Abstract
Background
Due to resistance to all classes of anti-HIV drugs and drug toxicity, there is a need for the discovery and development of new anti-HIV drugs.
Methods
HIV-1 inhibitors were identified and biologically characterized for mechanism of action.
Results
We identified a dibenzocyclooctadiene lignan, termed HDS2 that possessed anti-HIV activity against a wide variety of viral strains with EC50 values in the 1–3 µM range. HDS2 was shown to act as an NNRTI by qPCR and in vitro enzyme assays.
Conclusions
This compound provides a new scaffold for further optimization of activity through structure-guided design.
Introduction
Highly active anti-retroviral therapy (HAART) usually combines three or more different drugs that simultaneously target multiple stages of the HIV life cycle. To date, more than 25 Food and Drug Administration (FDA)-approved drugs are available for treatment of HIV patients. However, the emergence of drug resistance is still a major problem,1,2 as is the fact that many drugs have encountered significant dose-limiting and long-term toxicities. Thus, there is a need for the discovery and development of new agents that have novel structures or mechanisms of action different from currently approved drugs.
The discovery of new anti-HIV compounds largely depends on valid biochemical and cell-based assays 3 and a source of unique chemicals. High-throughput screening (HTS) has enabled the discovery of several HIV inhibitors, 4 but this approach usually requires large libraries of compounds, that increase costs. More recently, drug screens have moved away from large libraries of compounds to smaller focused collections (100–3000 compounds) that can be tested in biochemical or cell-based assays. 5
Recently, there has been renewed interest in natural products as sources for drug discovery 6 and natural products derived from medicinal plants have great potential as source of anti-HIV drugs that might possess activity against drug-resistant HIV strains.5,7–14 Plants of the Schisandraceae are a rich source of lignans and triterpenoids, that have given rise to compounds with anti-HBV, anti-HIV, and anti-tumor properties.8
Recently, cell-based phenotypic screening approaches have led to new drug discovery.15,16 A classic MT4 cell assay, that measures HIV-induced cytopathic effect (CPE), has been used successfully to identify new HIV inhibitors17,18 and can identify compounds that target any stage of the HIV life cycle. 3 Herein, we describe studies that have evaluated such a cell-based assay to study an in-house collection of natural products from plants of the Schisandraceae that possess unique structures. From a relatively small set of selected compounds, we first identified an interesting chemical scaffold containing the molecule dibenzocyclooctadiene lignan, also termed gomisin M1 (referred to hereafter as HDS2), that possessed high-level activity against a wide variety of HIV-1 strains as well as against HIV-2. We now also describe the mechanisms of action of this compound, both at a biochemical and molecular level, and show that it functions as a non-nucleoside reverse transcriptase inhibitor (NNRTI).
Materials and methods
Cells and reagents
TZM-bl, MT-2, MT-4 cells, the infectious molecular clones pNL4-3 and pBaL, the HIV-1 primary isolates HIV-2 (CBL-23), BK132 (X4), 92TH001 (R5), T-20, AMD3100, and VRC01 were all obtained through the NIH AIDS Research and Reference Reagent Program. The 293T cell line was obtained from the American Type Culture Collection (CRL-11268). TZM-bl and 293T cells were subcultured every 3–4 days in Dulbecco’s minimal essential medium (DMEM), while MT-2 and MT-4 cells were cultured in RPMI 1640 medium at 37°C and 5% CO2. Complete DMEM and RPMI 1640 media were supplemented with 10% fetal bovine serum (FBS), 2 mM
Raltegravir (RAL) was obtained from Merck-Frosst Canada (Kirkland, QC, Canada).
Measurements of viral p24 antigen levels and RT activity
p24 antigen levels in culture supernatants were assessed using a HIV-1 p24 antigen capture assay (ABL, Inc., Rockville, MD) according to the manufacturer's instructions and RT enzymatic activity in culture supernatants was measured using a tritiated thymidine triphosphate based assay as previously described. 19
HIV-1 RT in vitro enzyme assay
Recombinant wild-type HIV-1 reverse transcriptase (RT) and mutant RTs were expressed and purified as previously described. 19 RT enzymatic activity and kinetics studies were conducted as previously described by measuring the extension of Td100/Pd18 template-primer using a PicoGreen-based spectrophotometric assay. 20
Generation of replication-competent HIV-1 and virus production
Different replication-competent HIV-1 plasmids, i.e. pNL4.3INB(G140S), pNL4.3INB(E92Q), pNL4.3INB(Q148H), pNL4.3INB(N155H), and pNL4.3INB(G140S/Q148R), were generated from wild-type pNL4-3 using the Quick Change II XL site-directed mutagenesis kit 21 and were produced by transfecting 293T cells using calcium phosphate. Briefly, 2.5 × 106 293T cells were plated in a 10-cm dish containing 10 ml DMEM supplemented with 10% FBS. The next morning, medium was replaced with 8 ml fresh medium and in the afternoon cells were transfected with 10 µg of plasmid per cell culture dish. The transfection medium was replaced with 10 ml of fresh medium after 24 h. At 48 h posttransfection, cell-free supernatants were harvested and filtered through 0.45 µm filters, and aliquots were stored at −80°C. HIV-1 NL 4-3 was also produced in MT-4 cells infected with virus isolated from the transfections at a multiplicity of infection (MOI) of 0.001. Cell-free supernatants were harvested as viral stocks from infected MT-4 cell cultures at five days post-infection (pi).
