Abstract
Objective
Osteoarthritis (OA) is a degenerative joint disease. A growing number of studies have shown that microRNAs (miRNAs) play an important role in the pathogenesis of OA. However, the specific function of miR-322 in OA is unknown. This study was aimed to explore the ability of miR-322 in the cartilage matrix degradation and the mechanism in OA.
Methods
Quantitative reverse transcription polymerase chain reaction (qRT-PCR) was performed to detect miR-322 expression in cartilage and OA-associated gene expression in chondrocytes treated with miR-322 mimics/inhibitors or interleukin (IL)-1β, respectively. The targets of miR-322 were analyzed using software and the luciferase reporter experiment. In vivo, intra-articular injection of miR-322 mimics was administered at the knee of DMM mice. After 12 weeks, the knee joints of mice were collected for histological analysis.
Results
The expression of miR-322 was decreased in knee cartilage of DMM mice and was significantly reduced by IL-1β. miR-322 mimics inhibited IL-1β-induced extracellular matrix degradation, as evidenced by higher expression of Col2α1 and Aggrecan, and lower expression of Adamts5, MMP3, and MMP13. In contrast, miR-322 inhibitor promoted extracellular matrix degradation of chondrocytes. TRAF3 was the predicted target of miR-322 from databases. Luciferase reporter assay verified the targeting relationship between miR-322 and TRAF3. The effect of miR-322 on extracellular matrix degradation was partially reversed by overexpression of TRAF3. In addition, H&E and Safranin-O fast green staining assays in OA mouse models showed that miR-322 mimics attenuated the progression of OA in vivo.
Conclusions
miR-322 suppressed chondrocytes matrix degradation and alleviated OA cartilage injury via inhibition of the TRAF3.
Introduction
Osteoarthritis (OA), a chronic degenerative joint condition, is the major cause of disability in the older population, defined by joint inflammation, subchondral bone (SCB) alterations, and cartilage degradation. 1 Articular cartilage anabolism and catabolism imbalance, particularly an increase in catabolism, have been linked to these conditions. 2 Degradation of cartilage often starts with the breakdown of aggrecan, a proteoglycan in articular cartilage, and continues to destroy collagen fibers, ultimately resulting in fibrillation and laceration of the cartilage. 3 Chondrocytes are the sole kind of cell found in cartilage, and they are involved in regulating the extracellular matrix (ECM) and preserving the tissue’s normal structure and function. 4 On the one hand, chondrocytes may keep synthesizing proteoglycan, collagen, and other ECM components. 5 On the other hand, various matrix-degrading enzymes, ADAMTS, and other hydrolases are dynamically synthesized by chondrocytes. 6
Growing evidence over the last decade has pointed to the crucial involvement of epigenetic processes in altering cell phenotype in OA. 7 Epigenetic modifications in gene expression caused by environmental interactions contribute to the etiology of disease and are passed on to daughter cells in a stable manner through several cell divisions. 8 MicroRNAs (miRNAs) stand out among these epigenetic processes because of their potential impact on OA disease development via their involvement in maintaining a normal chondrocyte phenotype and modulating inflammatory response pathways. 9 miRNAs are a kind of noncoding, single-stranded RNA with an average length of 20 to 25 nucleotides involved in post-transcriptional gene expression regulation and affect cell destiny during proliferation and differentiation. 10 Sequence-specific interactions with the 3′-untranslated regions (3′-UTRs) of target mRNAs result in mRNA degradation and/or translational suppression through this regulatory mechanism. 11 In previous investigations, numerous miRNAs were associated with OA development and shown to have a variable expression in OA cartilage. Knockdown of miR-892b was shown by Zhao et al. 12 to suppress apoptosis in chondrocytes and slow OA development by targeting Cyclin D1 and Cyclin D2. miR-29b-3p was discovered to be upregulated in OA cartilage by Chen et al., 13 and further research found that miR-29b-3p increases the onset and progression of OA by targeting Progranulin (PGRN), which in turn promotes chondrocyte apoptosis. In order to maintain joint homeostasis and prevent the advancement of OA, Huang et al. 14 hypothesized that 2 essential miRNAs, miR-211 and miR-204, play significant roles by targeting Runx2. Therefore, it is crucial to investigate the function and mechanism of miRNAs in OA progression.
