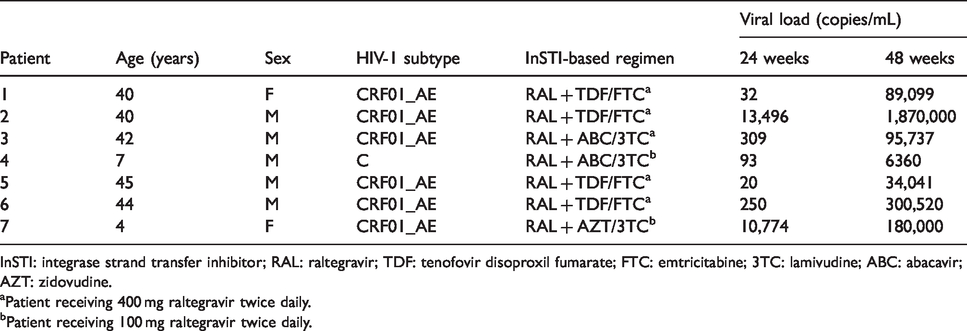

InSTI: integrase strand transfer inhibitor; RAL: raltegravir; TDF: tenofovir disoproxil fumarate; FTC: emtricitabine; 3TC: lamivudine; ABC: abacavir; AZT: zidovudine.
aPatient receiving 400 mg raltegravir twice daily.
bPatient receiving 100 mg raltegravir twice daily.
M184V mutation associated with high level resistance to lamivudine (3TC) and FTC was detected in six out of seven patients (Table 2). Among patients with M184V mutation, two had mutations at position 138 of reverse transcriptase that is associated with low-level resistance to the NNRTI, rilpivirine, and three had major mutations (Y143C, N155H, T66I, G118R, E138K) conferring high level resistance to RAL (Table 2). Furthermore, accessory mutations potentially associated with low level resistance to InSTIs accompanied the reverse transcriptase M184V mutation in three patients.
Mutations associated with resistance to reverse transcriptase or integrase inhibitors in patients with virologic failure.
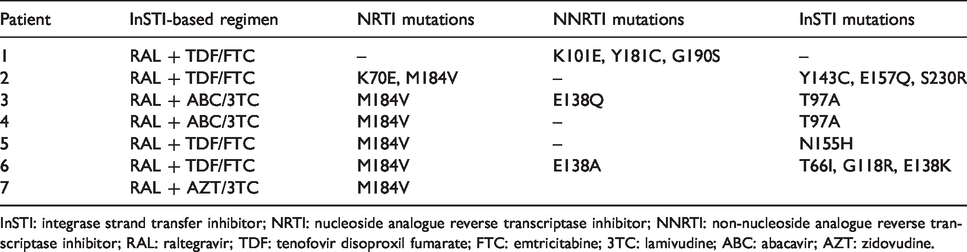
InSTI: integrase strand transfer inhibitor; NRTI: nucleoside analogue reverse transcriptase inhibitor; NNRTI: non-nucleoside analogue reverse transcriptase inhibitor; RAL: raltegravir; TDF: tenofovir disoproxil fumarate; FTC: emtricitabine; 3TC: lamivudine; ABC: abacavir; AZT: zidovudine.
Patients “1” through “3” were on NNRTI-based regimen before starting RAL-based regimen as salvage therapy. HIV-1 reverse transcriptase gene from patient “1” had developed mutations associated with high level resistance to all available NNRTIs (Table 2). The patient had a history of ART non-compliance, and showed viral rebound within 48 weeks of the RAL-based salvage therapy. Major and accessory mutations conferring high level resistance to RAL and to the NRTIs, 3TC and FTC, were detected in HIV-1 sequence of patient “2” who had also a history of low adherence to ART. Non-polymorphic mutations associated with resistance to InSTIs could not be detected in the HIV-1 sequence of patient “3”, despite a viral rebound detected four months after the start of RAL-based regimen. However, before starting the salvage therapy, the HIV-1 sequence had the polymorphic T97A mutation that has usually minimal effects on RAL susceptibility, and the E138Q mutation associated with low level resistance to the NNRTI, rilpivirine. Thereafter, the M184V mutation that is associated with resistance to NRTIs was detected within 32 weeks of treatment with RAL and Kivexa (abacavir/3TC), while the T97A and E138Q mutations disappeared at 48 weeks of treatment.