Anti-HIV and cytotoxicity assays
Anti-HIV activity in MT-4 cell cultures was measured as described previously, 17 with minor modifications. The assay is based on HIV-induced CPE, which was assessed by a 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) cell viability assay. For the sake of convenience, each virus stock was titrated by measuring protection against CPE using a MT-4/MTT assay to determine the viral dose that might kill 95% of MT-4 cells by three days pi, defined as the cell culture cytotoxicity infective dose (CCCID95). Compounds were diluted in cell culture medium (2 × working solution) and placed into 96-well plates (50 µl/well). Twenty-five microliters of exponentially growing MT-4 cells (1 × 106/ml) were added to 96-well plates containing serial dilutions of the compound. Then, 25 µl of HIV-1 NL 4-3 diluted in medium at the CCCID95, usually equal to a final MOI of ∼0.03, were added to each well, except for wells with cells only or wells without drug and virus, that served as controls. Wells with dimethyl sulfoxide (DMSO) alone, without drugs, served as virus controls. Plates were incubated in a humidified incubator at 37°C under 5% CO2. After three days pi, when cell death in the virus control wells just reached >90%, as monitored by Trypan Blue staining, 10 µl of MTT reagent (5 mg/ml in PBS) was added to each well and incubation continued for 3 h at 37°C. Then, 110 µl of lysis buffer containing 10% (v/v) Triton X-100 in acidified isopropanol (4 ml HCl per 500 ml) were added to each well to lyse cells and solubilize the formazan crystal. The plates were gently shaked at room temperature overnight and were read at a wavelength of 570 nm using a FLUOStar Optima plate reader (BMG Labtech). Antiviral activity was determined by cell viability as measured by absorbance. In these experiments, the cell control (no HIV and no drug) was used as a positive control (set as 100%) and the virus control (without drug) was used as a negative control (i.e. background). The percentage of cell viability of cultures treated with drugs was calculated by comparison with the cell control. The concentration of a compound achieving 50% protection against virus-induced CPE, defined as the 50% effective concentration (EC50), was determined using Prism 5.0 software (GraphPad, San Diego, CA).
Anti-HIV activity was also evaluated in MT-2 cells by observing inhibition of syncytium formation and levels of RT activity and p24 in MT-2 cells. Anti-HIV activity was also examined in CBMC cell cultures as described previously. 19 Briefly, CBMCs were plated onto 96-well plates (1 × 106/well) in the presence of each compound. Cells were infected with the HIV-1 primary isolates BK132 (X4) or 92TH001 (R5) (NIH AIDS Reagent Program; www.aidsreagent.org) at an MOI of 0.02. After three days in culture, the cells were refreshed with media containing the corresponding drug. After seven days, the culture supernatants were collected and analyzed for RT activity. HIV infection in TZM-bl cells was conducted to investigate the anti-HIV activity of the compounds as previously described. 21 In brief, TZM-bl cells were plated at 2 × 104 cells per well in a 96-well plate in 100 μl of complete RPMI 1640 media. After 24 h, cells were treated with drugs and followed after infection with HIV-1 NL 4-3 or BaL. At 48 h pi, luciferase activity was measured using a Micro-Beta2 luminometer (PerkinElmer) using the luciferase assay system (Promega). The EC50 was calculated as the concentration of an inhibitor that reduced relative luciferase activity to 50% of that of control wells containing HIV-infected cells without an inhibitor.
For cytotoxicity assays, cells were separately evaluated in the presence of serial dilutions of compounds to determine the concentration of the compound killing 50% of cells, i.e. the 50% cytotoxic concentration (CC50), using the MTT assay. The EC50s and CC50s were calculated using Prism 5.0 software.
Time-of-addition assay
TZM-bl cells were plated onto 96-well plates as described above. After incubation overnight, cells were infected with HIV-1 NL 4-3 at a MOI of 0.05. Reference and test compounds were added at 5–50 times their EC50 at different time points, i.e. several hours after infection (0, 2, 4, 6, 8, 12 and 24 h). Luciferase activities were measured at day 2 after infection.
Combination antiviral activity assay in MT-4 cells
For combination testing, HDS2 was serially diluted into 96-well master assay plates in complete RPMI 1640 medium. Representative approved anti-HIV drugs were diluted horizontally across separate master assay plates. Combinations of aliquots from both the horizontally and vertically diluted master plates were arranged in checkerboard-style dilutions into daughter plates. Anti-HIV activities were tested in the MT-4/MTT assay as described previously, and the interaction of compound combinations was analyzed by a combination index (CI) using CompuSyn software as described, 22 i.e. CI < 1 synergy, CI = 1–1.2 additivism, and CI > 1.2 antagonism.
Quantitative PCR analysis of viral DNA
HIV-1 early and late reverse transcripts were measured by qPCR as previously described. 23 Briefly, MT-4 cells (3.5 × 106 cells/well in a 24-well plate) were plated in the presence of DMSO or drugs. AZT (1 µM) and RAL (0.5 µM) were used as references as RT or integrase inhibitors. AMD3100 (0.25 µg/ml) was used as reference as entry inhibitors. Cells were infected with HIV-1 NL 4-3 at a MOI of 0.05. The viruses used in this experiment were produced in MT-4 cells. After 24 h pi, genomic DNA was extracted using the DNeasy Blood and Tissue Kit (Qiagen, Mississauga, ON, Canada) and was measured by a NanoDrop 1000 spectrophotometer (Thermo Fisher, Wilmington, DE). Quantification of HIV-1 reverse transcripts was performed on a Corbett Rotor-Gene 6000 thermocycler in a 25 µl reaction containing 70 ng of DNA, qPCR Supermix UDG (Invitrogen), and 0.16 µM each of primers and probes. The primers and probe for the early reverse transcripts were: strong-stop F: 5′-AACTAGGGAACCCACTGCTTAA-3′, strong-stop R: 5′-TGAGGGATCTCTAGTTACCAGAGTCA-3′, strong-stop probe: 5′-[FAM]-CCTCAATAAAGCTTGCCTTGAGTGCTTCAA-BHQ1-3′. The primers and probe used for the late reverse transcripts were primers: full-length F: 5′-AACTAGGGAACCCACTGCTTAA-3′, full-length R: 5′-CGAGTCCTGCGTCGAGAGA-3′, full-length probe: 5′-[FAM]-CCTCAATAAAGCTTGCCTTGAGTGCTTCAA-BHQ1-3′ (Biosearch Technologies, Novato, CA). The primers and probe used for GAPDH were: GAPDH F: 5′-ACCGGGAAGGAAATGAATGG-3′, GAPDH R: 5′-GCAGGAGCGCAGGGTTAGT-3′, GAPDH probe: 5′-(VIC) ACCGGCAGGCTTTCCTAACGGCT-3′. The cycling conditions were 95°C for 10 min, and 50 cycles at 95°C for 15 s and 60°C for 60 s. All samples were quantified against standards of HIV-1 NL 4-3 plasmid for late RT product and DNA from uninfected cells for GAPDH. The copy numbers per µg DNA for each sample were normalized by their GAPDH contents.