The miR-424, a miR-15/107 family member homologous to mmu-miR-322-5p, is only found in mammals. 15 The key physiological processes that miR-424(322) controls include the epithelial-to-mesenchymal transition, cell cycle, hypoxia, and other stress response, and its expression is more dynamic and tissue-restricted than that of other family members. 16 Pinger Wang et al. 17 screened blood samples from patients with OA and found that hsa-miR-424-5p was lowly expressed in patients with arthritis. To prevent the RAF/MEK/ERK pathway activation in cartilage, miR-322 is upregulated during transcriptome profiling for chondrocyte differentiation. 18 Therefore, we hypothesize that miR-322 may be involved in the process of cartilage matrix degradation in OA. In the present study, we aimed to explore the function of miR-322 in the disease pathogenesis and the therapeutic potential relevant to OA. Thereto we investigated the expression of miR-322 in articular cartilage of DMM model mice. We also explored the role of miR-322 and target genes in interleukin (IL)-1β-induced mouse chondrocytes in vitro. On this basis, we further examined the effect of intra-articular injection of miR-322 mimics on chondrocyte matrix degradation in the OA tissues of mice. Our results demonstrated that high-level miR-322 decreases OA chondrocyte matrix degradation by reducing TRAF level.
Materials and Methods
Cell Transfection
MiR-322 mimics and inhibitor or miR-Con were obtained from GenePharma (Shanghai, China). A TRAF3 overexpression plasmid (pcDNA3.1-TRAF3) and a control vector were designed and constructed by OBiO Technology (Shanghai, China). For transfection, cells (4 × 104 cells/cm2) in 6-well plates were transfected with miR-322 mimics, inhibitor, or pcDNA3.1-TRAF3 using HiPerFect Transfection according to the manufacturer’s protocol. Cells were then treated with IL-1β (10 ng/mL) for 24 hours as indicated.
Experimental OA in Mice
Eight-week-old C57BL/6 mice were obtained from Shanghai SLAC Laboratory Animal Co. (License No. SCXK 2003-0003; Shanghai, China). The procedure was approved by the Institutional Animal Care and Use Committee of Zhejiang Academy of Medical Sciences, and all animal breeding and housing was performed in the Animal Care Facility of Zhejiang Academy of Medical Sciences in accordance with the institution’s guidelines for laboratory animals. Mice were randomized into 4 groups: sham+miR-Con, DMM+miR-Con, sham+miR-322, and DMM+miR-322. A surgical procedure was then performed to establish an experimental OA model. DMM of the right knee was performed under general anesthesia using an operating microscope. The right knee was opened and the components exposed, without manipulation of the joint tissue; this was a sham operation. To treat experimental OA with miR-322, a 33G needle and microsyringe (Hamilton) were used to inject 10 μL volumes of miR-322 mimics and negative controls into the knee joint. The mice received the first injection 7 days after DMM and then once a week for 11 weeks. After 12 weeks of DMM, the mice were euthanized for histopathological examination.
Histological Evaluation
Mouse knee joints were decalcified in 0.5 M EDTA (pH 7.4) for 2 weeks after fixation in 4% paraformaldehyde. Cartilage tissue samples were embedded in paraffin and sectioned at a thickness of 5 µm. Hematoxylin and eosin (H&E), safranin O, and Fast Green staining were then performed on the cartilage tissue samples. Hyaline cartilage (HC) and calcified cartilage (CC) thicknesses were measured using H&E staining and determined by Image J software (National Institutes of Health, Bethesda, MD, USA). Briefly, the articular cartilage region was divided equally into 6 sections, the lengths of which were measured separately and then averaged. Three observers blinded to treatment allocation scored the extent of cartilage degradation in the OA mouse model using the Osteoarthritis Research Society International (OARSI) grading system (grades 0-6). OARSI scores were presented as the mean maximum score for each mouse.
Collection of Articular Cartilage
Mouse articular chondrocytes for culture were isolated from the cartilage of postnatal day 5 C57BL/6J mice. Femoral heads, femoral condyles, and tibial plateau were isolated and digested with proteinase and collagenase, 19 and then maintained in Dulbecco’s modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum (FBS; Gibco, USA), 100 μg/mL streptomycin, and 100 U/mL penicillin (Gibco).
