Abstract
Purpose
Creating an effective MRI protocol for examining the brachial plexus poses significant challenges, and despite the abundance of protocols in the literature, there is a lack of reference standards for basic sequences and essential parameters needed for replication. The aim of this study is to establish a reproducible 1.5 T brachial plexus imaging protocol, including patient positioning, coil selection, imaging planes, and essential sequence parameters.
Methods
We systematically investigated MRI sequences, testing each parameter through in vivo experiments, examining their effects on signal-to-noise ratio (SNR), contrast-to-noise ratio (CNR), visual quality scores, and acquisition time. Sequences were refined based on optimal quality and timing scores. The final protocol was tested on scanners from two other vendors for reliability.
Results
The final protocol included a combination of 2D turbo-spin-echo and 3D SPACE T1, SPACE STIR, and VIBE DIXON sequences. Recommendations for imaging planes, phase encoding, field of view, TR, TE, resolution, number of slices, slice thickness, fat and blood suppression, and acceleration strategies are provided. The protocol was successfully translated to other vendor's scanners with comparable quality.
Conclusion
We present an optimized protocol detailing the essential parameters for reproducibility. Our comprehensive list of experiments describes the impact of each parameter on image quality and scan time, addressing common artifacts and potential solutions. This protocol can benefit both young radiologists new to the field and experienced professionals seeking to refine their existing protocols.
Introduction
Imaging the brachial plexus is a complex and challenging task due to several factors. The inhomogeneity in the main magnetic field caused by air-neck-shoulder interfaces, breathing and pulsating artifacts from the proximity of the lungs and the vessels of the neck, and the requirement for fast 3D isotropic sequences with a large field of view make obtaining high-quality images difficult. Furthermore, the intricate and minute anatomical structures of the brachial plexus necessitate high-resolution images for accurate visualization and assessment, and achieving homogeneous fat and vessel suppression is crucial for accurate diagnosis. Consequently, numerous sequences and planes are required for the examination, which typically results in lengthy scan times.
Despite the extensive literature available on various protocols for brachial plexus imaging, there is a surprising degree of variability in the parameters used, leading to fragmentation and difficulties in reproducing the presented protocols.1–3 Even basic parameters display consistent variations, such as TR in SPACE STIR, a commonly employed sequence for achieving isolated neurography, with reported values ranging from 1500 to 4500,4–10 in-plane resolution for 2D TSE varying from 0.6 to 1.3 mm, or slice thickness ranging from 3 to 5 mm, even when using 3T scanners.9,11–14 These discrepancies can significantly impact image quality and scan time. Moreover, essential parameters for accurately reproducing a sequence, such as phase encoding (PE) direction, oversampling, concatenations, real and reconstructed matrices, bandwidth, turbo factor or acceleration techniques, are often omitted from published protocols.
The purpose of this study is to develop a comprehensive and accessible MRI protocol for brachial plexus imaging, specifically tailored for 1.5 Tesla scanners, with an emphasis on compatibility and reproducibility. We carried out a systematic investigation of various sequences for brachial plexus imaging, conducted in vivo experiments, addressed common technical challenges, and provided a detailed table with all the essential parameters needed to replicate the protocol, along with advice on patient positioning, coil selection, and imaging planes. For each sequence type, we assessed the impact of different parameters on signal-to-noise ratio, contrast-to-noise ratio, visual quality scores, and acquisition time, allowing readers to effectively optimize their own protocols. Additionally, we demonstrated the reproducibility of our protocol using scanners from two distinct manufacturers.
Materials
Subjects
This study adhered to HIPAA guidelines and received approval from the local ethics committee. All participants gave written informed consent. Imaging was conducted on 12 healthy subjects (aged 27–51 years) with no significant clinical history or symptoms. The same subject (M.T.) underwent testing with the finalized protocol on scanners from two other manufacturers. Due to ethical considerations, contrast agents were not administered to these healthy volunteers; therefore three patients were included in the study. We specifically selected patients with non-significant pathologic changes of the brachial plexus in order to minimize any confounding factors that could arise from pathological anomalies. These patients underwent a complete examination of the brachial plexus, and their images were used solely to demonstrate the superiority of contrast over other non-contrast blood suppression methods.
Scanner, coils and positioning