Determination of anti-HIV activity post-attachment
Inhibition of HIV-1 infection at times post-attachment was measured as described previously. 24 TZM-bl cells were plated in 96-well plates. For studies after-attachment, cells were spin-infected with HIV-1 NL 4-3 virus at 1250 × g for 2 h at 16°C. For pre-attachment (i.e. standard infection), cells were spun under the same conditions, but without infection by virus. The cells were placed on ice and supernatants and/or unbound virus were removed and cells were washed two times with PBS containing 0.2% bovine serum albumin. For post-attachment studies, cell culture media containing inhibitors (T-20, VRC01 or HDS2) were added to the cells and incubated for 30 min on ice. For pre-attachment studies, viruses were pretreated with the same inhibitors on ice for 30 min and were then added to the cells. The inhibitors VRC01, a monoclonal antibody targeting the CD4-binding site of HIV-1 gp120, and T-20, a fusion inhibitor, were used as references. Cells were then moved to a 37°C incubator, and luciferase activity was measured two days later. These results were then compared to determine whether there were shifts in anti-HIV activities in terms of pre- and post-cellular attachment.
Data analysis
Data were analyzed using Prism 5.0 and expressed as means ± standard deviation (SD). For each assay, three or more independent experiments, each in duplicate or triplicate, were performed, and the results were analyzed.
Results
Identification of HIV inhibitors from natural products
Since some natural products isolated from medicinal plants of the Schisandraceae have been shown to possess anti-HIV activity,8,25 we screened a collection of 60 compounds for their anti-HIV activity using the MT-4/MTT assay. This effort resulted in identification of 14 compounds with various degrees of anti-HIV activities and these were ranked on the basis of antiviral potency and toxicity. The most potent compound HDS2 (Figure 1(a)), a dibenzocyclooctadiene lignan previously termed gomisin M1, possessed good characteristics (MW ≤ 400, rings ≤ 4, hydrogen-bond donors ≤5, hydrogen-bond acceptors ≤9, and log P 1.9–5.5) and can serve as a starting point for further chemical modifications.
26
Biological evaluation of this compound and studies on its molecular mechanisms of action in cell culture and in biochemical assays were initiated.
Anti-HIV activity of HDS2. (a) Chemical structure of HDS2; (b) dose–response curve of HDS2 in MT-4 cells infected with the NL 4-3 strain, measured by MTT; (c) inhibition of RT activity in MT-2 cells infected with the NL 4-3 strain; (d) inhibition of p24 levels in MT-2 cells; (e) dose–response curve of HDS2 in TZM-bl cells infected with the NL 4-3 strain; (f) dose–response curve of HDS2 in TZM-bl cells infected with the BaL strain. Data (b–f) represent mean ± SD calculated from two or three independent experiments with duplicate samples and expressed as the relative percentage of the DMSO control (set at 100%).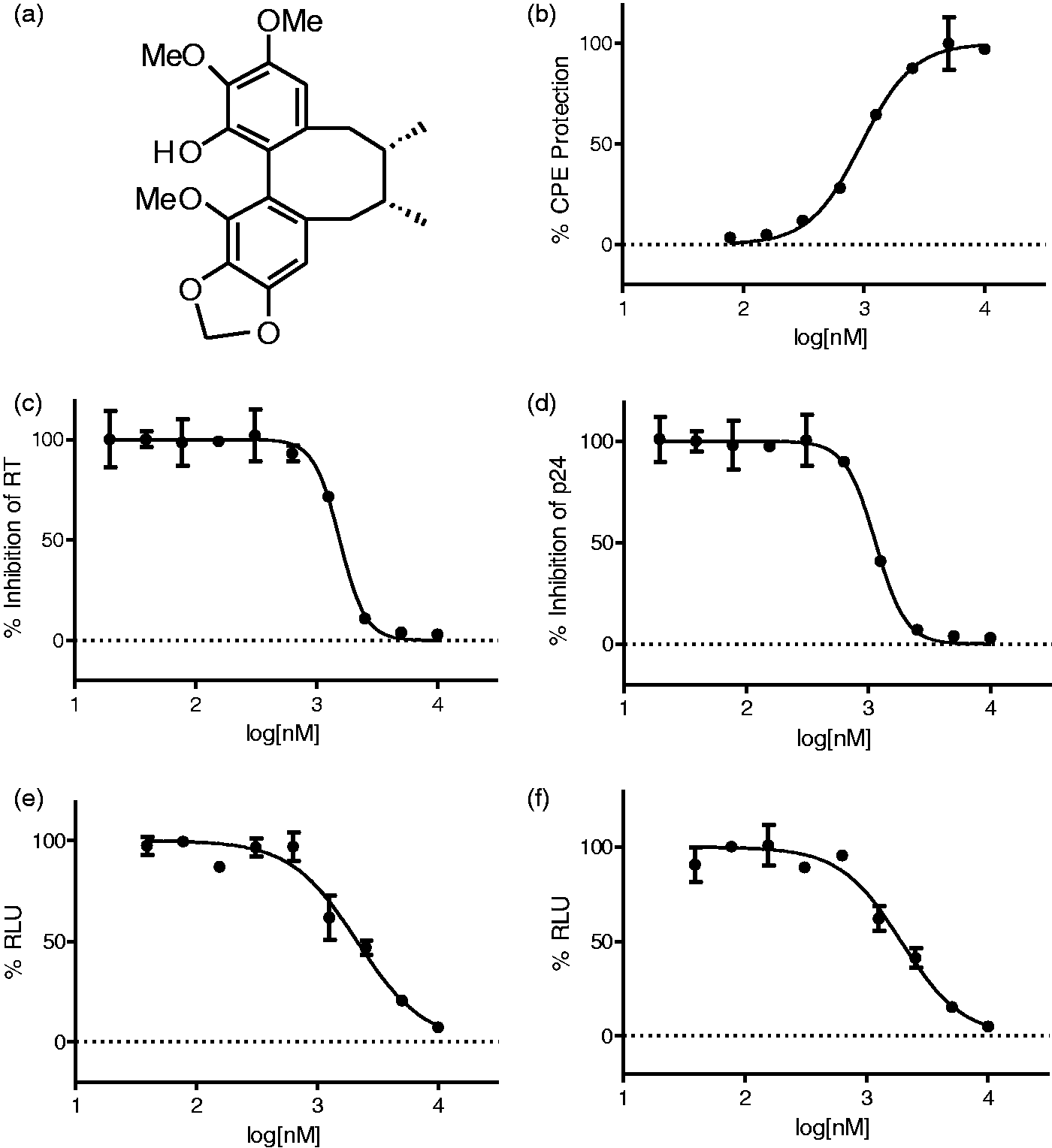
Anti-retroviral activity profile of HDS2
Antiretroviral activity of HDS2 a against laboratory strains and primary isolates in different cells.