In experimental OA mice, articular cartilage samples were obtained from the femoral head of mice as follows: exposure of the dislocated hip, stripping of the subchondral helmet-like articular cartilage, cutting of the articular cartilage into several vertical slices, and removal of the remaining ligamentous bulb, synovial tissue. 20
RNA Isolation and qRT-PCR
Chondrocytes’ total RNA was isolated using the TRIzol reagent (Invitrogen, USA). Then, using a PrimeScript RT reagent kit (Takara, Tokyo, Japan), cDNA was synthesized from 1 μg of total RNA. SYBR Real-time PCR Master Mix-Plus (Toyobo, Osaka, Japan) was used in the LightCycler96 real-time PCR system to quantify mRNA after cDNA synthesis was complete (Roche, Mannheim, Germany). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and small nuclear RNA U6 were used as internal controls for cDNA and miRNA, respectively. The 2−ΔΔCt method was used to determine the relative levels of gene expression.
Western Blot
Protein samples were prepared by washing chondrocytes in phosphate-buffered saline (PBS) and lysing them in radioimmunoprecipitation assay (RIPA) lysis buffer (Biosharp, Hefei, China). Total proteins were separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and then transferred onto polyvinylidene fluoride (PVDF) membranes (Merck-Millipore, MA, USA). After