Experiments were conducted on a Siemens Avanto Fit 1.5T scanner with a gradient amplitude of 45 mT/m/ms and a slew rate of 200 mT/m. The final protocol was tested on two scanners from other manufacturers: a Philips Ingenia Ambition (gradient amplitude 45 mT/m/ms and slew rate 200 mT/m) and a GE Signa Voyager AIR Edition (amplitude 45 mT/m and slew rate 200 T/m/s). Subjects were positioned supine with their heads entering the scanner first. Pads were placed beneath the knees and legs to alleviate back tension and enhance comfort. Although some authors recommend using pads, sandbags, or pillows around the neck to decrease field inhomogeneity due to air,
15
we opted against them to minimize patient discomfort and simplify the technologists’ tasks. A head/neck coil with both anterior and posterior elements was used, along with a body matrix coil and integrated spine coils. The body coil was positioned at the lower border of the head/neck coil, encompassing the neck, shoulder, and arms (see Figure 1). Positioning of the patient. Comparison of the Siemens (a), GE (b) and Philips (c) configurations. The patient lies supine with the head first entering the scanner. We use a head/neck coil with both anterior and posterior elements embedded with a body coil and the spine coil elements. A cushion was placed under the legs to relieve back tension.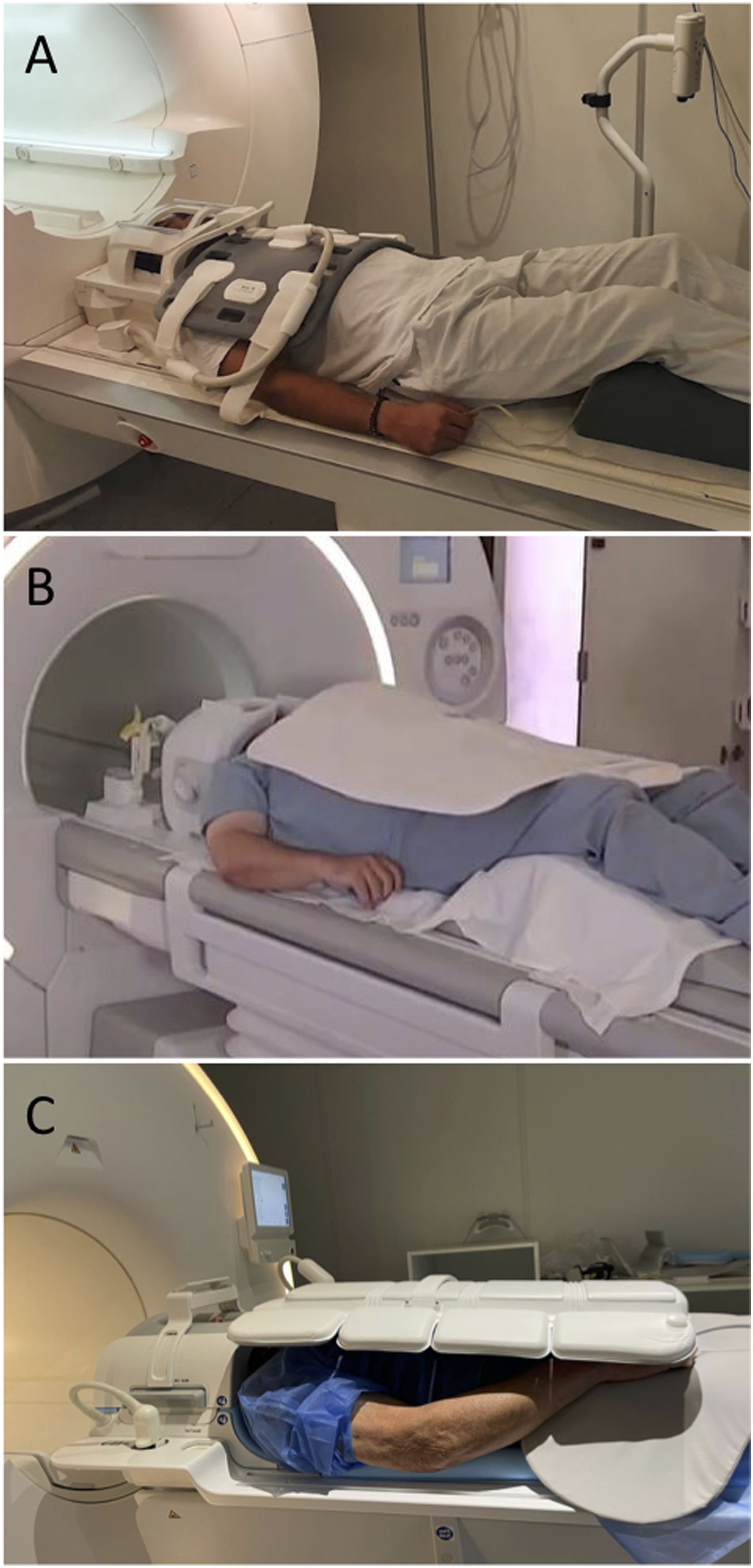
Imaging protocols
We specifically investigated widely available, commonly reported sequences for 1.5 Tesla scanners, including 2D turbo spin echo sequences, SPACE sequence (myelography, T1, and STIR), and T1 VIBE. Starting from the manufacturer’s basic preset, we acquired each sequence multiple times, modifying one parameter at a time and monitoring the changes in image quality and acquisition time, as detailed in the following section.
For 2D imaging, we examined STIR, T1, and T2-weighted sequences in axial, coronal, and sagittal planes. We determined the number of slices required to cover the relevant anatomical volume, with upper and lower limits of C1 and D6, respectively, and lateral limits including the deltoids, and the pectoralis muscles and upper back muscles anteriorly and posteriorly. Sagittal sequences were acquired unilaterally, extending from the spine to the axilla. Consequently, we tested different slice thicknesses and adjusted the number of slices accordingly. We also assessed different PE directions, TR, TE, in-plane resolutions, acceleration strategies (averages, iPAT), and fat suppression techniques (spectral fat saturation pulse, water excitation, SPAIR, and Dixon).
For 3D SPACE sequences (STIR-, T1-, and T2-SPACE), we evaluated coronal and axial planes of acquisition, along with different PE directions. For T2-SPACE Myelo, we used a smaller field of view, a coronal plane, and focused on the cervical spine. Excitation pulses were always performed with slab selection mode to reduce the need for oversampling (and acquisition time) in at least one dimension. In the coronal plane with a lateral PE direction, we acquired SPACE with varying TR, TE, and voxel sizes, fat saturation strategies (conventional fat saturation, water excitation, SPAIR), and acceleration strategies (iPAT 1, 2, and Caipirinha 4). For STIR-SPACE, we tested blood suppression, including blood suppression gradients and post-contrast acquisition with different inversion times (160 and 220).
VIBE imaging was acquired on axial and coronal planes with isotropic voxels, utilizing Dixon fat suppression. We assessed different voxel sizes, flip angles (FA), averages, and acceleration strategies.
Image quality and scan time assessment
Two radiologists (with more than 15 and 5 years of experience in the field, respectively) evaluated image quality in terms of signal-to-noise ratio (SNR), contrast-to-noise ratio (CNR), qualitative visual scales, and acquisition time. The less experienced radiologist drew circle-shaped regions of interest (ROIs) containing only air-voxels at one corner of the image (at least 50 cm2). Signal ROIs were drawn as circles with 8–15 mm2 areas, positioned on the radicular post-ganglionic tracts of the C6 and C7 nerves bilaterally, totaling four ROIs, and reporting their average. 16 For muscles, one ROI was placed on the left supraspinatus. CNR and SNR were calculated using the following formulas: SNR = (Signal ROI/SD of the noise); CNR = (SNR(muscle)−SNR(nerves))/SD noise.
Drawing on methods from previous studies,15,16 the delineation of relevant anatomical structures, such as spinal cord, nerve roots, trunks, chords, and main branches, was scored using a five-point grading system: 1, non-existent; 2, poor; 3, acceptable; 4, good; 5, excellent. The impact of artifacts on image quality and interpretation was assessed using a five-point grading system: 5, absent; 4, minimal; 3, mild; 2, moderate; 1, severe/limiting diagnostic value. Specific types of artifacts (e.g., motion, susceptibility, pulsation, ringing, etc.) were not recorded. Acquisition time for individual sequences was scored on a five-point scale: for 2D and VIBE imaging: 1, unacceptable (more than 5 min); 2, poor (4–5 min); 3, acceptable (3–4 min); 4, good (2:30–3 min); 5, excellent (less than 2:30 min). For 3D SPACE, acquisition time was scored as follows: 1 (more than 7 min); 2 (6–7 min); 3 (5–6 min); 4 (4–5 min); 5 (less than 4 min).