Patients “4” through “7” who had no previous exposure to ART, failed RAL-based regimen offered as first-line treatment. Baseline HIV-1 genotyping results showed the presence of integrase and reverse transcriptase gene polymorphism in only two patients, “4” and “6”. However, following RAL-based therapy, the polymorphic mutations disappeared, the NRTI mutation, M184V, occurred in all four patients, while RAL-associated mutations were observed in two patients, “5” and “6” (Table 2). Furthermore, patient “6” developed cytomegalovirus-associated retinitis.
Discussion
In this study, mutations in the HIV-1 integrase gene were analyzed before and after starting InSTI-based therapy. Only patients who showed suboptimal viral suppression or virologic failure during InsTI-based therapy were eligible for this study. Kuwait has a very low prevalence of HIV-1 infection, 22 and as expected, a few number of patients developed resistance to the antiretroviral drugs included in the regimen, during the study period.
G118R, Y143C, and N155H were the major integrase mutations accounting for the virologic failure detected in three out of seven patients, following 32 to 48 weeks of RAL-based regimen. In an earlier study, major mutations conferring resistance to RAL-based regimen were detected in 10 out of 69 patients on RAL-based regimen with median time period up to RAL therapy failure of 36 weeks. N155H was the main mutation pathway for virologic failure, followed by Q143R and Q148R/H mutation pathways. 23 M184V mutation in the reverse transcriptase conferring high level resistance to 3TC and FTC accompanied the major integrase mutations. This is consistent with previous studies reported in patients treated with RAL and two NRTIs.24,25 This resistance profile has been also reported in patients on EVG-based regimen, with the emergence of primary mutations associated with resistance to InSTIs (RAL and EVG) and NRTIs (abacavir (ABC), 3TC and FTC) in 9 and 10 out 348 patients, respectively. Interestingly, these mutations disappeared later at week 144 of therapy. 26
The polymorphic mutation, T97A, was detected at baseline in two patients, and disappeared later after starting the RAL-based regimen when only M184V mutation emerged. T97A mutation occurs in 1% to 4% of viruses from ART-naive patients depending on subtype. 27 It emerges in patients on RAL, 28 EVG, 29 and DTG. 2 Alone, pre-existing T97A has no or minimal effect on RAL susceptibility, 30 but in combination with other primary mutations in the integrase gene, it reduces susceptibility to RAL and other InSTIs.13,31 At 48 weeks of treatment, T97A mutation could not be detected in the HIV-1 sequences, possibly due to the selection of variants with higher fitness by RAL or NRTIs included in the regimen, as previously described, 23 resulting in detection missing due to the low sensitivity of Sanger bulk sequencing that is known to detect only variants with prevalence of more than 20%. 32
Primary mutations conferring resistance to RAL could not be identified in four patients with virologic failure. This may result from a combination of different factors that have been clearly explained elsewhere. 8 One of these factors is the presence of drug resistant minority viral populations that failed to be detected by a standard sequencing method as described above, and required more sensitive method like next generation sequencing.33–35 Another important factor is the low adherence to medications, and the prolonged interval between the time of antiretroviral drug discontinuation and genotypic testing. Indeed, two subjects had non-compliance history. An early study has shown that in more than half of subjects on RAL who experienced a virologic failure, HIV-1 variants harboring mutations conferring resistance to RAL become undetectable after drug interruption within a few weeks. 36
In conclusion, virologic failure was detected only in patients on RAL-based regimen at 16–48 weeks of treatment. G118R, Y143C, and N155H were the main pathways to virologic failure. A longer follow-up study (>96 weeks treatment regimen) is on progress to investigate whether the resistance pathways detected in this study will persist, disappear, or replaced by other mutational pathways. Furthermore, the association of different factors combined, e.g. pre-existing accessory mutations in the integrase and/or reverse transcriptase, emergence of primary mutations in the reverse transcriptase gene, and non-compliance history, with drug resistance, should be investigated in future studies.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