sealing for 1 hour, the membranes were incubated with primary antibodies against GAPDH (ab181602, Abcam), MMP13 (ab39012, Abcam), collagen II (ab12268, Abcam), or aggrecan (ab36861, Abcam), followed by incubation with secondary antibodies. Internal standards were established using GAPDH, and the negative control was set to a value of 1.
Luciferase Reporter Assay
GenePharma Co., Ltd. (Shanghai) designed and produced the plasmids harboring the mutant (TRAF3-MT) and wild-type (TRAF3-WT) TRAF3 3′-UTRs. The ATDC5 cells were co-transfected with wild-type or mutant TRAF3 luciferase reporter vectors and miR-322 mimics, miR-322 inhibitor, or miR-control plasmids using HiPerFect transfection. The Dual-Glo Luciferase Assay System (Promega, USA) was used to assess luciferase activity in cell lysates extracted 48 hours post-transfection, per the manufacturer’s protocol. The expression of the Renilla reporter gene served as an internal control. Triplicate samples prepared for the 3 independent experiments were analyzed to obtain data.
Statistical Analysis
Numerous data were presented as mean ± standard deviation (SD) and analyzed using 1-way analysis of variance (ANOVA) and Tukey-Kramer test for multiple comparisons (SPSS 13.0J; SPSS, Inc., USA). P < 0.05 and P < 0.01 were used to determine statistical significance. The experiments were replicated independently 3 times, and each set of findings was qualitatively the same. Representative experiments were shown.