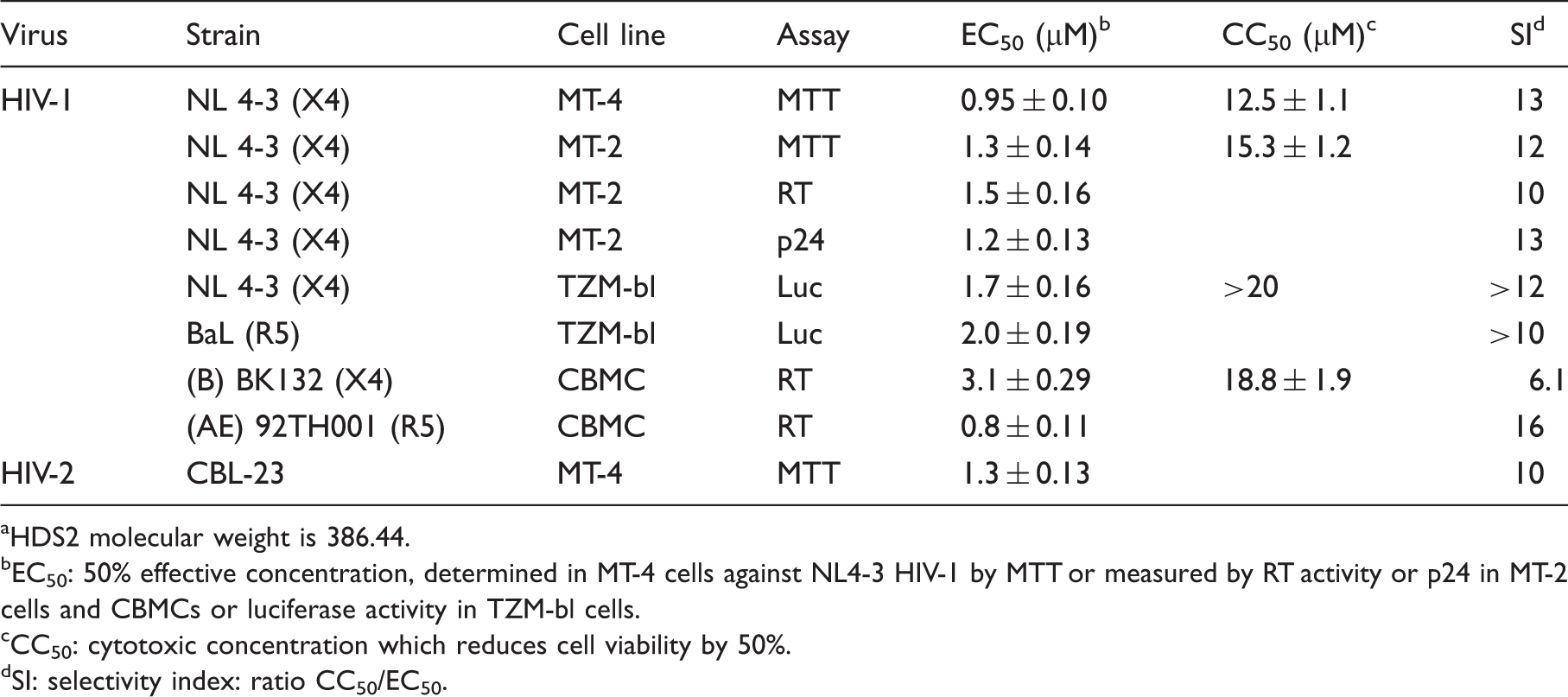
HDS2 molecular weight is 386.44.
EC50: 50% effective concentration, determined in MT-4 cells against NL4-3 HIV-1 by MTT or measured by RT activity or p24 in MT-2 cells and CBMCs or luciferase activity in TZM-bl cells.
CC50: cytotoxic concentration which reduces cell viability by 50%.
SI: selectivity index: ratio CC50/EC50.