Cohen’s Kappa coefficients were utilized to measure the agreement between the two observers concerning image quality. A value of 0 indicates poor agreement; 0.01–0.20, slight agreement; 0.21–0.40, fair agreement; 0.41–0.60, moderate agreement; 0.61–0.80, good agreement; and 0.81–1.00, excellent agreement. 17
Protocol optimization
In Figure 2 (graphs), we have illustrated the trends of SNR, CNR, visual scales, and acquisition time concerning some parameter under investigation. This allows for an easier visual evaluation of these trends. The ideal protocol was identified by selecting points on the trend that exhibited optimal SNR, CNR, visual scales, and time efficiency, as denoted by the dotted vertical line. We retained the parameters with the highest scores in a progressive manner, beginning with the imaging planes, followed by the PE direction, TR, TE, resolution, and finally, acceleration strategies. After establishing these parameters, we optimized the acquisition time by fine-tuning the bandwidth, echo train length, and concatenations, adjusting the TR to the closest values obtained from the experiments. Graphs displaying the impact of various parameters on image quality and acquisition time (AT). The gray dotted lines indicate the highest scores on the scales, assisting in selecting the optimal parameter values. The graphs show trends for STIR 2D and SPACE STIR sequences and demonstrate how different factors, such as TR, TE, resolution, slice thickness, and acceleration strategies, affect SNR, CNR, visual scale, artifact scale, and AT. The best trade-offs and recommendations are highlighted for each parameter to achieve optimal image quality and time efficiency. The complete list of experiments can be found on the supplementary material.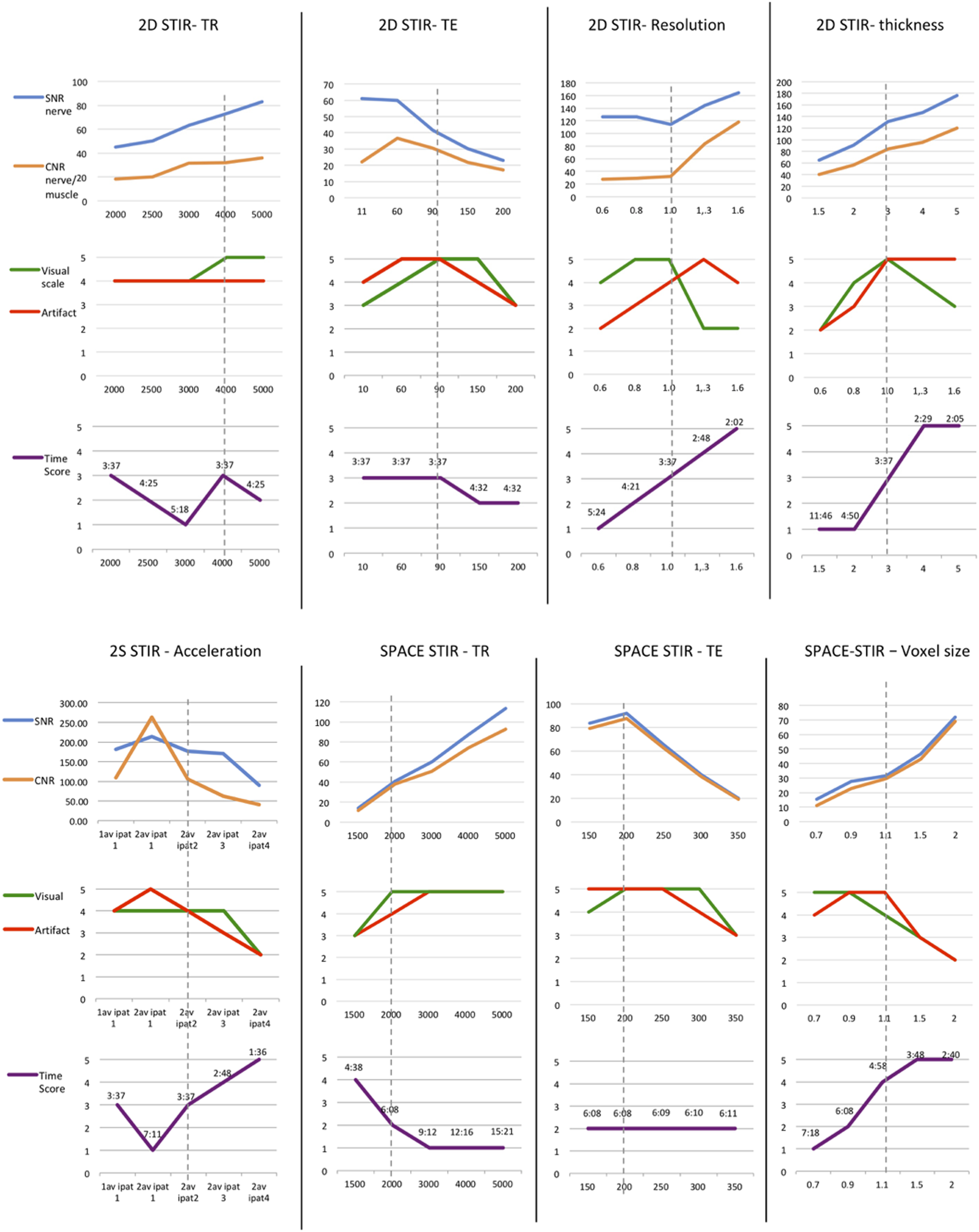
Results
Recommended protocol considerations
The Recommended Protocol Inclues a Standard Examination of the Cervical Spine and a Pre-/Post-contrastographic Study. 2D Sagittal sequences are Optional Because can be Easily Reconstructed From 3D imaging. T2 SPACE may Optionally Substitute 2D T2-Weighted sequences.
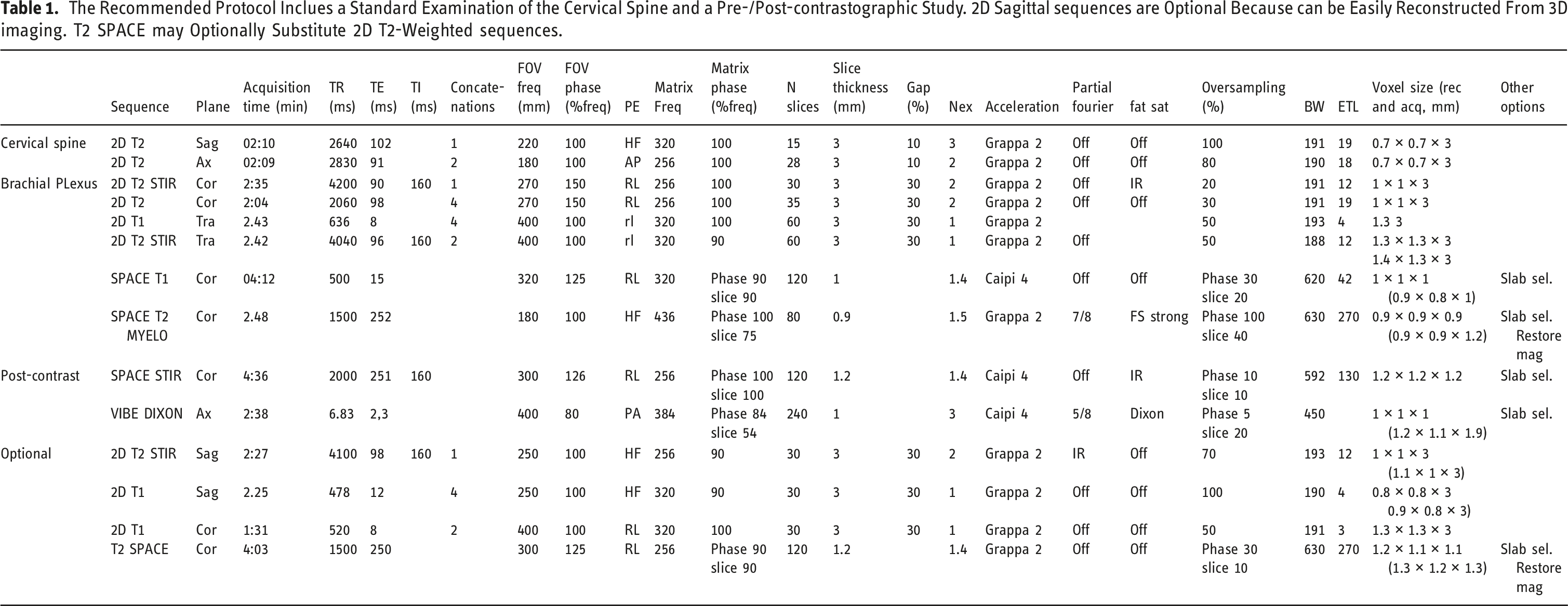
Cohen’s Kappa agreement between the two readers was excellent (0.827, 95% CI 0.783 to 0.870). All the results in detail are reported in the Supplementary Materials.