Results
MiR-322 Is Downregulated in OA Cartilage and Participates in Chondrocyte Matrix Synthesis
The expression of miR-424 (322) is substantially decreased in the blood of patients with knee osteoarthritis (KOA) compared to healthy controls, as detected by microarray expression profiling detection.
17
To test whether the expression of miR-322 is also decreased in a mouse model of OA, a model of surgical destabilization of the medial meniscus (DMM) in knee joints was established. Data from qRT-PCR experiments performed on knee joints extracted 8 and 12 weeks following DMM surgery in mice compared to tissues from animals that had sham surgery revealed substantially decreased miR-322 expression in the articular cartilage in the DMM group (
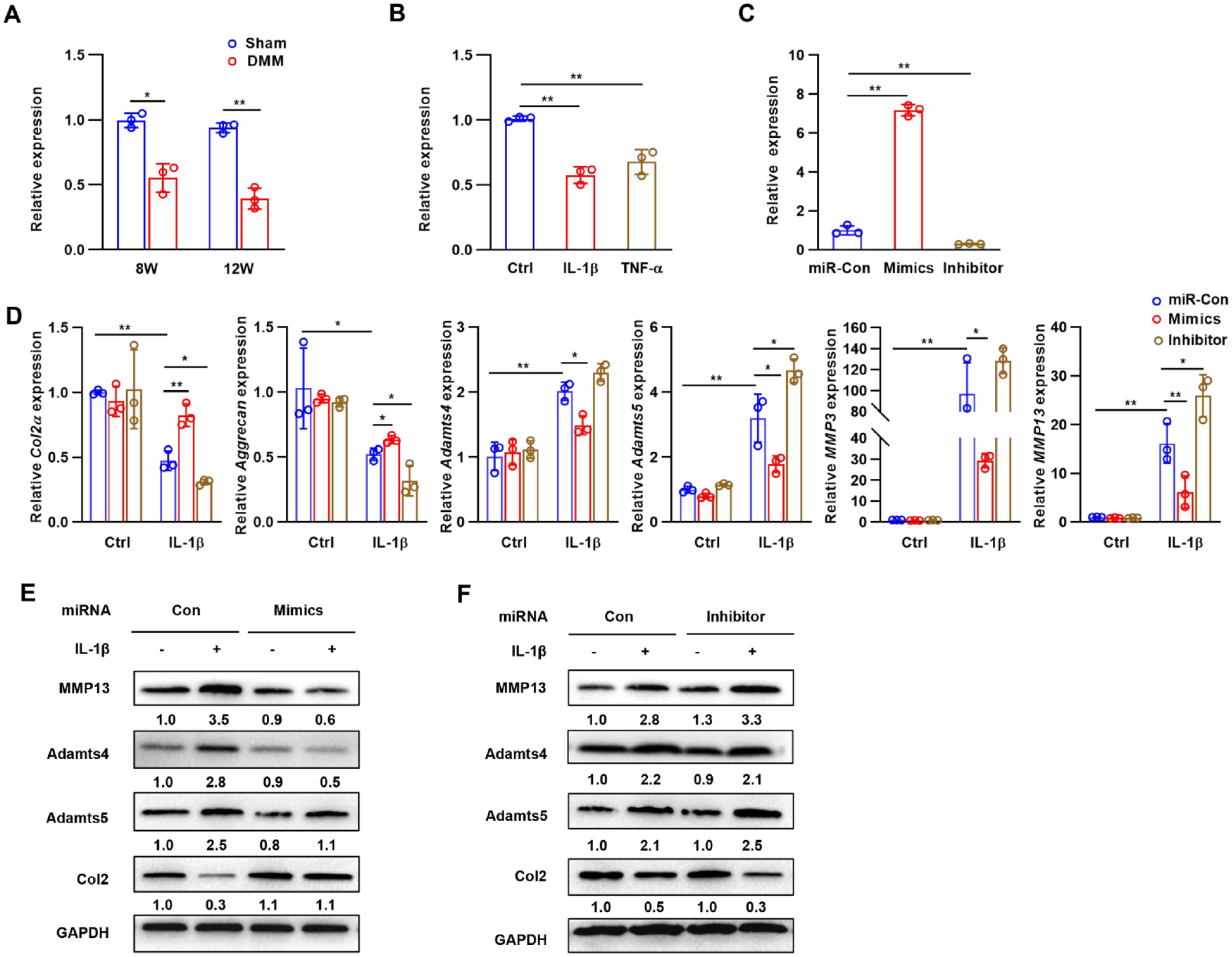
miR-322 is downregulated in OA and inhibits IL-1β-induced chondrocyte matrix degradation. (
Chondrocytes were transfected with miR-322 mimics or inhibitor to determine the role of miR-322 in controlling the IL-1β-induced production of matrix-degrading enzymes. Transfection of miR-322 mimics significantly increased the expression of miR-322, which was suppressed by miR-322 inhibitor (