Cross-resistance profile of HDS2
Resistance profile of HDS2 against a panel of HIV-1 drug-resistant recombinant viruses and primary isolates. a
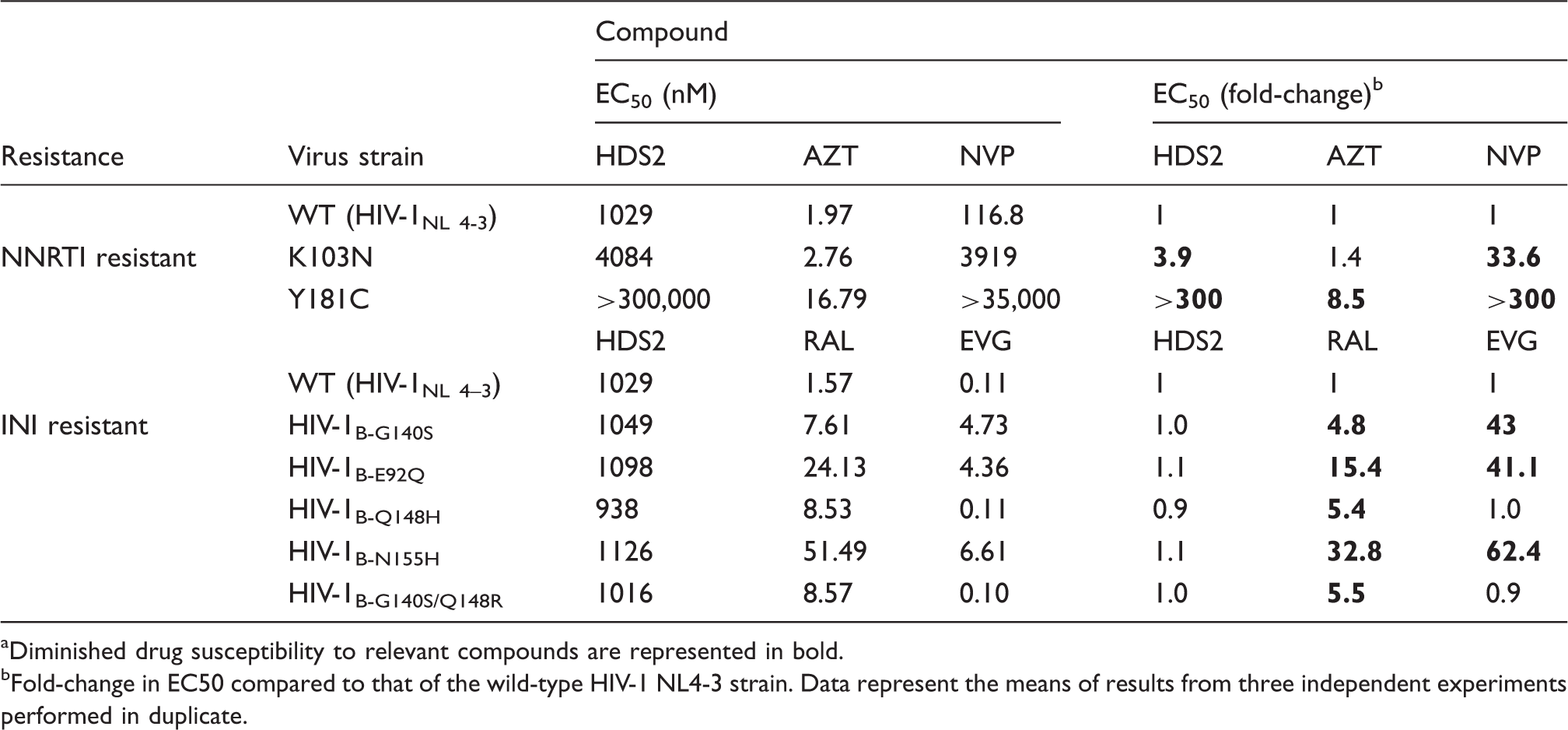
Diminished drug susceptibility to relevant compounds are represented in bold.
Fold-change in EC50 compared to that of the wild-type HIV-1 NL4-3 strain. Data represent the means of results from three independent experiments performed in duplicate.
Cellular combination studies
Combination studies of HDS2 with approved anti-HIV drugs using the MT-4/MTT assay.
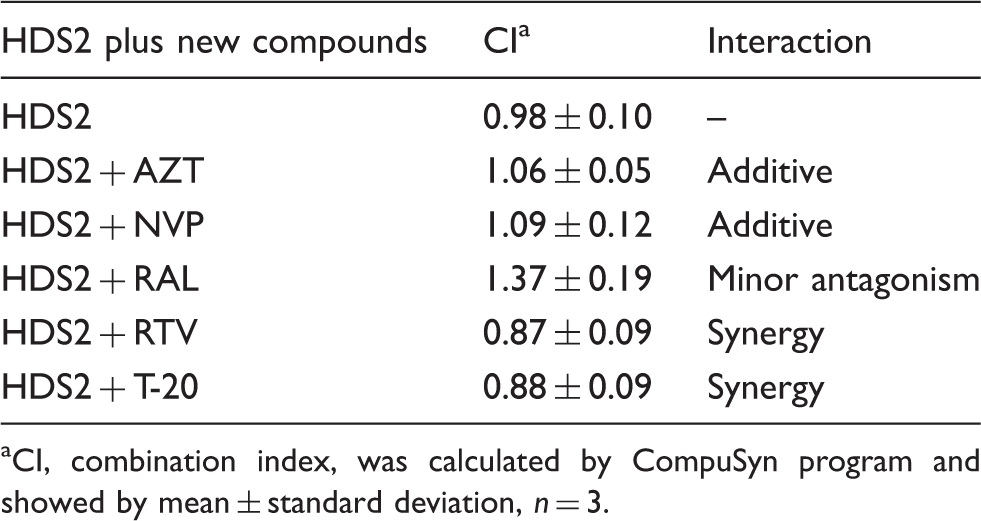
CI, combination index, was calculated by CompuSyn program and showed by mean ± standard deviation, n = 3.
Mechanism of action
In an initial effort to determine the stage of HIV-1 replication that was targeted, HDS2 was evaluated in a single-round of HIV-1 replication using TZM-bl cells, which can distinguish between early versus late events in the HIV replication cycle. 27 As shown in Figure 1(e) and (f), HDS2 exerted antiviral activity against both X4 and R5-tropic viruses in this assay, with similar EC50s being obtained as in the MT-4/MTT assay (Table 1), showing that HDS2 targets an early step in HIV-1 replication (between entry and integration).
Next, a time-of-addition assay was performed in which several different classes of drugs were used as reference compounds, including AZT (a NRTI), RAL (an integrase inhibitor), and T-20 (a fusion inhibitor). Inhibition of HIV-1 replication by HDS2 was significantly decreased when it was added at 4 h pi (Figure 2), again suggesting that it blocks an early step prior to reverse transcription. In contrast, pretreatment of HIV-1 virions with HDS2 at 5 µM for 24 h did not affect viral infectiousness (data not shown).