2D imaging
In the coronal plane, lateral PE direction yielded superior scores, whereas feet-head PE direction exhibited vertical propagation of respiratory and pulse artifacts that overlapped with the brachial plexus, resulting in reduced image quality. Furthermore, feet-head PE necessitated large values of oversampling to avoid wrap-around artifacts, with increased in acquisition time as illustrated in Figure 3. For axial and sagittal planes, better scores were achieved with PE in the AP direction. Differences in phase encoding direction. 2D TSE STIR, all sequences had the same duration and number or phase-encoding steps. (a) and (b) Coronal. Pulsatility from carotid and subclavian arteries and motion breathing artifacts that propagate along the phase encoding direction are more detrimental to image quality on the feet-head (b) rather than right-left (a) PE direction. (c) and (d) Axial. Motion related phase encoding artifacts are more detrimental with PE on the anterior-posterior direction. (e) and (f) Sagittal. Motion related phase encoding artifacts are more evident on the feet-head direction, superimposing over branches of the brachial plexus.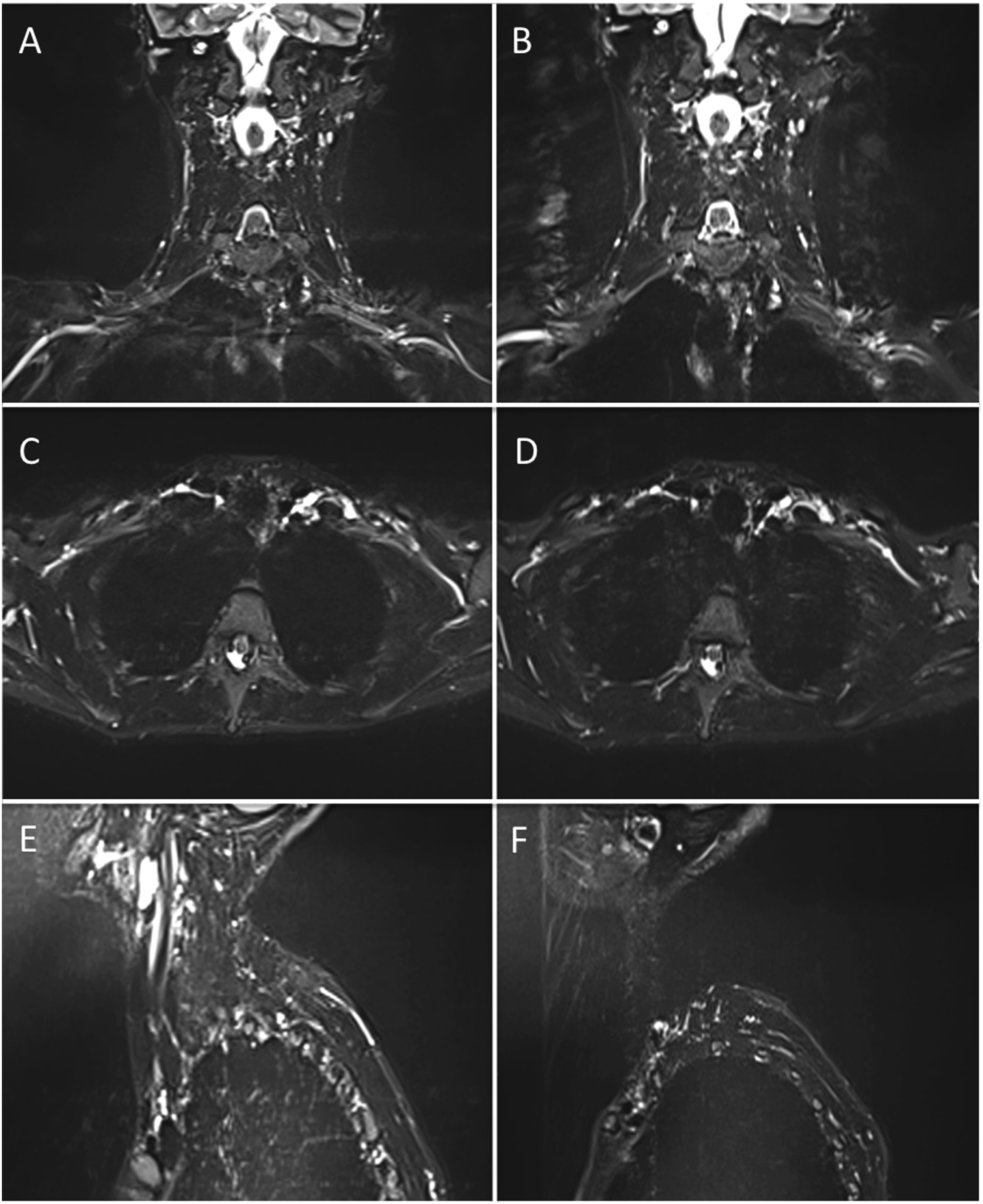
As expected, longer TRs in T2 and STIR imaging led to higher SNR, CNR, and visual scores. However, the relationship with acquisition time was less straightforward. Imaging at TR 2000 with two concatenations and TR 4000 with one concatenation both resulted in the same acquisition time. Consequently, we opted for the latter choice. The TE displayed a distinct pattern, with a progressive improvement in quality peaking around 60–90 ms and declining afterward due to increased noise at longer TEs. It is important to mention that there were no time differences in TEs up to 90 ms.
Regarding resolution, a 1 to 0.8 mm2 provided the best image quality and acquisition time in STIR, T2 and T1 imaging. A 2 mm slice thickness offered improved image sharpness compared to 3 mm, despite some trade-offs in image quality and acquisition time. As expected, thinner slices resulted in increased noise and acquisition time, while thicker slices had drawbacks such blurring and partial volume effects. With the selected parameters, two averages were effective for STIR and T2, while 1 average was sufficient for T1 imaging. Generally, acceleration strategies like iPAT2 yielded acceptable image quality. As expected, higher acceleration values led to a decrease in acquisition time but also resulted in image degradation.