Identification of TRAF3 as a Target Gene for MiR-322
To identify the potential targets of miR-322, bioinformatics tools TargetScan and miRWalk were used, and TRAF3 was predicted to be a potential target of miR-322, with a highly conserved complementary miR-322 binding site in its 3′-UTR (
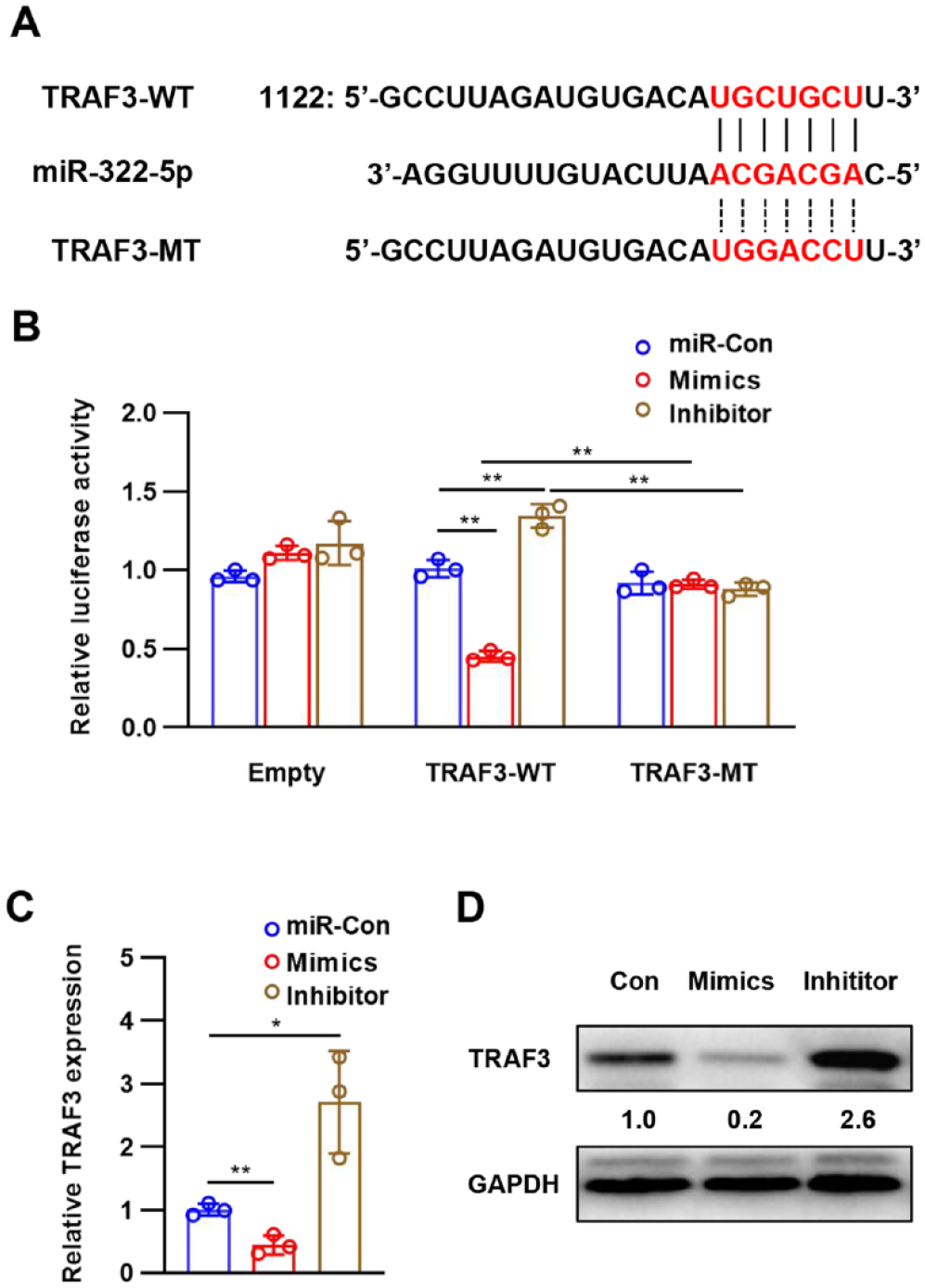
miR-322 directly targeted TARF3 to inhibit its expression. (
TRAF3 mRNA and protein expression levels in chondrocytes transfected with miR-322 mimics and inhibitor were analyzed to determine whether or not miR-322 can regulate TRAF3 expression. At 48 hours post-transfection, the expression of TRAF3 is decreased in the miR-322 mimics group but significantly increased in the miR-322 inhibitor group (
TRAF3 Participates in the MiR-322-Mediated Matrix Degradation of Chondrocytes
Given that miR-322 directly binds to TRAF3 to inhibit its expression, we further investigated the role of TRAF3 in influencing matrix degradation. IL-1β induced TRAF3 mRNA and protein levels in chondrocytes, and similarly TRAF3 expression was elevated in articular cartilage after DMM surgery (
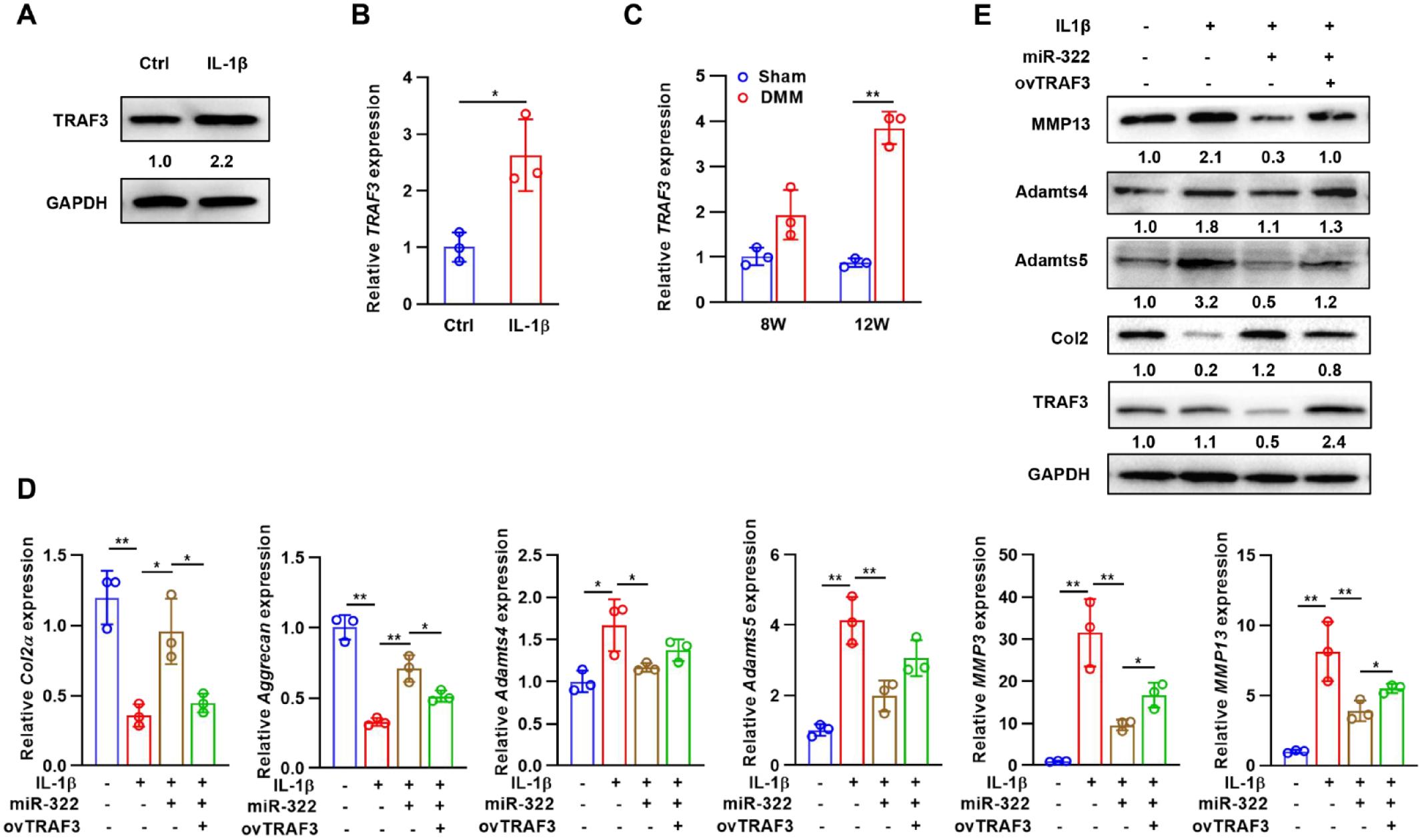
The effects of TRAF3 on miR-322-mediated expression of matrix-degrading enzymes. (
MiR-322 Mimics Attenuates the Progression of OA In Vivo