Time-of-addition analysis. TZM-bl cells were infected with HIV-1 NL 4-3, and test compounds were added at different times (0, 2, 4, 6, 8, 12, 24 h) at or after infection. Final compound concentrations were 5- to 100-fold higher than their EC50s: AZT (1 μM), RAL (0.5 μM), T-20 (2 µg/ml), and HDS2 (5 μM). The percent relative light unit (RLU) relative to the control was determined in TZM-bl assay.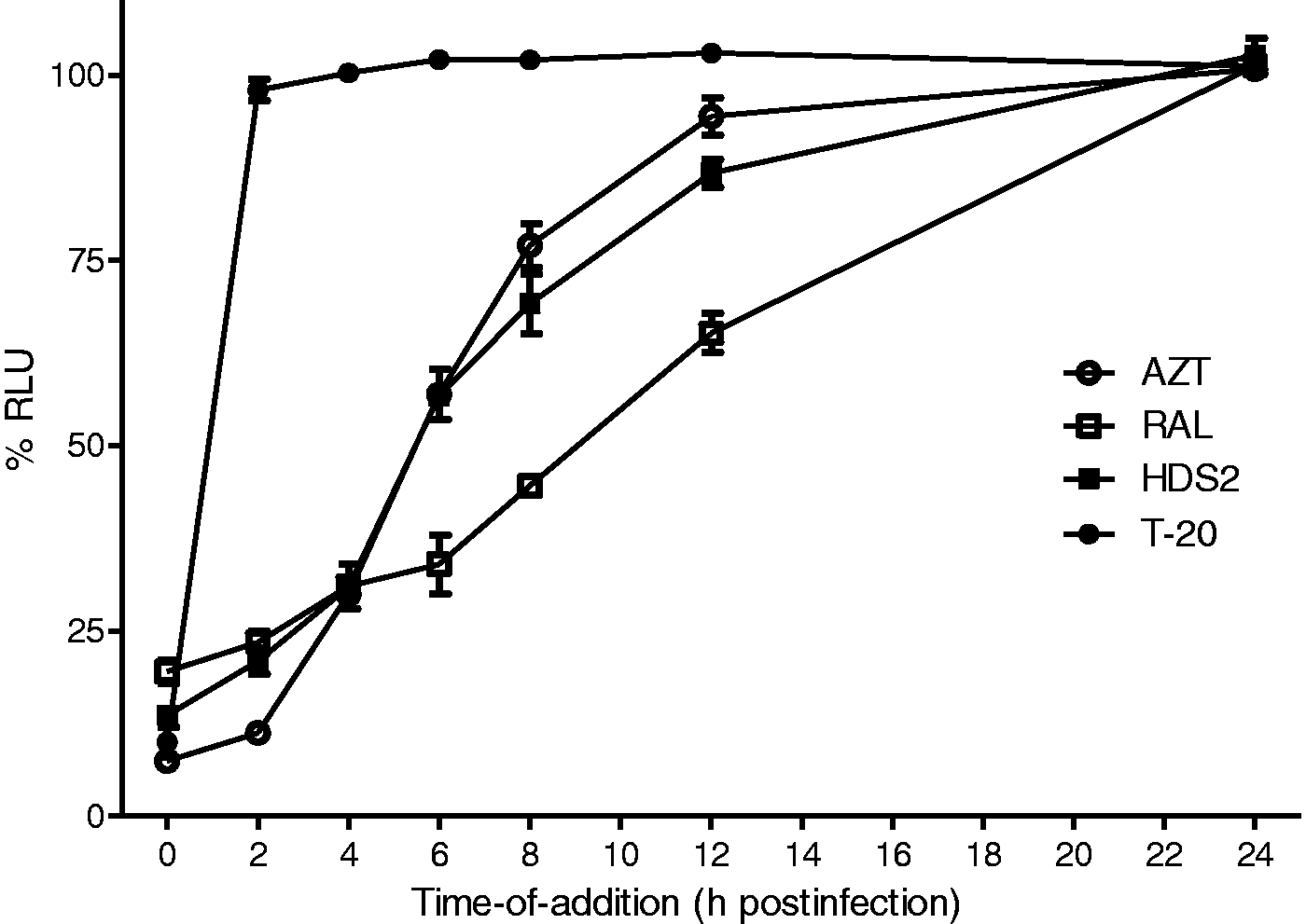
HDS2 was then evaluated for ability to inhibit HIV-1 infection at times post-attachment.
24
In such assays, VRC01,
28
a CD4 binding antibody, and T-20, a fusion inhibitor, were used as controls. As expected, a large decrease in the inhibitory activity of the HIV-1 attachment inhibitor VRC01 was observed in the post-attachment assay (Figure 3(a) and Table 4), whereas the bound virus remained completely susceptible to the fusion inhibitor T-20 (Figure 3(b) and Table 4). HDS2 also completely blocked HIV-1 infection in both the pre- and post-attachment assays (Figure 3(c) and Table 4), suggesting that it acts at a later step than virus attachment to CD4.