In terms of fat saturation techniques, STIR and Dixon offered uniform fat suppression with comparable acquisition times. Conventional fat saturation and Water excitation yielded similarly inconsistent fat suppression in the neck region (refer to Figure 4). SPAIR provided improved fat suppression but suffered from a significant signal drop around the neck, resulting in limited visualization of the proximal tracts of the brachial plexus. Utilizing a “weak” (longer) or “strong” saturation pulse did not resolve this artifact. Fat suppression techniques. (a) T2 TSE with frequency selective fat saturation pulse. There is inconsistent fat suppression of the subcutaneous and intermuscular fat tissues at the neck and the left axilla region. (b) T2 TSE with water excitation with frequency selective pulse. Similarly to A, inconsistent fat saturation occurs at the neck. (c) T2 TSE with SPAIR-strong pulse. Fat suppression appears more homogeneous compared to (a) and (b), however, there is a consistent signal drop at the neck. (d) T2 TSE with SPAIR-weak pulse. Similarly to C, there is a signal drop at the neck. (e) STIR provides consistent and homogeneous fat suppression. (f) T2 TSE with DIXON, Water image. Similarly to E, fat suppression is homogeneous.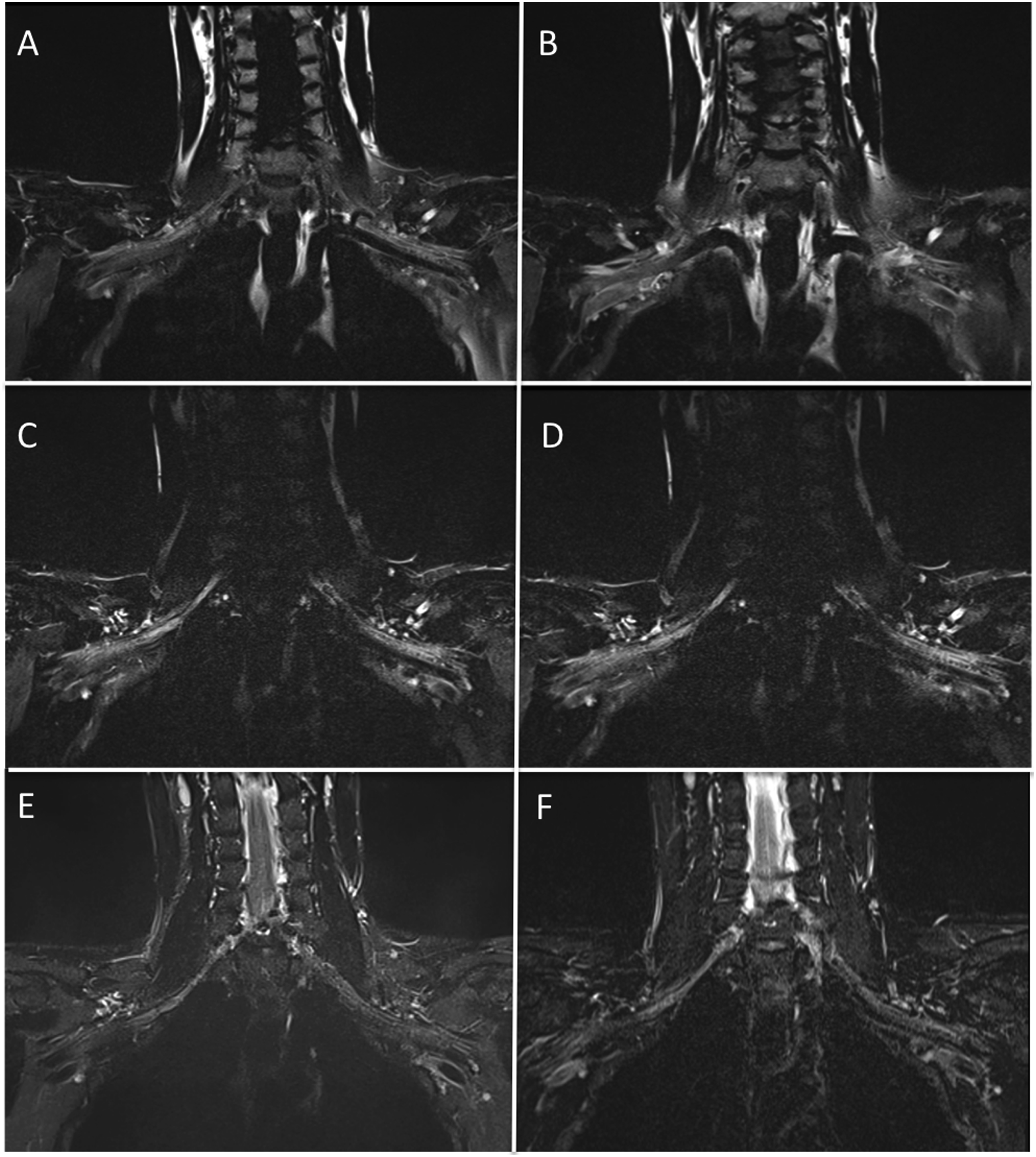
3D imaging
In 3D imaging, slice-selective excitation pulses were consistently employed to enable the reduction of oversampling to only one direction, optimizing acquisition time. 17 Similar to 2D imaging, the lateral PE direction minimized respiratory and pulsatory motion artifacts. To prevent wraparound of the shoulder, oversampling was required. However, the feet-head PE direction required even more oversampling and resulted in lower quality scores. While reconstructed coronal images from the axial acquisition had comparable scores to those from the coronal acquisition with lateral PE direction, they required a significant increase of acquisition time.