DMM surgery on mice was then performed, and their cartilage degeneration was analyzed using histopathology and the OARSI to establish whether or not miR-332 affected the severity of the degeneration. The surface of the articular cartilage in the sham-operated group was intact, with smooth junctions, clear and distinct tide lines, moderate cartilage thickness, normal chondrocyte numbers, and regular structure. Compared to sham mice, DMM animals showed surface disruption, discontinuities, significant proteoglycan loss, and even HC loss and CC exposure at 12 weeks postoperatively. Significant increase in articular cartilage thickness and reduction in cartilage damage in the miR-322 mimic group compared to the model group (
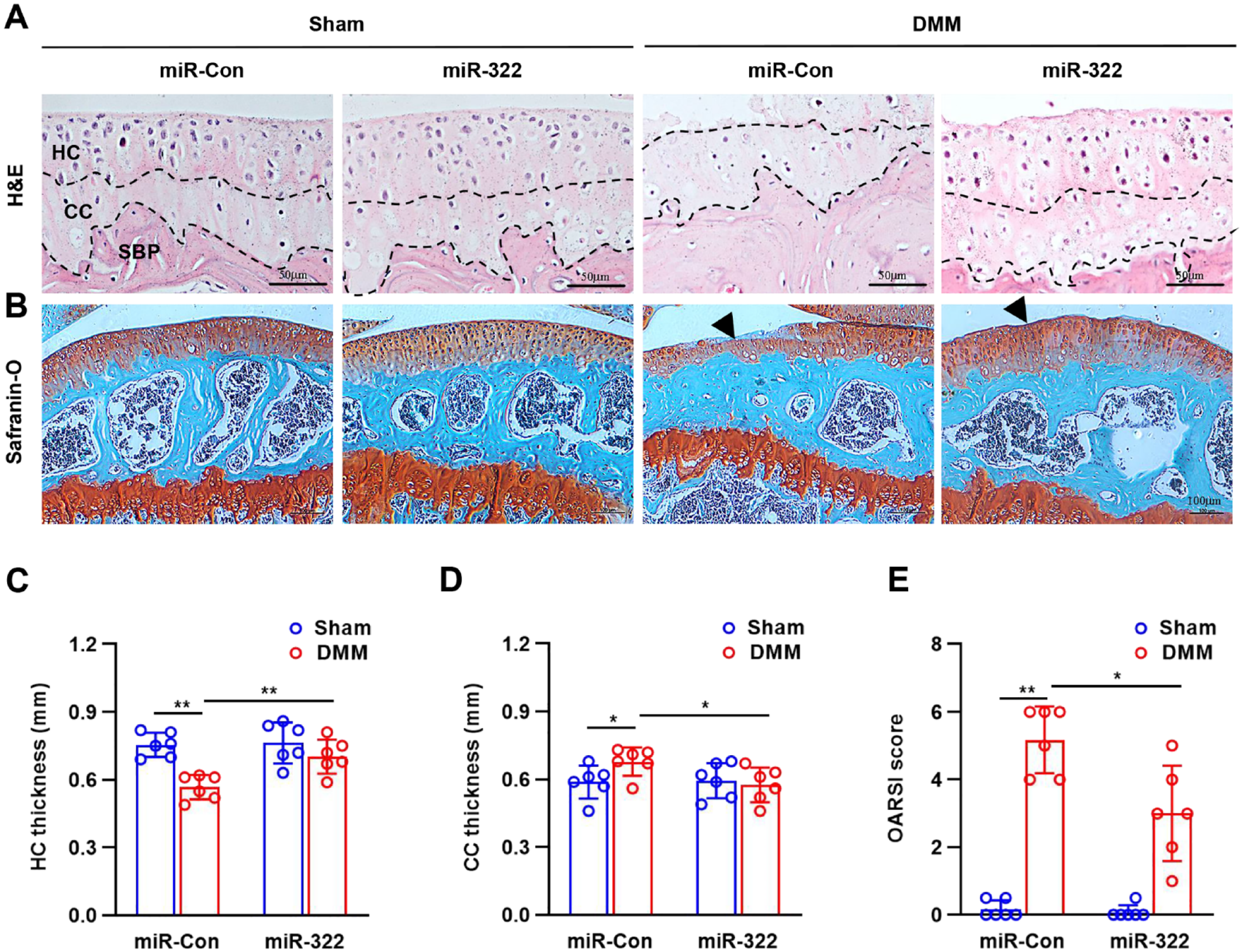
miR-322 attenuates the progression of OA in vivo. (
Discussion
The alarmingly rising prevalence of OA is one of the primary causes of pain, disability, and socioeconomic burden throughout the globe. 21 Despite the dearth of medications that may slow the course of osteoarthritis, research has been done to identify the underlying biological mechanisms contributing to the disease. 4 This research provided more evidence that OA is associated with decreased miR-322 levels and increased TRAF3 levels. Moreover, miR-322 reduced OA cartilage damage by inhibiting ECM breakdown in cartilage via TRAF3.