HDS2 primarily inhibits a step after HIV attachment to the host cells. TZM-bl cells were spin-infected at 16°C, a temperature that allows for attachment of virus but does not allow fusion events to occur. The unbound virus was removed, and the cells were incubated in media containing inhibitors (a) VRC01; (b) T-20; (c) HDS2 on ice for 30 min, and then the plates were transferred to 37°C, which allows for fusion and infection to be completed (referred to as post-attachment, ○). The results were compared with a standard infection procedure (pre-attachment), in which the virus and inhibitors were incubated together on ice for 30 min and then added to TZM-bl cells and incubated at 37°C (referred to as pre-attachment, •). Summary of the calculated EC50s of compounds in either pre- or post-attachment assays.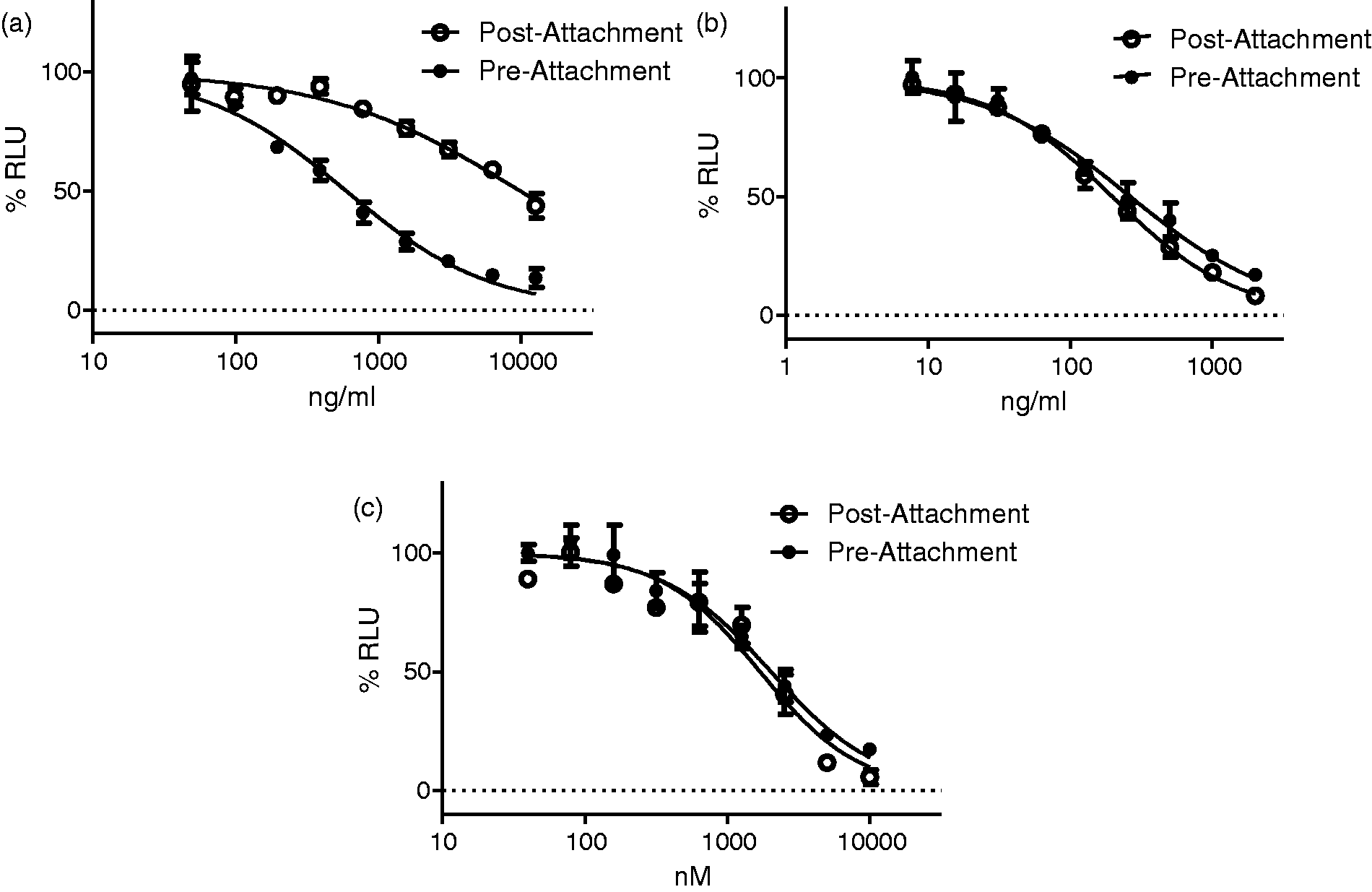

Next, a quantitative real-time PCR was used to show that treatment with HDS2 blocked the synthesis of both early and late viral reverse transcription products in MT-4 cells (Figure 4(a)). In control experiments, AMD3100, the entry inhibitors, blocked both early and late reverse transcription products, AZT largely blocked late reverse transcription product, whereas RAL did not affected both early and late reverse transcription products. Since strong-stop DNA can be detected soon after viral entry but just before viral uncoating takes place,
24
the results demonstrate that HDS2 inhibits a step in HIV-1 infection prior to the completion of reverse transcription but do not exclude the possibility that HDS2 could be an inhibitor of HIV-1 RT.
Effects of HDS2 on HIV-1 reverse transcription. (a) qPCR analysis of the early and late RT products in MT-4 cells. Cells were infected with HIV-1 NL 4-3 in the presence of DMSO or various drugs. DNA was extracted at 24 h post-infection and early and late reverse transcription products were quantified by qPCR. AZT (1 μM), RAL (0.5 μM), and AMD3100 (0.25 µg/ml) were used as reference compounds. (b) Effect of HDS2 on HIV-1 RT activity in vitro, determined by incorporation of dTTP. NVP was used as a reference and the DMSO was as a control (set at 100%). (c) Lineweaver-Burk plot analysis of purified recombinant HIV-1 RT inhibition by HDS2. (d) HDS2 is a non-competitive NNRTI. Inhibition curves were calculated for HDS2 and NVP in an HIV-1 RT assay with 1, 2, 4, 6, 8, and 10 μM dNTP. The fold-change in IC50 observed with increasing substrate concentration (compared to the IC50 measured at 1 μM) is shown. (e) Inhibitory effects of HDS2 on recombinant wild-type and mutant HIV-1 RTs.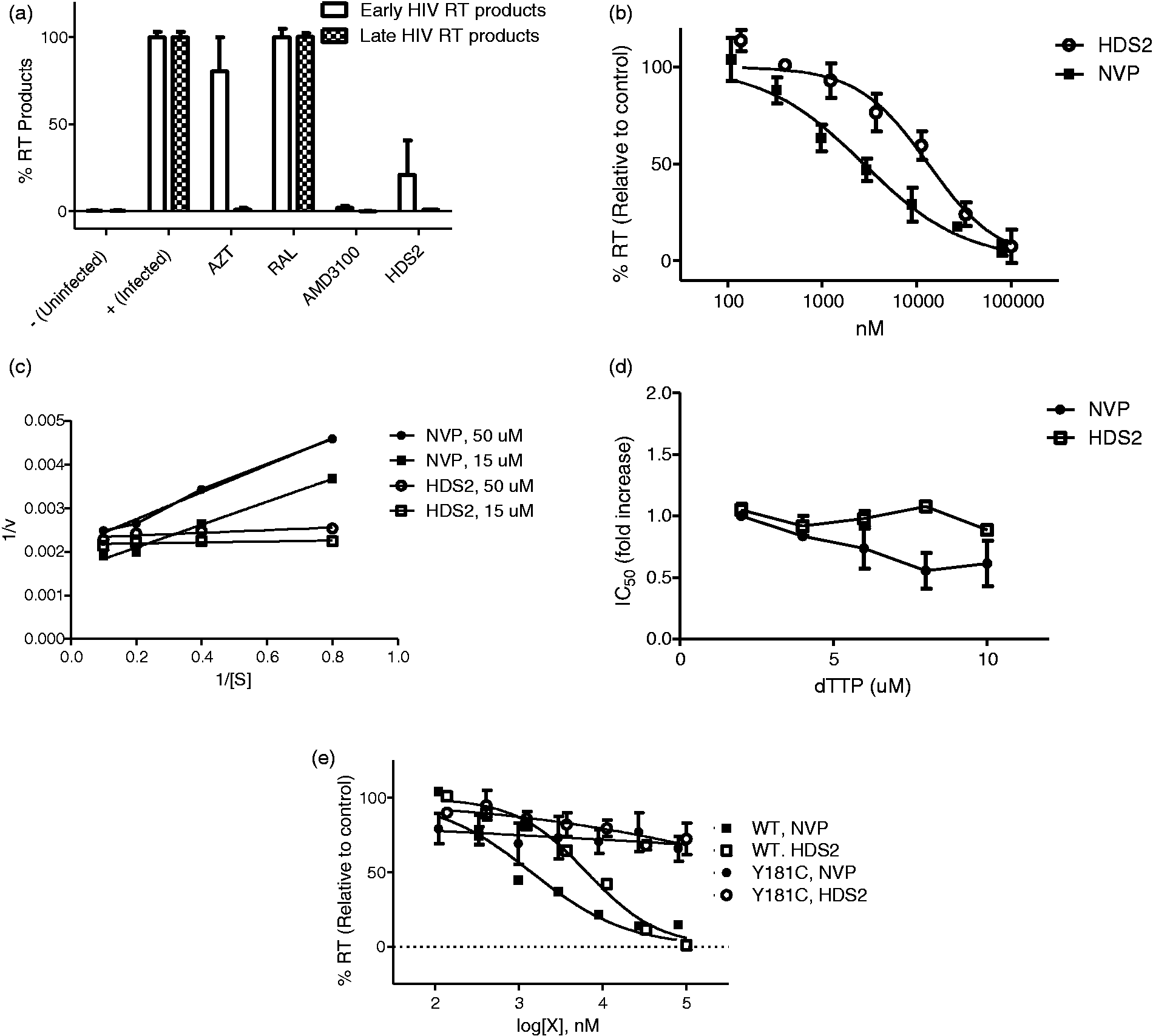
We then tested the effect of HDS2 on recombinant purified HIV-1 RT activity in a biochemical assay. In these experiments, NVP was used as a reference. As shown in Figure 4(b), HDS2 inhibited RT activity in a dose-dependent manner with an IC50 of 17.25 ± 1.8 µM. In addition, kinetic studies showed that HDS2 had a similar profile to NVP and appeared to be non-competitive with regard to the substrate (Figure 4(c)). The results also showed that an increase in dNTP concentration from 1 to 10 µM in the RT primer extension assay did not lead to an increase in the IC50 of HDS2 (Figure 4(d)). Biochemical analysis of wild-type and mutated Y181C containing RTs showed that the IC50 of HDS2 for the mutated Y181C RT was similar to that for NVP, i.e. about 300-fold higher than for wild-type (Figure 4(e)). Taken together, these results suggest that HDS2 acts as a non-competitive NNRTI. The result that the IC50 obtained biochemically for HDS2 was higher than in cellular assays is consistent with previous results for NNRTIs. 29