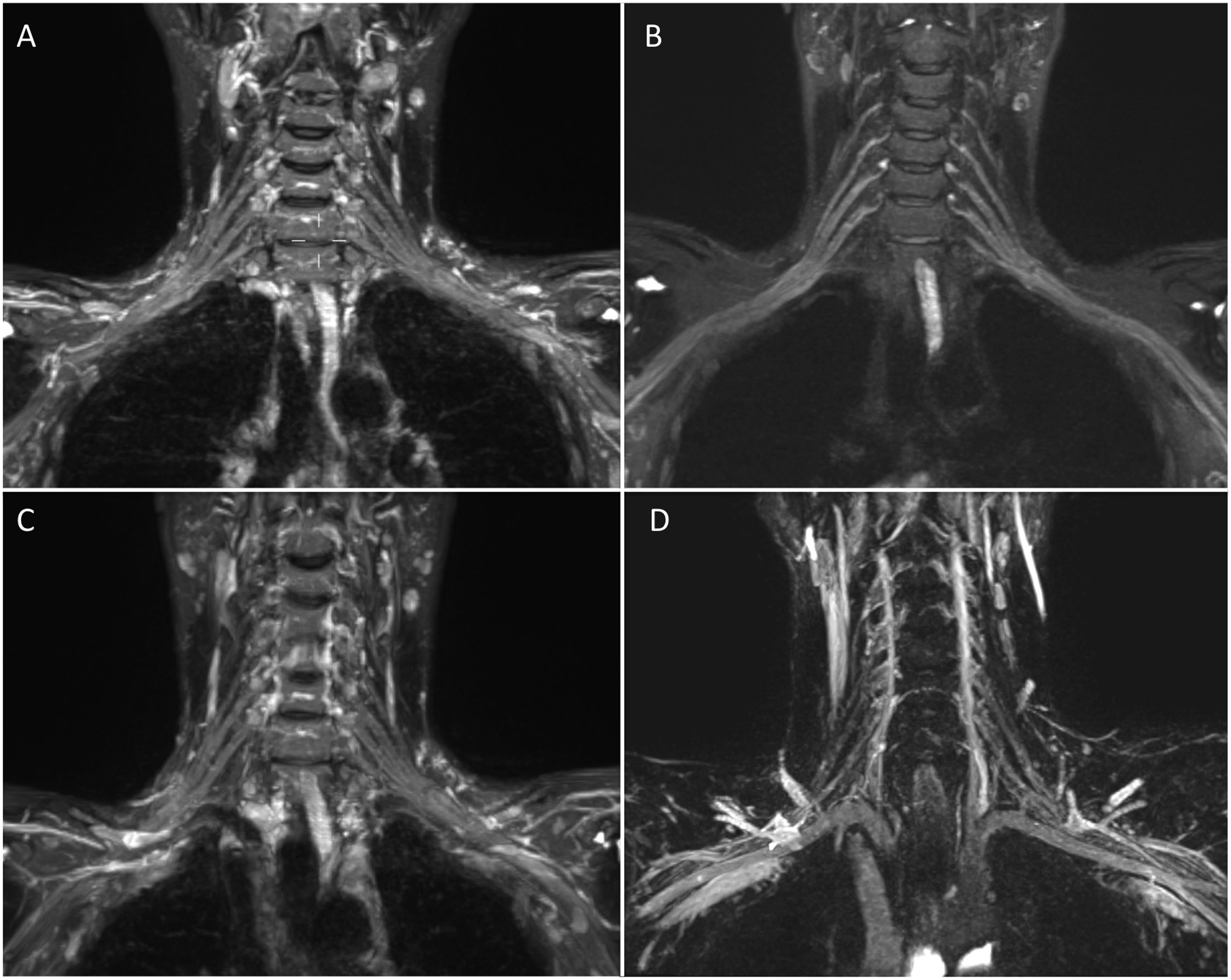
In 3D T2 and STIR SPACE, increasing TR resulted in higher image quality but, unlike 2D imaging, always required increased acquisition time. Nevertheless, a TR of 2000 STIR) or 1500 (T2-myelo) achieved excellent visual scores significantly reducing acquisition time. Optimal quality scores were obtained with longer TEs that did not significantly impact acquisition time. Voxel sizes between 0.9 (T2 Myelo) and 1.3 (SPACE STIR) produced the best scores. All 3D SPACE acquisitions employed acceleration methods like Caipirinha 4 which consistently aided in reducing acquisition time. In terms of fat suppression, 3D STIR ensured uniform and consistent fat suppression. However, similar to 2D imaging, T2 SPACE with fat suppression, water excitation, and SPAIR led to signal loss around the neck region and image degradation (refer to Figures 4 and 5). Blood suppression gradients offered inconsistent blood suppression. A TI of 160 ms reliably suppressed fat and vessels, while a TI of 220 ms, typically employed at 3T, demonstrated inconsistent blood suppression (see Figure 6). T1 imaging. (a) 2D T1 TSE Axial. B-C-D T1 SPACE (b) axial reconstruction, (c) original coronal acquisition (d) sagittal reconstruction (detail of the branches). 2D T1 and SPACE T1 provided similar image quality, but the latter has multiplanar capabilities for an acquisition time of circa 4 min. E-F-G Demonstration of inconsistent fat suppression T1 space with spectral saturation pulse (e), SPAIR (f) or water excitation (g). H-I Vibe Dixon in Axial (h) and Coronal (i) reconstructions provided excellent and homogeneous fat suppression, the acquisition time of circa 2.30 min, and multiplanar capabilities. Blood suppression techniques. SPACE STIR with MIP reconstructions (10 mm) (a) and (b). SPACE STIR with MIP reconstructions before and after gadolinium administration, providing homogeneous vessel suppression and clear brachial plexus visualization. (c). Blood suppression with spoiling gradients (black blood option) provide inadequate vessel suppression. (d). SPACE STIR with TI 220 ms after gadolinium administration. Inconsistent fat suppression occurs with the incorrect TI (220 ms is used at 3T). Of note, fat is still suppressed.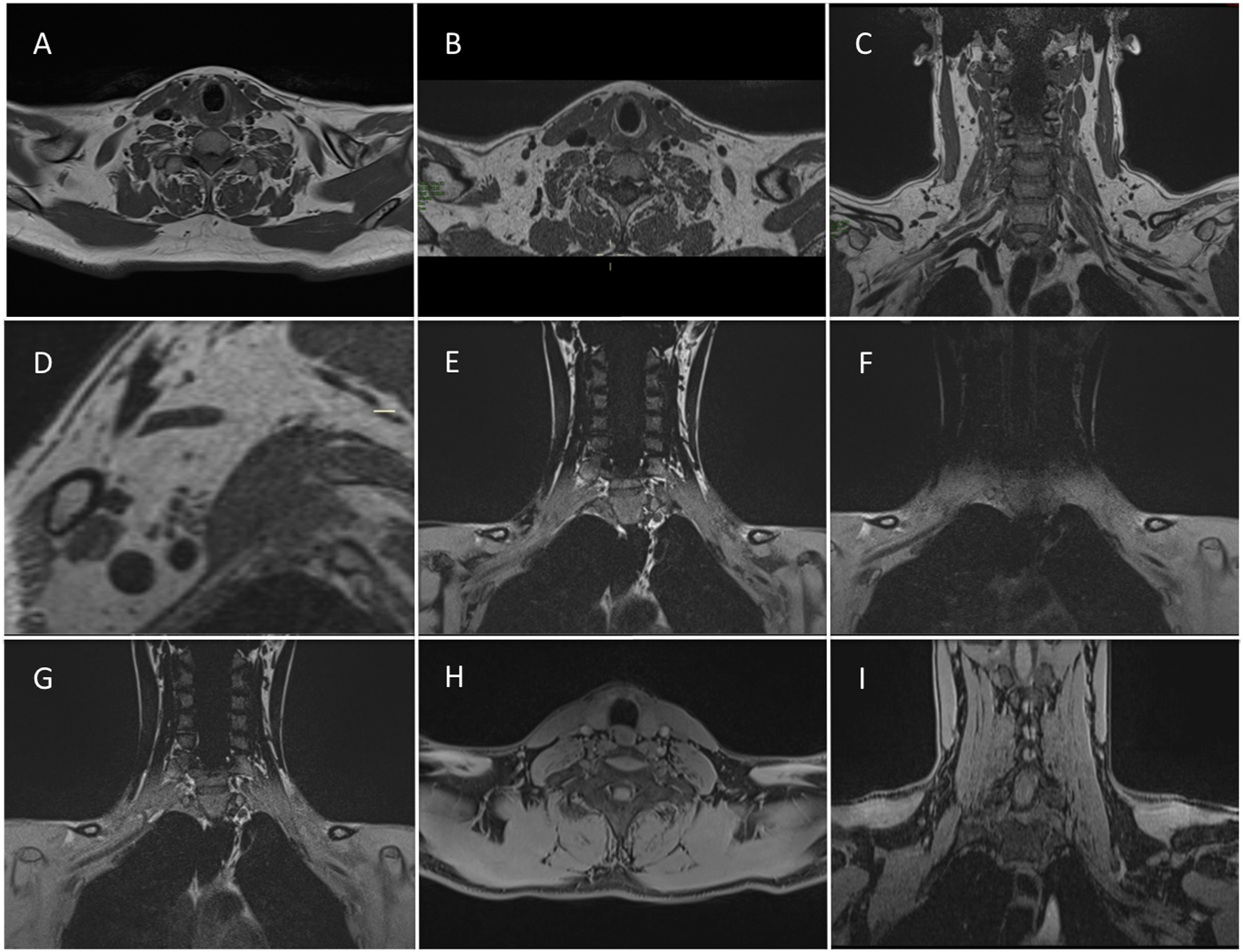
In VIBE imaging, resolution with 0.9 mm isotropic voxels yielded excellent quality scores and acquisition time. A higher flip angle resulted in reduced signal and contrast without impacting AT. Optimal scores were achieved with a flip angle (FA) of 10, while an FA of 30 or more caused significant image degradation.