An increasing body of research has highlighted the significance of miRNAs in the onset and progression of OA. 22 miR-373, miR-26a, miR-210, miR-337, miR-25, miR-140, and miR-29a were shown to be downregulated in osteoarthritic cartilage of the human knee joint compared to normal cartilage, miR-509, miR-23b, miR-30b, whereas miR-223, miR-16, miR-103, miR-377, miR-22, and miR-483 were upregulated. 23 In early OA cartilage, miR-146a expression was elevated by inflammatory cytokines and that the negative feedback of miR-146, including downregulation of TRAF6 and IRAK1, may adversely control the expression of catabolic proteins like MMP13. 24 Bo Yang et al. 25 found that miR-145 altered its downstream target gene expression and ECM degradation induced by IL-1β in OA chondrocytes by directly regulating mothers against decapentaplegic homolog 3. Several other miRNAs, including miR-140 and miR-483, have been linked to OA development.26,27
miR-322 expression is upregulated during chondrocyte differentiation and stabilizes MEK1 expression to inhibit activation of the RAF/MEK/ERK pathway in cartilage, as screened by transcriptome analysis. 18 In addition, it has been shown that miR-424 was downregulated in IL-1β-induced chondrocytes and OA chondrocytes. 28 Downregulation of hsa-miR-424-5p in KOA blood relative to non-KOA was validated by Wang et al., 17 and it was shown to be related to the pathophysiology of SCB sclerosis. MiR-322 was shown to be downregulated in DMM mice in this study. A key player in the progression of OA, IL-1β is a cytokine that may trigger a cascade of pathogenic responses in chondrocytes. Similar suppression of miR-322 expression was observed in IL-1β-treated chondrocytes and DMM mice.
Loss of aggrecan and its subsequent degradation is generally considered a crucial step in the onset and development of OA and is followed by the largely irreversible degradation of collagen, which causes cartilage dysfunction. 29 ECM homeostasis relies on several variables, including the enzymes MMP13 and ADAMTS5, which are involved in cartilage degradation. 30 Our data showed that miR-322 mimics decreased ADAMTS5, MMP3, and MMP13 expression in IL-1β-activated chondrocytes while increasing Col2α1 and Aggrecan expression. Previous research has shown that upregulating Sox9 increases miR-322-5p expression, facilitating BMP2-induced chondrogenesis. 31 We found that overexpressing miR-322 mitigated surgical-induced OA-related cartilage degradation in vivo. According to these results, miR-322 upregulation may slow the degradation of cartilage ECM in OA.
TRAF3 plays an important regulatory role in the immune response and has been reported to be involved in inflammation initiation and osteoclast bone resorption. Clinical studies have shown a strong correlation between the 3′-UTR region of TRAF3 and the clinical outcomes of OA patients. 32 Our research confirms these prior findings. Substantial increases in TRAF3 were seen in IL-1β-treated chondrocytes in DMM mice. Hu et al. 33 demonstrated that overexpression of TRAF3 using adenoviruses reduced the expression of catabolic genes in chondrocytes and inhibited the ECM loss and cartilage degradation in experimental mouse OA induced by IL-17. It has been previously reported that miR-671-3p regulates apoptosis and inflammation in chondrocytes via TRAF3 and may have a part in OA pathogenesis. 34 Chondrocytes in OA are regulated by miR-107, which affects autophagy and apoptosis by targeting TRAF3. 35 RIP2 regulates cartilage degradation and oxidative stress through regulation of TRAF3. 36 The current work identifies TRAF3 as a miR-322 target gene with high conservation. TRAF3 translation was severely impeded when miR-322 bound to its 3′-UTR, inhibiting IL-1β-mediated ECM degradation. The impact of miR-322 mimics on the expression of proteins involved in cartilage degradation was also significantly attenuated by overexpression of TRAF3.
In conclusion, we provide new insight into the treatment of OA by elucidating a unique regulatory mechanism: miR-322 acts as a critical negative regulator of cartilage matrix degradation through the TRAF3 during OA pathogenesis.
Footnotes
Author Contributions
Bo Ma supervised the study. Jirong Wang and Ying Tang performed the experiments. Jirong Wang and Yizhong Bao designed the experiments and wrote the paper. Lan Chai and Bo Ma participated in data analysis and interpretation. Xiujun Pei contributed to revise the manuscript. All authors read and approved the final manuscript.
Acknowledgments and Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. This work was supported by the Zhejiang Province Welfare Technology Applied Project (No. LGD20H070001), Zhejiang Provincial Administration of Traditional Chinese Medicine (No. 2021ZA003 and No. 2020ZB002).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Approval
The Animal Care and Use Committee of Zhejiang Academy of Medical Sciences approved the protocols for the animal experiments.
Availability of Data and Materials
The data that support the findings of this study are available from the corresponding author upon reasonable request.