Discussion
In this study, we report the identification and characterization of a new HIV inhibitor that has broad spectrum activity against wild-type HIV-1 and HIV-2 as well as against a variety of HIV-1 strains of varying tropism (Figure 1 and Table 1). This class of compound contains a dibenzocyclooctadiene lignan core structure.
The mechanism of action of HDS2 seems to be that of an NNRTI. First, the results of studies in TZM-bl indicator cells showed that HDS2 exerted inhibitory activity against both NL 4-3 and BaL (Figure 1(e) and (f)), suggesting that it targets an early step in the HIV life cycle. Second, time-of-addition studies showed that the activity of HDS2 was significantly reduced when it was added at 4 h post-HIV-1 infection (Figure 2). Third, quantitative real-time PCR demonstrated that HDS2 blocks both early and late HIV-1 reverse transcription products (Figure 4(a)). Finally, biochemical assays demonstrated that HDS2 acts as a non-competitive NNRTI (Figure 4(b) to (e)).
It has been reported that different types of plant lignans can exhibit anti-HIV activity.10,25,30–33 Some of these lignans can inhibit HIV-1 RT, such as anolignan 34 and tetrahydronaphthalene lignan. 35 However, the structures of these compounds are different from the dibenzocyclooctadiene skeleton and further structure–activity studies are needed to determine the nature of the pharmacophore that might be modified to further improve potency.
It may be unique that HDS2 possesses activity against HIV-2 as well as against HIV-1 and we cannot exclude that the inhibition of HIV-2 by HDS2 might be through inhibition of virus entry. This notion is supported by its inhibition of post-attachment entry event (Figure 3(c)) and early reverse transcription product, as determined by qPCR assay (Figure 4(a)). Recent in vitro studies have shown that HIV-2 is susceptible to entry inhibitors. 36 It was also reported that a few derivatives of SJ-3366, a potent NNRTI, have the unique feature of displaying a dual mechanism of action against HIV-1 and HIV-2, both as NNRTI and entry inhibitors. 36
Although HDS2 does not possess as low an IC50 as some other NNRTI molecules, its discovery now provides a novel scaffold as a starting point for further optimization of activity through structure-guided design. Our results also demonstrate that different chemical classes of HIV-1 inhibitors can be identified from a small chemical library containing selected natural products with unique structures. Further work will (i) use the cell-based screening system with other chemical libraries and (ii) optimize HDS2 for improved potency and toxicity, based on structure–activity studies. This will hopefully lead to the identification of more potent HIV-1 inhibitors in future studies.
Footnotes
Declaration of conflicting interests
The authors declare no competing interests.
Funding
This work was supported by the Canadian Institutes for Health Research (CIHR). A portion of this project was financially supported by a CAS grant (KSCX2-EW-Q-10) and by the Natural Science Foundation of China (grant 81373290). The content is solely the responsibility of the authors and does not necessarily represent the official views of the CIHR or the NSFC.