Final protocol translation
The final protocol was replicated on other manufacturers’ scanners using the same or equivalent sequences. SPACE equivalents included VISTA (Philips) and CUBE (GE). Vibe equivalents were THRIVE (Philips) and LAVA (GE), which utilized conventional fat suppression since Dixon was unavailable in those scanner configurations. Figure 7 displays images obtained from the various scanners. Comparison between different manufacturer’ scanners. The final protocol developed on a 1.5 T Siemens Magnetom Avanto was tested on a Philips Ingenia Ambition and a GE Signa Voyager in order to demonstrate reproducibility. The same volunteer was tested for the main non-contrast sequences on all the three scanners, while 3D STIR and 3D T1 were used on different patients.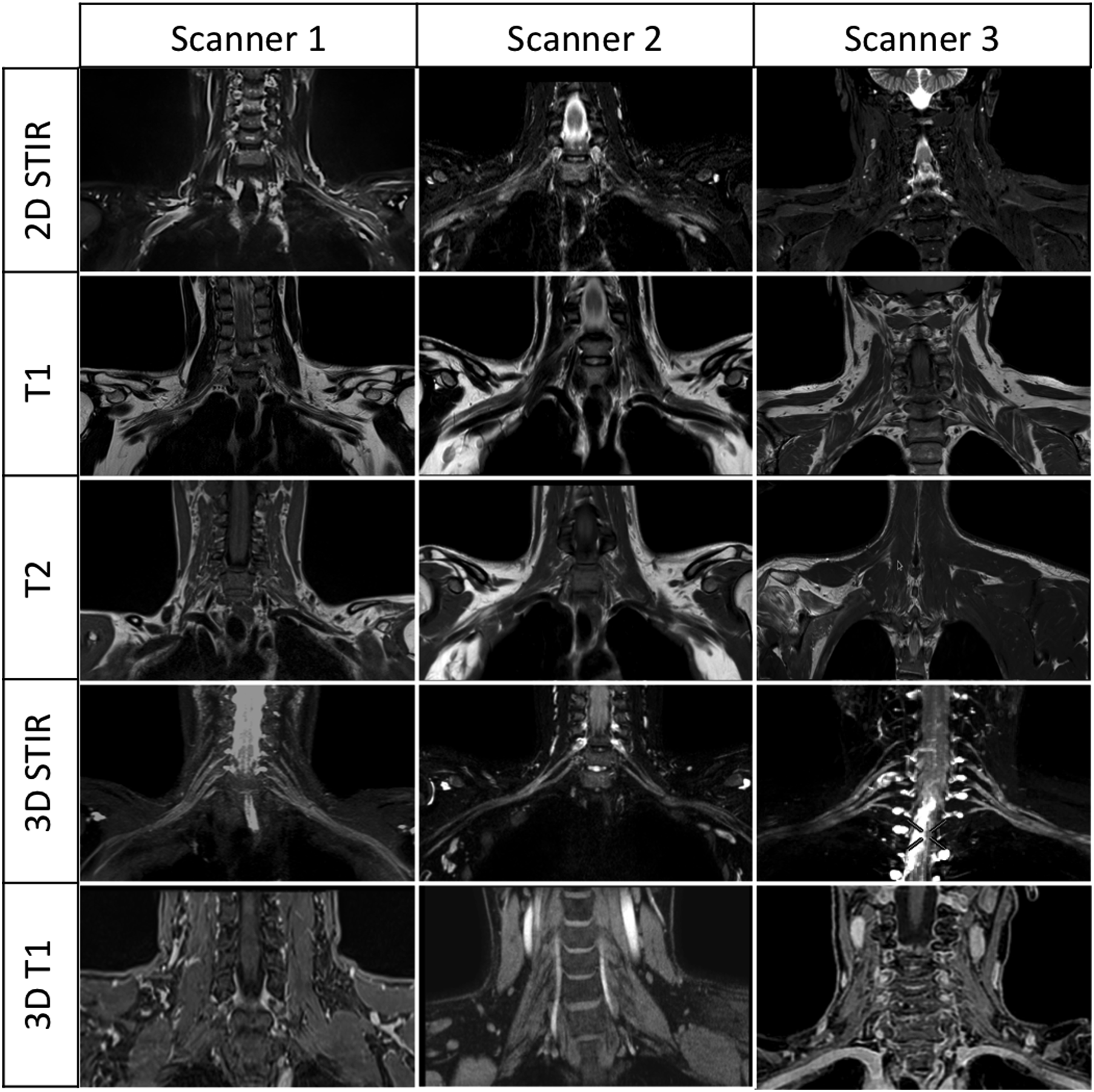
Discussion
In this study, we systematically investigated MRI sequences to establish a standardized MRI protocol for the brachial plexus. We present all the crucial parameters needed to replicate our recommended protocol, along with a comprehensive list of experiments that detail the effects of each parameter on image quality and scan time. Additionally, we tested the protocol on various manufacturers’ scanners to demonstrate its reproducibility.
Our final protocol includes a standard cervical spine examination, which is deemed fundamental to assess the neural foramina caliber and possible cervical disc herniation. 18 There is a set of 2D TSE sequences, 3D T1 SPACE, T2 SPACE-myelo, and, after contrast administration, 3D SPACE STIR and VIBE. A complete examination of the brachial plexus requires approximately 35 min.
There are no definitive recommendations regarding the optimal field strength for visualizing the brachial plexus, with 3T often considered ideal, but 1.5T scanners are still widely and effectively used. 1 There is no consensus on the fundamental sequences to utilize, such as the specific 2D and 3D or weightings to employ, and some researchers even choose to exclude 2D imaging entirely. 8
2D imaging offers advantages such as higher in-plane resolution and standard weighting but suffers from partial volume effects, inter-slice gaps, and the necessity for multiple plane acquisition. In contrast, 3D TSE sequences (e.g., SPACE, CUBE, VISTA) enable isotropic imaging with multiplanar reconstruction, similar weightings to 2D TSE, but with increased acquisition time, making them more susceptible to motion artifacts. SPACE STIR is almost universally recommended for selective neurography of the brachial plexus due to its homogeneous fat suppression, multiplanar and MIP reconstructions, and enhanced nerve visualization.5,14 It is necessary to acquire SPACE STIR after contrast administration for optimal blood suppression. We replaced 2D T1 imaging with T1 SPACE for its superior resolution and relatively quick acquisition, and T2 SPACE can potentially replace 2D T2 as well. We do not routinely acquire 2D sagittal slices, as they can be reconstructed from 3D imaging.
Bilateral examination is crucial for evaluating nerve thickness and signal symmetry, accounting for individual anatomical variations due to age, sex, and muscle/fat distribution. Field of view coverage should encompass the spine, shoulder girdle, and rotator cuff muscles, using fluid-sensitive fat-suppressed and T1-weighted sequences to assess muscle trophism and edema. We prefer true orthogonal coronal planes as oblique planes offer no significant improvement in nerve visualization. 19 Additionally, the brachial plexus’s slight anterior bend would require axial plane orientation, preventing simultaneous imaging of both sides. 12 This approach also enhances standardization and simplifies tasks for technologists.
Some sequence parameters, like PE direction, can significantly affect image quality in terms of pulsatory and respiratory artifacts and acquisition time, even though they are not frequently reported in the literature. We demonstrated this in our experiments.
It may seem obvious that increased TR or TE would enhance image contrast and quality, and that higher resolution images provide more detail, but also lower SNR and longer acquisition time. Nevertheless, by experimentally examining the changes in image quality when modifying each sequence parameter, we can better assess whether the trade-off in increased acquisition time is worth the improved image quality. Additionally, our experiments revealed non-obvious effects of certain parameters on image quality. In 2D STIR, increasing TR from 2000 to 4000 improved quality without affecting acquisition time. In contrast, in SPACE-STIR, TR strongly influenced acquisition time, but had a limited impact on image quality. Surprisingly, there is significant variability in the literature regarding these parameters. For instance, TR in SPACE-STIR is reported to range from 1500 to 4500, affecting acquisition time up to 3-fold.4–10 Although the best contrast was obtained with longer TRs, we demonstrated that excellent results can still be achieved by reducing it to 2000. Similar variability exists in TE for STIR SPACE, ranging from 70 to 250.7,8 Since it doesn’t affect acquisition time up to a certain value, TE should be set to the longest possible to increase contrast.
There is also substantial variability in resolution reported in the literature. In-plane resolution on 2D TSE ranges from 0.6 to 1.3 mm, and slice thickness varies from 3 to 5 mm, even at 3T scanners.9,11–14 Moreover, real and reconstructed matrices (obtained with partial k-space filling) are often reported interchangeably. The real matrix, which impacts acquisition time, should be considered instead. This variability may drastically affect image quality and acquisition time. Higher resolution can be achieved with T1 SPACE (up to 0.8 mm), which is sufficient to completely replace 2D T1 imaging.
3D imaging benefits from acceleration strategies, and we recommend strong utilization of these techniques, such as advanced k-space filling strategies like Caipirinha, which maximizes advantages by using at least three anterior and three posterior elements of activated coils.
Proper fat and water suppression is essential for evaluating nerve lesions. We have demonstrated that frequency-selective saturation strategies like fat saturation, water excitation, and SPAIR are less effective than non-frequency-dependent techniques like STIR or DIXON due to the typical B0 field inhomogeneities in the cervical region. Although SPAIR is more effective with 3T scanners, 14 SPACE Dixon can be a suitable alternative, though generally available only on recent scanners. 20
Blood suppression using preparation pulses, spoiling, or crushing gradients requires high B0 field homogeneity and is commonly reported for 3T.21–23 Gadolinium chelates shorten blood T1, allowing for signal suppression after applying an inversion pulse. STIR provides homogeneous blood suppression with excellent visualization of the brachial plexus when acquired after gadolinium administration, as long as the inversion time (TI) is correct (160–180 ms at 1.5 T, 220 ms at 3T). It is always worthwhile to verify the value of TI prior to administering contrast.
Owing to inconsistent fat suppression in 2D fat-sat T1w, we chose VIBE, a fast volumetric gradient-echo sequence that produces 3D T1w images
This study has some limitations. Subjects were healthy volunteers, and including patients with actual pathologies could have aided in evaluating image quality. However, examining healthy subjects allowed for a more systematic evaluation of technical aspects of the imaging sequences, without being constrained by acquisition time. Our focus was on optimizing imaging sequences in terms of signal, noise, and acquisition time, rather than assessing pathological patterns. Though, some patients were examined to assess the effect of gadolinium on blood suppression. We acknowledge that our protocol construction method, selecting parameter values based on higher scores, may not be as rigorous as other statistical methods. The determination of acquisition time quality using our scoring system involved subjective judgment, as the specific time thresholds for each score were somewhat arbitrary. While our approach provided a practical means of evaluating acquisition time, the lack of standardized criteria may introduce variability and limit direct comparability with other studies. Future research could explore the development of more objective and standardized methods for assessing acquisition time quality in order to enhance the consistency and reliability of such evaluations. There was some variability in the absolute values of the ROIs between subjects, and a larger sample size with varying age, sex, and weight would have provided broader standardization. More sequences and parameters could also have been tested.
Demonstrating the adaptability of our brachial plexus imaging protocol to different scanners was a study’s secondary objective. By showcasing its ease of translation, we hope to encourage wider adoption across various institutions and settings. While further validation may be needed, our study offers a valuable starting point for optimizing protocols on scanners from different manufacturers.
Conclusion
We have developed a basic protocol for the study of the brachial plexus utilizing 2D and 3D imaging techniques, incorporating post-contrast SPACE-STIR for neurography, optimizing sequences in terms of spatial resolution, contrast, and acquisition time. By providing detailed instructions and insights into artifact analysis and potential solutions, this protocol may serve as a valuable resource for both novice and experienced radiologists seeking to study the brachial plexus. We believe that this standardized approach will contribute to improved imaging quality, reproducibility, and further advancements in brachial plexus research and clinical practice.
Supplemental Material
Supplemental Material - An optimized 1.5 Tesla MRI protocol of the brachial plexus
Supplemental Material for An optimized 1.5 Tesla MRI protocol of the brachial plexus by Paolo Florent Felisaz, Angela Napolitano, Simone Terrani, Carmelo Parisi, Marco Toto-Brocchi, Maurizio Cè, Francesco Alessandrino, Giancarlo Oliva, Michaela Cellina and Simonetta Gerevini in The Neuroradiology Journal.
Footnotes
Acknowledgments
We thank Francesca Zaccone (Siemens) for providing unconditioned support for sequences design and optimization and Marcello Cadioli (Philips) for support and constant inspiration.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics approval
We confirm that we have read the Journal’s position on issues involved in ethical publication and affirm that this report is consistent with those guidelines.
Informed consent
all subjects provided written informed consent.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
